

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0042.13
(250 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
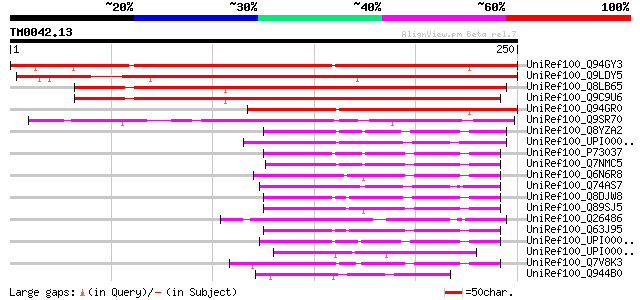
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q94GY3 Putative FK506-binding protein [Oryza sativa] 315 5e-85
UniRef100_Q9LDY5 T10F20.17 protein [Arabidopsis thaliana] 287 2e-76
UniRef100_Q8LB65 Putative FK506-binding protein [Arabidopsis tha... 237 2e-61
UniRef100_Q9C9U6 Hypothetical protein F25P22.7 [Arabidopsis thal... 235 9e-61
UniRef100_Q94GR0 Putative FK506-binding protein, 5'-partial [Ory... 192 9e-48
UniRef100_Q9SR70 T22K18.11 protein [Arabidopsis thaliana] 73 6e-12
UniRef100_Q8YZA2 Peptidyl-prolyl cis-trans isomerase [Anabaena sp.] 69 9e-11
UniRef100_UPI00002A4102 UPI00002A4102 UniRef100 entry 68 2e-10
UniRef100_P73037 Peptidyl-prolyl cis-trans isomerase [Synechocys... 66 9e-10
UniRef100_Q7NMC5 Peptidyl-prolyl cis-trans isomerase [Gloeobacte... 64 4e-09
UniRef100_Q6N6R8 FKBP-type peptidyl-prolyl cis-trans isomerase p... 64 4e-09
UniRef100_Q74AS7 FKBP-type peptidyl-prolyl cis-trans isomerase [... 64 5e-09
UniRef100_Q8DJW8 Peptidyl-prolyl cis-trans isomerase [Synechococ... 63 6e-09
UniRef100_Q89SJ5 Peptidyl-prolyl cis-trans isomerase [Bradyrhizo... 63 6e-09
UniRef100_Q26486 46 kDa FK506-binding nuclear protein [Spodopter... 63 8e-09
UniRef100_Q63J95 Peptidyl-prolyl cis-trans isomerase [Burkholder... 62 1e-08
UniRef100_UPI00002EEB60 UPI00002EEB60 UniRef100 entry 62 1e-08
UniRef100_UPI00002E8307 UPI00002E8307 UniRef100 entry 60 4e-08
UniRef100_Q7V8K3 FKBP-type peptidyl-prolyl cis-trans isomerase (... 60 4e-08
UniRef100_Q944B0 AT4g26550/M3E9_20 [Arabidopsis thaliana] 60 4e-08
>UniRef100_Q94GY3 Putative FK506-binding protein [Oryza sativa]
Length = 252
Score = 315 bits (808), Expect = 5e-85
Identities = 160/255 (62%), Positives = 200/255 (77%), Gaps = 8/255 (3%)
Query: 1 MATFFGSPPVF-SLPLTRNSHHIASSSPTPP---PIPPQQSQSPTASSSQSVQGTAKAQP 56
MATF GS P F + P+ H + SP PP P P QQ +PT + QS+Q +A+P
Sbjct: 1 MATFLGSSPAFLARPVAAKPHLSCAQSPRPPSAQPPPEQQPPTPTPTQQQSMQAQPQARP 60
Query: 57 QNPIKPVTSSTKVDSTDWIATSLTRRFGLGAGLAWVGFLAFGVVSEQIKTRLEVSQQEAN 116
+ P +++ DSTDW+ATSLTRRFG+GAGLAWVGFLAFGVVSEQ+KTR EV+QQ AN
