

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0041.7
(1610 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
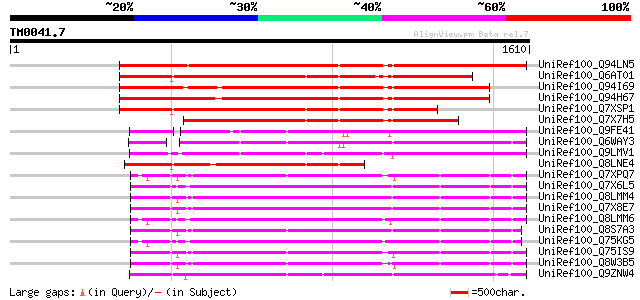
Sequences producing significant alignments: (bits) Value
UniRef100_Q94LN5 Putative retroelement pol polyprotein [Oryza sa... 1183 0.0
UniRef100_Q6AT01 Putative polyprotein [Oryza sativa] 1054 0.0
UniRef100_Q94I69 Putative retroelement [Oryza sativa] 1049 0.0
UniRef100_Q94H67 Putative gag-pol protein [Oryza sativa] 1048 0.0
UniRef100_Q7XSP1 OSJNBa0070M12.15 protein [Oryza sativa] 952 0.0
UniRef100_Q7X7H5 OSJNBa0085C10.2 protein [Oryza sativa] 875 0.0
UniRef100_Q9FE41 Oryza sativa (japonica cultivar-group) genomic ... 818 0.0
UniRef100_Q6WAY3 Gag/pol polyprotein [Pisum sativum] 753 0.0
UniRef100_Q9LMV1 F5M15.26 [Arabidopsis thaliana] 743 0.0
UniRef100_Q8LNE4 Putative retroelement [Oryza sativa] 700 0.0
UniRef100_Q7XPQ7 OSJNBa0053K19.16 protein [Oryza sativa] 684 0.0
UniRef100_Q7X6L5 OSJNBb0093G06.3 protein [Oryza sativa] 683 0.0
UniRef100_Q8LMM4 Putative gag-pol [Oryza sativa] 682 0.0
UniRef100_Q7X8E7 OSJNBa0042F21.5 protein [Oryza sativa] 681 0.0
UniRef100_Q8LMM6 Putative gag-pol [Oryza sativa] 676 0.0
UniRef100_Q8S7A3 Putative retroelement [Oryza sativa] 676 0.0
UniRef100_Q75KG5 Putative polyprotein [Oryza sativa] 676 0.0
UniRef100_Q75IS9 Putative polyprotein [Oryza sativa] 675 0.0
UniRef100_Q8W3B5 Putative gag-pol [Oryza sativa] 675 0.0
UniRef100_Q9ZNW4 Polyprotein [Sorghum bicolor] 674 0.0
>UniRef100_Q94LN5 Putative retroelement pol polyprotein [Oryza sativa]
Length = 1580
Score = 1183 bits (3060), Expect = 0.0
Identities = 597/1263 (47%), Positives = 829/1263 (65%), Gaps = 20/1263 (1%)
Query: 341 DFDESDGEDLNIICIDDKAIFQRPSEHMKSHLKPLFVWAKVEDKGVNKILVDGGAAINLM 400
D DE + E +I ++ +F++P HLKPL++ V K ++K++VDGGAA+NLM
Sbjct: 330 DVDEIEEESAKLILSPEQVVFEKPEGTENRHLKPLYINGYVNGKPMSKMMVDGGAAVNLM 389
Query: 401 PRFMMKRLGKTETDLIPHDMVLSDYEGKTSTSMGAIMLNITAGTVSRSTLFIVVPSKANY 460
P ++LG+ DLI +MVL D+ G S +MG + + +T G+ + T F V+ K +Y
Sbjct: 390 PYATFRKLGRNAEDLIKTNMVLKDFGGNPSETMGVLNVELTVGSKTIPTTFFVIDGKGSY 449
Query: 461 NLLLGREWIHGVGAVPSTLHQRISIWKPDGVVENVQADQSYYLAEAGYVDKKNFEKSPLV 520
+LLLGR+WIH +PST+HQ + W+ D + E V AD + Y + E S +
Sbjct: 450 SLLLGRDWIHANCCIPSTIHQCLIQWQGDKI-EIVPADSQLKMENPSYCFEGVMEGSNVY 508
Query: 521 DATKMEAQDPLEKVDLGDGSKKRPTYISSLIDPELRGRMIELLKEFKDCFA*DNDEMPGL 580
++ D + L IS I + R +IELLKEF+DCF + EMPG
Sbjct: 509 TKDTVDDLDDKQGQGLMSADDLEEIDISPAIG-QGRHLLIELLKEFRDCFTWEYYEMPGH 567
Query: 581 SRDLVELQLPIKEDKKLVK*LPRRFHPDVLVKIKEEIERLLRCKFIRAAKYVEWLANVVP 640
SR +VE +LP+K + + LPRR D+L +K EI+RL FIR +Y EW++++VP
Sbjct: 568 SRSIVEHRLPLKPGVRPHQQLPRRCKADMLEPVKAEIKRLYDAGFIRPCRYAEWVSSIVP 627
Query: 641 VIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAK 700
VIKKNGK+RVCIDFR LN ATPKDEY M +A+ +VD+A+GH+ LS +DG +GYNQIF+A+
Sbjct: 628 VIKKNGKVRVCIDFRYLNKATPKDEYPMLVADQLVDAASGHKILSFMDGNAGYNQIFMAE 687
Query: 701 EDVSKTAFRCSGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDLVVKS 760
ED+ KT FRC GA+G +EWVV+ F LK+AGATYQR MN I+HD I ++VYIDD+VVKS
Sbjct: 688 EDIHKTTFRCPGAIGLFEWVVITFVLKSAGATYQRAMNYIYHDLIGWLVEVYIDDVVVKS 747
Query: 761 PSRDEHLSHLRKSFERMRIHGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILD 820
++H++ LRK FER R +GLKMNP KCAFGV AG FLGF+VH++GIE+ + AI
Sbjct: 748 KEIEDHIADLRKVFERTRKYGLKMNPTKCAFGVSAGQFLGFLVHERGIEVTQRSINAIKK 807
Query: 821 TSPPTSKKQLQSLLGKVNFLRRFIDNLSDKTKPFSSLLRLKREDIFRWEA*HQKAFDELK 880
PP K +LQ ++GK+NF+RRFI LS K +PF+ LLRLK + F W A QKA D +K
Sbjct: 808 IKPPEDKTELQEMIGKINFVRRFISILSGKLEPFTPLLRLKADQQFAWGAEQQKALDNIK 867
Query: 881 NYLAIPPVMIPPIKGKPMRLYISATDETIGSMLAQEDEDGIERAVFYLSRVLNDAKTRYT 940
YL+ PPV+IPP KG P RLY+SA D++I S+L QE E ER VFYLSR L +A+TRY+
Sbjct: 868 EYLSSPPVLIPPQKGIPFRLYLSAGDKSISSVLIQELERK-ERVVFYLSRRLLNAETRYS 926
Query: 941 MIEKLCLCLYFSCTKLKYYIKPIDVMVFSHFDIIKHMLSKPILHSRIGKWSLALTEYSLT 1000
+EKLCLCLYFSCT+L++Y+ + V D++K+MLS PIL RIGKW +LTE+ L
Sbjct: 927 PVEKLCLCLYFSCTRLRHYLLSNECTVICKADVVKYMLSAPILKGRIGKWIFSLTEFDLR 986
Query: 1001 YAPLKTIKGQAVADFLADHTLPEEIAYVGLQPWKLFFDDSSHKEGSGIGMFIVSPQGIPT 1060
Y K IKGQA+A+F+ D+ + I V + PW LFFD S G GIG+ I+SP+G
