

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0044a.6
(617 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
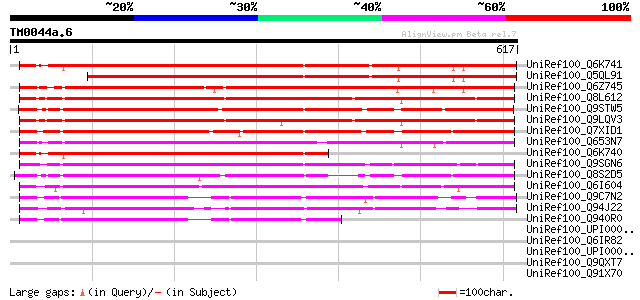
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6K741 Hypothetical protein P0419C03.10-1 [Oryza sativa] 751 0.0
UniRef100_Q5QL91 Hypothetical protein P0419C03.10-3 [Oryza sativa] 680 0.0
UniRef100_Q6Z745 Hypothetical protein P0487D09.21 [Oryza sativa] 528 e-148
UniRef100_Q8L612 Hypothetical protein At1g14780 [Arabidopsis tha... 518 e-145
UniRef100_Q9STW5 Hypothetical protein T22A6.120 [Arabidopsis tha... 511 e-143
UniRef100_Q9LQV3 F10B6.18 [Arabidopsis thaliana] 504 e-141
UniRef100_Q7XID1 Hypothetical protein P0039H02.131 [Oryza sativa] 499 e-139
UniRef100_Q653N7 Hypothetical protein P0431E05.15 [Oryza sativa] 489 e-136
UniRef100_Q6K740 Hypothetical protein P0419C03.10-2 [Oryza sativa] 480 e-134
UniRef100_Q9SGN6 F3M18.18 [Arabidopsis thaliana] 446 e-123
UniRef100_Q8S2D5 P0401G10.6 protein [Oryza sativa] 438 e-121
UniRef100_Q6I604 Hypothetical protein OJ1214_E03.15 [Oryza sativa] 437 e-121
UniRef100_Q9C7N2 Hypothetical protein F15D2.24 [Arabidopsis thal... 421 e-116
UniRef100_Q94J22 P0481E12.26 protein [Oryza sativa] 368 e-100
UniRef100_Q940R0 At1g29690/F15D2_24 [Arabidopsis thaliana] 286 1e-75
UniRef100_UPI00004386F2 UPI00004386F2 UniRef100 entry 44 0.014
UniRef100_Q6IR82 MGC82168 protein [Xenopus laevis] 43 0.024
UniRef100_UPI00004381D4 UPI00004381D4 UniRef100 entry 43 0.031
UniRef100_Q9QXT7 Complement component 6 [Mus musculus] 42 0.041
UniRef100_Q91X70 Complement component 6 [Mus musculus] 42 0.041
>UniRef100_Q6K741 Hypothetical protein P0419C03.10-1 [Oryza sativa]
Length = 634
Score = 751 bits (1938), Expect = 0.0
Identities = 390/622 (62%), Positives = 475/622 (75%), Gaps = 32/622 (5%)
Query: 12 ALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQNKRDILIPGAG-----GV 66
A +LG GFDL SDFRL+FAK EGR LV LDE RD+ +PG G
Sbjct: 7 AAMALGAGFDLTSDFRLKFAK---EGR--------LVELDEAGARDVPVPGGGVGGGAAA 55
Query: 67 TIKGVSENIRCDKGDLLRFKSDVLEFNQMSELLNQKSAVQGKIPSGYFNALFDLSGDWFR 126
++GV ++ DKGD +RF+SDVLEFNQMSELLNQKS+VQGK+PSGYFN LFDLSG W
Sbjct: 56 VLRGVPRDVGVDKGDRIRFRSDVLEFNQMSELLNQKSSVQGKVPSGYFNTLFDLSGAWMT 115
Query: 127 DAADTKYLAFDGYFISLYYLHLTASPIVLQEEVKKSVPAQWDPAALSRFIQTYGTHLIVG 186
DA +TK+LAFDGYFISLY LHL SP+VL++EV+ +VP +WDPAALSRFI+TYGTH+IV
Sbjct: 116 DAKETKHLAFDGYFISLYKLHLKTSPLVLRDEVRSAVPPKWDPAALSRFIKTYGTHIIVE 175
Query: 187 MAVGGQDVICVKQKHSSKIPPGDLRRHLEDLGDFLFSDVRSPPLLQRKAADGKQKVPEVF 246
MAVGGQDVICVKQ SS I DL+ HLEDLGDFLFSD R+ + RK DGK KVP+VF
Sbjct: 176 MAVGGQDVICVKQSPSSTISSADLKLHLEDLGDFLFSDGRNHSPIHRKTRDGKSKVPDVF 235
Query: 247 NRV-MQSNTMSFTSISETSSKDGLTIICSKRGGDMLKHSHSSWLQTVPSNPEAILFKFVP 305
R+ Q N + +S SE+S+KDGLTI CSKRGGD SHS WLQTVP P+AI+FKFVP
Sbjct: 236 VRMEQQPNNLHLSSYSESSTKDGLTITCSKRGGDASIASHSKWLQTVPRVPDAIMFKFVP 295
Query: 306 ISSLLAGIPGSGYLSHAINLYLRYKPPAEDLQYFLEFQIPRQWAPMFCELPLRHHQRRKS 365
I+SLL GIPGSGYLSHAINLYLRYKP EDLQ+FLEFQ+P QWAP+F EL L Q+RK
Sbjct: 296 ITSLLTGIPGSGYLSHAINLYLRYKPDPEDLQHFLEFQVPLQWAPLFNELIL-GPQKRKG 354
Query: 366 SSLSLQFNFLGPKLHVSSTQVLSEQKPVVGLRLYLEGRKCDRLALHIHHLSSLPNKMILS 425
S SLQF FLGPKL VS++QV S KPVVGLRLYLEGRKC+RLA+H+ HLSS P+ M+
Sbjct: 355 SYPSLQFRFLGPKLQVSTSQVSSSHKPVVGLRLYLEGRKCNRLAIHVQHLSSAPS-MLGD 413
Query: 426 SETSTPSMWRGSDDNESSNQFLEPVRWKRFSNVCTAVVKHDPNWLN---SSTGVYIVTGA 482
S +S+ S WR S+ E ++EP++WK +S VCT+ V ++P WL GV++VTGA
Sbjct: 414 SLSSSMSEWRESE--EVGAGYIEPIQWKSYSCVCTSKVDYNPEWLKRVPGGRGVFVVTGA 471
Query: 483 QLVSRCSWPKNVLHLRLLFTNIPNCSIRKTEWAAAPEASRKSSFLTNLSTTFS--FTQ-- 538
QLV++ +W + VLHLRL +T++P C+I++TEWAAAP AS++ SFLT +STT S FTQ
Sbjct: 472 QLVTKGTWSRKVLHLRLHYTHVPGCAIQRTEWAAAPAASQRGSFLTTISTTLSSPFTQLQ 531
Query: 539 -QSATSGPPKQAP---AMLNSGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVT 594
+A + PP+ P A+LNSG+YPDGPPVP+++ KLLK+V+ +EVV+GPHD PGHWLVT
Sbjct: 532 AAAAPAAPPRNEPAPAALLNSGVYPDGPPVPLQSRKLLKFVDMSEVVKGPHDVPGHWLVT 591
Query: 595 AAKLVTEGGKIGLQVKFALLDY 616
AAKLV +GGKIGL VKFALL+Y
Sbjct: 592 AAKLVKDGGKIGLNVKFALLNY 613
>UniRef100_Q5QL91 Hypothetical protein P0419C03.10-3 [Oryza sativa]
Length = 551
Score = 680 bits (1754), Expect = 0.0
Identities = 346/534 (64%), Positives = 421/534 (78%), Gaps = 16/534 (2%)
Query: 95 MSELLNQKSAVQGKIPSGYFNALFDLSGDWFRDAADTKYLAFDGYFISLYYLHLTASPIV 154
MSELLNQKS+VQGK+PSGYFN LFDLSG W DA +TK+LAFDGYFISLY LHL SP+V
Sbjct: 1 MSELLNQKSSVQGKVPSGYFNTLFDLSGAWMTDAKETKHLAFDGYFISLYKLHLKTSPLV 60
Query: 155 LQEEVKKSVPAQWDPAALSRFIQTYGTHLIVGMAVGGQDVICVKQKHSSKIPPGDLRRHL 214
L++EV+ +VP +WDPAALSRFI+TYGTH+IV MAVGGQDVICVKQ SS I DL+ HL
Sbjct: 61 LRDEVRSAVPPKWDPAALSRFIKTYGTHIIVEMAVGGQDVICVKQSPSSTISSADLKLHL 120
Query: 215 EDLGDFLFSDVRSPPLLQRKAADGKQKVPEVFNRV-MQSNTMSFTSISETSSKDGLTIIC 273
EDLGDFLFSD R+ + RK DGK KVP+VF R+ Q N + +S SE+S+KDGLTI C
Sbjct: 121 EDLGDFLFSDGRNHSPIHRKTRDGKSKVPDVFVRMEQQPNNLHLSSYSESSTKDGLTITC 180
Query: 274 SKRGGDMLKHSHSSWLQTVPSNPEAILFKFVPISSLLAGIPGSGYLSHAINLYLRYKPPA 333