Sbjct: 61 RRA--PAAAASGADSTDWVATSLTRRFGIGAGLAWVGFLAFGVVSEQLKTRFEVAQQLAN 118
Query: 117 TRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKP 176
T+DVE+E+EV LPNGIRY E++VGGG PRPGDLVVID+ G+V + GE FV+TF K+P
Sbjct: 119 TKDVEQEQEVVLPNGIRYYEMRVGGGDVPRPGDLVVIDLKGRV-TGGEAFVDTFGDGKRP 177
Query: 177 LALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLG-SGVQIPPLAT 235
LALVMGSRPY++G+CEG+EYVL++M+AGGKR+V+VPP LGF ++GAD G + Q+PP AT
Sbjct: 178 LALVMGSRPYTRGMCEGVEYVLRSMRAGGKRRVVVPPALGFGDDGADFGDAAAQVPPGAT 237
Query: 236 LEYIVEVDKVSIAPA 250
LEY+VEVDKVSIAPA
Sbjct: 238 LEYVVEVDKVSIAPA 252
>UniRef100_Q9LDY5 T10F20.17 protein [Arabidopsis thaliana]
Length = 247
Score = 287 bits (735), Expect = 2e-76
Identities = 152/258 (58%), Positives = 192/258 (73%), Gaps = 26/258 (10%)
Query: 4 FFGSPPVFSL--PLTRN-SHHIASSSPTPPPIPPQQSQSPTASSSQSVQGTAKAQPQNPI 60
F + P SL P T+ S+H +S + PP P S P P P
Sbjct: 5 FTATAPFLSLSKPFTKTASYHQCYASSSNPPEPESSSPPP---------------PPPPP 49
Query: 61 KPVTSSTK----VDSTDWIATSLTRRFGLGAGLAWVGFLAFGVVSEQIKTRLEVSQQEAN 116
+P+ S K V++TDW+A+SLTRRFG+GAGLAW GFLAFGV+SEQIKTR+EVSQ+ AN
Sbjct: 50 QPLASQQKRKKNVETTDWVASSLTRRFGIGAGLAWAGFLAFGVISEQIKTRIEVSQEVAN 109
Query: 117 TRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTF----EG 172
TRDVE+E+E+ LPNGIRY + +VGGGA+PR GDLVVID+ G+V+ TG+VFV+TF +
Sbjct: 110 TRDVEEEKEIVLPNGIRYYDQRVGGGATPRAGDLVVIDLKGQVQGTGQVFVDTFGTKDKK 169
Query: 173 DKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPP 232
KPLALV+GS+PYSKG+CEGI+YVL++MKAGGKR+VIVPP LGF +GA+L SG+QIPP
Sbjct: 170 KMKPLALVVGSKPYSKGLCEGIDYVLRSMKAGGKRRVIVPPSLGFGVDGAELESGLQIPP 229
Query: 233 LATLEYIVEVDKVSIAPA 250
A+LEYIVE+D+VSIAPA
Sbjct: 230 NASLEYIVEIDRVSIAPA 247
>UniRef100_Q8LB65 Putative FK506-binding protein [Arabidopsis thaliana]
Length = 234
Score = 237 bits (605), Expect = 2e-61
Identities = 120/214 (56%), Positives = 154/214 (71%), Gaps = 5/214 (2%)
Query: 33 PPQQSQSPTASSSQSVQGTAKAQPQNPIKPVTSSTKVDSTDWIATSLTRRFGLGAGLAWV 92
P Q + S S SV T QP V + + TDWI + +TRRFG+GAG W
Sbjct: 18 PSQHPKQSLLSQSLSVTFTENPQPT----AVVTLQEQQLTDWITSPVTRRFGIGAGFTWA 73
Query: 93 GFLAFGVVSEQIK-TRLEVSQQEANTRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLV 151
GFLAFGVVSEQ+K +RL+V Q+E NTR +EK+EE+ LPNGIRY +L+VG GA+P G LV
