

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0041.8
(562 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
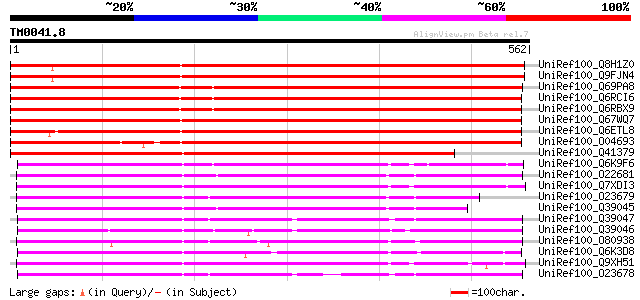
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8H1Z0 Cuticle protein [Arabidopsis thaliana] 822 0.0
UniRef100_Q9FJN4 Lipid transfer protein; glossy1 homolog [Arabid... 822 0.0
UniRef100_Q69PA8 Putative Gl1 protein [Oryza sativa] 787 0.0
UniRef100_Q6RCI6 Gl1 protein [Zea mays] 785 0.0
UniRef100_Q6RBX9 Glossy1 protein [Zea mays] 785 0.0
UniRef100_Q67WQ7 Putative Gl1 [Oryza sativa] 781 0.0
UniRef100_Q6ETL8 Putative glossy1 protein [Oryza sativa] 769 0.0
UniRef100_O04693 Glossy1 homolog [Oryza sativa] 723 0.0
UniRef100_Q41379 Lipid transfer protein [Senecio odorus] 692 0.0
UniRef100_Q6K9F6 Putative CER1 protein [Oryza sativa] 406 e-112
UniRef100_O22681 Arabidopsis thaliana gl1 homolog [Arabidopsis t... 389 e-107
UniRef100_Q7XDI3 Putative CER1 [Oryza sativa] 368 e-100
UniRef100_O23679 CER1 protein [Arabidopsis thaliana] 356 9e-97
UniRef100_Q39045 Possible aldehyde decarbonylase [Arabidopsis th... 348 3e-94
UniRef100_Q39047 CER1-like protein [Arabidopsis thaliana] 347 5e-94
UniRef100_Q39046 CER1-like protein [Arabidopsis thaliana] 345 2e-93
UniRef100_O80938 CER1-like protein [Arabidopsis thaliana] 345 2e-93
UniRef100_Q6K3D8 Putative CER1 [Oryza sativa] 332 2e-89
UniRef100_Q9XH51 CER1 [Oryza sativa] 315 2e-84
UniRef100_O23678 CER1-like protein [Arabidopsis thaliana] 315 2e-84
>UniRef100_Q8H1Z0 Cuticle protein [Arabidopsis thaliana]
Length = 632
Score = 822 bits (2122), Expect = 0.0
Identities = 393/564 (69%), Positives = 459/564 (80%), Gaps = 8/564 (1%)
Query: 1 MLFLTRNRRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLP----FLQNLPSWNVK 56
MLF+TR RI +G+DFKQID EW WDN++ILQ II S+ YM P + +LP WN K
Sbjct: 67 MLFVTRTLRINPKGIDFKQIDHEWHWDNYIILQAIIVSLICYMSPPLMMMINSLPLWNTK 126
Query: 57 GVIVALVLHVGISEPLYYWVHRKFH-GDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFI 115
G+I +VLHV SEPLYY++HR FH +Y FT+YHS HHSSPVP +T G+AT++E+ +
Sbjct: 127 GLIALIVLHVTFSEPLYYFLHRSFHRNNYFFTHYHSFHHSSPVPHPMTAGNATLLENIIL 186
Query: 116 SVVIGIPILGASVMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYT 175
VV G+P++G + G GS S +YGY + FDF+RCLGHCNVE+ H LF+ P ++YL+YT
Sbjct: 187 CVVAGVPLIGCCLFGVGSLSAIYGYAVMFDFMRCLGHCNVEIFSHKLFEILPVLRYLIYT 246
Query: 176 PTYHSLHHTDNKDTNFCLFMPLFDALGHTLNTKSWELHESF--SSGKCSRVPDFVFLAHI 233
PTYHSLHH + TNFCLFMPLFD LG T N SWEL + S+G+ RVP+FVFLAH
Sbjct: 247 PTYHSLHHQE-MGTNFCLFMPLFDVLGDTQNPNSWELQKKIRLSAGERKRVPEFVFLAHG 305
Query: 234 VDVTSSMHAQFCLRSFASLPFRTRFFLIPFWPIAITTLLAMWVRSKAFLFSFYYLRDRLH 293
VDV S+MHA F RSFAS+P+ TR FL+P WP +L MW SK FLFSFY LR+ L
Sbjct: 306 VDVMSAMHAPFVFRSFASMPYTTRIFLLPMWPFTFCVMLGMWAWSKTFLFSFYTLRNNLC 365
Query: 294 QTWVVPRCGFQYFLPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVD 353
QTW VPR GFQYFLPFA +GIN +IE AIL ADKIGVKVISLAALNKNEALNGGG LFV+
Sbjct: 366 QTWGVPRFGFQYFLPFATKGINDQIEAAILRADKIGVKVISLAALNKNEALNGGGTLFVN 425
Query: 354 KHPNLRVRVVHGNTLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTL 413
KHP+LRVRVVHGNTLTAAVI+ EIPKDV EVFLTGATSKLGRAIALYLC++ V+VLMLTL
Sbjct: 426 KHPDLRVRVVHGNTLTAAVILYEIPKDVNEVFLTGATSKLGRAIALYLCRRGVRVLMLTL 485
Query: 414 SADRFKRIQKEAPQEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVV 473
S +RF++IQKEAP E+Q+ LVQVTKY AAQHCKTWIVGKW+TPREQSWAP GTHFHQFVV
Sbjct: 486 SMERFQKIQKEAPVEFQNNLVQVTKYNAAQHCKTWIVGKWLTPREQSWAPAGTHFHQFVV 545
Query: 474 PPILPFRRDCTYGELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVG 533
PPIL FRR+CTYG+LAAM+LP+DVEGLG+CEYTMERGVVHACHAGGVVH LEGW HHEVG
Sbjct: 546 PPILKFRRNCTYGDLAAMKLPKDVEGLGTCEYTMERGVVHACHAGGVVHMLEGWKHHEVG 605
Query: 534 AIDVDRIDVVWKAALKHGLRPVSS 557
AIDVDRID+VW+AA+K+GL VSS
Sbjct: 606 AIDVDRIDLVWEAAMKYGLSAVSS 629
>UniRef100_Q9FJN4 Lipid transfer protein; glossy1 homolog [Arabidopsis thaliana]
Length = 566
Score = 822 bits (2122), Expect = 0.0
Identities = 393/564 (69%), Positives = 459/564 (80%), Gaps = 8/564 (1%)
Query: 1 MLFLTRNRRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLP----FLQNLPSWNVK 56
MLF+TR RI +G+DFKQID EW WDN++ILQ II S+ YM P + +LP WN K
Sbjct: 1 MLFVTRTLRINPKGIDFKQIDHEWHWDNYIILQAIIVSLICYMSPPLMMMINSLPLWNTK 60
Query: 57 GVIVALVLHVGISEPLYYWVHRKFH-GDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFI 115
G+I +VLHV SEPLYY++HR FH +Y FT+YHS HHSSPVP +T G+AT++E+ +
Sbjct: 61 GLIALIVLHVTFSEPLYYFLHRSFHRNNYFFTHYHSFHHSSPVPHPMTAGNATLLENIIL 120
Query: 116 SVVIGIPILGASVMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYT 175
VV G+P++G + G GS S +YGY + FDF+RCLGHCNVE+ H LF+ P ++YL+YT
Sbjct: 121 CVVAGVPLIGCCLFGVGSLSAIYGYAVMFDFMRCLGHCNVEIFSHKLFEILPVLRYLIYT 180
Query: 176 PTYHSLHHTDNKDTNFCLFMPLFDALGHTLNTKSWELHESF--SSGKCSRVPDFVFLAHI 233
PTYHSLHH + TNFCLFMPLFD LG T N SWEL + S+G+ RVP+FVFLAH
Sbjct: 181 PTYHSLHHQE-MGTNFCLFMPLFDVLGDTQNPNSWELQKKIRLSAGERKRVPEFVFLAHG 239
Query: 234 VDVTSSMHAQFCLRSFASLPFRTRFFLIPFWPIAITTLLAMWVRSKAFLFSFYYLRDRLH 293
VDV S+MHA F RSFAS+P+ TR FL+P WP +L MW SK FLFSFY LR+ L
Sbjct: 240 VDVMSAMHAPFVFRSFASMPYTTRIFLLPMWPFTFCVMLGMWAWSKTFLFSFYTLRNNLC 299
Query: 294 QTWVVPRCGFQYFLPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVD 353
QTW VPR GFQYFLPFA +GIN +IE AIL ADKIGVKVISLAALNKNEALNGGG LFV+
Sbjct: 300 QTWGVPRFGFQYFLPFATKGINDQIEAAILRADKIGVKVISLAALNKNEALNGGGTLFVN 359
Query: 354 KHPNLRVRVVHGNTLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTL 413
KHP+LRVRVVHGNTLTAAVI+ EIPKDV EVFLTGATSKLGRAIALYLC++ V+VLMLTL
Sbjct: 360 KHPDLRVRVVHGNTLTAAVILYEIPKDVNEVFLTGATSKLGRAIALYLCRRGVRVLMLTL 419
Query: 414 SADRFKRIQKEAPQEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVV 473
S +RF++IQKEAP E+Q+ LVQVTKY AAQHCKTWIVGKW+TPREQSWAP GTHFHQFVV