SKRGGD SHS WLQTVP P+AI+FKFVPI+SLL GIPGSGYLSHAINLYLRYKP
Sbjct: 181 SKRGGDASIASHSKWLQTVPRVPDAIMFKFVPITSLLTGIPGSGYLSHAINLYLRYKPDP 240
Query: 334 EDLQYFLEFQIPRQWAPMFCELPLRHHQRRKSSSLSLQFNFLGPKLHVSSTQVLSEQKPV 393
EDLQ+FLEFQ+P QWAP+F EL L Q+RK S SLQF FLGPKL VS++QV S KPV
Sbjct: 241 EDLQHFLEFQVPLQWAPLFNELIL-GPQKRKGSYPSLQFRFLGPKLQVSTSQVSSSHKPV 299
Query: 394 VGLRLYLEGRKCDRLALHIHHLSSLPNKMILSSETSTPSMWRGSDDNESSNQFLEPVRWK 453
VGLRLYLEGRKC+RLA+H+ HLSS P+ M+ S +S+ S WR S+ E ++EP++WK
Sbjct: 300 VGLRLYLEGRKCNRLAIHVQHLSSAPS-MLGDSLSSSMSEWRESE--EVGAGYIEPIQWK 356
Query: 454 RFSNVCTAVVKHDPNWLN---SSTGVYIVTGAQLVSRCSWPKNVLHLRLLFTNIPNCSIR 510
+S VCT+ V ++P WL GV++VTGAQLV++ +W + VLHLRL +T++P C+I+
Sbjct: 357 SYSCVCTSKVDYNPEWLKRVPGGRGVFVVTGAQLVTKGTWSRKVLHLRLHYTHVPGCAIQ 416
Query: 511 KTEWAAAPEASRKSSFLTNLSTTFS--FTQ---QSATSGPPKQAP---AMLNSGIYPDGP 562
+TEWAAAP AS++ SFLT +STT S FTQ +A + PP+ P A+LNSG+YPDGP
Sbjct: 417 RTEWAAAPAASQRGSFLTTISTTLSSPFTQLQAAAAPAAPPRNEPAPAALLNSGVYPDGP 476
Query: 563 PVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIGLQVKFALLDY 616
PVP+++ KLLK+V+ +EVV+GPHD PGHWLVTAAKLV +GGKIGL VKFALL+Y
Sbjct: 477 PVPLQSRKLLKFVDMSEVVKGPHDVPGHWLVTAAKLVKDGGKIGLNVKFALLNY 530
>UniRef100_Q6Z745 Hypothetical protein P0487D09.21 [Oryza sativa]
Length = 620
Score = 528 bits (1359), Expect = e-148
Identities = 293/623 (47%), Positives = 400/623 (64%), Gaps = 36/623 (5%)
Query: 12 ALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQNKRDILIPGAGGVTIKGV 71
A+ LG+GFD+A D RL++ KG G G R+ + +PG G I V
Sbjct: 10 AVRCLGRGFDMAGDLRLKYCKG--GGAGCLVERRG-------ETTPLTVPGVG--VIADV 58
Query: 72 SENIRCDKGDLLRFKSDVLEFNQMSELLNQKSAVQGKIPSGYFNALFDL-SGDWFRDAAD 130
++RCDKGD +RFKSDVLEFN+MSEL NQ+S+V+GKIPSG FNA FDL SG W DA
Sbjct: 59 PADVRCDKGDRVRFKSDVLEFNKMSELFNQRSSVEGKIPSGQFNASFDLDSGSWAHDAPH 118
Query: 131 TKYLAFDGYFISLYYLHLTASPIVLQEEVKKSVPAQWDPAALSRFIQTYGTHLIVGMAVG 190
T+ LA DGYFISL+ L L + L V VP WDP+A++RFI+ YGTH+IVG+++G
Sbjct: 119 TRCLAMDGYFISLFDLRLDHRHLALDAGVLADVPPAWDPSAIARFIEKYGTHVIVGLSMG 178
Query: 191 GQDVICVKQKHSSKIPPGDLRRHLEDLGDFLFSDVRSPPLLQRKAADGKQKVPEVFN--- 247
GQDV+ VKQ SS + P +++ HL+ LGD LF+ + P L ++ D K K+PE FN
Sbjct: 179 GQDVVYVKQDKSSSLSPSEIKEHLDRLGDQLFTGTCAMPPLHCRSKD-KFKIPEAFNVFD 237
Query: 248 -RVMQSNTMSFTSISETSSKDGLTIICSKRGGDMLKHSHSSWLQTVPSNPEAILFKFVPI 306
+V Q T++ SSK+G+T+I SKRGG+ SHS WL TVP+ P+ I K VPI
Sbjct: 238 AQVAQQRLHGITTL--VSSKEGVTVIYSKRGGNTTVSSHSEWLLTVPAMPDVINVKLVPI 295
Query: 307 SSLLAGIPGSGYLSHAINLYLRYKPPAEDLQYFLEFQIPRQWAPMFCELPLRHHQRRKSS 366
+SL+ G+PG+G+LSHAINLYLRYKPP DL+YFL+FQ WAP+ ELPL R+ S
Sbjct: 296 TSLIRGVPGTGFLSHAINLYLRYKPPVADLRYFLDFQHHCVWAPVLGELPLGPCSHRQGS 355
Query: 367 SLSLQFNFLGPKLHVSSTQVLSEQKPVVGLRLYLEGRKCDRLALHIHHLSSLPNKMILSS 426
S +L F+ LG KL+VSST+V+ + PV G+RL+LEG+K +RL +H+ HLS+ P + ++
Sbjct: 356 SPALHFSLLGSKLYVSSTEVVVPKLPVTGMRLHLEGKKNNRLGIHLQHLSTTPT-FVAAA 414
Query: 427 ETSTPSMWRGSDDNESSNQFLEPVRWKRFSNVCTAVVKHDPNWL----NSSTGVYIVTGA 482
P +WRG+ + + +++ EPV+W+ + VCTA VK+DP W +V GA
Sbjct: 415 RADKPPVWRGT-EAVTDDRYYEPVQWRMLARVCTAPVKYDPRWCAGDRRRRPAACVVAGA 473
Query: 483 QL-VSRCSWPKNVLHLRLLFTNIPNCSIRKTEW---AAAPEASRKSSFLTNLSTTFSFTQ 538
QL V NVLHLRLL++ +P ++ +++W AA P + R SSFL+ + T
Sbjct: 474 QLHVVAHDAANNVLHLRLLYSQLPGYAVVQSKWARGAARPPSGRSSSFLSIPFSGSPSTS 533
Query: 539 QSAT--SGPPKQAP-----AMLNSGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHW 591
A G P+Q A +NSG++ GPPVPV A KLLK+V+ ++V GP D+PG+W
Sbjct: 534 SGAAEKGGRPEQGASPVGVANVNSGVFAGGPPVPVGAQKLLKFVDTSQVTMGPQDSPGYW 593
Query: 592 LVTAAKLVTEGGKIGLQVKFALL 614
LVT A+L + GKI L VKF+LL
Sbjct: 594 LVTGARLDVDKGKIMLHVKFSLL 616
>UniRef100_Q8L612 Hypothetical protein At1g14780 [Arabidopsis thaliana]
Length = 627
Score = 518 bits (1335), Expect = e-145
Identities = 277/617 (44%), Positives = 397/617 (63%), Gaps = 24/617 (3%)
Query: 12 ALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQNKRDILIPGAGGVTIKGV 71
A++SLGKGFDL +DFRL++ K +G GS+ RLVVLD+ R++ IPG G + V
Sbjct: 12 AVKSLGKGFDLTADFRLKYCK---DGDGSAGD-DRLVVLDQTQNRELHIPGFG--VFQNV 65
Query: 72 SENIRCDKGDLLRFKSDVLEFNQMSELLNQKSAVQGKIPSGYFNALFDL-SGDWFRDAAD 130
S +I CDKG+ RF+SD+L+FN+MSE NQ+S+V GKIPSG FNA F SG W DAA+
Sbjct: 66 SADINCDKGERTRFRSDILDFNKMSEYFNQRSSVTGKIPSGNFNATFGFQSGSWATDAAN 125
Query: 131 TKYLAFDGYFISLYYLHL-TASPIVLQEEVKKSVPAQWDPAALSRFIQTYGTHLIVGMAV 189
K L D ++L+ LH+ + + L + V+ +VP+ WDP L+RFI+ YGTH+I G++V
Sbjct: 126 VKSLGLDASVVTLFNLHIHNPNRLRLTDRVRNAVPSSWDPQLLARFIERYGTHVITGVSV 185
Query: 190 GGQDVICVKQKHSSKIPPGDLRRHLEDLGDFLF--SDVRSPPLLQRKA--ADGKQKVPEV 245
GGQDV+ V+Q SS + LR HL DLGD LF S + S L + + + K PE
Sbjct: 186 GGQDVVVVRQDKSSDLDNDLLRHHLYDLGDQLFTGSCLLSTRRLNKAYHHSHSQPKFPEA 245
Query: 246 FNRVMQSNTMSFTSISETSSKDGLTIICSKRGGDMLKHSHSSWLQTVPSNPEAILFKFVP 305
FN T++F + S +S++G+T+IC+KRGGD SHS WL TVP P+AI F F+P
Sbjct: 246 FNVFDDKQTVAFNNFS-INSQNGITVICAKRGGDGRAKSHSEWLITVPDKPDAINFNFIP 304
Query: 306 ISSLLAGIPGSGYLSHAINLYLRYKPPAEDLQYFLEFQIPRQWAPMFCELPLRHHQRRKS 365