Sbjct: 74 GFLAFGVVSEQMKKSRLDVFQEEDNTRGLEKQEEIILPNGIRYYDLQVGSGATPSSGYLV 133
Query: 152 VIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIV 211
V D+ G+V T +VFV+TF G K LA+VM SRPYSKG+C+GIE+VL++MKAGGKR+VI+
Sbjct: 134 VFDVKGQVHGTEQVFVDTFGGKGKSLAMVMDSRPYSKGLCQGIEHVLRSMKAGGKRRVII 193
Query: 212 PPQLGFRENGADLGSGVQIPPLATLEYIVEVDKV 245
PP LGF + + G G++IPP ATL+YI+EVD V
Sbjct: 194 PPSLGFGDRNVEFGQGLEIPPSATLDYIIEVDTV 227
>UniRef100_Q9C9U6 Hypothetical protein F25P22.7 [Arabidopsis thaliana]
Length = 451
Score = 235 bits (599), Expect = 9e-61
Identities = 119/211 (56%), Positives = 152/211 (71%), Gaps = 5/211 (2%)
Query: 33 PPQQSQSPTASSSQSVQGTAKAQPQNPIKPVTSSTKVDSTDWIATSLTRRFGLGAGLAWV 92
P Q + S S SV T QP V + + TDWI + +TRRFG+GAG W
Sbjct: 18 PSQHPKQSLLSQSLSVTFTENPQPT----AVVTLQEQQLTDWITSPVTRRFGIGAGFTWA 73
Query: 93 GFLAFGVVSEQIK-TRLEVSQQEANTRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLV 151
GFLAFGVVSEQ+K +RL+V Q+E NTR +EK+EE+ LPNGIRY +L+VG GA+P G LV
Sbjct: 74 GFLAFGVVSEQMKKSRLDVFQEEDNTRGLEKQEEIILPNGIRYYDLQVGSGATPSSGYLV 133
Query: 152 VIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIV 211
V D+ G+V T +VFV+TF G K LA+VM SRPYSKG+C+GIE+VL++MKAGGKRKVI+
Sbjct: 134 VFDVKGQVHGTEQVFVDTFGGKGKSLAMVMDSRPYSKGLCQGIEHVLRSMKAGGKRKVII 193
Query: 212 PPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
PP LGF + + G G++IPP ATL+YI+EV
Sbjct: 194 PPSLGFGDRNVEFGQGLEIPPSATLDYIIEV 224
>UniRef100_Q94GR0 Putative FK506-binding protein, 5'-partial [Oryza sativa]
Length = 149
Score = 192 bits (487), Expect = 9e-48
Identities = 93/134 (69%), Positives = 115/134 (85%), Gaps = 2/134 (1%)
Query: 118 RDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPL 177
RDVE+E+EV LPNGIRY E++VGGG PRPGDLVVID+ G+V GE FV+TF K+PL
Sbjct: 17 RDVEQEQEVVLPNGIRYYEMRVGGGDVPRPGDLVVIDLKGRVTG-GEAFVDTFGDGKRPL 75
Query: 178 ALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLG-SGVQIPPLATL 236
ALVMGSRPY++G+CEG+EYVL++M+AGGKR+V+VPP LGF ++GAD G + Q+PP ATL
Sbjct: 76 ALVMGSRPYTRGMCEGVEYVLRSMRAGGKRRVVVPPALGFGDDGADFGDAAAQVPPGATL 135
Query: 237 EYIVEVDKVSIAPA 250
EY+VEVDKVSIAPA
Sbjct: 136 EYVVEVDKVSIAPA 149
>UniRef100_Q9SR70 T22K18.11 protein [Arabidopsis thaliana]
Length = 230
Score = 73.2 bits (178), Expect = 6e-12