Sbjct: 987 YESPKAIKGQAIANFIVDYR-DDSIGSVEVVPWTLFFDGSVCTHGCGIGLVIISPRGACF 1045
Query: 1061 KFMFRIRESCSNNESEYEALVSGLEILLALGAKNVEVKGDSELVVKQLTKEYKCISKNLA 1120
+F + I+ +NN++EYEA++ GL++L + A +E+ GDS LV+ QL EY+C + L
Sbjct: 1046 EFAYTIKPYATNNQAEYEAVLKGLQLLKEVEADTIEIMGDSLLVISQLAGEYECKNDTLI 1105
Query: 1121 KYYVKAMSLLANFDQAGVSYIPRVSNQEANELAQIASGYMIDKQKLKELIRIKEKLSPLD 1180
Y K L+ F + ++ R N EAN+LAQ ASGY K + +
Sbjct: 1106 VYNEKCQELMREFRLVTLKHVSREQNIEANDLAQGASGY-------------KLMIKHVQ 1152
Query: 1181 LEVMVIDNLTPNDWRKPIVEYLQNPVGSTDIKVKYRALNYTIIGNELFKKNVDGTLLKCL 1240
+EV VI T +DWR + +YLQ+P S K++Y+AL Y ++ +EL+ + +DG LLKCL
Sbjct: 1153 IEVAVI---TADDWRYYVYQYLQDPSQSASRKLRYKALKYILLDDELYYRTIDGVLLKCL 1209
Query: 1241 SEVDAFIAVSAAHGGLCGAHQAGAKMKWILFRQGMYWPTIMKDCIEYAKGCQDCQKHSGI 1300
S A +A+ H G+CG HQ+ KMKW+L G +WPT+++DC Y KGCQDCQK I
Sbjct: 1210 STDQAKVAIGEVHEGICGTHQSAHKMKWLLRCAGYFWPTMLEDCFRYYKGCQDCQKFGAI 1269
Query: 1301 QHVPASELHSIIKSWPFRGWAIDLIGEIHPALSRQHKYIIVAIDYFIKWVEAIPLQSVTQ 1360
Q PAS ++ IIK WPFRGW ID+IG I+P S+ HK+I+VA DYF KWVEAIPL+ V
Sbjct: 1270 QRAPASAMNPIIKPWPFRGWGIDMIGMINPPSSKGHKFILVATDYFTKWVEAIPLKKVDS 1329
Query: 1361 ETVIEFIQNHIVYRFGLPESLTTDQGTMFVGRKVAAFAESWGIKLLTSIPYYAQANGQVE 1420
I+F+Q HI+YR G+P+++TTDQG++FV + FA+S GIKLL S PYYAQANGQ E
Sbjct: 1330 GDAIQFVQEHIIYRVGIPQTITTDQGSIFVSDEFIQFADSMGIKLLNSSPYYAQANGQAE 1389
Query: 1421 AANKILISLIKKHVGRKPKSWHESLSQVLWAYRNSPREATGTTPFRLAYGQEAVLPAEVY 1480
A+NK LI LIK+ + P+ WH L++ LW+YR + + P++L Y +A+LP EV
Sbjct: 1390 ASNKSLIKLIKRKISDYPRQWHTRLAEALWSYRMACHGSIQVPPYKLVYRHKAILPWEVR 1449
Query: 1481 LQSFRIQRQEDIPSEVYWNMMLDELVNLDEERVLALDVLTRQKDRIAKAYNKKVKNRSFV 1540
+ S R + Q D+ ++ Y N+M DE +L + R+ AL + + K+R+ + YNKKV + F
Sbjct: 1450 IGSRRTELQNDLTADEYSNLMADEREDLVQSRLRALAKVIKDKERVTRHYNKKVVPKDFS 1509
Query: 1541 TGDYVWKVILPTDKKDRAYGKWAPNWEGPFKVKKVLSNNAYVIKELSGQRQFVTINDKYL 1600
G+ +WK+ILP +D +GKW+PNWEGPF++ KV+S AY+++ L G+ +N KYL
Sbjct: 1510 EGELIWKLILPIGTRDSKFGKWSPNWEGPFQIHKVVSKGAYMLQGLDGEVYDRALNGKYL 1569
Query: 1601 KAY 1603
K Y
Sbjct: 1570 KKY 1572
>UniRef100_Q6AT01 Putative polyprotein [Oryza sativa]
Length = 1739
Score = 1054 bits (2725), Expect = 0.0
Identities = 538/1107 (48%), Positives = 739/1107 (66%), Gaps = 43/1107 (3%)
Query: 341 DFDESDGEDLNIICIDDKAIFQRPSEHMKSHLKPLFVWAKVEDKGVNKILVDGGAAINLM 400
D DE + E ++ ++A+FQ+P HLKPL++ V K ++K++VDGGAA+NLM
Sbjct: 661 DVDEVEEESAKLVLSPEQAVFQKPEGTENRHLKPLYINGYVNGKLMSKMMVDGGAAVNLM 720
Query: 401 PRFMMKRLGKTETDLIPHDMVLSDYEGKTSTSMGAIMLNITAGTVSRSTLFIVVPSKANY 460
P ++LG+ DLI +MVL D+ G S + G + + +T G+ + T F+V+ K +Y
Sbjct: 721 PYATFRKLGRNAEDLIKTNMVLKDFGGNPSETKGVLNVGLTVGSKTIPTTFLVIDGKGSY 780
Query: 461 NLLLGREWIHGVGAVPSTLHQRISIWKPDGVVENVQADQ-------SYYLAEAGYVDKKN 513
+L LGR+WIH +PST+HQ + W+ D V E V AD SYY G V+ +
Sbjct: 781 SLFLGRDWIHANCCIPSTMHQCLIQWQGDKV-EIVPADSQLKMENPSYYFE--GVVEGSS 837
Query: 514 -FEKSPLVDATKMEAQ-----DPLEKVDLGDGSKKRPTYISSLIDPELRGRMIELLKEFK 567
+ K + D + Q D LE++D+G G + RPT++S + PE R ++IELLKEF+
Sbjct: 838 VYTKDTVDDLYDKQGQGFMSADDLEEIDIGPGDRPRPTFVSKNLSPEFRTKLIELLKEFR 897
Query: 568 DCFA*DNDEMPGLSRDLVELQLPIKEDKKLVK*LPRRFHPDVLVKIKEEIERLLRCKFIR 627
DCFA + EMPGLSR +VE +LPIK + + RR D+L +K EI+RL FIR
Sbjct: 898 DCFAWEYYEMPGLSRSIVEHRLPIKPGVRPRQQPLRRCKADMLEPVKAEIKRLYDAGFIR 957
Query: 628 AAKYVEWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLL 687
+Y +W++++VPVIKKNGK+RVCIDFRDLN ATPKDEY MP+A+ +VD+A+G++ LS +
Sbjct: 958 PWRYAKWVSSIVPVIKKNGKVRVCIDFRDLNKATPKDEYPMPVADQLVDAASGNKILSFM 1017
Query: 688 DGYSGYNQIFIAKEDVSKTAFRCSGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIET 747
DG +GYNQIF+A+ED+ KTAFRC A+G +EWVVM FGLK+AGATYQR MN I+HD I
Sbjct: 1018 DGNAGYNQIFMAEEDIHKTAFRCPCAIGLFEWVVMTFGLKSAGATYQRAMNYIYHDLIGW 1077
Query: 748 FMQVYIDDLVVKSPSRDEHLSHLRKSFERMRIHGLKMNPLKCAFGVIAGDFLGFVVHKKG 807
+ VYIDD+VVKS ++ ++ LRK FER R +GLKMNP KCAFGV AG FLGF+VH +G
Sbjct: 1078 LVGVYIDDVVVKSKEIEDQIADLRKVFERTRKYGLKMNPTKCAFGVSAGQFLGFLVHDRG 1137
Query: 808 IEINKNKAKAILDTSPPTSKKQLQSLLGKVNFLRRFIDNLSDKTKPFSSLLRLKREDIFR 867
IE+ + AI PP +K +LQ ++GK++F+RRFI NLS + +PF+ LLRLK + F
Sbjct: 1138 IEVTQRSVNAIKKIQPPENKTELQEMIGKIHFVRRFISNLSGRLEPFTPLLRLKADQQFT 1197