I+SLL +PGSG LSHA++LYLRYKPP DLQYFL+F PR WAP+ +LP S
Sbjct: 305 ITSLLKDVPGSGLLSHAMSLYLRYKPPLMDLQYFLDFSGPRAWAPVHNDLPFGAAPNMAS 364
Query: 366 SSLSLQFNFLGPKLHVSSTQVLSEQKPVVGLRLYLEGRKCDRLALHIHHLSSLPNKMILS 425
+ +L NF+GPKL+V++T V SE+ PV G+R +LEG+KC+RLA+H+ HL + + +
Sbjct: 365 AYPALHINFMGPKLYVNTTPVTSEKNPVTGMRFFLEGKKCNRLAIHLQHLDN--TRTTVG 422
Query: 426 SETSTPSMWRGSDDNESSNQFLEPVRWKRFSNVCTAVVKHDPNWLNSSTG-------VYI 478
+ + +WRGSD ++++ EP+ K+FS+VCT VK+DPNW+ +++ +I
Sbjct: 423 EKITDEHIWRGSDQITDNDRYFEPLNGKKFSHVCTVPVKYDPNWIKTTSNHKSQNDVAFI 482
Query: 479 VTGAQLVSRCSWPKNVLHLRLLFTNIPNCSIRKTEWAAAP-EASRKSSFLTNLSTTFSFT 537
VTGAQL + K+VLHLRL +T + + + + W P S+KS +++S +
Sbjct: 483 VTGAQLEVKKHGSKSVLHLRLRYTKVSDHYVVQNSWVHGPIGTSQKSGIFSSMSMPLTSG 542
Query: 538 QQSATSGPPKQAPAMLNSGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAK 597
+ +L+SG++P GPPVP K++K+V+ +++ RGP +PGHWLVT +
Sbjct: 543 SVHHNMIQKDKNEVVLDSGVFPGGPPVPAN-NKIVKFVDLSQLCRGPQHSPGHWLVTGVR 601
Query: 598 LVTEGGKIGLQVKFALL 614
L + GK+ L VKFALL
Sbjct: 602 LYLDKGKLCLHVKFALL 618
>UniRef100_Q9STW5 Hypothetical protein T22A6.120 [Arabidopsis thaliana]
Length = 606
Score = 511 bits (1315), Expect = e-143
Identities = 276/605 (45%), Positives = 393/605 (64%), Gaps = 29/605 (4%)
Query: 11 VALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQNKRDILIPGAGGVTIKG 70
VA+ S+G G+DLA D RL++ KG GS SR L + + + +I++PG G++I
Sbjct: 13 VAIGSIGCGYDLAIDLRLKYCKG-----GSKDSRL-LDIKEGDDNCEIVLPG--GISIPN 64
Query: 71 VSENIRCDKGDLLRFKSDVLEFNQMSELLNQKSAVQGKIPSGYFNALFDLSGDWFRDAAD 130
VS++I+CDKG+ +RF+SD+L F QM+E NQ+ ++ GKIPSG FNA+F+ S W +DAA
Sbjct: 65 VSKSIKCDKGERMRFRSDILPFQQMAEQFNQELSLAGKIPSGLFNAMFEFSSCWQKDAAY 124
Query: 131 TKYLAFDGYFISLYYLHLTASPIVLQEEVKKSVPAQWDPAALSRFIQTYGTHLIVGMAVG 190
TK LAFDG FISLY + L S ++L+E VK++VP+ WDPAAL+RFI YGTH+IV + +G
Sbjct: 125 TKNLAFDGVFISLYSVALDKSQVLLREHVKQAVPSTWDPAALARFIDIYGTHIIVSVKMG 184
Query: 191 GQDVICVKQKHSSKIPPGDLRRHLEDLGDFLFSDVRSPPLLQRKAADGKQKVPEVFNRVM 250
G+DVI KQ+HSSK+ P DL++ L+++ D F + + KV R+
Sbjct: 185 GKDVIYAKQQHSSKLQPEDLQKRLKEVADKRFVEASVVHNTGSERVQASSKVETKEQRLR 244
Query: 251 QSNTMSFTSISETSSKDGLTIICSKRGG-DMLKHSHSSWLQTVPSNPEAILFKFVPISSL 309
++T +S+ ++K+ +C +RGG D H+ WLQTV P+ I F+PI+SL
Sbjct: 245 FADT---SSLGSYANKEDYVFMCKRRGGNDNRNLMHNEWLQTVQMEPDVISMSFIPITSL 301
Query: 310 LAGIPGSGYLSHAINLYLRYKPPAEDLQYFLEFQIPRQWAPMFCELPLRHHQRRKSSSLS 369
L G+PGSG+LSHAINLYLRYKPP E+L FLEFQ+PRQWAP+F ELPL QR++ S S
Sbjct: 302 LNGVPGSGFLSHAINLYLRYKPPIEELHQFLEFQLPRQWAPVFSELPL-GPQRKQQSCAS 360
Query: 370 LQFNFLGPKLHVSSTQVLSEQKPVVGLRLYLEGRKCDRLALHIHHLSSLPNKMILSSETS 429
LQF+F GPKL+V++T V ++P+ G+RLYLEGR+ +RLA+H+ HLSSLP L + +
Sbjct: 361 LQFSFFGPKLYVNTTPVDVGKRPITGMRLYLEGRRSNRLAIHLQHLSSLPKIYQLEDDLN 420
Query: 430 TPSMWRGSDDNESSNQFLEPVRWKRFSNVCTAVVKHDPNWLNSSTGVYIVTGAQLVSRCS 489
R ++ E V WK +S+VCT V+ D + + +VTGAQL
Sbjct: 421 -----RSIRQESHDRRYYEKVNWKNYSHVCTEPVESDDD-------LSVVTGAQLHVESH 468
Query: 490 WPKNVLHLRLLFTNIPNCS-IRKTEWAAAPEASRKSSFLTNLSTTFSFTQQSATSGPPKQ 548
KNVL LRL F+ + + ++ +EW A + KS +ST S +A PP+
Sbjct: 469 GFKNVLFLRLCFSRVVGATLVKNSEWDEAVGFAPKSGL---ISTLISHHFTAAQKPPPRP 525
Query: 549 APAMLNSGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIGLQ 608
A +NS IYP GPPVP +A KLLK+V+ +E+ RGP ++PG+W+V+ A+L+ E GKI L+
Sbjct: 526 ADVNINSAIYPGGPPVPTQAPKLLKFVDTSEMTRGPQESPGYWVVSGARLLVEKGKISLK 585
Query: 609 VKFAL 613
VK++L
Sbjct: 586 VKYSL 590
>UniRef100_Q9LQV3 F10B6.18 [Arabidopsis thaliana]
Length = 645
Score = 504 bits (1297), Expect = e-141
Identities = 276/635 (43%), Positives = 396/635 (61%), Gaps = 42/635 (6%)
Query: 12 ALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQNKRDILIPGAGGVTIKGV 71
A++SLGKGFDL +DFRL++ K +G GS+ RLVVLD+ R++ IPG G + V
Sbjct: 12 AVKSLGKGFDLTADFRLKYCK---DGDGSAGD-DRLVVLDQTQNRELHIPGFG--VFQNV 65
Query: 72 SENIRCDKGDLLRFKSDVLEFNQMSELLNQKSAVQGKIPSGYFNALFDL-SGDWFRDAAD 130
S +I CDKG+ RF+SD+L+FN+MSE NQ+S+V GKIPSG FNA F SG W DAA+
Sbjct: 66 SADINCDKGERTRFRSDILDFNKMSEYFNQRSSVTGKIPSGNFNATFGFQSGSWATDAAN 125
Query: 131 TKYLAFDGYFISLYYLHL-TASPIVLQEEVKKSVPAQWDPAALSRFIQTYGTHLIVGMAV 189
K L D ++L+ LH+ + + L + V+ +VP+ WDP L+RFI+ YGTH+I G++V
Sbjct: 126 VKSLGLDASVVTLFNLHIHNPNRLRLTDRVRNAVPSSWDPQLLARFIERYGTHVITGVSV 185
Query: 190 GGQDVICVKQKHSSKIPPGDLRRHLEDLGDFLF--SDVRSPPLLQRKA--ADGKQKVPEV 245
GGQDV+ V+Q SS + LR HL DLGD LF S + S L + + + K PE
Sbjct: 186 GGQDVVVVRQDKSSDLDNDLLRHHLYDLGDQLFTGSCLLSTRRLNKAYHHSHSQPKFPEA 245
Query: 246 FNRVMQSNTMSFTSISETSSKDGLTIICSKRGGDMLKHSHSSWLQTVPSNPEAILFKFVP 305
FN T++F + S +S++G+T+IC+KRGGD SHS WL TVP P+AI F F+P
Sbjct: 246 FNVFDDKQTVAFNNFS-INSQNGITVICAKRGGDGRAKSHSEWLITVPDKPDAINFNFIP 304
Query: 306 ISSLLAGIPGSGYLSHAINLYLRY------------------KPPAEDLQYFLEFQIPRQ 347
I+SLL +PGSG LSHA++LYLR KPP DLQYFL+F PR
Sbjct: 305 ITSLLKDVPGSGLLSHAMSLYLRCNYSSCLILTTEMSSNTFDKPPLMDLQYFLDFSGPRA 364
Query: 348 WAPMFCELPLRHHQRRKSSSLSLQFNFLGPKLHVSSTQVLSEQKPVVGLRLYLEGRKCDR 407
WAP+ +LP S+ +L NF+GPKL+V++T V SE+ PV G+R +LEG+KC+R