Identities = 67/249 (26%), Positives = 110/249 (43%), Gaps = 39/249 (15%)
Query: 10 VFSLPLTRNSHHIASSSPTPPPIPPQQSQSPTASSSQSVQGTAKA-QPQNPIKPVTSSTK 68
+ ++ L HH SSS P P ++ QS S G K + + + P S +
Sbjct: 2 ILTMKLVHPLHHSLSSSI---PFPSRKRQSKPYRCSLPSPGCEKVIRTETVLPPAPVSCE 58
Query: 69 VDSTDWIATSLTRRFGLGAGLAWVGFLAFGVVSEQIKTRLEVSQQEANTRDVEKEEEVTL 128
RR LG LA A G++S + S++ + + + TL
Sbjct: 59 -----------GRRVLLGCLLA----TASGILSTGSAEAVSTSRRALRASKLPESDFTTL 103
Query: 129 PNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYS- 187
PNG++Y ++KVG GA G V + + K + G F+ + +G V G PY
Sbjct: 104 PNGLKYYDIKVGNGAEAVKGSRVAVHYVAKWK--GITFMTSRQG-----LGVGGGTPYGF 156
Query: 188 -------KGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIV 240
V +G++ ++ M+ GG+R VIVPP+L + + G +IPP AT+E +
Sbjct: 157 DVGQSERGNVLKGLDLGVEGMRVGGQRLVIVPPELAYGKKGVQ-----EIPPNATIELDI 211
Query: 241 EVDKVSIAP 249
E+ + +P
Sbjct: 212 ELLSIKQSP 220
>UniRef100_Q8YZA2 Peptidyl-prolyl cis-trans isomerase [Anabaena sp.]
Length = 165
Score = 69.3 bits (168), Expect = 9e-11
Identities = 45/120 (37%), Positives = 70/120 (57%), Gaps = 10/120 (8%)
Query: 126 VTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRP 185
VT +G++Y E++ G GA+P+ G VV+ G +E G F ++ + ++ P + +G
Sbjct: 55 VTTDSGLKYVEIEEGTGATPKSGQTVVVHYTGTLED-GTKFDSSRDRNR-PFSFTIGVGQ 112
Query: 186 YSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEVDKV 245
KG EG L TMK GG+R++I+P +LG+ GA G IPP ATL + VE+ +V
Sbjct: 113 VIKGWDEG----LSTMKVGGRRQLIIPSELGYGARGA----GGVIPPYATLLFDVELLEV 164
>UniRef100_UPI00002A4102 UPI00002A4102 UniRef100 entry
Length = 131
Score = 67.8 bits (164), Expect = 2e-10
Identities = 45/130 (34%), Positives = 68/130 (51%), Gaps = 9/130 (6%)
Query: 116 NTRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKK 175
N D K T+ NG+ +L VG G + + VV++ G +E G +F ++ ++
Sbjct: 8 NNCDSNKRSVKTMDNGLVIEDLVVGEGQEAKDYNKVVVNYTGTLED-GSIFDSSLNPGRE 66
Query: 176 PLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLAT 235
P +G KG G+ K MK GGKRK+ +PP+LG+ D G+G IPP AT
Sbjct: 67 PFTFTLGVGSVIKGWDLGV----KDMKVGGKRKLTIPPELGY----GDKGAGNVIPPGAT 118
Query: 236 LEYIVEVDKV 245
L + VE+ +V
Sbjct: 119 LIFEVELLEV 128
>UniRef100_P73037 Peptidyl-prolyl cis-trans isomerase [Synechocystis sp.]