Sbjct: 420 SMERFQKIQKEAPVEFQNNLVQVTKYNAAQHCKTWIVGKWLTPREQSWAPAGTHFHQFVV 479
Query: 474 PPILPFRRDCTYGELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVG 533
PPIL FRR+CTYG+LAAM+LP+DVEGLG+CEYTMERGVVHACHAGGVVH LEGW HHEVG
Sbjct: 480 PPILKFRRNCTYGDLAAMKLPKDVEGLGTCEYTMERGVVHACHAGGVVHMLEGWKHHEVG 539
Query: 534 AIDVDRIDVVWKAALKHGLRPVSS 557
AIDVDRID+VW+AA+K+GL VSS
Sbjct: 540 AIDVDRIDLVWEAAMKYGLSAVSS 563
>UniRef100_Q69PA8 Putative Gl1 protein [Oryza sativa]
Length = 619
Score = 787 bits (2032), Expect = 0.0
Identities = 367/555 (66%), Positives = 448/555 (80%), Gaps = 2/555 (0%)
Query: 1 MLFLTRNRRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIV 60
MLF TR RRI++ GVDF QID+EWDWDNFLILQ +A+ A+Y P L++LP W+ +G+ V
Sbjct: 67 MLFATRRRRIVRDGVDFGQIDREWDWDNFLILQVHMAAAAFYAFPSLRHLPLWDARGLAV 126
Query: 61 ALVLHVGISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVVIG 120
A +LHV +EPL+Y HR FH +LF+ YH HHS+ VP+ T G AT +E + ++
Sbjct: 127 AALLHVAATEPLFYAAHRAFHRGHLFSCYHLQHHSAKVPQPFTAGFATPLEQLVLGALMA 186
Query: 121 IPILGASVMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTPTYHS 180
+P+ A G+GS +L + YVLGFD LR +GHCNVEV P GLF++ P +KYL+YTPTYH+
Sbjct: 187 VPLAAACAAGHGSVALAFAYVLGFDNLRAMGHCNVEVFPGGLFQSLPVLKYLIYTPTYHT 246
Query: 181 LHHTDNKDTNFCLFMPLFDALGHTLNTKSWELHESFSSGKCSRVPDFVFLAHIVDVTSSM 240
+HHT +D NFCLFMPLFD +G TL+ +SWE+ + S+G VP+FVFLAH+VDV S+
Sbjct: 247 IHHT-KEDANFCLFMPLFDLIGGTLDAQSWEMQKKTSAG-VDEVPEFVFLAHVVDVMQSL 304
Query: 241 HAQFCLRSFASLPFRTRFFLIPFWPIAITTLLAMWVRSKAFLFSFYYLRDRLHQTWVVPR 300
H F LR+FAS PF + FL+P WP A +L MW SK F+ S Y LR RLHQ W VPR
Sbjct: 305 HVPFVLRTFASTPFSVQPFLLPMWPFAFLVMLMMWAWSKTFVISCYRLRGRLHQMWAVPR 364
Query: 301 CGFQYFLPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVDKHPNLRV 360
GF YFLPFA +GIN +IE AIL ADK+G KV+SLAALNKNEALNGGG LFV+KHP LRV
Sbjct: 365 YGFHYFLPFAKDGINNQIELAILRADKMGAKVVSLAALNKNEALNGGGTLFVNKHPGLRV 424
Query: 361 RVVHGNTLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTLSADRFKR 420
RVVHGNTLTAAVI++EIP+ EVF+TGATSKLGRAIALYLC+KKV+V+M+TLS +RF++
Sbjct: 425 RVVHGNTLTAAVILNEIPQGTTEVFMTGATSKLGRAIALYLCRKKVRVMMMTLSTERFQK 484
Query: 421 IQKEAPQEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVVPPILPFR 480
IQ+EA E+Q YLVQVTKY++AQHCKTWIVGKW++PREQ WAP GTHFHQFVVPPI+ FR
Sbjct: 485 IQREATPEHQQYLVQVTKYRSAQHCKTWIVGKWLSPREQRWAPPGTHFHQFVVPPIIGFR 544
Query: 481 RDCTYGELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVGAIDVDRI 540
RDCTYG+LAAMRLP+DV+GLG+CEY++ERGVVHACHAGGVVH LEG+THHEVGAIDVDRI
Sbjct: 545 RDCTYGKLAAMRLPKDVQGLGACEYSLERGVVHACHAGGVVHFLEGYTHHEVGAIDVDRI 604
Query: 541 DVVWKAALKHGLRPV 555
DVVW+AAL+HGLRPV
Sbjct: 605 DVVWEAALRHGLRPV 619
>UniRef100_Q6RCI6 Gl1 protein [Zea mays]
Length = 621
Score = 785 bits (2028), Expect = 0.0
Identities = 369/556 (66%), Positives = 445/556 (79%), Gaps = 4/556 (0%)
Query: 1 MLFLTRNRRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIV 60
MLF TR RR+++ GVDF QIDKEWDWDNFLIL ++A+ A P L++LP+W+ +G V
Sbjct: 67 MLFATRRRRVVRDGVDFDQIDKEWDWDNFLILHALMAAAALCAFPSLRHLPAWDGRGFAV 126
Query: 61 ALVLHVGISEPLYYWVHRKFHGDY--LFTNYHSLHHSSPVPESLTVGHATIVEHAFISVV 118
ALV H +EPL Y HR HG L+ YHSLHHSS VP+ T G AT +EH + +
Sbjct: 127 ALVAHAAATEPLSYLAHRALHGSSGRLYARYHSLHHSSRVPQPFTAGLATPLEHVALGAL 186
Query: 119 IGIPILGASVMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTPTY 178
+ +P+ A G S +L + YVL FD LR +GHCNVEVVP LF+ P ++Y+LYTPTY
Sbjct: 187 MSLPLAAARAAGCASVALAFAYVLAFDSLRAMGHCNVEVVPASLFRAIPALRYVLYTPTY 246
Query: 179 HSLHHTDNKDTNFCLFMPLFDALGHTLNTKSWELHESFSSGKCSRVPDFVFLAHIVDVTS 238
H++HHT K+ NFCLFMPLFD LG T++ +SW++ S+G VPDFVFLAH+VDV
Sbjct: 247 HAIHHT-KKEANFCLFMPLFDLLGGTIDRRSWDMQRKMSAG-VDEVPDFVFLAHVVDVMQ 304
Query: 239 SMHAQFCLRSFASLPFRTRFFLIPFWPIAITTLLAMWVRSKAFLFSFYYLRDRLHQTWVV 298
S+H F +R+FAS PF + FL+P WP A +LAMWV SK F+ S Y LR RLHQ W V
Sbjct: 305 SLHVPFVMRTFASTPFSVQLFLLPMWPFAFLVMLAMWVWSKTFVISCYNLRGRLHQIWAV 364
Query: 299 PRCGFQYFLPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVDKHPNL 358
PR GFQYFLPFA +GIN++IE AIL ADK+GVKV+SLAALNKNEALNGGG LFV+KHP+L
Sbjct: 365 PRYGFQYFLPFAKDGINRQIELAILRADKMGVKVLSLAALNKNEALNGGGTLFVNKHPDL 424
Query: 359 RVRVVHGNTLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTLSADRF 418
RVRVVHGNTLTAAVI++EIPK EVFLTGATSKLGRAIALYLC+K+V+V+M+TLS +RF
Sbjct: 425 RVRVVHGNTLTAAVILNEIPKGTAEVFLTGATSKLGRAIALYLCKKRVRVMMMTLSTERF 484
Query: 419 KRIQKEAPQEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVVPPILP 478
++IQKEAP E+Q YLVQVTKY++AQHC+TWIVGKW++PREQ WAP GTHFHQFVVPPI+
Sbjct: 485 QKIQKEAPAEFQQYLVQVTKYRSAQHCRTWIVGKWLSPREQRWAPPGTHFHQFVVPPIIG 544
Query: 479 FRRDCTYGELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVGAIDVD 538
FRRDCTYG+LAAMRLP+DV GLG+CEY++ERG+VHACHAGGVVH LEG+THHEVGAIDVD
Sbjct: 545 FRRDCTYGKLAAMRLPKDVRGLGACEYSLERGLVHACHAGGVVHFLEGYTHHEVGAIDVD 604
Query: 539 RIDVVWKAALKHGLRP 554
RIDVVW+AALKHGLRP
Sbjct: 605 RIDVVWEAALKHGLRP 620
>UniRef100_Q6RBX9 Glossy1 protein [Zea mays]
Length = 621
Score = 785 bits (2026), Expect = 0.0
Identities = 369/556 (66%), Positives = 444/556 (79%), Gaps = 4/556 (0%)
Query: 1 MLFLTRNRRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIV 60
MLF TR RR+++ GVDF QIDKEWDWDNFLIL ++A+ A P L++LP+W+ +G V
Sbjct: 67 MLFATRRRRVVRDGVDFDQIDKEWDWDNFLILHALMAAAALCAFPSLRHLPAWDGRGFAV 126
Query: 61 ALVLHVGISEPLYYWVHRKFHGDY--LFTNYHSLHHSSPVPESLTVGHATIVEHAFISVV 118
AL H +EPL Y HR HG L+ YHSLHHSS VP+ T G AT +EH + +
Sbjct: 127 ALDAHAAATEPLSYLAHRALHGSSGRLYARYHSLHHSSRVPQPFTAGLATALEHVALGAL 186
Query: 119 IGIPILGASVMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTPTY 178
+ +P+ A G S +L + YVL FD LR +GHCNVEVVP LF+ P ++Y+LYTPTY
Sbjct: 187 MSLPLAAARAAGCASVALAFAYVLAFDSLRAMGHCNVEVVPASLFRAIPALRYVLYTPTY 246