Sbjct: 365 WAPVHNDLPFGAAPNMASAYPALHINFMGPKLYVNTTPVTSEKNPVTGMRFFLEGKKCNR 424
Query: 408 LALHIHHLSSLPNKMILSSETSTPSMWRGSDDNESSNQFLEPVRWKRFSNVCTAVVKHDP 467
LA+H+ HL + + + + + +WRGSD ++++ EP+ K+FS+VCT VK+DP
Sbjct: 425 LAIHLQHLDN--TRTTVGEKITDEHIWRGSDQITDNDRYFEPLNGKKFSHVCTVPVKYDP 482
Query: 468 NWLNSSTG-------VYIVTGAQLVSRCSWPKNVLHLRLLFTNIPNCSIRKTEWAAAP-E 519
NW+ +++ +IVTGAQL + K+VLHLRL +T + + + + W P
Sbjct: 483 NWIKTTSNHKSQNDVAFIVTGAQLEVKKHGSKSVLHLRLRYTKVSDHYVVQNSWVHGPIG 542
Query: 520 ASRKSSFLTNLSTTFSFTQQSATSGPPKQAPAMLNSGIYPDGPPVPVRAGKLLKYVEAAE 579
S+KS +++S + + +L+SG++P GPPVP K++K+V+ ++
Sbjct: 543 TSQKSGIFSSMSMPLTSGSVHHNMIQKDKNEVVLDSGVFPGGPPVPAN-NKIVKFVDLSQ 601
Query: 580 VVRGPHDTPGHWLVTAAKLVTEGGKIGLQVKFALL 614
+ RGP +PGHWLVT +L + GK+ L VKFALL
Sbjct: 602 LCRGPQHSPGHWLVTGVRLYLDKGKLCLHVKFALL 636
>UniRef100_Q7XID1 Hypothetical protein P0039H02.131 [Oryza sativa]
Length = 608
Score = 499 bits (1284), Expect = e-139
Identities = 267/608 (43%), Positives = 397/608 (64%), Gaps = 34/608 (5%)
Query: 12 ALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQNKRDILIPGAGGVTIKGV 71
A++S+G G+D+A D RL++ K SS L+ LD +DI++PG G+T+ GV
Sbjct: 14 AIQSIGLGYDIAHDIRLKYCKQ------RSSPDPLLIELDHDEVQDIVLPG--GLTVAGV 65
Query: 72 SENIRCDKGDLLRFKSDVLEFNQMSELLNQKSAVQGKIPSGYFNALFDLSGDWFRDAADT 131
S++I+CDKG+ RF+SDVL F QMSE NQ+ ++ GKIPSG FN +F+ +G W +DAA+T
Sbjct: 66 SKSIKCDKGERTRFRSDVLSFQQMSEQFNQELSLSGKIPSGLFNTMFEFTGCWQKDAANT 125
Query: 132 KYLAFDGYFISLYYLHLTASPIVLQEEVKKSVPAQWDPAALSRFIQTYGTHLIVGMAVGG 191
K LAFDG+ I+LY + L+ + IVL++ VK++VP+ W+PAAL+RFI+ +GTH++VG+ +GG
Sbjct: 126 KSLAFDGWCITLYTVALSKAQIVLRDHVKQAVPSTWEPAALARFIRKFGTHVVVGIKMGG 185
Query: 192 QDVICVKQKHSSKIPPGDLRRHLEDLGDFLFSDVRSPPLLQRKAADGKQKVPEVFNRVMQ 251
+D+I +KQ+HSS + D+++ L+++ D F D K + GK K+ R +
Sbjct: 186 KDIIYLKQQHSSTLQAVDVQKRLKEMSDRRFLDANGQSDFSFKDSYGKDKID---TREHR 242
Query: 252 SNTMSFTSISETSSKDGLTIICSKRGG---DMLKHSHSSWLQTVPSNPEAILFKFVPISS 308
+ + ++ SSK+ L ++ +RGG D+L SHS WL TV + P+ I F+PI+S
Sbjct: 243 LRFVDSSPLNSYSSKEDLVMMPKRRGGRDKDIL--SHSEWLNTVQAEPDVISMSFIPITS 300
Query: 309 LLAGIPGSGYLSHAINLYLRYKPPAEDLQYFLEFQIPRQWAPMFCELPLRHHQRRKSSSL 368
LL G+PG G+L+HAINLYLRYKP E+L FLEFQ+PRQWAP++ +LPL QR++ S++
Sbjct: 301 LLNGVPGCGFLNHAINLYLRYKPQIEELHQFLEFQLPRQWAPVYSDLPL-GPQRKRQSTV 359
Query: 369 SLQFNFLGPKLHVSSTQVLSEQKPVVGLRLYLEGRKCDRLALHIHHLSSLPNKMILSSET 428
SL N +GPKL+V + V ++PV G+RL+LEG++ ++LA+H+ HL SLP + L +
Sbjct: 360 SLPVNLIGPKLYVCTNMVDVGKRPVTGIRLFLEGKRSNKLAIHLQHLCSLPQILQLEDDP 419
Query: 429 STPSMWRGSDDNESSNQFLEPV-RWKRFSNVCTAVVKHDPNWLNSSTGVYIVTGAQLVSR 487
+ ++ EP+ WKRFS+VCTA V+ D + IVTGAQL
Sbjct: 420 -----YNDQTPEAYDRKYYEPIGSWKRFSHVCTAPVESDDS--------SIVTGAQLEVV 466
Query: 488 CSWPKNVLHLRLLFTNIPNC-SIRKTEWAAAPEASRKSSFLTNLSTTFSFTQQSATSGPP 546
K +L LRL F+ + N S+R EW +P ++KS ++ L +T T +A P
Sbjct: 467 SHGFKKILFLRLHFSKVCNATSVRNPEWEGSPNLAQKSGLISTLISTHFST--AAQKPAP 524
Query: 547 KQAPAMLNSGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIG 606
+ A +NS +YP GPPVPV+ KLLK+V+ E++RGP D PG+W+V+ AKL E GKI
Sbjct: 525 RPADVNINSAVYPGGPPVPVQTPKLLKFVDPTEMMRGPQDLPGYWVVSGAKLQLERGKIS 584
Query: 607 LQVKFALL 614
L+VK++LL
Sbjct: 585 LRVKYSLL 592
>UniRef100_Q653N7 Hypothetical protein P0431E05.15 [Oryza sativa]
Length = 614
Score = 489 bits (1258), Expect = e-136
Identities = 278/621 (44%), Positives = 376/621 (59%), Gaps = 44/621 (7%)
Query: 12 ALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQNKRDILIPGAGGVTIKGV 71
A+ LG+G D+A D RL+ K EG LV + + +PG G + GV
Sbjct: 16 AVRCLGRGVDMAGDLRLKHCKD--EGGC-------LVARSGEKAAAVAVPGVG--VVAGV 64
Query: 72 SENIRCDKGDLLRFKSDVLEFNQMSELLNQKSAVQGKIPSGYFNALFDLSGD-WFRDAAD 130
+++ KGD +RFKSDVLEFN+MS+L N +S++ GKIPSG FN+ FD D W DA D
Sbjct: 65 PADVKFGKGDRIRFKSDVLEFNKMSDLFNHRSSLPGKIPSGLFNSCFDFGSDSWASDAGD 124
Query: 131 TKYLAFDGYFISLYYLHLTASPIVLQEEVKKSVPAQWDPAALSRFIQTYGTHLIVGMAVG 190
T+ LAFDGYFISL L L P+ L V VPA WDP+A++ FI+ YGTH+IVG+++G
Sbjct: 125 TRCLAFDGYFISLLDLRLDCRPLALAGHVVADVPAAWDPSAIASFIEKYGTHIIVGLSMG 184
Query: 191 GQDVICVKQKHSSKIPPGDLRRHLEDLGDFLFSDVRSPPLLQRKAADGKQKVPEVFNRVM 250
GQDV+ VKQ SS + P ++ HL+ LGD LF+ + P K+ D K KVPE FN
Sbjct: 185 GQDVVYVKQDKSSPLSPSVIKEHLDKLGDQLFTGTCTLPPSHCKSRDHKFKVPEAFNVFD 244
Query: 251 QSNTMSFTS--ISETSSKDGLTIICSKRGGDMLKHSHSSWLQTVPSNPEAILFKFVPISS 308
T + S K+G+T+I SKRGGD +HS WL TVP P+AI FK VPI+S
Sbjct: 245 AQMTRQRIEGMTAPMSCKEGVTVIYSKRGGDTAASNHSEWLPTVPLMPDAINFKLVPITS 304
Query: 309 LLAGIPGSGYLSHAINLYLRYKPPAEDLQYFLEFQIPRQWAPMFCELPLRHHQRRKSSSL 368
LL G+ G G+LSHAINLYLRYKPP +L+YFL+FQ R WAP+ +LPL R+ ++
Sbjct: 305 LLKGVAGVGFLSHAINLYLRYKPPVAELRYFLDFQHHRLWAPVLSDLPLGLCSNRQGTNP 364
Query: 369 SLQFNFLGPKLHVSSTQVLSEQKPVVGLRLYLEGRKCDRLALHIHHLSSLPNKMILSSET 428
+L F+ V+ + P+ G+RL+LEG+K +RL +H+ HLS+ P I +
Sbjct: 365 ALHFSL-----------VIVPKLPITGMRLHLEGKKNNRLGIHLQHLSTTPT-FIAGGWS 412
Query: 429 STPSMWRGSDDNESSNQFLEPVRWKRFSNVCTAVVKHDPNWLNSSTG------VYIVTGA 482
P WRGS + + ++ EPV+ + F++VCT VKHDP WL + G Y+V+GA
Sbjct: 413 GRPPAWRGS-EAIADERYYEPVQRRMFAHVCTVPVKHDPRWLAAGDGGGGRPAAYVVSGA 471