Query: 868 WEA*HQKAFDELKNYLAIPPVMIPPIKGKPMRLYISATDETIGSMLAQEDEDGIERAVFY 927
A QKA D++K YL+ PPV+IPP KG P RLY+SA +++IGS+L QE E G ER VFY
Sbjct: 1198 CGAEQQKALDDIKEYLSSPPVLIPPHKGIPFRLYLSAGEKSIGSVLIQELE-GKERVVFY 1256
Query: 928 LSRVLNDAKTRYTMIEKLCLCLYFSCTKLKYYIKPIDVMVFSHFDIIKHMLSKPILHSRI 987
LSR L +A+TRY+ ++KLCLCLYFSCT+L++Y+ + V D++K+MLS PIL R+
Sbjct: 1257 LSRRLLNAETRYSPVKKLCLCLYFSCTRLRHYLLSNECTVICKADVVKYMLSAPILKGRV 1316
Query: 988 GKWSLALTEYSLTYAPLKTIKGQAVADFLADHTLPEEIAYVGLQPWKLFFDDSSHKEGSG 1047
GKW +LTE+ Y K IKGQA+ADF+ +H + I V + PW LFFD S G G
Sbjct: 1317 GKWIFSLTEFDHRYESPKAIKGQAIADFIVEHR-DDSIGSVEIVPWTLFFDGSVCTHGCG 1375
Query: 1048 IGMFIVSPQGIPTKFMFRIRESCSNNESEYEALVSGLEILLALGAKNVEVKGDSELVVKQ 1107
IG+ I+SP+G +F + I+ +NN++EYEA++ GL++L + A +E+ GDS LV+ Q
Sbjct: 1376 IGLVIISPRGACFEFAYTIKPYATNNQAEYEAVLKGLQLLKKVEADTIEIMGDSLLVISQ 1435
Query: 1108 LTKEYKCISKNLAKYYVKAMSLLANFDQAGVSYIPRVSNQEANELAQIASGYMIDKQKLK 1167
L EY+C + L Y K L+ F R+ N EAN+LAQ ASGY
Sbjct: 1436 LAGEYECKNDTLMVYNEKCQELMKEF---------RLQNIEANDLAQGASGY-------- 1478
Query: 1168 ELIRIKEKLSPLDLEVMVIDNLTPNDWRKPIVEYLQNPVGSTDIKVKYRALNYTIIGNEL 1227
K + + +E+ +T +DWR + YL NP S K++Y+AL YT++ +EL
Sbjct: 1479 -----KPMIKDVKVEIAA---MTADDWRYDVHRYLSNPSQSASRKLRYKALKYTLLDDEL 1530
Query: 1228 FKKNVDGTLLKCLSEVDAFIAVSAAHGGLCGAHQAGAKMKWILFRQGMYWPTIMKDCIEY 1287
+ + +DG LLKCLS A +A+ H G+CG HQ+ KMKW+L R G +W T+++DC Y
Sbjct: 1531 YYRTIDGVLLKCLSADQAKVAIGEVHEGICGTHQSAHKMKWLLRRAGYFWSTMLEDCFRY 1590
Query: 1288 AKGCQDCQKHSGIQHVPASELHSIIKSWPFRGWAIDLIGEIHPALSRQHKYIIVAIDYFI 1347
KGCQDCQK IQ PAS ++ IIK WPFRGW ID+IG I+P S+ HK+I+VA DYF
Sbjct: 1591 YKGCQDCQKFGAIQRAPASAMNPIIKPWPFRGWGIDMIGMINPPSSKGHKFILVATDYFT 1650
Query: 1348 KWVEAIPLQSVTQETVIEFIQNHIVYRFGLPESLTTDQGTMFVGRKVAAFAESWGIKLLT 1407
KWVEAIPL+ V I+F+Q +I+YRFG P+++TTDQG++FV + FA+S IKLL
Sbjct: 1651 KWVEAIPLKKVDSGDAIQFVQEYIIYRFGTPQTITTDQGSIFVSDEFVQFADSMSIKLLN 1710
Query: 1408 SIPYYAQANGQVEAANKILISLIKKHV 1434
S PYYAQANGQ EA+NK LI LIK+ +
Sbjct: 1711 SSPYYAQANGQAEASNKSLIKLIKRKI 1737
>UniRef100_Q94I69 Putative retroelement [Oryza sativa]
Length = 2014
Score = 1049 bits (2713), Expect = 0.0
Identities = 546/1158 (47%), Positives = 741/1158 (63%), Gaps = 55/1158 (4%)
Query: 341 DFDESDGEDLNIICIDDKAIFQRPSEHMKSHLKPLFVWAKVEDKGVNKILVDGGAAINLM 400
D DE + E +I + ++AIF++P HLKPL++ V K ++K++VDGGA +NLM
Sbjct: 883 DVDEVEEESAKLILLPEQAIFEKPEGTENRHLKPLYINGYVNGKPMSKMMVDGGAEVNLM 942
Query: 401 PRFMMKRLGKTETDLIPHDMVLSDYEGKTSTSMGAIMLNITAGTVSRSTLFIVVPSKANY 460
P ++LG+ DLI +MVL D+ G S + G + +T G + T F V+ +Y
Sbjct: 943 PYATFRKLGRNVEDLIKTNMVLIDFGGNLSETKGVSNVELTVGNKTIPTTFFVIDGNGSY 1002
Query: 461 NLLLGREWIHGVGAVPSTLHQRISIWKPDGVVENVQADQSYYLAEA-----GYVDKKN-F 514
+LLLGR+WIH +PST+HQ + W+ D + E V AD + G V+ N +
Sbjct: 1003 SLLLGRDWIHANCCIPSTMHQCLIQWQDDKI-EIVPADSQLKMENPSCYFEGVVEGSNVY 1061
Query: 515 EKSPLVDATKMEAQ-----DPLEKVDLGDGSKKRPTYISSLIDPELRGRMIELLKEFKDC 569
K + D + Q D LE++D+G PE R ++IELLKEF+DC
Sbjct: 1062 IKDTVDDLDDKQGQGFISADDLEEIDIG---------------PEFRAKLIELLKEFRDC 1106
Query: 570 FA*DNDEMPGLSRDLVELQLPIKEDKKLVK*LPRRFHPDVLVKIKEEIERLLRCKFIRAA 629
FA + EMPGLSR +VE +LPIK + + RR D+L +K EI+RL FIR
Sbjct: 1107 FAWEYYEMPGLSRSIVEHRLPIKPGVRPHQQPLRRCKADMLEPVKVEIKRLYDACFIRPC 1166
Query: 630 KYVEWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDG 689
+Y EW++++VPVIKKN DLN ATPKDEY MP+A+ +VD+A+G++ LS +DG
Sbjct: 1167 RYAEWVSSIVPVIKKN----------DLNKATPKDEYPMPVADQLVDAASGNKILSYMDG 1216
Query: 690 YSGYNQIFIAKEDVSKTAFRCSGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFM 749
GYNQIF+A+ED+ KTAFRC GA+G +EWVVM FGLK+AGATYQR MN I+HD I +
Sbjct: 1217 NVGYNQIFMAEEDIHKTAFRCPGAIGLFEWVVMTFGLKSAGATYQRAMNYIYHDLIGWLV 1276
Query: 750 QVYIDDLVVKSPSRDEHLSHLRKSFERMRIHGLKMNPLKCAFGVIAGDFLGFVVHKKGIE 809
+VYIDD+VVKS ++H++ LRK FER R +GLKMNP KCAFGV G FLGF+VH++GIE
Sbjct: 1277 EVYIDDVVVKSKEIEDHIADLRKVFERTRKYGLKMNPTKCAFGVSVGQFLGFLVHERGIE 1336
Query: 810 INKNKAKAILDTSPPTSKKQLQSLLGKVNFLRRFIDNLSDKTKPFSSLLRLKREDIFRWE 869
+ + AI PP +K +LQ + GK+NF+RRFI LS + +PF+ LLRLK + F W
Sbjct: 1337 VTQRSVNAIKKIQPPENKTELQEMNGKINFVRRFISYLSGRLEPFTPLLRLKADQQFTWG 1396
Query: 870 A*HQKAFDELKNYLAIPPVMIPPIKGKPMRLYISATDETIGSMLAQEDEDGIERAVFYLS 929