Query: 179 HSLHHTDNKDTNFCLFMPLFDALGHTLNTKSWELHESFSSGKCSRVPDFVFLAHIVDVTS 238
H++HHT K+ NFCLFMPLFD LG T++ +SW++ S+G VPDFVFLAH+VDV
Sbjct: 247 HAIHHT-KKEANFCLFMPLFDLLGGTIDRRSWDMQRKMSAG-VDEVPDFVFLAHVVDVMQ 304
Query: 239 SMHAQFCLRSFASLPFRTRFFLIPFWPIAITTLLAMWVRSKAFLFSFYYLRDRLHQTWVV 298
S+H F +R+FAS PF + FL+P WP A +LAMWV SK F+ S Y LR RLHQ W V
Sbjct: 305 SLHVPFVMRTFASTPFSVQLFLLPMWPFAFLVMLAMWVWSKTFVISCYNLRGRLHQIWAV 364
Query: 299 PRCGFQYFLPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVDKHPNL 358
PR GFQYFLPFA +GIN++IE AIL ADK+GVKV+SLAALNKNEALNGGG LFV+KHP+L
Sbjct: 365 PRYGFQYFLPFAKDGINRQIELAILRADKMGVKVLSLAALNKNEALNGGGTLFVNKHPDL 424
Query: 359 RVRVVHGNTLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTLSADRF 418
RVRVVHGNTLTAAVI++EIPK EVFLTGATSKLGRAIALYLC+K+V+V+M+TLS +RF
Sbjct: 425 RVRVVHGNTLTAAVILNEIPKGTAEVFLTGATSKLGRAIALYLCKKRVRVMMMTLSTERF 484
Query: 419 KRIQKEAPQEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVVPPILP 478
++IQKEAP E+Q YLVQVTKY++AQHC+TWIVGKW++PREQ WAP GTHFHQFVVPPI+
Sbjct: 485 QKIQKEAPAEFQQYLVQVTKYRSAQHCRTWIVGKWLSPREQRWAPPGTHFHQFVVPPIIG 544
Query: 479 FRRDCTYGELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVGAIDVD 538
FRRDCTYG+LAAMRLP+DV GLG+CEY++ERGVVHACHAGGVVH LEG+THHEVGAIDVD
Sbjct: 545 FRRDCTYGKLAAMRLPKDVRGLGACEYSLERGVVHACHAGGVVHFLEGYTHHEVGAIDVD 604
Query: 539 RIDVVWKAALKHGLRP 554
RIDVVW+AALKHGLRP
Sbjct: 605 RIDVVWEAALKHGLRP 620
>UniRef100_Q67WQ7 Putative Gl1 [Oryza sativa]
Length = 627
Score = 781 bits (2018), Expect = 0.0
Identities = 373/556 (67%), Positives = 442/556 (79%), Gaps = 3/556 (0%)
Query: 1 MLFLTRNRRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYY--MLPFLQNLPSWNVKGV 58
MLF TR RR++ GVDF+QID EWDWDN +I+QT+IA++ + P +L +W+++G
Sbjct: 72 MLFFTRRRRVVDDGVDFRQIDTEWDWDNMVIMQTLIAAVLVTSRVFPATSDLSAWDLRGW 131
Query: 59 IVALVLHVGISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVV 118
+A+VLHV +SEP +YW HR H LF+ YHSLHHS ++LT G T +E +++V
Sbjct: 132 AIAVVLHVAVSEPAFYWAHRALHLGPLFSRYHSLHHSFQATQALTAGFVTPLESLILTLV 191
Query: 119 IGIPILGASVMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTPTY 178
P+ GA + G+GS SLVYG++L FD+LR +G+ NVEV+ H F+ FPF++YL+YTP+Y
Sbjct: 192 AWAPLAGAFMAGHGSVSLVYGHILLFDYLRSMGYSNVEVISHKTFQDFPFLRYLIYTPSY 251
Query: 179 HSLHHTDNKDTNFCLFMPLFDALGHTLNTKSWELHESFSSGKCSRVPDFVFLAHIVDVTS 238
SLHH + KD+NFCLFMPLFDALG TLN KSW+L + GK RVPDFVFL H+VDV S
Sbjct: 252 LSLHHRE-KDSNFCLFMPLFDALGGTLNPKSWQLQKEVDLGKNHRVPDFVFLVHVVDVVS 310
Query: 239 SMHAQFCLRSFASLPFRTRFFLIPFWPIAITTLLAMWVRSKAFLFSFYYLRDRLHQTWVV 298
SMH F R+ +SLPF T L+P WPIA +L W SK F SFY LR LHQTW V
Sbjct: 311 SMHVPFAFRACSSLPFATHLVLLPLWPIAFGFMLLQWFCSKTFTVSFYKLRGFLHQTWSV 370
Query: 299 PRCGFQYFLPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVDKHPNL 358
PR GFQYF+P A +GIN+ IE AIL ADK+GVKV+SLAALNKNEALNGGG LFV KHP+L
Sbjct: 371 PRYGFQYFIPSAKKGINEMIELAILRADKMGVKVLSLAALNKNEALNGGGTLFVRKHPDL 430
Query: 359 RVRVVHGNTLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTLSADRF 418
RVRVVHGNTLTAAVI++EIP DV EVFLTGATSKLGRAIALYLC+KK++VLMLTLS +RF
Sbjct: 431 RVRVVHGNTLTAAVILNEIPGDVAEVFLTGATSKLGRAIALYLCRKKIRVLMLTLSTERF 490
Query: 419 KRIQKEAPQEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVVPPILP 478
IQ+EAP E+Q YLVQVTKYQAAQ+CKTWIVGKW++PREQ WAP GTHFHQFVVPPI+
Sbjct: 491 MNIQREAPAEFQQYLVQVTKYQAAQNCKTWIVGKWLSPREQRWAPAGTHFHQFVVPPIIG 550
Query: 479 FRRDCTYGELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVGAIDVD 538
FRRDCTYG+LAAMRLPEDVEGLG+CEYTM RGVVHACHAGGVVH LEGW HHEVGAIDVD
Sbjct: 551 FRRDCTYGKLAAMRLPEDVEGLGTCEYTMGRGVVHACHAGGVVHFLEGWDHHEVGAIDVD 610
Query: 539 RIDVVWKAALKHGLRP 554
RID VW AAL+HGL P
Sbjct: 611 RIDAVWNAALRHGLTP 626
>UniRef100_Q6ETL8 Putative glossy1 protein [Oryza sativa]
Length = 628
Score = 770 bits (1987), Expect = 0.0
Identities = 368/563 (65%), Positives = 444/563 (78%), Gaps = 12/563 (2%)
Query: 1 MLFLTRNRRILQQGVDFKQIDKEWDWDNFLILQTIIASMAY---------YMLPFLQNLP 51
MLF TR RR++ VDF+Q+D EWDWDNFL+LQT+I + +LP L+
Sbjct: 68 MLFFTRRRRVVPDSVDFRQVDAEWDWDNFLLLQTLIGATLVGSPAVARQQLLLPSLKQ-- 125
Query: 52 SWNVKGVIVALVLHVGISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVE 111
+W+ +G +AL+LHV ++EPL+YW HR H LF+ YH+ HH + V LT G T +E
Sbjct: 126 AWDPRGWAIALLLHVLVAEPLFYWAHRALHRAPLFSRYHAAHHHASVTTPLTAGFGTPLE 185
Query: 112 HAFISVVIGIPILGASVMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKY 171
++VVIG+P+ GA +MG GS LVYG+VL FDFLR +G+ NVEV+ +F+ P ++Y
Sbjct: 186 SLLLTVVIGVPLAGAFLMGVGSVGLVYGHVLLFDFLRSMGYSNVEVISPRVFQAVPLLRY 245
Query: 172 LLYTPTYHSLHHTDNKDTNFCLFMPLFDALGHTLNTKSWELHESFSSGKCSRVPDFVFLA 231
L+YTPTY SLHH + KD+NFCLFMP+FD LG TLN KSWEL + GK + PDFVFLA
Sbjct: 246 LIYTPTYLSLHHRE-KDSNFCLFMPIFDLLGGTLNHKSWELQKEVYLGKNDQAPDFVFLA 304
Query: 232 HIVDVTSSMHAQFCLRSFASLPFRTRFFLIPFWPIAITTLLAMWVRSKAFLFSFYYLRDR 291
H+VD+ +SMH F LRS +S PF F L+PFWP+A +L MW SK FL S Y LR
Sbjct: 305 HVVDIMASMHVPFVLRSCSSTPFANHFVLLPFWPVAFGFMLLMWCCSKTFLVSSYRLRGN 364
Query: 292 LHQTWVVPRCGFQYFLPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLF 351
LHQ W VPR GFQYF+P A +GIN++IE AIL AD++GVKV+SLAALNKNEALNGGG LF
Sbjct: 365 LHQMWTVPRYGFQYFIPAAKKGINEQIELAILRADRMGVKVLSLAALNKNEALNGGGTLF 424
Query: 352 VDKHPNLRVRVVHGNTLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLML 411
V+KHP LRVRVVHGNTLTAAVI++EIP +V++VFLTGATSKLGRAIALYLC+KK++VLML
Sbjct: 425 VNKHPELRVRVVHGNTLTAAVILNEIPSNVKDVFLTGATSKLGRAIALYLCRKKIRVLML 484
Query: 412 TLSADRFKRIQKEAPQEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQF 471
TLS++RF +IQ+EAP E+Q YLVQVTKYQ AQ+CKTW+VGKW++PREQ WAP GTHFHQF
Sbjct: 485 TLSSERFLKIQREAPAEFQQYLVQVTKYQPAQNCKTWLVGKWLSPREQRWAPAGTHFHQF 544
Query: 472 VVPPILPFRRDCTYGELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHE 531