Query: 483 QLVSRCSWPKNVLHLRLLFTNIPNCSIRKTEWA-------AAPEASRKSSFLTNLSTTFS 535
QL + +VLHLRLL+T +P S+ ++ WA AA + K SFL+ + S
Sbjct: 472 QLHVKAHESTSVLHLRLLYTELPGHSVVQSRWAHGGGGGGAARMSGVKGSFLS--MSFAS 529
Query: 536 FTQQSATSGPPKQAPAMLN--SGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLV 593
+A KQA A LN SG++ GPP PV A +LLK+VE ++V GP D PG+WLV
Sbjct: 530 MAAAAAEKEQQKQAAARLNVDSGVFAGGPPAPVGAQRLLKFVETSQVTMGPQDCPGYWLV 589
Query: 594 TAAKLVTEGGKIGLQVKFALL 614
T AKL + G+I L VKF+LL
Sbjct: 590 TGAKLDVDKGRISLHVKFSLL 610
>UniRef100_Q6K740 Hypothetical protein P0419C03.10-2 [Oryza sativa]
Length = 406
Score = 480 bits (1236), Expect = e-134
Identities = 252/382 (65%), Positives = 292/382 (75%), Gaps = 18/382 (4%)
Query: 12 ALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQNKRDILIPGAG-----GV 66
A +LG GFDL SDFRL+FAK EGR LV LDE RD+ +PG G
Sbjct: 7 AAMALGAGFDLTSDFRLKFAK---EGR--------LVELDEAGARDVPVPGGGVGGGAAA 55
Query: 67 TIKGVSENIRCDKGDLLRFKSDVLEFNQMSELLNQKSAVQGKIPSGYFNALFDLSGDWFR 126
++GV ++ DKGD +RF+SDVLEFNQMSELLNQKS+VQGK+PSGYFN LFDLSG W
Sbjct: 56 VLRGVPRDVGVDKGDRIRFRSDVLEFNQMSELLNQKSSVQGKVPSGYFNTLFDLSGAWMT 115
Query: 127 DAADTKYLAFDGYFISLYYLHLTASPIVLQEEVKKSVPAQWDPAALSRFIQTYGTHLIVG 186
DA +TK+LAFDGYFISLY LHL SP+VL++EV+ +VP +WDPAALSRFI+TYGTH+IV
Sbjct: 116 DAKETKHLAFDGYFISLYKLHLKTSPLVLRDEVRSAVPPKWDPAALSRFIKTYGTHIIVE 175
Query: 187 MAVGGQDVICVKQKHSSKIPPGDLRRHLEDLGDFLFSDVRSPPLLQRKAADGKQKVPEVF 246
MAVGGQDVICVKQ SS I DL+ HLEDLGDFLFSD R+ + RK DGK KVP+VF
Sbjct: 176 MAVGGQDVICVKQSPSSTISSADLKLHLEDLGDFLFSDGRNHSPIHRKTRDGKSKVPDVF 235
Query: 247 NRV-MQSNTMSFTSISETSSKDGLTIICSKRGGDMLKHSHSSWLQTVPSNPEAILFKFVP 305
R+ Q N + +S SE+S+KDGLTI CSKRGGD SHS WLQTVP P+AI+FKFVP
Sbjct: 236 VRMEQQPNNLHLSSYSESSTKDGLTITCSKRGGDASIASHSKWLQTVPRVPDAIMFKFVP 295
Query: 306 ISSLLAGIPGSGYLSHAINLYLRYKPPAEDLQYFLEFQIPRQWAPMFCELPLRHHQRRKS 365
I+SLL GIPGSGYLSHAINLYLRYKP EDLQ+FLEFQ+P QWAP+F EL L Q+RK
Sbjct: 296 ITSLLTGIPGSGYLSHAINLYLRYKPDPEDLQHFLEFQVPLQWAPLFNELIL-GPQKRKG 354
Query: 366 SSLSLQFNFLGPKLHVSSTQVL 387
S SLQF FLGPKL VS++QVL
Sbjct: 355 SYPSLQFRFLGPKLQVSTSQVL 376
>UniRef100_Q9SGN6 F3M18.18 [Arabidopsis thaliana]
Length = 612
Score = 446 bits (1146), Expect = e-123
Identities = 255/606 (42%), Positives = 361/606 (59%), Gaps = 23/606 (3%)
Query: 12 ALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQNKRDILIPGAGGVTIKGV 71
A+ +G G+DL SD R K +G RLV +D RD++ PG G+ + V
Sbjct: 18 AVSVIGLGYDLCSDVRFSACKTTPDG-------SRLVEIDPTRNRDLIFPG--GIVVNNV 68
Query: 72 SENIRCDKGDLLRFKSDVLEFNQMSELLNQKSAVQGKIPSGYFNALFDLSGDWFRDAADT 131
S +I+CDKG+ R +SD+L FNQMSE NQ + GKIPSG FN +F S W +DA+
Sbjct: 69 SSSIKCDKGERTRLRSDILSFNQMSEKFNQDMCLSGKIPSGMFNNMFAFSKCWPKDASSV 128
Query: 132 KYLAFDGYFISLYYLHLTASPIVLQEEVKKSVPAQWDPAALSRFIQTYGTHLIVGMAVGG 191
K LA+DG+FISLY + + + L++EVK+ VP+ WD AAL+ FI+ YGTH++VG+ +GG
Sbjct: 129 KTLAYDGWFISLYSVEIVRKQLTLRDEVKREVPSSWDSAALAGFIEKYGTHVVVGVTMGG 188
Query: 192 QDVICVKQKHSSKIPPGDLRRHLEDLGDFLFSDVRSPPLLQRKAADGKQKVPEVFNRVMQ 251
+DVI VKQ S P ++++ L+ GD F GK K + +Q
Sbjct: 189 KDVIHVKQMRKSNHEPEEIQKMLKHWGDERFCVDPVESKSPASVYSGKPKEENLLQWGLQ 248
Query: 252 SNTMSFTSISETSSK-DGLTIICSKRGGDMLKHSHSSWLQTVPSNPEAILFKFVPISSLL 310
S +S +K + + +C +RGG L SH WL TV P I FVPI+SLL
Sbjct: 249 PFGTSVSSAVVMHTKNEEIMRVCIRRGGVDLGQSHERWLSTVSQAPNVISMCFVPITSLL 308
Query: 311 AGIPGSGYLSHAINLYLRYKPPAEDLQYFLEFQIPRQWAPMFCELPLRHHQRRKSSSLSL 370
+G+PG+G+LSHA+NLYLRYKPP E+L FLEFQ+PRQWAP++ +LPL +R K SS SL
Sbjct: 309 SGLPGTGFLSHAVNLYLRYKPPIEELHQFLEFQLPRQWAPVYGDLPL-GLRRSKQSSPSL 367
Query: 371 QFNFLGPKLHVSSTQVLSEQKPVVGLRLYLEGRKCDRLALHIHHLSSLPNKMILSSETST 430
QF+ +GPKL+V++++V S ++PV GLR +LEG+K + LA+H+ HLS+ P + LS + +
Sbjct: 368 QFSLMGPKLYVNTSKVDSGERPVTGLRFFLEGKKGNHLAIHLQHLSACPPSLHLSHDDTY 427
Query: 431 PSMWRGSDDNESSNQFLEPVRWKRFSNVCTAVVKHDPNWLNSSTGVYIVTGAQLVSRCSW 490
+ + + PV+W FS+VCT V++ N S IVT A L +
Sbjct: 428 EPI-----EEPVEKGYYVPVKWGIFSHVCTYPVQY--NGARSDDTASIVTKAWLEVKGMG 480
Query: 491 PKNVLHLRLLFTNIPNCSIRKTEWAAAPEASRKSSFLTNLSTTFS--FTQQSATSGPPKQ 548
+ VL LRL F+ + RK+ W SRKS + +ST S + AT+ P Q
Sbjct: 481 MRKVLFLRLGFSLDASAVTRKSCWDNLSTNSRKSGVFSMISTRLSTGLSPNPATTKP--Q 538
Query: 549 APAMLNSGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIGLQ 608
+ +NS +YP GP PV+ KLL V+ EV+RGP + PG+W+VT AKL E GKI ++
Sbjct: 539 SKIDINSAVYPRGPSPPVKP-KLLSLVDTKEVMRGPEEQPGYWVVTGAKLCVEAGKISIK 597
Query: 609 VKFALL 614
K++LL
Sbjct: 598 AKYSLL 603
>UniRef100_Q8S2D5 P0401G10.6 protein [Oryza sativa]
Length = 573
Score = 438 bits (1126), Expect = e-121
Identities = 250/615 (40%), Positives = 369/615 (59%), Gaps = 70/615 (11%)
Query: 6 EGIEMVALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQNKRDILIPGAGG 65
+ + A+ S+G G+D+A+D RL+ K RGS L+ LD +DI++PG
Sbjct: 8 QSVSESAIRSIGLGYDIANDIRLKNCKQ----RGSPDPL--LIELDHDKVQDIVLPG--N 59
Query: 66 VTIKGVSENIRCDKGDLLRFKSDVLEFNQMSELLNQKSAVQGKIPSGYFNALFDLSGDWF 125
+T+ GVS++I+CDKG+ +RF+SDVL F QMSE N++ ++ GKIPSG+FNA+F+ +G W