A QKA D++K YL+ PPV+IPP KG RLY+SA +++IGS+L QE DG ER VFYLS
Sbjct: 1397 AEQQKALDDIKEYLSSPPVLIPPQKGISFRLYLSAGEKSIGSVLIQE-LDGKERVVFYLS 1455
Query: 930 RVLNDAKTRYTMIEKLCLCLYFSCTKLKYYIKPIDVMVFSHFDIIKHMLSKPILHSRIGK 989
R L D +TRY+ +EKLCLCLYFSCT+L +Y+ + V D+IK+MLS PIL R+GK
Sbjct: 1456 RRLLDVETRYSPMEKLCLCLYFSCTRLSHYLLSNECTVICKADVIKYMLSAPILKGRVGK 1515
Query: 990 WSLALTEYSLTYAPLKTIKGQAVADFLADHTLPEEIAYVGLQPWKLFFDDSSHKEGSGIG 1049
W +LTE+ L Y K IKGQA+ADF+ +H + I V + PW FFD S GIG
Sbjct: 1516 WIFSLTEFDLRYESPKAIKGQAIADFIVEHC-DDSIGSVEVVPWTSFFDGSVCTHDCGIG 1574
Query: 1050 MFIVSPQGIPTKFMFRIRESCSNNESEYEALVSGLEILLALGAKNVEVKGDSELVVKQLT 1109
+ I+SP+G KF + I+ +NN++EYEA++ GL++L + A +E+ GDS LV+ QL
Sbjct: 1575 LVIISPRGACFKFAYTIKPYATNNQAEYEAVLKGLQLLKEVEADAIEIMGDSLLVISQLA 1634
Query: 1110 KEYKCISKNLAKYYVKAMSLLANFDQAGVSYIPRVSNQEANELAQIASGYMIDKQKLKEL 1169
EY+C S L Y K L+ F + ++ R N EAN+LAQ ASGY
Sbjct: 1635 GEYECKSDTLMVYNEKCQELMQEFRLVTLKHVSREQNIEANDLAQGASGY---------- 1684
Query: 1170 IRIKEKLSPLDLEVMVIDNLTPNDWRKPIVEYLQNPVGSTDIKVKYRALNYTIIGNELFK 1229
K + + E+ I T DWR + +YL NP S K++Y+AL YT++ +EL+
Sbjct: 1685 ---KPMIKDVKAEIAAI---TAGDWRYDVHQYLHNPSQSASRKLRYKALKYTLLDDELYY 1738
Query: 1230 KNVDGTLLKCLSEVDAFIAVSAAHGGLCGAHQAGAKMKWILFRQGMYWPTIMKDCIEYAK 1289
+ +DG LLKCLS A +A+ H G+CG HQ+ KMKW+L R +WPT+++DC +Y K
Sbjct: 1739 RTIDGVLLKCLSADQAKVAIGEVHEGICGTHQSAHKMKWLLRRARYFWPTMLEDCFKYYK 1798
Query: 1290 GCQDCQKHSGIQHVPASELHSIIKSWPFRGWAIDLIGEIHPALSRQHKYIIVAIDYFIKW 1349
GCQDCQK IQ PAS ++ IIK WPFRGW ID+IG I S+ HK+I+VA DYF K
Sbjct: 1799 GCQDCQKFGAIQRAPASAMNPIIKPWPFRGWGIDMIGMISRPSSKGHKFILVATDYFTKL 1858
Query: 1350 VEAIPLQSVTQETVIEFIQNHIVYRFGLPESLTTDQGTMFVGRKVAAFAESWGIKLLTSI 1409
VEAIPL+ V I+F+Q HI+YRFG+P+++TTDQG++FV + F +S GIKLL S
Sbjct: 1859 VEAIPLKKVDSGDAIQFVQEHIIYRFGIPQTITTDQGSIFVSDEFVQFVDSMGIKLLNSS 1918
Query: 1410 PYYAQANGQVEAANKILISLIKKHVGRKPKSWHESLSQVLWAYRNSPREATGTTPFRLAY 1469
PYYAQANGQ EA+NK LI LIK+ P+ WH L + LW+YR + + P++L Y
Sbjct: 1919 PYYAQANGQAEASNKSLIKLIKRKNSDYPRQWHTRLDEALWSYRMACHGSIQVPPYKLVY 1978
Query: 1470 GQEAVLPAEVYLQSFRIQ 1487
G EA+LP EV + S R +
Sbjct: 1979 GHEAILPWEVRIGSRRTE 1996
>UniRef100_Q94H67 Putative gag-pol protein [Oryza sativa]
Length = 2273
Score = 1048 bits (2711), Expect = 0.0
Identities = 543/1158 (46%), Positives = 739/1158 (62%), Gaps = 49/1158 (4%)
Query: 341 DFDESDGEDLNIICIDDKAIFQRPSEHMKSHLKPLFVWAKVEDKGVNKILVDGGAAINLM 400
D DE + E +I + ++AIF++P HLKPL++ V K ++K++VDGGA +NLM
Sbjct: 1136 DVDEVEEESAKLILLPEQAIFEKPEGTENRHLKPLYINGYVNGKPMSKMMVDGGAEVNLM 1195
Query: 401 PRFMMKRLGKTETDLIPHDMVLSDYEGKTSTSMGAIMLNITAGTVSRSTLFIVVPSKANY 460
P ++LG+ DLI +MVL D+ G S + G + +T G + T F V+ +Y
Sbjct: 1196 PYATFRKLGRNVEDLIKTNMVLIDFGGNLSETKGVSNVELTVGNKTIPTTFFVIDGNGSY 1255
Query: 461 NLLLGREWIHGVGAVPSTLHQRISIWKPDGVVENVQADQSYYLAEA-----GYVDKKN-F 514
+LLLGR+WIH +PST+HQ + W+ D + E V AD + G V+ N +
Sbjct: 1256 SLLLGRDWIHANCCIPSTMHQCLIQWQDDKI-EIVPADSQLKMENPSCYFEGVVEGSNVY 1314
Query: 515 EKSPLVDATKMEAQ-----DPLEKVDLGDGSKKRPTYISSLIDPELRGRMIELLKEFKDC 569
K + D + Q D LE++D+G G + RPT+I + E R ++IELLKEF+DC
Sbjct: 1315 IKDTVDDLDDKQGQGFISADDLEEIDIGPGDRPRPTFICKNLSTEFRAKLIELLKEFRDC 1374
Query: 570 FA*DNDEMPGLSRDLVELQLPIKEDKKLVK*LPRRFHPDVLVKIKEEIERLLRCKFIRAA 629
FA + EMPGLSR +VE +LPIK + + RR D+L +K EI+RL FIR
Sbjct: 1375 FAWEYYEMPGLSRSIVEHRLPIKPGVRPHQQPLRRCKADMLEPVKVEIKRLYDACFIRPC 1434
Query: 630 KYVEWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDG 689
+Y EW++N LN ATPKDEY MP+A+ +VD+A+G++ LS +DG
Sbjct: 1435 RYAEWVSN-------------------LNKATPKDEYPMPVADQLVDAASGNKILSYMDG 1475
Query: 690 YSGYNQIFIAKEDVSKTAFRCSGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFM 749
GYNQIF+A+ED+ KTAFRC GA+G +EWVVM FGLK+AGATYQR MN I+HD I +
Sbjct: 1476 NVGYNQIFMAEEDIHKTAFRCPGAIGLFEWVVMTFGLKSAGATYQRAMNYIYHDLIGWLV 1535
Query: 750 QVYIDDLVVKSPSRDEHLSHLRKSFERMRIHGLKMNPLKCAFGVIAGDFLGFVVHKKGIE 809
+VYIDD+VVKS ++H++ LRK FER R +GLKMNP KCAFGV G FLGF+VH++GIE
Sbjct: 1536 EVYIDDVVVKSKEIEDHIADLRKVFERTRKYGLKMNPTKCAFGVSVGQFLGFLVHERGIE 1595
Query: 810 INKNKAKAILDTSPPTSKKQLQSLLGKVNFLRRFIDNLSDKTKPFSSLLRLKREDIFRWE 869
+ + AI PP +K +LQ + GK+NF+RRFI LS + +PF+ LLRLK + F W