VVPPI+ FRRDCTYG+LAAMRLP+DV+GLG CEYTMERGVVHACHAGGVVH LEGW HHE
Sbjct: 545 VVPPIIGFRRDCTYGKLAAMRLPKDVQGLGYCEYTMERGVVHACHAGGVVHFLEGWEHHE 604
Query: 532 VGAIDVDRIDVVWKAALKHGLRP 554
VGAIDVDRIDVVWKAALKHGL P
Sbjct: 605 VGAIDVDRIDVVWKAALKHGLTP 627
>UniRef100_O04693 Glossy1 homolog [Oryza sativa]
Length = 555
Score = 723 bits (1867), Expect = 0.0
Identities = 358/561 (63%), Positives = 422/561 (74%), Gaps = 15/561 (2%)
Query: 1 MLFLTRNRRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYY--MLPFLQNLPSWNVKGV 58
MLF TR RR++ GVDF+QID EWDWDN +I+QT+IA++ + P +L +W+++G
Sbjct: 2 MLFFTRRRRVVDDGVDFRQIDTEWDWDNMVIMQTLIAAVLVTSRVFPATSDLSAWDLRGW 61
Query: 59 IVALVLHVGISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVV 118
+A+VLHV +SEP +YW HR H LF+ YHSLHHS ++LT G T + +++V
Sbjct: 62 AIAVVLHVAVSEPAFYWAHRALHLGPLFSRYHSLHHSFQATQALTAGFVTPLXXLILTLV 121
Query: 119 IGIPILGASVMGYGSASLVYGYVLG-----FDFLRCLGHCNVEVVPHGLFKTFPFMKYLL 173
P L G LVYG++ + GH + F+ FPF++YL+
Sbjct: 122 AW-PHLQGLHGGTRLRELVYGHISSSTTPVHGVQQRRGHLTQD------FQDFPFLRYLI 174
Query: 174 YTPTYHSLHHTDNKDTNFCLFMPLFDALGHTLNTKSWELHESFSSGKCSRVPDFVFLAHI 233
YTP+Y SLHH + KD+NFCLFMPLFDALG TLN KSW+L + GK RVPDFVFL H+
Sbjct: 175 YTPSYLSLHHRE-KDSNFCLFMPLFDALGGTLNPKSWQLQKEVDLGKNHRVPDFVFLVHV 233
Query: 234 VDVTSSMHAQFCLRSFASLPFRTRFFLIPFWPIAITTLLAMWVRSKAFLFSFYYLRDRLH 293
VDV SSMH F R+ +SLPF T L+P WPIA +L W SK F SFY LR LH
Sbjct: 234 VDVVSSMHVPFAFRACSSLPFATHLVLLPLWPIAFGFMLLQWFCSKTFTVSFYKLRGFLH 293
Query: 294 QTWVVPRCGFQYFLPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVD 353
QTW VPR GFQYF+P A +GIN+ IE AIL ADK+GVKV+SLAALNKNEALNGGG LFV
Sbjct: 294 QTWSVPRYGFQYFIPSAKKGINEMIELAILRADKMGVKVLSLAALNKNEALNGGGTLFVR 353
Query: 354 KHPNLRVRVVHGNTLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTL 413
KHP+LRVRVVHGNTLTAAVI++EIP DV EVFLTGATSKLGRAIALY C+KK++VLMLTL
Sbjct: 354 KHPDLRVRVVHGNTLTAAVILNEIPGDVAEVFLTGATSKLGRAIALYFCRKKIRVLMLTL 413
Query: 414 SADRFKRIQKEAPQEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVV 473
S +RF IQ+EAP E+Q YLVQVTKYQAAQ+CKTWIVGKW++PREQ WAP GTHFHQFVV
Sbjct: 414 STERFMNIQREAPAEFQQYLVQVTKYQAAQNCKTWIVGKWLSPREQRWAPAGTHFHQFVV 473
Query: 474 PPILPFRRDCTYGELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVG 533
PPI+ FRRDCTYG+LAAMRLPEDVEGLG+CEYTM RGVVHACHAGGVVH LEGW HHEVG
Sbjct: 474 PPIIGFRRDCTYGKLAAMRLPEDVEGLGTCEYTMGRGVVHACHAGGVVHFLEGWDHHEVG 533
Query: 534 AIDVDRIDVVWKAALKHGLRP 554
AIDVDRID VW AAL+HGL P
Sbjct: 534 AIDVDRIDAVWNAALRHGLTP 554
>UniRef100_Q41379 Lipid transfer protein [Senecio odorus]
Length = 524
Score = 692 bits (1787), Expect = 0.0
Identities = 323/484 (66%), Positives = 394/484 (80%), Gaps = 4/484 (0%)
Query: 1 MLFLTRNRRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPF-LQNLPSWNVKGVI 59
MLFLTRNRRIL Q +DF QIDKEW+WDNF+ILQ +IAS+A YM P NLP W KG++
Sbjct: 33 MLFLTRNRRILHQSIDFNQIDKEWNWDNFVILQALIASLAIYMFPQEFANLPVWKTKGLV 92
Query: 60 VALVLHVGISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVVI 119
+V+HV +SEPLYYW+HR H +YLFT YHS HHSS VP+ +TVG T +E ++ V+
Sbjct: 93 AIVVIHVVVSEPLYYWLHRLLHTNYLFTPYHSFHHSSAVPQPVTVGSTTFLEELLVTAVL 152
Query: 120 GIPILGASVMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTPTYH 179
G+PILG S+ GYGS S++YGYVL FDFLRCLGH NVE++PH +F FPF ++++YTPTY+
Sbjct: 153 GLPILGCSLSGYGSKSIIYGYVLVFDFLRCLGHSNVEIMPHWIFDYFPFFRFIIYTPTYY 212
Query: 180 SLHHTDNKDTNFCLFMPLFDALGHTLNTKSWELHESFS--SGKCSRVPDFVFLAHIVDVT 237
SLHH++ K +N+CLFMPL+D + +TLNTKSW LH+ S SGK +RVPDFVFLAH+VD+T
Sbjct: 213 SLHHSEMK-SNYCLFMPLYDTMWNTLNTKSWGLHKKISLDSGKSTRVPDFVFLAHVVDIT 271
Query: 238 SSMHAQFCLRSFASLPFRTRFFLIPFWPIAITTLLAMWVRSKAFLFSFYYLRDRLHQTWV 297
S++H F +RSF+++ + R FL+P WP ++ MW RSK FL S Y LR RLHQTWV
Sbjct: 272 SALHVPFVIRSFSAMAYSARLFLLPLWPFTFAVMIVMWARSKTFLLSSYNLRGRLHQTWV 331
Query: 298 VPRCGFQYFLPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVDKHPN 357
VPR GFQYFLPFA +GIN IEEAIL ADK+GVKVISLAALNKNE+LN GG LFV KHPN
Sbjct: 332 VPRFGFQYFLPFACQGINNHIEEAILRADKLGVKVISLAALNKNESLNRGGTLFVKKHPN 391
Query: 358 LRVRVVHGNTLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTLSADR 417
L+VRVVHGNTLTAAVI++EI +DV+EVFLTGATSKLGRAIALYLC++ V VLMLTLS +R
Sbjct: 392 LKVRVVHGNTLTAAVILNEINEDVKEVFLTGATSKLGRAIALYLCRRGVHVLMLTLSTER 451
Query: 418 FKRIQKEAPQEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVVPPIL 477
F+ IQ+EAP + + LVQVTKYQAA++CKTW++GKWITP +Q WAP GTHFHQFVVPPIL
Sbjct: 452 FQNIQEEAPSKCRKNLVQVTKYQAAKNCKTWVIGKWITPGQQRWAPSGTHFHQFVVPPIL 511
Query: 478 PFRR 481
FRR
Sbjct: 512 AFRR 515
>UniRef100_Q6K9F6 Putative CER1 protein [Oryza sativa]
Length = 619
Score = 406 bits (1044), Expect = e-112
Identities = 215/550 (39%), Positives = 323/550 (58%), Gaps = 12/550 (2%)
Query: 9 RILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIVALVLHVGI 68
RI+ +G++F Q+D+E WD+ ++ ++ Y +P ++ +P W G +V ++H G
Sbjct: 78 RIVDRGIEFDQVDRERGWDDQILFNGLVFYAGYLAMPSVRRMPVWRTDGAVVTALVHTGP 137
Query: 69 SEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVVIGIPILGASV 128
E LYYW HR H +L++ YHS HH+S V E +T EH ++ IPIL
Sbjct: 138 VEFLYYWFHRALHHHFLYSRYHSHHHASIVTEPITSVIHPFAEHVVYFILFAIPILSTIY 197
Query: 129 MGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTPTYHSLHHTDNKD 188
+G SA + GY+ DF+ +GHCN E+VP +F+ FP +KYL+YTP++HSLHHT +
Sbjct: 198 LGNVSAMGIVGYIAYIDFMNNMGHCNFELVPEWIFQIFPPLKYLIYTPSFHSLHHTQFR- 256
Query: 189 TNFCLFMPLFDALGHTLNTKSWELHESFSSGKCSRVPDFVFLAHIVDVTSSMHAQFCLRS 248
TN+ LFMP +D + +T++ S EL+ES G PD V L H+ ++ S+ H + + S
Sbjct: 257 TNYSLFMPFYDYIYNTMDKSSDELYESSLKG-TEETPDLVHLTHMTNLQSAYHLRIGIAS 315
Query: 249 FASLPFR-TRFFLIPFWPIAITTLLAMWV-RSKAFLFSFYYLRDRLHQTWVVPRCGFQYF 306
AS P+ + +++ WP+A +++ W+ S AF+ L QTW +PR FQY
Sbjct: 316 IASKPYSDSAWYMWTLWPLAWLSMVLAWIYGSSAFVVERIKLNKMKMQTWALPRYNFQYG 375