Sbjct: 60 LTVTGVSKSIKCDKGERMRFRSDVLSFQQMSEQFNRELSLSGKIPSGFFNAMFEFTGCWQ 119
Query: 126 RDAADTKYLAFDGYFISLYYLHLTASPIVLQEEVKKSVPAQWDPAALSRFIQTYGTHLIV 185
+DA+ TK LAFDG+ I+LY + L+ + I+L++ VK++VP+ W+PAAL+RFI+ +GTH++V
Sbjct: 120 KDASITKSLAFDGWCITLYTVALSKAHIILKDHVKQAVPSTWEPAALARFIKKFGTHIVV 179
Query: 186 GMAVGGQDVICVKQKHSSKIPPGDLRRHLEDLGDFLFSDVRSPP---LLQRKAADGKQKV 242
G+ +GG+DVI +KQ+HSS + D+++ L+++ D F D L A D K +
Sbjct: 180 GVKMGGKDVIYLKQQHSSSLQAVDVQKRLKEMSDQRFLDANGHSDISLADSYAKDNKVEA 239
Query: 243 PEVFNRVMQSNTMSFTSISETSSKDGLTIICSKRGG-DMLKHSHSSWLQTVPSNPEAILF 301
E R ++SN ++ SS + L ++ +RGG D SHS WL TV + P+ I
Sbjct: 240 REQRLRFVESNPLN-----SYSSNEELVMMPKRRGGRDKDIISHSEWLNTVQAEPDVISM 294
Query: 302 KFVPISSLLAGIPGSGYLSHAINLYLRYKPPAEDLQYFLEFQIPRQWAPMFCELPLRHHQ 361
F+PI+SLL G+PG G+L+HAINLYLRYKP E+L FLEFQ+PRQWAP++ +LPL Q
Sbjct: 295 SFIPITSLLNGVPGCGFLNHAINLYLRYKPRVEELHQFLEFQLPRQWAPVYSDLPLGP-Q 353
Query: 362 RRKSSSLSLQFNFLGPKLHVSSTQVLSEQKPVVGLRLYLEGRKCDRLALHIHHLSSLPNK 421
R++ SS SL N +GPKL+V + ++
Sbjct: 354 RKRQSSASLPVNLIGPKLYVCTNMIIQ--------------------------------- 380
Query: 422 MILSSETSTPSMWRGSDDNESSNQFLEPV-RWKRFSNVCTAVVKHDPNWLNSSTGVYIVT 480
L +T P ++ EP+ WKRFS+VCTA V D + IVT
Sbjct: 381 --LEDDTYNPQT-----PEAEIRKYYEPIGSWKRFSHVCTAPVDSDDS--------SIVT 425
Query: 481 GAQLVSRCSWPKNVLHLRLLFTNIPNC-SIRKTEWAAAPEASRKSSFLTNLSTTFSFTQQ 539
GA L K +L LRL F+ + N S++ EW +P +KS ++ L +T T
Sbjct: 426 GAHLEVVSHGFKKILFLRLHFSKVCNATSVKNPEWDGSPNLGQKSGLISTLISTHFST-- 483
Query: 540 SATSGPPKQAPAMLNSGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAKLV 599
+A P+ A +NS +YP GPPVPV+ KLL++V+ E++RGP D PG+W+V+ AKL
Sbjct: 484 AALKPAPRPAEVNINSAVYPGGPPVPVQTPKLLRFVDTTEMLRGPQDLPGYWVVSGAKLH 543
Query: 600 TEGGKIGLQVKFALL 614
E GKI L+VK++LL
Sbjct: 544 LERGKISLRVKYSLL 558
>UniRef100_Q6I604 Hypothetical protein OJ1214_E03.15 [Oryza sativa]
Length = 628
Score = 437 bits (1125), Expect = e-121
Identities = 256/618 (41%), Positives = 366/618 (58%), Gaps = 40/618 (6%)
Query: 12 ALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQN---KRDILIPGAGGVTI 68
A+ ++G G+DL SD RL K + RLV +D + +R++++P G +
Sbjct: 27 AVGAIGCGYDLTSDLRLSRVK----------AGGRLVDIDGASGAARRELVLPW--GAVV 74
Query: 69 KGVSENIRCDKGDLLRFKSDVLEFNQMSELLNQKSAVQGKIPSGYFNALFDLSGDWFRDA 128
GV I DKG+ RF+SDVL F QM+E +NQ +V GKIPSG FNA+FD G W +DA
Sbjct: 75 GGVPVGIVADKGERTRFRSDVLSFAQMAEQVNQTMSVAGKIPSGAFNAMFDYHGCWHKDA 134
Query: 129 ADTKYLAFDGYFISLYYLHLTASPIVLQEEVKKSVPAQWDPAALSRFIQTYGTHLIVGMA 188
A T L FDG FI LY + + + L + VK+ VP WDPAAL+ FI YGTH+I G+
Sbjct: 135 AATGSLCFDGRFIELYAVEAPRAHLALLDRVKRDVPPFWDPAALAEFIDKYGTHVIAGVK 194
Query: 189 VGGQDVICVKQKHSSKIPPGDLRRHLEDLGDFLFSDVRSPPLLQRKAADGKQKVPEVFNR 248
+GG+DV+C+KQ S + D++ L+ L D + SP L A D K + +
Sbjct: 195 MGGKDVVCIKQLKGSNLTQSDVQSRLKKLSDDKLAQ-DSPESL--TARDDKFLLGLNGSL 251
Query: 249 VMQSNTMSFTSI--SETSSKDGLTIICSKRGGDMLKHSHSSWLQTVPSNPEAILFKFVPI 306
++ + ++ S S S KD + I +RGG HS+WL T+ +P+ I FVPI
Sbjct: 252 LLGPGSAAWRSFRPSVVSHKDDILSIHIRRGGVDNGQGHSNWLSTISGSPDVISMAFVPI 311
Query: 307 SSLLAGIPGSGYLSHAINLYLRYKPPAEDLQYFLEFQIPRQWAPMFCELPLRHHQRRKSS 366
+SLL G+ G G+L+HA+NLYLRYKPP E+L FLEFQ+PRQWAP F ELPL R+K +
Sbjct: 312 TSLLTGVRGCGFLNHAVNLYLRYKPPIEELHQFLEFQVPRQWAPEFGELPLALGPRKKKN 371
Query: 367 SL-SLQFNFLGPKLHVSSTQVLSEQKPVVGLRLYLEGRKCDRLALHIHHLSSLPNKMILS 425
SL SLQF +GPKLHV++ + S +PV G+RL+LEG+K +RL +H+ HLS+ P + ++
Sbjct: 372 SLPSLQFTLMGPKLHVTTAKADSGNRPVTGIRLFLEGKKNNRLGVHLQHLSATPGTITIA 431
Query: 426 SETSTPSMWRGSDDNESSNQFLEPVRWKRFSNVCTAVVKHDPNWLNSSTGVYIVTGAQLV 485
E ++ D ++EP++ S+VCTA V+++ ++ IVT A L
Sbjct: 432 GEVAS-----AEDATVRERDYIEPIKSPLLSHVCTAPVQYNGARIDDCAA--IVTRAWLE 484
Query: 486 SRCSWPKNVLHLRLLFTNIPNCSIRKTEWAAAPEASRKSSFLTNLSTTFSFTQQSATSG- 544
+ + K VL LRL F+ + + IR++EW SRKS +LS FS +A +G
Sbjct: 485 VQETCLKKVLFLRLGFSGVASTKIRRSEWDGPFVVSRKSG---SLSALFSARLSAAGAGG 541
Query: 545 --------PPKQAPAMLNSGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAA 596
P +NS I+P GPPVP+ ++ +YV+ EV+RGP D PG+W+VT A
Sbjct: 542 SAQMMQQQQPVGEKVEVNSAIFPKGPPVPLPVQRMARYVDTTEVMRGPADLPGYWVVTGA 601
Query: 597 KLVTEGGKIGLQVKFALL 614
KL EGGK+ L+VK++LL
Sbjct: 602 KLCIEGGKVALKVKYSLL 619
>UniRef100_Q9C7N2 Hypothetical protein F15D2.24 [Arabidopsis thaliana]
Length = 561
Score = 421 bits (1083), Expect = e-116
Identities = 236/608 (38%), Positives = 354/608 (57%), Gaps = 73/608 (12%)
Query: 12 ALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQNKRDILIPGAGGVTIKGV 71
A+++LG+GFD+ SD RL + KG + RLV ++E RD+ + + G + V
Sbjct: 24 AIQALGRGFDVTSDVRLLYCKG--------APGSRLVRIEEGQNRDLEL--SHGFLLPNV 73
Query: 72 SENIRCDKGDLLRFKSDVLEFNQMSELLNQKSAVQGKIPSGYFNALFDLSGDWFRDAADT 131
+I C +G+ + V F++M+E N +S V+G IP G FNA+F+ +G W DAA T
Sbjct: 74 PADIDCSRGNSGTQRISVCSFHEMAEEFNVRSGVKGNIPLGCFNAMFNYTGSWQVDAAST 133
Query: 132 KYLAFDGYFISLYYLHLTASPIVLQEEVKKSVPAQWDPAALSRFIQTYGTHLIVGMAVGG 191
K LA GYFI LY + L +VL E++++VP+ WDPA+L+ FI+ YGTH++ + +GG
Sbjct: 134 KSLALVGYFIPLYDVKLAKLTLVLHNEIRRAVPSSWDPASLASFIENYGTHIVTSVTIGG 193