Sbjct: 1596 VTQRSVNAIKKIQPPENKTELQEMNGKINFVRRFISYLSGRLEPFTPLLRLKADQQFTWG 1655
Query: 870 A*HQKAFDELKNYLAIPPVMIPPIKGKPMRLYISATDETIGSMLAQEDEDGIERAVFYLS 929
A QKA D++K YL+ PPV+IPP KG RLY+SA +++IGS+L QE DG ER VFYLS
Sbjct: 1656 AEQQKALDDIKEYLSSPPVLIPPQKGISFRLYLSAGEKSIGSVLIQE-LDGKERVVFYLS 1714
Query: 930 RVLNDAKTRYTMIEKLCLCLYFSCTKLKYYIKPIDVMVFSHFDIIKHMLSKPILHSRIGK 989
R L D +TRY+ +EKLCLCLYFSCT+L +Y+ + V D+IK+MLS PIL R+GK
Sbjct: 1715 RRLLDVETRYSPMEKLCLCLYFSCTRLSHYLLSNECTVICKADVIKYMLSAPILKGRVGK 1774
Query: 990 WSLALTEYSLTYAPLKTIKGQAVADFLADHTLPEEIAYVGLQPWKLFFDDSSHKEGSGIG 1049
W +LTE+ L Y K IKGQA+ADF+ +H + I V + PW FFD S GIG
Sbjct: 1775 WIFSLTEFDLRYESPKAIKGQAIADFIVEHC-DDSIGSVEVVPWTSFFDGSVCTHDCGIG 1833
Query: 1050 MFIVSPQGIPTKFMFRIRESCSNNESEYEALVSGLEILLALGAKNVEVKGDSELVVKQLT 1109
+ I+SP+G KF + I+ +NN++EYEA++ GL++L + A +E+ GDS LV+ QL
Sbjct: 1834 LVIISPRGACFKFAYTIKPYATNNQAEYEAVLKGLQLLKEVEADAIEIMGDSLLVISQLA 1893
Query: 1110 KEYKCISKNLAKYYVKAMSLLANFDQAGVSYIPRVSNQEANELAQIASGYMIDKQKLKEL 1169
EY+C S L Y K L+ F + ++ R N EAN+LAQ ASGY
Sbjct: 1894 GEYECKSDTLMVYNEKCQELMQEFRLVTLKHVSREQNIEANDLAQGASGY---------- 1943
Query: 1170 IRIKEKLSPLDLEVMVIDNLTPNDWRKPIVEYLQNPVGSTDIKVKYRALNYTIIGNELFK 1229
K + + E+ I T DWR + +YL NP S K++Y+AL YT++ +EL+
Sbjct: 1944 ---KPMIKDVKAEIAAI---TAGDWRYDVHQYLHNPSQSASRKLRYKALKYTLLDDELYY 1997
Query: 1230 KNVDGTLLKCLSEVDAFIAVSAAHGGLCGAHQAGAKMKWILFRQGMYWPTIMKDCIEYAK 1289
+ +DG LLKCLS A +A+ H G+CG HQ+ KMKW+L R +WPT+++DC +Y K
Sbjct: 1998 RTIDGVLLKCLSADQAKVAIGEVHEGICGTHQSAHKMKWLLRRARYFWPTMLEDCFKYYK 2057
Query: 1290 GCQDCQKHSGIQHVPASELHSIIKSWPFRGWAIDLIGEIHPALSRQHKYIIVAIDYFIKW 1349
GCQDCQK IQ PAS ++ IIK WPFRGW ID+IG I S+ HK+I+VA DYF K
Sbjct: 2058 GCQDCQKFGAIQRAPASAMNPIIKPWPFRGWGIDMIGMISRPSSKGHKFILVATDYFTKL 2117
Query: 1350 VEAIPLQSVTQETVIEFIQNHIVYRFGLPESLTTDQGTMFVGRKVAAFAESWGIKLLTSI 1409
VEAIPL+ V I+F+Q HI+YRFG+P+++TTDQG++FV + F +S GIKLL S
Sbjct: 2118 VEAIPLKKVDSGDAIQFVQEHIIYRFGIPQTITTDQGSIFVSDEFVQFVDSMGIKLLNSS 2177
Query: 1410 PYYAQANGQVEAANKILISLIKKHVGRKPKSWHESLSQVLWAYRNSPREATGTTPFRLAY 1469
PYYAQANGQ EA+NK LI LIK+ P+ WH L + LW+YR + + P++L Y
Sbjct: 2178 PYYAQANGQAEASNKSLIKLIKRKNSDYPRQWHTRLDEALWSYRMACHGSIQVPPYKLVY 2237
Query: 1470 GQEAVLPAEVYLQSFRIQ 1487
G EA+LP EV + S R +
Sbjct: 2238 GHEAILPWEVRIGSRRTE 2255
>UniRef100_Q7XSP1 OSJNBa0070M12.15 protein [Oryza sativa]
Length = 1685
Score = 952 bits (2460), Expect = 0.0
Identities = 485/999 (48%), Positives = 663/999 (65%), Gaps = 34/999 (3%)
Query: 341 DFDESDGEDLNIICIDDKAIFQRPSEHMKSHLKPLFVWAKVEDKGVNKILVDGGAAINLM 400
D DE + E ++ ++A+F++P HLKPL++ V K ++K++VDGGAA+NLM
Sbjct: 706 DVDELEEESAKLVLSPERAVFEKPEGTENRHLKPLYINGYVNGKPMSKMMVDGGAAVNLM 765
Query: 401 PRFMMKRLGKTETDLIPHDMVLSDYEGKTSTSMGAIMLNITAGTVSRSTLFIVVPSKANY 460
P ++LG+T DL+ +MVL D+ G S + G + + +T G+ + T F V+ K +Y
Sbjct: 766 PYTTFRKLGRTTEDLMKTNMVLKDFGGNPSETKGVLNVELTVGSKTIPTTFFVIDGKGSY 825
Query: 461 NLLLGREWIHGVGAVPSTLHQRISIWKPDGVVENVQADQ-------SYYLAEAGYVDKKN 513
+LLLGR+WIH +PST+HQ + W+ D V E V AD SYY G V+ +
Sbjct: 826 SLLLGRDWIHANCCIPSTMHQCLIQWQGDKV-EIVPADSRLKMKNPSYYFE--GVVEGSS 882
Query: 514 -FEKSPLVDATKMEAQ-----DPLEKVDLGDGSKKRPTYISSLIDPELRGRMIELLKEFK 567
+ K + D + Q D L++VD+G G + RPT+IS + E R +++ELLKEF+
Sbjct: 883 IYAKDTVDDLDDKQGQGFMSADDLDEVDIGPGDRPRPTFISKNLSSEFRTKLMELLKEFR 942
Query: 568 DCFA*DNDEMPGLSRDLVELQLPIKEDKKLVK*LPRRFHPDVLVKIKEEIERLLRCKFIR 627
DCFA + EMPGLSR +VE +LP+K + + PRR D+L +K EI+RL FI
Sbjct: 943 DCFAWEYYEMPGLSRSIVEHRLPLKPGIRPHQQPPRRCKADMLEPVKAEIKRLYDAGFIH 1002
Query: 628 AAKYVEWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLL 687
+Y EW++++VPVIKKNGK+RVCIDFRDLN ATPKDEY MP+A+ +VD+A+GH+ LS +
Sbjct: 1003 PCRYAEWVSSIVPVIKKNGKVRVCIDFRDLNKATPKDEYPMPVADQLVDAASGHKILSFM 1062
Query: 688 DGYSGYNQIFIAKEDVSKTAFRCSGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIET 747
DG +GYNQIF+ +ED+ KT FRC GA+G +EWVVM FGLK+AGATYQR MN I+HD I
Sbjct: 1063 DGNAGYNQIFMTEEDIHKTVFRCPGAIGLFEWVVMTFGLKSAGATYQRAMNYIYHDLIGW 1122
Query: 748 FMQVYIDDLVVKSPSRDEHLSHLRKSFERMRIHGLKMNPLKCAFGVIAGDFLGFVVHKKG 807
++VYIDD+VVKS ++H++ LRK FER R +GLKMNP KCAFGV AG FLGF+VH++G
Sbjct: 1123 LVEVYIDDVVVKSREIEDHIADLRKVFERTRKYGLKMNPTKCAFGVSAGQFLGFLVHERG 1182