Length = 201
Score = 65.9 bits (159), Expect = 9e-10
Identities = 44/117 (37%), Positives = 64/117 (54%), Gaps = 10/117 (8%)
Query: 126 VTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRP 185
VT +G++Y + VG G SP G V + G++ + G F ++ + +K P +G
Sbjct: 91 VTTESGLQYIDEVVGEGPSPTKGQKVEVHYTGRL-TDGTKFDSSVDRNK-PFTFTIGVGQ 148
Query: 186 YSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
KG EG+ TM+ GGKRK+I+PP L + GA G IPP ATLE+ VE+
Sbjct: 149 VIKGWDEGVA----TMQVGGKRKLIIPPDLAYGSRGA----GGVIPPNATLEFEVEL 197
>UniRef100_Q7NMC5 Peptidyl-prolyl cis-trans isomerase [Gloeobacter violaceus]
Length = 161
Score = 63.9 bits (154), Expect = 4e-09
Identities = 43/116 (37%), Positives = 64/116 (55%), Gaps = 10/116 (8%)
Query: 127 TLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPY 186
T +G++Y + VG GASP+ G V + G +E G+ F ++ + + P + +G
Sbjct: 52 TTTSGLKYLDETVGNGASPQKGQRVTVHYTGTLED-GKKFDSSRDRGQ-PFSFTIGVGQV 109
Query: 187 SKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
+G EG+ TMK GGKRK++VP LG+ GA G IPP ATL + VE+
Sbjct: 110 IQGWDEGVA----TMKVGGKRKLVVPANLGYGARGA----GGVIPPNATLLFDVEL 157
>UniRef100_Q6N6R8 FKBP-type peptidyl-prolyl cis-trans isomerase precursor
[Rhodopseudomonas palustris]
Length = 152
Score = 63.9 bits (154), Expect = 4e-09
Identities = 44/125 (35%), Positives = 63/125 (50%), Gaps = 12/125 (9%)
Query: 121 EKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGD---KKPL 177
E + VT P+G++ + +VG GASP G + V+ G + G F+ +P
Sbjct: 33 ESAKSVTTPSGLQIVDTQVGTGASPARGQICVMHYTGWLYENGAK-TRKFDSSVDRNEPF 91
Query: 178 ALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLE 237
+G KG EG+ +MK GGKR +I+PP LG+ GA G IPP ATL
Sbjct: 92 EFPIGMGRVIKGWDEGVA----SMKVGGKRTLIIPPDLGYGARGA----GGVIPPNATLV 143
Query: 238 YIVEV 242
+ VE+
Sbjct: 144 FDVEL 148
>UniRef100_Q74AS7 FKBP-type peptidyl-prolyl cis-trans isomerase [Geobacter
sulfurreducens]
Length = 138
Score = 63.5 bits (153), Expect = 5e-09
Identities = 44/119 (36%), Positives = 65/119 (53%), Gaps = 10/119 (8%)
Query: 124 EEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGS 183
+ VT +G+ Y +L G GA+P G V + G +E+ + + G+ P +G+
Sbjct: 25 DAVTTASGLSYVDLAAGSGAAPVAGKPVKVHYTGWLENGTKFDSSVDRGE--PFVFTIGA 82
Query: 184 RPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
G EG+ +MK GGKR++IVPPQLG+ GA G+G IPP ATL + VE+
Sbjct: 83 GEVIPGWDEGV----MSMKVGGKRRLIVPPQLGY---GA-AGAGGVIPPNATLIFEVEL 133
>UniRef100_Q8DJW8 Peptidyl-prolyl cis-trans isomerase [Synechococcus elongatus]
Length = 162
Score = 63.2 bits (152), Expect = 6e-09
Identities = 41/117 (35%), Positives = 64/117 (54%), Gaps = 10/117 (8%)
Query: 126 VTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRP 185
VT P+G++Y ++ VG GA P+ GD V + G + + G +F ++ +P +G
Sbjct: 52 VTTPSGLQYEDIVVGSGAQPQVGDRVTVHYTGML-TDGRIF-DSSRDRGQPFQFQIGVGQ 109
Query: 186 YSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