Query: 192 QDVICVKQKHSSKIPPGDLRRHLEDLGDFLFSDVRSPPLLQRKAADGKQKVPEVFNRVMQ 251
+DV+ ++Q SS +P ++ ++ D+ + +R +
Sbjct: 194 RDVVYIRQHQSSPLPVSEIENYVNDM---------------------------IKHRFHE 226
Query: 252 SNTMSFTSISETSSKDGLTIICSKRGGDMLKHSHSSWLQTVPSNPEAILFKFVPISSLLA 311
+ + S T + KD +T+I +RGGD L+ SH+ W +TVP+ P+ I F PI SLL
Sbjct: 227 AESQSITGPLKYKDKD-ITVIFRRRGGDDLEQSHARWAETVPAAPDIINMTFTPIVSLLE 285
Query: 312 GIPGSGYLSHAINLYLRYKPPAEDLQYFLEFQIPRQWAPMFCELPLRHHQRRKSSSLSLQ 371
G+PG +L+ AI LYL YKPP EDLQYFL++QI R WAP L QR++ SLQ
Sbjct: 286 GVPGLRHLTRAIELYLEYKPPIEDLQYFLDYQIARAWAPEQSNL-----QRKEPVCSSLQ 340
Query: 372 FNFLGPKLHVSSTQVLSEQKPVVGLRLYLEGRKCDRLALHIHHLSSLPNKMILSSETSTP 431
F+ +GPKL +S+ QV +KPV GLRL LEG K +RL++H+ HL SLP + ++ P
Sbjct: 341 FSLMGPKLFISADQVTVGRKPVTGLRLSLEGSKQNRLSIHLQHLVSLPKILQPHWDSHVP 400
Query: 432 ---SMWRGSDDNESSNQFLEPVRWKRFSNVCTAVVKHDPNWLNSSTGVYIVTGAQLVSRC 488
W+G ++ +S ++ EP++WK FS+V T+ ++H + +GV+IVTGAQL
Sbjct: 401 IGAPKWQGPEEQDS--RWFEPIKWKNFSHVSTSPIEHTETHIGDLSGVHIVTGAQLGVWD 458
Query: 489 SWPKNVLHLRLLFTNIPNCSIRKTEWAAAPEASRKSSFLTNLSTTFSFTQQSATSGPPKQ 548
KNVLHL+LLF+ +P C+IR++ W P AS + + GP
Sbjct: 459 FGSKNVLHLKLLFSKVPGCTIRRSVWDHTPVAS---------------SGRLEPGGPSTS 503
Query: 549 APAMLNSGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIGLQ 608
+ SG ++GKL K V+++E+++GP D PGHWLVT AKL E GKI L+
Sbjct: 504 SSTEEVSG----------QSGKLAKIVDSSEMLKGPQDLPGHWLVTGAKLGVEKGKIVLR 553
Query: 609 VKFALLDY 616
VK++LL+Y
Sbjct: 554 VKYSLLNY 561
>UniRef100_Q94J22 P0481E12.26 protein [Oryza sativa]
Length = 553
Score = 368 bits (945), Expect = e-100
Identities = 221/620 (35%), Positives = 342/620 (54%), Gaps = 92/620 (14%)
Query: 12 ALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQNKRDILIPGAGGVTIKGV 71
AL+++G+G D A D RL + KG RL++LDE RD+ I G ++GV
Sbjct: 11 ALQAVGRGLDAAGDHRLLYCKGTG----------RLLMLDESRARDLTINGG---VLRGV 57
Query: 72 SENIRCDKGD--LLRFKSD---------VLEFNQMSELLNQKSAV-QGKIPSGYFNALFD 119
++ ++G L R + V F +M+E N+K+ + + +P G FN+LF
Sbjct: 58 PPDVVVEEGHGILERIRQVPGPPTDEPVVCSFPKMAECFNRKAGLLETTVPLGSFNSLFS 117
Query: 120 LSGDWFRDAADTKYLAFDGYFISLYYLHLTASPIVLQEEVKKSVPAQWDPAALSRFIQTY 179
+G W D A TK LA DGY + L+ + +T+ + L E VK+++P WDP+AL+ FI+ Y
Sbjct: 118 FTGSWKNDEAATKSLAIDGYSVPLFKVKITSGELFLHESVKRAIPHSWDPSALASFIENY 177
Query: 180 GTHLIVGMAVGGQDVICVKQKHSSKIPPGDLRRHLEDLGDFLFSDVRSPPLLQRKAADGK 239
GTH+I + VGG+D + +KQ SS++ + R +++++G FSD D K
Sbjct: 178 GTHIITSVTVGGKDEVYIKQHSSSQLSELEFRNYVKEIGSERFSD-----------GDSK 226
Query: 240 QKVPEVFNRVMQSNTMSFTSISETSSKDGLTIICSKRGGDMLKHSHSSWLQTVPSNPEAI 299
+ S KD +T+I +RGG L + + W++TV S P+ I
Sbjct: 227 LNATPI----------------NYSEKD-MTVIFRRRGGCDLVQNFNDWIKTVQSAPDVI 269
Query: 300 LFKFVPISSLLAGIPGSGYLSHAINLYLRYKPPAEDLQYFLEFQIPRQWAPMFCELPLRH 359
F+PI SL+ +PG +L+ AI LYL+YKP E+LQYFL+FQ+ WAP+ + +H
Sbjct: 270 GMTFLPIVSLVGDMPGKKHLARAIELYLKYKPQIEELQYFLDFQVQLVWAPVPPGIAGQH 329
Query: 360 HQRRKSSSLSLQFNFLGPKLHVSSTQVLSEQKPVVGLRLYLEGRKCDRLALHIHHLSSLP 419
R++ SLQF+ +GPKL VS+ Q+ ++PV GL+L LEG K +RLA+H+ HL SLP
Sbjct: 330 --RKEPVCPSLQFSLMGPKLFVSTEQISVGRRPVTGLKLCLEGAKQNRLAIHLQHLGSLP 387
Query: 420 NKMIL---SSETSTPSMWRGSDDNESSNQFLEPVRWKRFSNVCTAVVKHDPNWLNSSTGV 476
+ S T P W+G ++ +S ++ EP++W+ F++V TA +++ + +GV
Sbjct: 388 KIFVPHWDSHITIGPPKWQGPEEQDS--RWFEPIKWRNFAHVSTAPIEYTETSITDLSGV 445
Query: 477 YIVTGAQLVSRCSWPKNVLHLRLLFTNIPNCSIRKTEWAAAPEASRKSSFLTNLSTTFSF 536
YIVTGAQL K+VLHL+LLF+ +P C+IR++ W +P S+
Sbjct: 446 YIVTGAQLGVWDFGAKSVLHLKLLFSRVPGCTIRRSVWDHSPS-----------SSLVHR 494
Query: 537 TQQSATSGPPKQAPAMLNSGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAA 596
T ++++S KL+K V+ E ++GP D PGHWLVT A
Sbjct: 495 TDEASSSSSDN---------------------AKLVKIVDMTETLKGPQDAPGHWLVTGA 533
Query: 597 KLVTEGGKIGLQVKFALLDY 616
KL E GKI ++ K++LL+Y
Sbjct: 534 KLGVEKGKIVVRAKYSLLNY 553
>UniRef100_Q940R0 At1g29690/F15D2_24 [Arabidopsis thaliana]
Length = 383
Score = 286 bits (733), Expect = 1e-75
Identities = 156/393 (39%), Positives = 231/393 (58%), Gaps = 43/393 (10%)
Query: 12 ALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQNKRDILIPGAGGVTIKGV 71
A+++LG+GFD+ SD RL + KG + RLV ++E RD+ + + G + V
Sbjct: 24 AIQALGRGFDVTSDVRLLYCKG--------APGSRLVRIEEGQNRDLEL--SHGFLLPNV 73
Query: 72 SENIRCDKGDLLRFKSDVLEFNQMSELLNQKSAVQGKIPSGYFNALFDLSGDWFRDAADT 131
+I C +G+ + V F++M+E N +S V+G IP G FNA+F+ +G W DAA T
Sbjct: 74 PADIDCSRGNSGTQRISVCSFHEMAEEFNVRSGVKGNIPLGCFNAMFNYTGSWQVDAAST 133
Query: 132 KYLAFDGYFISLYYLHLTASPIVLQEEVKKSVPAQWDPAALSRFIQTYGTHLIVGMAVGG 191
K LA GYFI LY + L +VL E++++VP+ WDPA+L+ FI+ YGTH++ + +GG
Sbjct: 134 KSLALVGYFIPLYDVKLAKLTLVLHNEIRRAVPSSWDPASLASFIENYGTHIVTSVTIGG 193
Query: 192 QDVICVKQKHSSKIPPGDLRRHLEDLGDFLFSDVRSPPLLQRKAADGKQKVPEVFNRVMQ 251
+DV+ ++Q SS +P ++ ++ D+ + +R +
Sbjct: 194 RDVVYIRQHQSSPLPVSEIENYVNDM---------------------------IKHRFHE 226
Query: 252 SNTMSFTSISETSSKDGLTIICSKRGGDMLKHSHSSWLQTVPSNPEAILFKFVPISSLLA 311
+ + S T + KD +T+I +RGGD L+ SH+ W +TVP+ P+ I F PI SLL