Query: 307 LPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVDKHPNLRVRVVHGN 366
L + E IN IE+AIL AD GVKVISL LN+ + LNG G+LF K+P L VR++ G+
Sbjct: 376 LTWEREPINDLIEKAILDADMKGVKVISLGLLNQAKQLNGNGELFRQKYPKLGVRIIDGS 435
Query: 367 TLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTLSADRFKRIQKEAP 426
L AV++ IP D ++VFL TSK+ RAIA+ LC + V+V+M + + ++ + P
Sbjct: 436 GLATAVVLKSIPSDAKKVFLRTGTSKIARAIAIALCDRGVQVIM--NEKEVYHMLKSQIP 493
Query: 427 QEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVVPPILPFRRDCTYG 486
+ SYL K + + WIV I EQ AP+GT F P+ R+DCTY
Sbjct: 494 ENRASYL----KLSSDNVPQLWIVHN-IDDNEQKMAPKGTIFIPISQFPLKKLRKDCTYM 548
Query: 487 ELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVGAIDVDRIDVVWKA 546
AMR+PE+++ + SCE + R V+ A H G++H+LEGW HE G +D I+ W A
Sbjct: 549 STPAMRIPEEMKNIHSCENWLPRRVMSAWHIAGILHALEGWNMHECGDEMMD-IEKSWSA 607
Query: 547 ALKHGLRPVS 556
A++HG P++
Sbjct: 608 AIRHGFLPLT 617
>UniRef100_O22681 Arabidopsis thaliana gl1 homolog [Arabidopsis thaliana]
Length = 625
Score = 389 bits (1000), Expect = e-107
Identities = 208/549 (37%), Positives = 314/549 (56%), Gaps = 6/549 (1%)
Query: 8 RRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIVALVLHVG 67
RRI+ +G+DF Q+D+E +WD+ ++ ++ + +L + LP W GV++ ++H G
Sbjct: 77 RRIVDKGIDFNQVDRETNWDDQILFNGVLFYIGINLLAEGKQLPWWRTDGVLMGALIHTG 136
Query: 68 ISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVVIGIPILGAS 127
E LYYWVH+ H +L++ YHS HHSS V E +T EH ++ IP+L
Sbjct: 137 PVEFLYYWVHKALHHHFLYSRYHSHHHSSIVTEPITSVIHPFAEHIAYFILFAIPLLTTL 196
Query: 128 VMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTPTYHSLHHTDNK 187
V S GY++ DF+ +GHCN E++P LF FP +K+L YTP+YHSLHHT +
Sbjct: 197 VTKTASIISFAGYIIYIDFMNNMGHCNFELIPKRLFHLFPPLKFLCYTPSYHSLHHTQFR 256
Query: 188 DTNFCLFMPLFDALGHTLNTKSWELHESFSSGKCSRVPDFVFLAHIVDVTSSMHAQFCLR 247
TN+ LFMPL+D + T++ + L+E RV D V L H+ S H + L
Sbjct: 257 -TNYSLFMPLYDYIYGTMDESTDTLYEKTLERGDDRV-DVVHLTHLTTPESIYHLRIGLP 314
Query: 248 SFASLPFRTRFFLIPFWPIAITTLLAMWVRSKAFLFSFYYLRDRLHQTWVVPRCGFQYFL 307
SFAS PF R+F+ WP +++ ++ F+ Q+WV+PR QY L
Sbjct: 315 SFASYPFAYRWFMRLLWPFTSLSMIFTLFYARLFVAERNSFNKLNLQSWVIPRYNLQYLL 374
Query: 308 PFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVDKHPNLRVRVVHGNT 367
+ E IN IE+AIL ADK GVKV+SL +N+ E LN G++++ HP+++VR+V G+
Sbjct: 375 KWRKEAINNMIEKAILEADKKGVKVLSLGLMNQGEELNRNGEVYIHNHPDMKVRLVDGSR 434
Query: 368 LTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTLSADRFKRIQKEAPQ 427
L AAV+I+ +PK V +TG +K+ IA LCQ+ V+V TL D +++I+ PQ
Sbjct: 435 LAAAVVINSVPKATTSVVMTGNLTKVAYTIASALCQRGVQV--STLRLDEYEKIRSCVPQ 492
Query: 428 EYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVVPPILPFRRDCTYGE 487
E + +LV +T +A K W+VG+ T EQ A +GT F F P+ R DC Y
Sbjct: 493 ECRDHLVYLTS-EALSSNKVWLVGEGTTREEQEKATKGTLFIPFSQFPLKQLRSDCIYHT 551
Query: 488 LAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVG-AIDVDRIDVVWKA 546
A+ +P+ + + SCE + R + A G++H+LEGW HE G ++ + +D VW+A
Sbjct: 552 TPALIVPKSLVNVHSCENWLPRKAMSATRVAGILHALEGWETHECGTSLLLSDLDKVWEA 611
Query: 547 ALKHGLRPV 555
L HG +P+
Sbjct: 612 CLSHGFQPL 620
>UniRef100_Q7XDI3 Putative CER1 [Oryza sativa]
Length = 621
Score = 368 bits (944), Expect = e-100
Identities = 196/553 (35%), Positives = 309/553 (55%), Gaps = 10/553 (1%)
Query: 8 RRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIVALVLHVG 67
R+I+++G++F Q+D+E +WD+ +IL I+ + +P Q+LP W G + +LH G
Sbjct: 77 RQIVRRGIEFDQVDRERNWDDQIILSGILLYLGALYVPGGQHLPLWRTDGAGLIALLHAG 136
Query: 68 ISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVVIGIPILGAS 127
E LYYW HR H +L+T+YHS HHSS V E +T E ++ IP++ +
Sbjct: 137 PVEFLYYWFHRALHHHFLYTHYHSHHHSSIVTEPITSVIHPFAELVAYELLFSIPLIACA 196
Query: 128 VMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTPTYHSLHHTDNK 187
+ G S Y++ DF+ +GHCN E+VP LF FP +KYL+YTP++HSLHHT +
Sbjct: 197 LTGTASIIAFEMYLIYIDFMNNMGHCNFELVPSWLFTWFPPLKYLMYTPSFHSLHHTQFR 256
Query: 188 DTNFCLFMPLFDALGHTLNTKSWELHE-SFSSGKCSRVPDFVFLAHIVDVTSSMHAQFCL 246
TN+ LFMP +D + +T++ S L+E S + + D V L H+ + S H +
Sbjct: 257 -TNYSLFMPFYDYIYNTMDKSSDTLYENSLKNNEEEEAVDVVHLTHLTTLHSIYHMRPGF 315
Query: 247 RSFASLPFRTRFFLIPFWPIAITTLLAMWVRSKAFLFSFYYLRDRLHQTWVVPRCGFQYF 306
FAS P+ +R+++ WP++ +++ W +F ++ Q+W +PR F Y
Sbjct: 316 AEFASRPYVSRWYMRMMWPLSWLSMVLTWTYGSSFTVERNVMKKIRMQSWAIPRYSFHYG 375
Query: 307 LPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVDKHPNLRVRVVHGN 366
L + E IN IE+A+ ADK G KV+SL LN+ LN G+ ++ K+P L R+V G
Sbjct: 376 LDWEKEAINDLIEKAVCEADKNGAKVVSLGLLNQAHTLNKSGEQYLLKYPKLGARIVDGT 435
Query: 367 TLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTLSADRFKRIQKEAP 426
+L AAV+++ IP+ ++V L G SK+ RA+A LC+K +KV M + + ++ E P
Sbjct: 436 SLAAAVVVNSIPQGTDQVILAGNVSKVARAVAQALCKKNIKVTM--TNKQDYHLLKPEIP 493
Query: 427 QEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQF-VVPPILPFRRDCTY 485
+ L + K W++G + EQ A +GT F + PP + + C+Y
Sbjct: 494 ETVADNL----SFSKTGTAKVWLIGDGLDSAEQFRAQKGTLFIPYSQFPPKMVRKDSCSY 549
Query: 486 GELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVGAIDVDRIDVVWK 545
AM +P+ ++ + SCE + R V+ A G++H+LEGW HE G +D +D VW
Sbjct: 550 STTPAMAVPKTLQNVHSCENWLPRRVMSAWRIAGILHALEGWNEHECGDKVLD-MDKVWS 608
Query: 546 AALKHGLRPVSSG 558
AA+ HG PV+ G
Sbjct: 609 AAIMHGFCPVAQG 621
>UniRef100_O23679 CER1 protein [Arabidopsis thaliana]
Length = 580
Score = 356 bits (914), Expect = 9e-97
Identities = 188/501 (37%), Positives = 290/501 (57%), Gaps = 5/501 (0%)
Query: 8 RRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIVALVLHVG 67
RRI+ +G+DF Q+D+E +WD+ ++ ++ + +LP + LP W GV++A ++H G
Sbjct: 77 RRIVDKGIDFNQVDRETNWDDQILFNGVLFYIGINLLPEAKQLPWWRTDGVLMAALIHTG 136
Query: 68 ISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVVIGIPILGAS 127