KG EG+ +M GG+R++I+PP LG+ G G IPP ATL + VE+
Sbjct: 110 VIKGWDEGV----GSMHVGGQRRLIIPPNLGYGARGV----GGVIPPNATLIFDVEL 158
>UniRef100_Q89SJ5 Peptidyl-prolyl cis-trans isomerase [Bradyrhizobium japonicum]
Length = 154
Score = 63.2 bits (152), Expect = 6e-09
Identities = 43/120 (35%), Positives = 63/120 (51%), Gaps = 12/120 (10%)
Query: 126 VTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGD---KKPLALVMG 182
+T +G++ + VG GASP+PG + V+ G + G+ F+ K+P +G
Sbjct: 40 MTTASGLQIIDTAVGTGASPQPGQICVMHYTGWLYENGQKG-KKFDSSVDRKEPFEFPIG 98
Query: 183 SRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
G EG+ +MK GGKR +I+PPQLG+ GA G IPP ATL + VE+
Sbjct: 99 KGRVIAGWDEGVA----SMKVGGKRTLIIPPQLGYGARGA----GGVIPPNATLMFDVEL 150
>UniRef100_Q26486 46 kDa FK506-binding nuclear protein [Spodoptera frugiperda]
Length = 412
Score = 62.8 bits (151), Expect = 8e-09
Identities = 45/141 (31%), Positives = 73/141 (50%), Gaps = 13/141 (9%)
Query: 105 KTRLEVSQQEANTRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGE 164
K + E ++EA VEK+E+ + G+ +LKVG G + G +V++ G+++ +
Sbjct: 284 KKKPEAKKEEA---PVEKKEKKQIAGGVSIEDLKVGSGPVAKAGKVVMVYYEGRLKQNNK 340
Query: 165 VFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADL 224
+F N +G L SK V G + + MK GGKRK++ PP + + GA
Sbjct: 341 MFDNCVKGPGFKFRL------GSKEVISGWDVGIAGMKVGGKRKIVCPPAMAY---GAK- 390
Query: 225 GSGVQIPPLATLEYIVEVDKV 245
GS IPP +TL + V++ V
Sbjct: 391 GSPPVIPPNSTLVFEVDLKNV 411
>UniRef100_Q63J95 Peptidyl-prolyl cis-trans isomerase [Burkholderia pseudomallei]
Length = 113
Score = 62.4 bits (150), Expect = 1e-08
Identities = 43/117 (36%), Positives = 63/117 (53%), Gaps = 10/117 (8%)
Query: 126 VTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRP 185
VT +G++Y +L G GA R G V + G + + G+ F ++ + P A V+G
Sbjct: 4 VTTESGLKYEDLTEGSGAEARAGQTVSVHYTGWL-TDGQKF-DSSKDRNDPFAFVLGGGM 61
Query: 186 YSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
KG EG++ MK GG R++ +PPQLG+ GA G IPP ATL + VE+
Sbjct: 62 VIKGWDEGVQ----GMKVGGVRRLTIPPQLGYGARGA----GGVIPPNATLVFEVEL 110
>UniRef100_UPI00002EEB60 UPI00002EEB60 UniRef100 entry
Length = 243
Score = 62.0 bits (149), Expect = 1e-08
Identities = 45/119 (37%), Positives = 62/119 (51%), Gaps = 10/119 (8%)
Query: 124 EEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGS 183
EE+ L +G+RY E G G G++VV+ G + S G F ++ + +P +G
Sbjct: 131 EEIVLESGLRYLEHIPGSGDVTTKGNIVVVHYSGFL-SDGTKFDSSHDR-ARPFNFTLGE 188
Query: 184 RPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
KG EG L MK GGKR +I+PP L + E GA G IPP ATL + VE+
Sbjct: 189 NRVIKGWEEG----LLNMKKGGKRTLIIPPDLAYGERGA----GGVIPPNATLLFEVEL 239
Score = 43.1 bits (100), Expect = 0.007
Identities = 36/124 (29%), Positives = 60/124 (48%), Gaps = 13/124 (10%)
Query: 119 DVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLA 178
++ +E +G++Y +K+G G P D V + G + G VF ++ + +
Sbjct: 4 ELNDNKEFITESGLKYEIIKMGNGEKPVATDKVEVHYHGTLLD-GTVFDSSVDRGQ---T 59