Query: 808 IEINKNKAKAILDTSPPTSKKQLQSLLGKVNFLRRFIDNLSDKTKPFSSLLRLKREDIFR 867
IEI + AI PP +K +LQ ++GK+NF+RRFI NLS K +PF+ LLRL+ + F
Sbjct: 1183 IEIPQRSINAIKKIKPPGNKTELQEMIGKINFVRRFISNLSGKLEPFTPLLRLRADQQFT 1242
Query: 868 WEA*HQKAFDELKNYLAIPPVMIPPIKGKPMRLYISATDETIGSMLAQEDEDGIERAVFY 927
W A QKA D +K YL+ PPV+IPP KG P LY+SA D++IGS+L QE E G ER VFY
Sbjct: 1243 WGAEQQKALDNIKEYLSSPPVLIPPQKGIPFWLYLSAGDKSIGSVLIQELE-GKERVVFY 1301
Query: 928 LSRVLNDAKTRYTMIEKLCLCLYFSCTKLKYYIKPIDVMVFSHFDIIKHMLSKPILHSRI 987
LSR L DA+TRY+ +EKLCLCLYFSCTKL++Y+ + V D+IK+MLS PIL R+
Sbjct: 1302 LSRRLLDAETRYSPVEKLCLCLYFSCTKLRHYLLSNECTVVCKADVIKYMLSAPILKVRV 1361
Query: 988 GKWSLALTEYSLTYAPLKTIKGQAVADFLADHTLPEEIAYVGLQPWKLFFDDSSHKEGSG 1047
GKW +LTE+ L + K IKGQAVADF+ H I V + PW LFFD S G G
Sbjct: 1362 GKWIFSLTEFDLRHESPKAIKGQAVADFIVGHR-DGSIGLVDIMPWVLFFDGSVCSHGYG 1420
Query: 1048 IGMFIVSPQGIPTKFMFRIRESCSNNESEYEALVSGLEILLALGAKNVEVKGDSELVVKQ 1107
IG+ I++P G +F + I +NN++EYEA++ GL++L + A +E+ GDS LV+ Q
Sbjct: 1421 IGLVIIAPWGASFEFAYAITHHVTNNQAEYEAVLKGLQLLQEVAADAIEIMGDSLLVISQ 1480
Query: 1108 LTKEYKCISKNLAKYYVKAMSLLANFDQAGVSYIPRVSNQEANELAQIASGYMIDKQKLK 1167
L EY+C L YY K +L++ F + +I R N EAN+L+Q ASGY
Sbjct: 1481 LAGEYECKDDTLMVYYEKCRTLISEFRLVTLRHISREQNIEANDLSQGASGY-------- 1532
Query: 1168 ELIRIKEKLSPLDLEVMVIDNLTPNDWRKPIVEYLQNPVGSTDIKVKYRALNYTIIGNEL 1227
K +++E+ I T +DWR + +YLQNP S K++Y+AL YT++ ++L
Sbjct: 1533 -----KPMTKDIEIEIATI---TADDWRYDVFQYLQNPSQSALRKLRYKALKYTLLDDDL 1584
Query: 1228 FKKNVDGTLLKCLSEVDAFIAVSAAHGGLCGAHQAGAKMKWILFRQGMYWPTIMKDCIEY 1287
+ + +DG LLKCLS A +A+ H G+CG HQ+ KMKW+L R G +WPT+++DC Y
Sbjct: 1585 YYRTIDGVLLKCLSADQAKVAIGEIHEGICGTHQSAHKMKWLLRRAGYFWPTMLEDCFRY 1644
Query: 1288 AKGCQDCQKHSGIQHVPASELHSIIKSWPFRGWAIDLIG 1326
KGCQDCQK IQ PAS ++ II+ WPFRGW ID+IG
Sbjct: 1645 YKGCQDCQKFGAIQRAPASAMNPIIQPWPFRGWGIDMIG 1683
>UniRef100_Q7X7H5 OSJNBa0085C10.2 protein [Oryza sativa]
Length = 861
Score = 875 bits (2261), Expect = 0.0
Identities = 431/852 (50%), Positives = 588/852 (68%), Gaps = 18/852 (2%)
Query: 539 GSKKRPTYISSLIDPELRGRMIELLKEFKDCFA*DNDEMPGLSRDLVELQLPIKEDKKLV 598
G + RPT+IS + E R ++IELLKEF+DCFA + EMPGLSR +VE +LPIK +
Sbjct: 26 GDQPRPTFISKNLSSEFRTKLIELLKEFRDCFAWEYYEMPGLSRSIVEHRLPIKPGVRPR 85
Query: 599 K*LPRRFHPDVLVKIKEEIERLLRCKFIRAAKYVEWLANVVPVIKKNGKMRVCIDFRDLN 658
+ PRR D+L +K EI+RL FI +Y EW++++VPVIKKNGK+RVCIDFRDLN
Sbjct: 86 QQPPRRCKADMLEPVKAEIKRLYDAGFIHPYRYAEWVSSIVPVIKKNGKVRVCIDFRDLN 145
Query: 659 AATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAKEDVSKTAFRCSGALGTYE 718
ATPKDEY MP+A+ + D+A+G++ LS +D +GYNQIF+A+ED+ KTAFRC A+G +E
Sbjct: 146 KATPKDEYPMPVADQLFDAASGNKILSFMDRNTGYNQIFMAEEDIHKTAFRCPSAIGLFE 205
Query: 719 WVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDLVVKSPSRDEHLSHLRKSFERMR 778
WVVM F LK+AGATYQR MN I+HD I + ++V IDD+VVKS D+H++ LRK FER R
Sbjct: 206 WVVMTFDLKSAGATYQRAMNYIYHDLIGSLVEVDIDDVVVKSKEVDDHIADLRKVFERTR 265
Query: 779 IHGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKVN 838
+GLKMNP KCAFGV A FLGF+VH++GIE+ + AI PP +K +LQ ++GK+N
Sbjct: 266 KYGLKMNPTKCAFGVSARQFLGFLVHERGIEVTQRSVNAIQKIKPPENKTELQEMIGKIN 325
Query: 839 FLRRFIDNLSDKTKPFSSLLRLKREDIFRWEA*HQKAFDELKNYLAIPPVMIPPIKGKPM 898
F+RRFI NLS + +PF+ LLRLK + F W A QKA D +K YL+ PPV+IPP KG P
Sbjct: 326 FVRRFISNLSGRLEPFTPLLRLKADQQFTWGAEQQKALDNIKEYLSSPPVLIPPQKGIPF 385
Query: 899 RLYISATDETIGSMLAQEDEDGIERAVFYLSRVLNDAKTRYTMIEKLCLCLYFSCTKLKY 958
RLY+SA +++ GS+L QE E G ER VFY+SR L DA+TRY+ +EKLCLCLYF CT+L++
Sbjct: 386 RLYLSAGEKSNGSVLIQELE-GKERVVFYISRRLLDAETRYSPMEKLCLCLYFLCTRLRH 444
Query: 959 YIKPIDVMVFSHFDIIKHMLSKPILHSRIGKWSLALTEYSLTYAPLKTIKGQAVADFLAD 1018
Y+ + V D++++MLS PIL +IG W +LTE+ L Y K IKGQA+ADF+ +
Sbjct: 445 YLLSNECTVICKADVVRYMLSAPILKGQIGNWIFSLTEFDLRYESPKAIKGQAIADFIVE 504
Query: 1019 HTLPEEIAYVGLQPWKLFFDDSSHKEGSGIGMFIVSPQGIPTKFMFRIRESCSNNESEYE 1078
H + I V + PW LFFD S G GIG+ I+SP+G +F + I+ +NN++EYE
Sbjct: 505 HR-DDSIGSVEIVPWTLFFDGSVCTHGCGIGLVIISPRGACFEFAYTIKSYATNNQAEYE 563
Query: 1079 ALVSGLEILLALGAKNVEVKGDSELVVKQLTKEYKCISKNLAKYYVKAMSLLANFDQAGV 1138
A++ GL++L + A +E+ GDS LV+ QL EY+C S Y K + L+ F +