E LYYW+H+ H +L++ YHS HHSS V E +T EH ++ IP+L
Sbjct: 137 PVEFLYYWLHKALHHHFLYSRYHSHHHSSIVTEPITSVIHPFAEHIAYFILFAIPLLTTL 196
Query: 128 VMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTPTYHSLHHTDNK 187
+ S GY++ DF+ +GHCN E++P LF FP +K+L YTP+YHSLHHT +
Sbjct: 197 LTKTASIISFAGYIIYIDFMNNMGHCNFELIPKRLFHLFPPLKFLCYTPSYHSLHHTQFR 256
Query: 188 DTNFCLFMPLFDALGHTLNTKSWELHESFSSGKCSRVPDFVFLAHIVDVTSSMHAQFCLR 247
TN+ LFMPL+D + T++ + L+E + + + D V L H+ S H + L
Sbjct: 257 -TNYSLFMPLYDYIYGTMDESTDTLYEK-TLERGDDIVDVVHLTHLTTPESIYHLRIGLA 314
Query: 248 SFASLPFRTRFFLIPFWPIAITTLLAMWVRSKAFLFSFYYLRDRLHQTWVVPRCGFQYFL 307
SFAS PF R+F+ WP +++ ++ F+ Q+WV+PR QY L
Sbjct: 315 SFASYPFAYRWFMRLLWPFTSLSMIFTLFYARLFVAERNSFNKLNLQSWVIPRYNLQYLL 374
Query: 308 PFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVDKHPNLRVRVVHGNT 367
+ E IN IE+AIL ADK GVKV+SL +N+ E LN G++++ HP+++VR+V G+
Sbjct: 375 KWRKEAINNMIEKAILEADKKGVKVLSLGLMNQGEELNRNGEVYIHNHPDMKVRLVDGSR 434
Query: 368 LTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTLSADRFKRIQKEAPQ 427
L AAV+I+ +PK V +TG +K+ IA LCQ+ V+V TL D +++I+ PQ
Sbjct: 435 LAAAVVINSVPKATTSVVMTGNLTKVAYTIASALCQRGVQV--STLRLDEYEKIRSCVPQ 492
Query: 428 EYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVVPPILPFRRDCTYGE 487
E + +LV +T +A K W+VG+ T EQ A +GT F F P+ RRDC Y
Sbjct: 493 ECRDHLVYLTS-EALSSNKVWLVGEGTTREEQEKATKGTLFIPFSQFPLKQLRRDCIYHT 551
Query: 488 LAAMRLPEDVEGLGSCEYTME 508
A+ +P+ + + SCE +++
Sbjct: 552 TPALIVPKSLVNVHSCEVSIQ 572
>UniRef100_Q39045 Possible aldehyde decarbonylase [Arabidopsis thaliana]
Length = 567
Score = 348 bits (892), Expect = 3e-94
Identities = 186/488 (38%), Positives = 280/488 (57%), Gaps = 5/488 (1%)
Query: 8 RRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIVALVLHVG 67
RRI+ +G+DF Q+D+E +WD+ ++ ++ + +LP + LP W GV++A ++H G
Sbjct: 77 RRIVDKGIDFNQVDRETNWDDQILFNGVLFYIGINLLPEAKQLPWWRTDGVLMAALIHTG 136
Query: 68 ISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVVIGIPILGAS 127
E LYYW+H+ H +L++ YHS HHSS V E +T EH ++ IP+L
Sbjct: 137 PVEFLYYWLHKALHHHFLYSRYHSHHHSSIVTEPITSVIHPFAEHIAYFILFAIPLLTTL 196
Query: 128 VMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTPTYHSLHHTDNK 187
+ S GY++ DF+ +GHCN E++P LF FP +K+L YTP+YHSLHHT +
Sbjct: 197 LTKTASIISFAGYIIYIDFMNNMGHCNFELIPKRLFHLFPPLKFLCYTPSYHSLHHTQFR 256
Query: 188 DTNFCLFMPLFDALGHTLNTKSWELHESFSSGKCSRVPDFVFLAHIVDVTSSMHAQFCLR 247
TN+ LFMPL+D + T++ + L+E RV D V L H+ S H + L
Sbjct: 257 -TNYSLFMPLYDYIYGTMDESTDTLYEKTLERGDDRV-DVVHLTHLTTPESIYHLRIGLA 314
Query: 248 SFASLPFRTRFFLIPFWPIAITTLLAMWVRSKAFLFSFYYLRDRLHQTWVVPRCGFQYFL 307
SFAS PF R+F+ WP +++ ++ F+ Q+WV+PR QY L
Sbjct: 315 SFASYPFAYRWFMRLLWPFTSLSMIFTLFYARLFVAERNSFNKLNLQSWVIPRYNLQYLL 374
Query: 308 PFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVDKHPNLRVRVVHGNT 367
+ E IN IE+AIL ADK GVKV+SL +N+ E LN G++++ HP+++VR+V G+
Sbjct: 375 KWRKEAINNMIEKAILEADKKGVKVLSLGLMNQGEELNRNGEVYIHNHPDMKVRLVDGSR 434
Query: 368 LTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTLSADRFKRIQKEAPQ 427
L AAV+I+ +PK V +TG +K+ IA LCQ+ V+V TL D +++I+ PQ
Sbjct: 435 LAAAVVINSVPKATTSVVMTGNLTKVAYTIASALCQRGVQV--STLRLDEYEKIRSCVPQ 492
Query: 428 EYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVVPPILPFRRDCTYGE 487
E + +LV +T +A K W+VG+ T EQ A +GT F F P+ R DC Y
Sbjct: 493 ECRDHLVYLTS-EALSSNKVWLVGEGTTREEQEKATKGTLFIPFSQFPLKQLRSDCIYHT 551
Query: 488 LAAMRLPE 495
A+ +P+
Sbjct: 552 TPALIVPK 559
>UniRef100_Q39047 CER1-like protein [Arabidopsis thaliana]
Length = 623
Score = 347 bits (890), Expect = 5e-94
Identities = 203/553 (36%), Positives = 305/553 (54%), Gaps = 18/553 (3%)
Query: 9 RILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIVALVLHVGI 68
+I+ + ++F+Q+D+E WD+ +I T++ +A LP +LP W V G I+ ++LH G
Sbjct: 78 KIVDKPIEFEQVDRERTWDDQVIFNTLLMYLANIKLPGASHLPPWRVDGGILMVLLHAGP 137
Query: 69 SEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVVIGIPILGASV 128
E LYYW HR H +L++ YHS HHSS V E +T EH +++ IP++ AS+
Sbjct: 138 VEFLYYWFHRGLHHHFLYSRYHSHHHSSIVTEPITSVVHPFGEHIVYTLLCDIPMVTASL 197
Query: 129 MGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTPTYHSLHHTDNKD 188
G S + GY+ DF+ +GHCN E+ P LF FP +K+L YTP++HSLHHT +
Sbjct: 198 CGILSIVSIMGYITYIDFMNNMGHCNFELFPKRLFHLFPPLKFLCYTPSFHSLHHTQFR- 256
Query: 189 TNFCLFMPLFDALGHTLNTKSWELHESFSSGKCSRVPDFVFLAHIVDVTSSMHAQFCLRS 248
TN+ LFMP++D + T + + L+E S PD + L H+ S + S
Sbjct: 257 TNYSLFMPIYDFIYGTTDNLTDSLYER-SLEIEEESPDVIHLTHLTTHNSIYQMRLGFPS 315
Query: 249 FASLPFRTR--FFLIPF-WPIAITTLLAMW--VRSKAFLFSFYYLRDRLHQTWVVPRCGF 303
+S P +R ++L F WP + A+ + + F+F LRD + ++P+ F
Sbjct: 316 LSSCPLWSRPPWYLTCFMWPFTLLCSFALTSAIPLRTFVFERNRLRDLTVHSHLLPKFSF 375
Query: 304 QYFLPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVDKHPNLRVRVV 363
E IN IEEAIL AD+ GVKV+SL +N E LNG G+++V K+P L++R+V
Sbjct: 376 HRH----HESINTIIEEAILEADEKGVKVMSLGLMNNREELNGSGEMYVQKYPKLKIRLV 431
Query: 364 HGNTLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTLSADRFKRIQK 423
G+++ A V+I+ IPK+ E+ G +K+ A+ LCQK VKV++L R + K
Sbjct: 432 DGSSMAATVVINNIPKEATEIVFRGNLTKVASAVVFALCQKGVKVVVL-----REEEHSK 486
Query: 424 EAPQEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVVPPILPFRRDC 483
LV T + K W+VG I EQ A GT F F P R+DC
Sbjct: 487 LIKSGVDKNLVLSTS-NSYYSPKVWLVGDGIENEEQMKAKEGTLFVPFSHFPPNKLRKDC 545
Query: 484 TYGELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVG-AIDVDRIDV 542
Y AMR+P+ + + SCE + R V+ A GG+VH+LEGW H+ G +V R+
Sbjct: 546 FYQSTPAMRVPKSAQNIDSCENWLGRRVMSAWKIGGIVHALEGWEEHDCGNTCNVLRLHA 605
Query: 543 VWKAALKHGLRPV 555
+W+AAL+H +P+
Sbjct: 606 IWEAALRHDFQPL 618
>UniRef100_Q39046 CER1-like protein [Arabidopsis thaliana]
Length = 622
Score = 345 bits (885), Expect = 2e-93