Query: 179 LVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEY 238
+ G KG EG L+ M G K K +PP+LG+ + D+GS IP +TL +
Sbjct: 60 ITFGLNQVIKGWTEG----LQLMPIGSKFKFTIPPELGYGDR--DIGS---IPANSTLIF 110
Query: 239 IVEV 242
VE+
Sbjct: 111 EVEL 114
>UniRef100_UPI00002E8307 UPI00002E8307 UniRef100 entry
Length = 154
Score = 60.5 bits (145), Expect = 4e-08
Identities = 35/105 (33%), Positives = 59/105 (55%), Gaps = 6/105 (5%)
Query: 131 GIRYCELKVGGGASPRPGDLVVIDIMGKV----ESTGEVFVNTFEGDKKPLALVMGSR-P 185
G+++ +L VG G P+ GD+VV+ + G + + VF++T +PL MG+ P
Sbjct: 35 GVQFSDLHVGSGEPPKYGDIVVLQVRGLLLDRRKQDDTVFLDT-AASGRPLVHQMGTAGP 93
Query: 186 YSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQI 230
+ V G+E ++TM+ GG+R V VPP++G+ G + QI
Sbjct: 94 RAAVVTLGLEQSIETMRGGGRRLVAVPPEMGYGSAGVSRFTAAQI 138
>UniRef100_Q7V8K3 FKBP-type peptidyl-prolyl cis-trans isomerase (PPIase) precursor
[Prochlorococcus marinus]
Length = 210
Score = 60.5 bits (145), Expect = 4e-08
Identities = 43/138 (31%), Positives = 70/138 (50%), Gaps = 15/138 (10%)
Query: 109 EVSQQEANTR----DVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGE 164
E+ +AN +E E +G+ +LK+G G G V ++ G +E+ G+
Sbjct: 80 EIDAAQANASALGGPIEAEPSQLTASGLSITDLKIGDGPEATAGQTVSVNYRGTLEN-GQ 138
Query: 165 VFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADL 224
F ++++ + P +G+ KG EG+ MK GGKRK+++PP+LG+ GA
Sbjct: 139 EFDSSYK--RGPFEFPLGAGRVIKGWDEGVA----GMKVGGKRKLVIPPELGYGSRGA-- 190
Query: 225 GSGVQIPPLATLEYIVEV 242
G IP ATL + VE+
Sbjct: 191 --GRVIPGNATLIFEVEL 206
>UniRef100_Q944B0 AT4g26550/M3E9_20 [Arabidopsis thaliana]
Length = 207
Score = 60.5 bits (145), Expect = 4e-08
Identities = 39/106 (36%), Positives = 58/106 (53%), Gaps = 18/106 (16%)
Query: 122 KEEEVT----LPNGIRYCELKVGGGASPRPGDLVVIDIMGK------VESTGEVFVNTFE 171
KE EV LP+G+RY E+ G G GDLV ++ + + V ST V+ F
Sbjct: 74 KEPEVIRTLKLPSGVRYQEIIEGEGREAHEGDLVELNYVCRRANGYFVHST----VDQFS 129
Query: 172 GDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGF 217
G+ P+ L++ V EG++ VL MKAGGKR+ ++PP +G+
Sbjct: 130 GESSPVKLILDEND----VIEGLKEVLVGMKAGGKRRALIPPSVGY 171
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.313 0.133 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 432,340,610
Number of Sequences: 2790947
Number of extensions: 18658051
Number of successful extensions: 123204
Number of sequences better than 10.0: 959
Number of HSP's better than 10.0 without gapping: 258
Number of HSP's successfully gapped in prelim test: 729
Number of HSP's that attempted gapping in prelim test: 120755
Number of HSP's gapped (non-prelim): 2282
length of query: 250
length of database: 848,049,833
effective HSP length: 124
effective length of query: 126
effective length of database: 501,972,405
effective search space: 63248523030
effective search space used: 63248523030
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0042.13