Sbjct: 564 AVLKGLQLLKEVEANIIEIMGDSLLVISQLAGEYECKSDTFIVYNEKCLELMKEFRLVTL 623
Query: 1139 SYIPRVSNQEANELAQIASGYMIDKQKLKELIRIKEKLSPLDLEVMVIDNLTPNDWRKPI 1198
++ R N EAN+LAQ ASGY K + + +E+ ++ +DWR +
Sbjct: 624 KHVSREQNLEANDLAQGASGY-------------KPMIKDVKVEIAA---MSADDWRYDV 667
Query: 1199 VEYLQNPVGSTDIKVKYRALNYTIIGNELFKKNVDGTLLKCLSEVDAFIAVSAAHGGLCG 1258
+YL NP S K++Y+AL YT++G+EL+ + +DG LLKCLS A +A+ H G+CG
Sbjct: 668 HQYLSNPSQSASRKLRYKALKYTLLGDELYYRVIDGVLLKCLSADQAKVAIGEVHEGICG 727
Query: 1259 AHQAGAKMKWILFRQGMYWPTIMKDCIEYAKGCQDCQKHSGIQHVPASELHSIIKSWPFR 1318
HQ+ KMKW+L R G +WPT+++ Y KGCQDCQK IQ PAS ++ IIK WPFR
Sbjct: 728 THQSAHKMKWLLRRTGYFWPTMLEVYFRYYKGCQDCQKFGAIQRAPASAMNPIIKPWPFR 787
Query: 1319 GWAIDLIGEIHPALSRQHKYIIVAIDYFIKWVEAIPLQSVTQETVIEFIQNHIVYRFGLP 1378
GW ID+IG I+P S+ HK+I+VA DYF KWVEAIPL+ V I+F+Q HI+Y+FGLP
Sbjct: 788 GWEIDMIGMINPPSSKGHKFILVATDYFTKWVEAIPLKKVDSGDAIQFVQEHIIYQFGLP 847
Query: 1379 ESLTTDQGTMFV 1390
+++T DQG++F+
Sbjct: 848 QTITMDQGSIFM 859
>UniRef100_Q9FE41 Oryza sativa (japonica cultivar-group) genomic DNA, chromosome 1, PAC
clone:P0433F09 [Oryza sativa]
Length = 2876
Score = 818 bits (2112), Expect = 0.0
Identities = 435/1110 (39%), Positives = 644/1110 (57%), Gaps = 46/1110 (4%)
Query: 529 DPLEKVDLGDGSKKRPTYISSLIDPELRGRMIELLKEFKDCFA*DNDEMPGLSRDLVELQ 588
D L +++LG RP ++S ++ E R L EF+DCFA EMPGL + +
Sbjct: 1776 DELVEMNLGTEDDPRPIFVSGMLTEEEREDYRSFLMEFRDCFAWTYKEMPGLDSRVATHK 1835
Query: 589 LPIKEDKKLVK*LPRRFHPDVLVKIKEEIERLLRCKFIRAAKYVEWLANVVPVIKKNGKM 648
L I + VK PRR P+ ++ E++RL+ FI+ +Y WLAN+VPV KKNG++
Sbjct: 1836 LAIDPQFRPVKQPPRRLRPEFQDQVIAEVDRLINVGFIKEIQYPRWLANIVPVEKKNGQV 1895
Query: 649 RVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAKEDVSKTAF 708
RVC+DFRDLN A PKD++ +PI EM+VDS G+ LS GYNQI + D TAF
Sbjct: 1896 RVCVDFRDLNRACPKDDFPLPITEMVVDSTTGYGALS------GYNQIKMDLLDAFDTAF 1949
Query: 709 RCSGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDLVVKSPSRDEHLS 768
R G + + VMPFGLKNAGATYQR M + D I ++ Y+DD+VVK+ + H
Sbjct: 1950 RTPK--GNFYYTVMPFGLKNAGATYQRAMQFVLDDLIHHSVECYVDDMVVKTKDHEHHQE 2007
Query: 769 HLRKSFERMRIHGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKK 828
LR FER+R H LKMNPLKCAF V +G FLGFV+ +GIEI K KAIL+ PP K
Sbjct: 2008 DLRIVFERLRRHQLKMNPLKCAFAVQSGVFLGFVIRHRGIEIEPKKIKAILNMPPPQELK 2067
Query: 829 QLQSLLGKVNFLRRFIDNLSDKTKPFSSLLRLKREDIFRWEA*HQKAFDELKNYLAIPPV 888
L+ L GK+ ++RRFI NLS + +PFS L+ K+ F W+ Q FD +K YL PPV
Sbjct: 2068 DLRKLQGKLAYIRRFISNLSGRIQPFSKLM--KKGTPFVWDEECQNGFDSIKRYLLNPPV 2125
Query: 889 MIPPIKGKPMRLYISATDETIGSMLAQEDEDGIERAVFYLSRVLNDAKTRYTMIEKLCLC 948
+ P+KG+P+ LYI+ +IG++LAQ +++G E A +YLSR + A+ Y+ IEKLCL
Sbjct: 2126 LAAPVKGRPLILYIATQPASIGALLAQHNDEGKEVACYYLSRTMVGAEQNYSPIEKLCLA 2185
Query: 949 LYFSCTKLKYYIKPIDVMVFSHFDIIKHMLSKPILHSRIGKWSLALTEYSLTYAPLKTIK 1008
L F+ KL++Y+ + + + D I+++LS+P+L R+GKW+L + EY +T+ P K IK
Sbjct: 2186 LIFALKKLRHYMLAHQIQLIARADPIRYVLSQPVLTGRLGKWALLMMEYDITFVPQKAIK 2245
Query: 1009 GQAVADFLADHTLPEEIAYVGLQP------------WKLFFDDSSHKE---------GSG 1047
GQA+A+FLA H +P++ + P W+L+FD +S K+ +G
Sbjct: 2246 GQALAEFLATHPMPDDSPLIANLPDEEIFTAELQEQWELYFDGASRKDINPDGTPRRRAG 2305
Query: 1048 IGMFIVSPQGIPTKFMFRI-RESCSNNESEYEALVSGLEILLALGAKNVEVKGDSELVVK 1106
G+ +PQG F + +E CSNNE+EYEAL+ GL + L++ +++ GDS L+++
Sbjct: 2306 AGLVFKTPQGGVIYHSFSLLKEECSNNEAEYEALIFGLLLALSMEVRSLRAHGDSRLIIR 2365
Query: 1107 QLTKEYKCISKNLAKYYVKAMSLLANFDQAGVSYIPRVSNQEANELAQIASGYMIDKQKL 1166
Q+ Y+ L YY A L+ F+ V ++PR N A+ LA++A+ +
Sbjct: 2366 QINNIYEVRKPELVPYYTVARRLMDKFEHIEVIHVPRSKNAPADALAKLAAALVFQGDNP 2425
Query: 1167 KELIRIKEK--------LSPLDLEVMVIDNLTPNDWRKPIVEYLQN---PVGSTDIKVKY 1215
+++ ++E+ L P ++ +++ ++ DWR+P ++Y ++ P + +
Sbjct: 2426 AQIV-VEERWLLPAVLELIPEEVNIIITNSAEEEDWRQPFLDYFKHGSLPEDPVERRQLQ 2484
Query: 1216 RAL-NYTIIGNELFKKNV-DGTLLKCLSEVDAFIAVSAAHGGLCGAHQAGAKMKWILFRQ 1273
R L +Y L+K++ LL+C+ +A + H G+CG HQ+G KM +
Sbjct: 2485 RRLPSYIYKAGVLYKRSYGQEVLLRCVDRSEANRVLQEVHHGVCGGHQSGPKMYHSIRLV 2544
Query: 1274 GMYWPTIMKDCIEYAKGCQDCQKHSGIQHVPASELHSIIKSWPFRGWAIDLIGEIHPALS 1333
G YWP IM DC++ AK C CQ H +H P + LH + SWPF W ID+IG I+P S
Sbjct: 2545 GYYWPGIMADCLKTAKTCHGCQIHDNFKHQPPAPLHPTVPSWPFDAWGIDVIGLINPPSS 2604