Identities = 199/554 (35%), Positives = 306/554 (54%), Gaps = 21/554 (3%)
Query: 9 RILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIVALVLHVGI 68
+I+ + ++F+Q+D+E WD+ +I T++ +A LP +LP W V G I+ ++LH G
Sbjct: 78 KIVDKPIEFEQVDRERTWDDQVIFNTLLMYLANIKLPGASHLPPWRVDGGILMVLLHAGP 137
Query: 69 SEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVVIGIPILGASV 128
E LYYW HR H +L++ YHS HHSS V E +TV H EH +++ IP++ AS+
Sbjct: 138 VEFLYYWFHRGLHHHFLYSRYHSHHHSSIVTEPITVVHP-FGEHIVYTLLCDIPMVTASL 196
Query: 129 MGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTPTYHSLHHTDNKD 188
G S + GY+ DF+ +GHCN E+ P LF FP +K+L YTP++HSLHHT +
Sbjct: 197 CGILSIVSIMGYITYIDFMNNMGHCNFELFPKRLFHLFPPLKFLCYTPSFHSLHHTQFR- 255
Query: 189 TNFCLFMPLFDALGHTLNTKSWELHESFSSGKCSRVPDFVFLAHIVDVTSSMHAQFCLRS 248
TN+ LFMP++D + T + + L+E + PD + L H+ S + S
Sbjct: 256 TNYSLFMPIYDFIYGTTDNLTDSLYERSLEIE-EESPDVIHLTHLTTHNSIYQMRLGFPS 314
Query: 249 FASLPFRTR------FFLIPFWPIAITTLLAMWVRSKAFLFSFYYLRDRLHQTWVVPRCG 302
+S P +R F+ PF + + L + + F+F LRD + ++P+
Sbjct: 315 LSSCPLWSRPPWYLTCFMXPF-TLLCSFALTSAIPLRTFVFERNRLRDLTVHSHLLPKFS 373
Query: 303 FQYFLPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVDKHPNLRVRV 362
F E IN IEEAIL AD+ GVKV+SL +N E LNG G+++V K+P L++R+
Sbjct: 374 FHRH----HESINTIIEEAILEADEKGVKVMSLGLMNNREELNGSGEMYVQKYPKLKIRL 429
Query: 363 VHGNTLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTLSADRFKRIQ 422
V G+++ A V+I+ IPK+ E+ G +K+ A+ LCQK VKV++L + K I+
Sbjct: 430 VDGSSMAATVVINNIPKEATEIVFRGNLTKVASAVVFALCQKGVKVVVLR-EEEHXKLIK 488
Query: 423 KEAPQEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVVPPILPFRRD 482
+ + ++ + K W+VG I EQ GT F F P R+D
Sbjct: 489 SGVDKN-----LVLSTSNSYYSPKVWLVGDGIENEEQMKPKEGTLFVPFSHFPPNKLRKD 543
Query: 483 CTYGELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVG-AIDVDRID 541
C Y AMR+P+ + + SCE + R V+ A GG+VH+LEGW H+ G +V R+
Sbjct: 544 CFYQSTPAMRVPKSAQNIDSCENWLGRRVMSAWKIGGIVHALEGWEEHDCGNTCNVLRLH 603
Query: 542 VVWKAALKHGLRPV 555
+W+AAL+H +P+
Sbjct: 604 AIWEAALRHDFQPL 617
>UniRef100_O80938 CER1-like protein [Arabidopsis thaliana]
Length = 635
Score = 345 bits (885), Expect = 2e-93
Identities = 199/566 (35%), Positives = 304/566 (53%), Gaps = 28/566 (4%)
Query: 8 RRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIVALVLHVG 67
+RIL + ++F Q+D+E WD+ +I T+I + + +P W GVI+ +LH G
Sbjct: 73 KRILNKSIEFDQVDRERTWDDQIIFNTLIVYLTKVYVSGTSTIPFWRTDGVILVALLHAG 132
Query: 68 ISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHAT-------------IVEHAF 114
E +YYW HR H +L++ YHS HHSS V E +T+ EH
Sbjct: 133 PVEFIYYWFHRALHHHFLYSRYHSHHHSSIVTEPITLCATNSKPWVLIVAVVHPFAEHIG 192
Query: 115 ISVVIGIPILGASVMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLY 174
++++G+P++ + G S + Y+ DF+ +GHCN E++P LF P +K+L Y
Sbjct: 193 YTLILGLPLITTFMCGTVSVVSIALYLTYIDFMNNMGHCNFELIPKFLFSLLPPLKFLCY 252
Query: 175 TPTYHSLHHTDNKDTNFCLFMPLFDALGHTLNTKSWELHESFSSGKCSRVPDFVFLAHIV 234
TP++HSLHHT + TN+ LFMP++D + T + S L+E+ S K PD + L H+
Sbjct: 253 TPSFHSLHHTQFR-TNYSLFMPMYDYIYGTTDECSDSLYET-SLEKEEEKPDAIHLTHLT 310
Query: 235 DVTSSMHAQFCLRSFASLPFRTRFFLIPFWPIAITTLLAMWVRS---KAFLFSFYYLRDR 291
+ S H + S +S P +R +L P A+ +L+ +RS + F+ RD
Sbjct: 311 SLDSIYHLRLGFASLSSHPLSSRCYLFLMKPFAL--ILSFILRSFSFQTFVVERNRFRDL 368
Query: 292 LHQTWVVPRCGFQYFLPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLF 351
+ ++P+ Y E INK IE AIL ADK GVKV+SL LN+ E LNG G+++
Sbjct: 369 TLHSHLLPKFSSHYMSHQQKECINKMIEAAILEADKKGVKVMSLGLLNQGEELNGYGEMY 428
Query: 352 VDKHPNLRVRVVHGNTLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLML 411
V +HP L++R+V G +L A V++ IP +EV G +K+ RAI LCQ +KV++
Sbjct: 429 VRRHPKLKIRIVDGGSLAAEVVLHSIPVGTKEVLFRGQITKVARAIVFSLCQNAIKVMV- 487
Query: 412 TLSADRFKRIQKEAPQEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQF 471
L + + + + + LV T Y + W+VG ++ +EQ A GT F F
Sbjct: 488 -LRKEEHSMLAEFLDDKCKENLVLTTNY----YPMIWLVGDGLSTKEQKMAKDGTLFLPF 542
Query: 472 VVPPILPFRRDCTYGELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHE 531
P R+DC Y AM +P + + SCE + R V+ A GG+VH+LEGW HE
Sbjct: 543 SQFPPKTLRKDCFYHTTPAMIIPHSAQNIDSCENWLGRRVMSAWRVGGIVHALEGWKEHE 602
Query: 532 VGAIDVDRIDV--VWKAALKHGLRPV 555
G D I+ VW+AAL++G +P+
Sbjct: 603 CGLDDNSIINPPRVWEAALRNGFQPL 628
>UniRef100_Q6K3D8 Putative CER1 [Oryza sativa]
Length = 635
Score = 332 bits (851), Expect = 2e-89
Identities = 195/554 (35%), Positives = 300/554 (53%), Gaps = 22/554 (3%)
Query: 9 RILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIVALVLHVGI 68
+I+ + +DF+QID+E +WD+ +IL ++ + +P Q P W+ KG++V VLH G
Sbjct: 79 KIVNKSLDFEQIDRERNWDDQIILTALVFYLVSATMPQAQVAPWWSTKGMVVTAVLHAGP 138
Query: 69 SEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVVIGIPILGASV 128
E LYYW+HR H +L+ YHS HH+S V E +T E V++ IPIL
Sbjct: 139 VEFLYYWLHRALHHHWLYARYHSHHHASIVTEPITSVIHPFAEEVVYFVLLAIPILSTVA 198
Query: 129 MGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTPTYHSLHHTDNKD 188
G S GY++ DF+ LGHCN E+VP LF FP +KYLLYTP++HSLHHT +
Sbjct: 199 TGTVSVVTANGYLVYIDFMNYLGHCNFELVPKCLFHVFPPLKYLLYTPSFHSLHHTQFR- 257
Query: 189 TNFCLFMPLFDALGHTLNTKSWELHESFSSGKCSRV--PDFVFLAHIVDVTSSMHAQFCL 246
TN+ LFMP++D + T + S EL+E G+ PD V L H+ S H +
Sbjct: 258 TNYSLFMPVYDYIYGTTDKSSDELYERTLQGRDEAAWRPDVVHLTHLTTPESVFHNRLGF 317
Query: 247 RSFASLPF---RTRFFLIPFWPIA--ITTLLAMWVRSKAFLFSFYYLRDRLH-QTWVVPR 300
+ AS P + L +A + +L A RS+A D+L+ +TWV+PR
Sbjct: 318 AAVASNPLGAAASGHLLRAASAVASPLLSLFASTFRSEANRL------DKLNIETWVIPR 371
Query: 301 CGFQYFLPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVDKHPNLRV 360
Y +++ IE+A+ A+ G +V++L LN+ LN G+L+V + P+L+
Sbjct: 372 FTSHYTSKSDGYKVSRLIEKAVSDAEASGARVLTLGLLNQGYDLNRNGELYVVRKPSLKT 431
Query: 361 RVVHGNTLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTLSADRFKR 420