Sbjct: 227 AESQSITGPLKYKDKD-ITVIFRRRGGDDLEQSHARWAETVPAAPDIINMTFTPIVSLLE 285
Query: 312 GIPGSGYLSHAINLYLRYKPPAEDLQYFLEFQIPRQWAPMFCELPLRHHQRRKSSSLSLQ 371
G+PG +L+ AI LYL YKPP EDLQYFL++QI R WAP L QR++ SLQ
Sbjct: 286 GVPGLWHLTRAIELYLEYKPPIEDLQYFLDYQIARAWAPEQSNL-----QRKEPVCSSLQ 340
Query: 372 FNFLGPKLHVSSTQVLSEQKPVVGLRLYLEGRK 404
F+ +GPKL +S+ QV +KPV GLRL LEG K
Sbjct: 341 FSLMGPKLFISADQVTVGRKPVTGLRLSLEGSK 373
>UniRef100_UPI00004386F2 UPI00004386F2 UniRef100 entry
Length = 517
Score = 43.9 bits (102), Expect = 0.014
Identities = 51/196 (26%), Positives = 80/196 (40%), Gaps = 19/196 (9%)
Query: 142 SLYYLHLTASPIVLQE--EVKKSVPAQWDPAALSRFIQTYGTHLIVGMAVGGQDVICVKQ 199
S Y ++ +P + E E KS+PA +D A I TYGTH + +GGQ
Sbjct: 145 SFYRYNVAENPPLHAEFLEAIKSLPASYDRDAYRSLIYTYGTHYTTNVKLGGQIKAITAI 204
Query: 200 KHSSKIPPGDLRRHLEDLGDFLFSDVRSPPLLQRKAADGKQKVPEVFNRVMQSNTMSFTS 259
K G ++D D S S ++ +A K++ + M +N +
Sbjct: 205 KTCQAAVSGLTDTAIKDCLDVEASGSYSVATVKAEAHFCKEQ-----QKKMGTNQKFSSM 259
Query: 260 ISETSSKDGLTIICSKRGGDMLKHSHS-------SWLQTVPSNPEAILFKFVPISSLL-A 311
SE + II G+ L S S SWL+++ S P+ + + P+ LL A
Sbjct: 260 FSERQT----DIIGGNINGEHLLFSGSSHPNALNSWLESLKSLPDVVQYSLKPLHFLLSA 315
Query: 312 GIPGSGYLSHAINLYL 327
P L A+ Y+
Sbjct: 316 KHPARKGLKKAVEEYI 331
>UniRef100_Q6IR82 MGC82168 protein [Xenopus laevis]
Length = 935
Score = 43.1 bits (100), Expect = 0.024
Identities = 33/159 (20%), Positives = 65/159 (40%), Gaps = 11/159 (6%)
Query: 163 VPAQWDPAALSRFIQTYGTHLIVGMAVGGQ-DVICVKQKHSSKIPPGDLRRHLEDLGDFL 221
+P +++ SR +GTH I ++GG DV+ Q + + L +++ +
Sbjct: 348 LPLEYNYPLYSRIFDDFGTHYITEGSMGGSYDVLF--QYSTENVQSSGLTD--QEMAHCV 403
Query: 222 FSDVRSPPLL------QRKAADGKQKVPEVFNRVMQSNTMSFTSISETSSKDGLTIICSK 275
F ++R+ R + +QS+ S + I ++ + K
Sbjct: 404 FEEIRTRRFFFFVKTTHRTTCTTNKMTERYSGSFLQSSEKSISFIKGGRAEFAAKLAWQK 463
Query: 276 RGGDMLKHSHSSWLQTVPSNPEAILFKFVPISSLLAGIP 314
G + ++ W+ + NP+ I +K PI L+ GIP
Sbjct: 464 EGASPDEKAYKDWVASTVDNPDLINYKLAPILDLITGIP 502
>UniRef100_UPI00004381D4 UPI00004381D4 UniRef100 entry
Length = 525
Score = 42.7 bits (99), Expect = 0.031
Identities = 51/207 (24%), Positives = 86/207 (40%), Gaps = 32/207 (15%)
Query: 143 LYYLHLTASPIVLQEEVK---KSVPAQWDPAALSRF---IQTYGTHLIVGMAVGG--QDV 194
+YY + AS L E++ ++PA +D + ++ +GTH I+ + +GG Q V
Sbjct: 164 VYYSYRIASSAPLHHELRDDFSNLPATYDNQTKESYLSLVEKFGTHYIIQVKLGGGVQSV 223
Query: 195 ICVKQKHSSKIPPGDLRRHLEDLGDFLFSDVRSPPLLQRKAAD-GKQKVPEVFNRVMQSN 253
VKQ ++ L D +V++ ++ A+ G+ V + QS
Sbjct: 224 TSVKQCMAT-------------LQDLSVDEVKTCLDVEASASVMGRISVDTAYRHCKQSQ 270
Query: 254 TMSFTSISETSS-KDGLTIICSKRGGDML--------KHSHSSWLQTVPSNPEAILFKFV 304
++ S +S D LT I D ++ WL +VP P+ I F
Sbjct: 271 DKKLSAHSFANSFSDRLTEITGGHTQDSALLFSASNDPGAYKQWLSSVPQQPDVISFSLK 330
Query: 305 PISSLL-AGIPGSGYLSHAINLYLRYK 330
P+ L+ A P L AI+ Y+ K
Sbjct: 331 PLHMLMPAKNPKRDQLRRAIHDYVLQK 357
>UniRef100_Q9QXT7 Complement component 6 [Mus musculus]
Length = 769
Score = 42.4 bits (98), Expect = 0.041
Identities = 29/154 (18%), Positives = 65/154 (41%), Gaps = 2/154 (1%)
Query: 163 VPAQWDPAALSRFIQTYGTHLIVGMAVGGQDVICVKQKHSSKIPPGDLRRHLEDLGDFLF 222
+P +++ A SR +GTH ++GG+ + + G ++ +
Sbjct: 345 LPLEYNSAVYSRIFDDFGTHYFTSGSLGGKYDLIYQFSRQELQNSGLTEEEAQNCVQYET 404
Query: 223 SDVRSPPL-LQRKAADGKQKVPEVFN-RVMQSNTMSFTSISETSSKDGLTIICSKRGGDM 280
++ + + ++ K K+ E + +Q + S + + S+ + K
Sbjct: 405 KKLKFLYMEIHKEDTCTKNKLSEKYGGSFLQGSEKSISLVQGGRSQQAAALAWEKGTSGP 464
Query: 281 LKHSHSSWLQTVPSNPEAILFKFVPISSLLAGIP 314
++ +S WL++V NP + +K PI+ L+ IP
Sbjct: 465 EENVYSEWLESVKENPAVVDYKLAPITDLVRNIP 498
>UniRef100_Q91X70 Complement component 6 [Mus musculus]
Length = 769
Score = 42.4 bits (98), Expect = 0.041
Identities = 29/154 (18%), Positives = 65/154 (41%), Gaps = 2/154 (1%)
Query: 163 VPAQWDPAALSRFIQTYGTHLIVGMAVGGQDVICVKQKHSSKIPPGDLRRHLEDLGDFLF 222
+P +++ A SR +GTH ++GG+ + + G ++ +
Sbjct: 345 LPLEYNSAVYSRIFDDFGTHYFTSGSLGGKYDLIYQFSRQELQNSGLTEEEAQNCVQYET 404
Query: 223 SDVRSPPL-LQRKAADGKQKVPEVFN-RVMQSNTMSFTSISETSSKDGLTIICSKRGGDM 280
++ + + ++ K K+ E + +Q + S + + S+ + K
Sbjct: 405 KKLKFLYMEIHKEDTCTKNKLSEKYGGSFLQGSEKSISLVQGGRSQQAAALAWEKGTSGP 464
Query: 281 LKHSHSSWLQTVPSNPEAILFKFVPISSLLAGIP 314
++ +S WL++V NP + +K PI+ L+ IP
Sbjct: 465 EENVYSEWLESVKENPAVVDYKLAPITDLVRNIP 498
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.135 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,054,784,811
Number of Sequences: 2790947
Number of extensions: 45099340
Number of successful extensions: 110117
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 24
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 109972
Number of HSP's gapped (non-prelim): 41
length of query: 617
length of database: 848,049,833
effective HSP length: 133
effective length of query: 484
effective length of database: 476,853,882
effective search space: 230797278888
effective search space used: 230797278888
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 78 (34.7 bits)
Lotus: description of TM0044a.6