++V G +L A +++ IP+ ++V L G +K+ + L LC+++++V M ++ + ++
Sbjct: 432 KIVDGTSLAVAAVLNMIPQGTKDVLLLGNANKISLVLTLSLCKREIQVRM--VNKELYEC 489
Query: 421 IQKEAPQEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVVPPILPFR 480
++++ E Q +LV Y + K W+VG +T EQ A +G+HF + P R
Sbjct: 490 LKQQLQPEMQEHLVLSCSYSS----KVWLVGDGVTDEEQMKAQKGSHFVPYSQFPPNKAR 545
Query: 481 RDCTYGELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVGAIDVDRI 540
DC Y A+ +PE E L CE + R V+ A A G+VH+LE W HE G V +
Sbjct: 546 NDCVYHCTPALLVPESFENLHVCENWLPRRVMSAWRAAGIVHALEKWDGHECGG-RVTGV 604
Query: 541 DVVWKAALKHGLRP 554
W AAL G RP
Sbjct: 605 QKAWSAALARGFRP 618
>UniRef100_Q9XH51 CER1 [Oryza sativa]
Length = 621
Score = 315 bits (807), Expect = 2e-84
Identities = 184/556 (33%), Positives = 287/556 (51%), Gaps = 16/556 (2%)
Query: 8 RRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIVALVLHVG 67
R+I+++G++F Q+D+E +WD+ +IL I+ + +P Q+LP W G + +LH G
Sbjct: 77 RQIVRRGIEFDQVDRERNWDDQIILSGILLYLGALYVPGGQHLPLWRTDGAGLIALLHAG 136
Query: 68 ISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVVIGIPILGAS 127
E LYYW HR H +L+T YHS HHSS V E +T E ++ IP++ +
Sbjct: 137 PVEFLYYWFHRALHHHFLYTRYHSHHHSSIVTEPITSVIHPFAELVAYELLFSIPLIACA 196
Query: 128 VMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTPTYHSLHHTDNK 187
+ G S Y++ DF+ +GHCN E+VP LF FP +KYL+YTP++HSLHHT +
Sbjct: 197 LTGTASIIAFEMYLIYIDFMNNMGHCNFELVPSWLFTWFPPLKYLMYTPSFHSLHHTQFR 256
Query: 188 DTNFCLFMPLFDALGHTLNTKSWELHE-SFSSGKCSRVPDFVFLAHIVDVTSSMHAQFCL 246
TN+ LFMP +D + +T++ S L+E S + D V L H+ + S H +
Sbjct: 257 -TNYSLFMPFYDYIYNTMDKSSDTLYENSLKNNDEEEAVDVVHLTHLTTLHSIYHMRPGF 315
Query: 247 RSFASLPFRTRFFLIPFWPIAITTLLAMWVRSKAFLFSFYYLRDRLHQTWVVPRCGFQYF 306
FAS P+ +R+++ WP++ +++ W +F +RD+ F Y
Sbjct: 316 AEFASRPYVSRWYMRMMWPLSWLSMVLTWTYGSSFTVERNVMRDQDAVMGHYQDTSFHYG 375
Query: 307 LPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVDKHPNLRVRVVHGN 366
L + E IN IE+A+ ADK G KV+SL LN+ LN G+ ++ K+P L R+V G
Sbjct: 376 LDWEKEAINDLIEKAVCEADKNGAKVVSLGLLNQAHTLNKSGEQYLLKYPKLGARIVDGT 435
Query: 367 TLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTLSADRFKRIQKEAP 426
+L AAV+++ IP+ ++V L G SK+ RA+A LC+K +KV M + + ++ E P
Sbjct: 436 SLAAAVVVNSIPQGTDQVILAGNVSKVARAVAQALCKKNIKVTM--TNKQDYHLLKPEIP 493
Query: 427 QEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQF-VVPPILPFRRDCTY 485
+ L + K W++G + EQ A +GT F + PP + + C+Y
Sbjct: 494 ETVADNL----SFSKTGTAKVWLIGDGLDSAEQFRAQKGTLFIPYSQFPPKMVRKDSCSY 549
Query: 486 GELAAMRLPEDVEGLGSCEYTMERGVVHA---CHAGGVVHSLEGWTHHEVGAIDVDRIDV 542
A+ ++ C + E GG + GW HE G V +
Sbjct: 550 STTPAIGCTKNA---AECAFMRELAAKEGYGRMANGGNSSCVGGWNEHECGD-KVLGMAK 605
Query: 543 VWKAALKHGLRPVSSG 558
VW ++HGL PV G
Sbjct: 606 VWTDTIEHGLCPVDQG 621
>UniRef100_O23678 CER1-like protein [Arabidopsis thaliana]
Length = 604
Score = 315 bits (807), Expect = 2e-84
Identities = 193/553 (34%), Positives = 290/553 (51%), Gaps = 37/553 (6%)
Query: 9 RILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIVALVLHVGI 68
+I+ + ++F+Q+D+E WD+ +I T++ +A LP +LP W + G I+ +LH G
Sbjct: 78 KIVDKPIEFEQVDRERTWDDQVIFNTLLMYLANIKLPGASHLPPWRLDGAILMALLHAGP 137
Query: 69 SEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVVIGIPILGASV 128
E LYYW HR H +L++ YHS HHSS V E +T EH +++ IP++ AS+
Sbjct: 138 VEFLYYWFHRALHHHFLYSRYHSHHHSSIVTEPITSVVHPFAEHIAYTLLFAIPMVTASL 197
Query: 129 MGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTPTYHSLHHTDNKD 188
G S + GY+ DF+ +GHCN E+ P LF FP +K+L YTP++HSLHHT +
Sbjct: 198 CGILSIVSIMGYITYIDFMNNMGHCNFELFPKRLFHLFPPLKFLCYTPSFHSLHHTQFR- 256
Query: 189 TNFCLFMPLFDALGHTLNTKSWELHESFSSGKCSRVPDFVFLAHIVDVTSSMHAQFCLRS 248
TN+ LFMP++D + T + + L+E S PD + L H+ S + S
Sbjct: 257 TNYSLFMPIYDFIYGTTDNLTDSLYER-SLEIEEESPDVIHLTHLTTHNSIYQMRLGFPS 315
Query: 249 FASLPFRTR--FFLIPF-WPIAITTLLAMW--VRSKAFLFSFYYLRDRLHQTWVVPRCGF 303
+S P +R ++L F WP + A+ + + F+F LRD + ++P+ F
Sbjct: 316 LSSCPLWSRPPWYLTCFMWPFTLLCSFALTSAIPLRTFVFERNRLRDLTVHSHLLPKFSF 375
Query: 304 QYFLPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVDKHPNLRVRVV 363
E IN IEEAIL AD+ GVKV+SL +N +R+V
Sbjct: 376 HRH----HESINTIIEEAILEADEKGVKVMSLGLMN-------------------NIRLV 412
Query: 364 HGNTLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTLSADRFKRIQK 423
G+++ A V+I+ IPK+ E+ G +K+ A+ LCQK VKV++L R + K
Sbjct: 413 DGSSMAATVVINNIPKEATEIVFRGNLTKVASAVVFALCQKGVKVVVL-----REEEHSK 467
Query: 424 EAPQEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVVPPILPFRRDC 483
LV T + K W+VG I EQ A GT F F P R+DC
Sbjct: 468 LIKSGVDKNLVLSTS-NSYYSPKVWLVGDGIENEEQMKAKEGTLFVPFSHFPPNKLRKDC 526
Query: 484 TYGELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVG-AIDVDRIDV 542
Y AMR+P+ + + SCE + R V+ A GG+VH+LEGW H+ G +V R+
Sbjct: 527 FYQSTPAMRVPKSAQNIDSCENWLGRRVMSAWKIGGIVHALEGWEEHDCGNTCNVLRLHA 586
Query: 543 VWKAALKHGLRPV 555
+W+AAL+H +P+
Sbjct: 587 IWEAALRHDFQPL 599
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.326 0.141 0.450
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 982,386,408
Number of Sequences: 2790947
Number of extensions: 42458680
Number of successful extensions: 137201
Number of sequences better than 10.0: 312
Number of HSP's better than 10.0 without gapping: 116
Number of HSP's successfully gapped in prelim test: 196
Number of HSP's that attempted gapping in prelim test: 136825
Number of HSP's gapped (non-prelim): 335
length of query: 562
length of database: 848,049,833
effective HSP length: 133
effective length of query: 429
effective length of database: 476,853,882
effective search space: 204570315378
effective search space used: 204570315378
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 77 (34.3 bits)
Lotus: description of TM0041.8