Query: 1334 RQHKYIIVAIDYFIKWVEAIPLQSVTQETVIEFIQNHIVYRFGLPESLTTDQGTMFVGRK 1393
R H++I+ A DYF KW EA+PL+ V VI F++ HI+YRFG+P +T+D F +K
Sbjct: 2605 RGHRFILTATDYFSKWAEAVPLREVKSSDVINFLERHIIYRFGVPHRITSDNAKAFKSQK 2664
Query: 1394 VAAFAESWGIKLLTSIPYYAQANGQVEAANKILISLIKKHVGRKPKSWHESLSQVLWAYR 1453
+ F E + IK S YY QANG EA NK L ++KK V + + WH+ L + LWAYR
Sbjct: 2665 IYRFMEKYKIKWNYSTGYYPQANGMAEAFNKTLGKILKKTVDKHRRDWHDRLYEALWAYR 2724
Query: 1454 NSPREATGTTPFRLAYGQEAVLPAEVYLQSFRIQRQEDIPSEVYWNMMLDELVNLDEERV 1513
+ R T TP+ L YG EAVLP E+ L S R+ +++ + + EL ++EER+
Sbjct: 2725 VTVRTPTQATPYSLVYGNEAVLPLEIQLPSLRVAIHDELTKDEQIRLRFQELDAVEEERL 2784
Query: 1514 LALDVLTRQKDRIAKAYNKKVKNRSFVTGDYVWKVILPTDKKDRAYGKWAPNWEGPFKVK 1573
AL L + + +AY+K VK R F G+ V + P + GK+ P WEGP+ ++
Sbjct: 2785 GALQNLELYRQNMVRAYDKLVKQRVFRKGELVLVLRRPIVVTHKMKGKFEPKWEGPYVIE 2844
Query: 1574 KVLSNNAYVIKELSGQRQFVTINDKYLKAY 1603
+ AY + + G + IN ++LK Y
Sbjct: 2845 QAYDGGAYQLIDHQGSQPMPPINGRFLKKY 2874
Score = 94.0 bits (232), Expect = 3e-17
Identities = 46/138 (33%), Positives = 77/138 (55%)
Query: 371 HLKPLFVWAKVEDKGVNKILVDGGAAINLMPRFMMKRLGKTETDLIPHDMVLSDYEGKTS 430
H +PL++ + + +IL+D G+A+N++P + R G T DL P D+V+ ++ +
Sbjct: 1378 HNRPLYIEGNIGSAHLRRILIDLGSAVNILPVRSLTRAGFTTKDLEPIDVVICGFDNQGK 1437
Query: 431 TSMGAIMLNITAGTVSRSTLFIVVPSKANYNLLLGREWIHGVGAVPSTLHQRISIWKPDG 490
++GAI + I T S F V+ + +Y+ LLGR WIH VPSTLHQ + +G
Sbjct: 1438 PTLGAITIKIQMSTFSFKVRFFVIEANTSYSALLGRPWIHKYRVVPSTLHQCLKFLDGNG 1497
Query: 491 VVENVQADQSYYLAEAGY 508
V + + ++ S Y + Y
Sbjct: 1498 VQQRITSNFSPYTIQESY 1515
>UniRef100_Q6WAY3 Gag/pol polyprotein [Pisum sativum]
Length = 2262
Score = 753 bits (1943), Expect = 0.0
Identities = 410/1113 (36%), Positives = 638/1113 (56%), Gaps = 47/1113 (4%)
Query: 528 QDPLEKVDLGDGSKKRPTYISSLIDPELRGRMIELLKEFKDCFA*DNDEMPGLSRDLVEL 587
Q+ LE V+LG R I + ++ ++ R+IE+L+E+ + FA +MPGL D+V
Sbjct: 1158 QEELEVVNLGTEDATREIKIGAALEDSVKRRLIEMLREYVEIFAWSYQDMPGLDTDIVVH 1217
Query: 588 QLPIKEDKKLVK*LPRRFHPDVLVKIKEEIERLLRCKFIRAAKYVEWLANVVPVIKKNGK 647
+LP++E VK RR PD+ KIKEE+++ F+ Y W+AN+VPV KK+GK
Sbjct: 1218 RLPLREGCPSVKQKLRRTSPDMATKIKEEVQKQWDAGFLAVTSYPPWMANIVPVPKKDGK 1277
Query: 648 MRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAKEDVSKTA 707
+R+C+D+RDLN A+PKD++ +P +++VD+ A S +DG+SGYNQI +A ED+ KT
Sbjct: 1278 VRMCVDYRDLNRASPKDDFPLPHIDVLVDNTAQSSVFSFMDGFSGYNQIKMAPEDMEKTT 1337
Query: 708 FRCSGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDLVVKSPSRDEHL 767
F GT+ + VMPFGLKNAGATYQR M T+FHD + ++VY+DD++ KS + +EHL
Sbjct: 1338 FITPW--GTFCYKVMPFGLKNAGATYQRAMTTLFHDMMHKEIEVYVDDMIAKSQTEEEHL 1395
Query: 768 SHLRKSFERMRIHGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSK 827
+L+K F+R+R L++NP KC FGV +G LGF+V +KGIE++ K KAI + P ++
Sbjct: 1396 VNLQKLFDRLRKFKLRLNPNKCTFGVRSGKLLGFIVSEKGIEVDPAKVKAIQEMPEPKTE 1455
Query: 828 KQLQSLLGKVNFLRRFIDNLSDKTKPFSSLLRLKREDIFRWEA*HQKAFDELKNYLAIPP 887
KQ++ LG++N++ RFI +L+ +P LLR + +W QKAFD++K YL PP
Sbjct: 1456 KQVRGFLGRLNYIARFISHLTATCEPIFKLLR--KNQAIKWNDDCQKAFDKIKEYLQKPP 1513
Query: 888 VMIPPIKGKPMRLYISATDETIGSMLAQEDEDGI-ERAVFYLSRVLNDAKTRYTMIEKLC 946
++IPP+ G+P+ +Y+S T+ ++G +L + DE G E A++YLS+ D +TRY+++EK C
Sbjct: 1514 ILIPPVPGRPLIMYLSVTENSMGCVLGRHDESGRKEHAIYYLSKKFTDCETRYSLLEKTC 1573
Query: 947 LCLYFSCTKLKYYIKPIDVMVFSHFDIIKHMLSKPILHSRIGKWSLALTEYSLTYAPLKT 1006
L ++ +L+ Y+ ++ S D +K++ KP L R+ +W + LTEY + Y K
Sbjct: 1574 CALAWAARRLRQYMLNHTTLLISKMDPVKYIFEKPALTGRVARWQMILTEYDIQYTSQKA 1633
Query: 1007 IKGQAVADFLADHTLP-----------EEIAYVGLQP---------------WKLFFDDS 1040
IKG ++D+LA+ + E+I Y+ ++ W L FD +
Sbjct: 1634 IKGSILSDYLAEQPIEDYQPMMFEFPDEDIMYLKMKDCKEPLVEEGPDPDDKWTLMFDGA 1693
Query: 1041 SHKEGSGIGMFIVSPQGIPTKFMFRIRESCSNNESEYEALVSGLEILLALGAKNVEVKGD 1100
+ G+G+G +++P+G F R+ +NNE+EYEA + G+E + L K +++ GD
Sbjct: 1694 VNMNGNGVGAVLINPKGAHMPFSARLTFDVTNNEAEYEACIMGIEEAIDLRIKTLDIFGD 1753
Query: 1101 SELVVKQLTKEYKCISKNLAKYYVKAMSLLANFDQAGVSYIPRVSNQEANELAQIASGYM 1160
S LVV Q+ ++ +L Y +L F + + ++PR NQ A+ LA ++S
Sbjct: 1754 SALVVNQVNGDWNTNQPHLIPYRDYTRRILTFFKKVKLYHVPRDENQMADALATLSSMIK 1813
Query: 1161 IDKQKLKELIRIKEKLSPLDL---EVMVIDNLTPNDWRKPIVEYLQN---PVG-STDIKV 1213
++ + + P + E +VID W I +L+ P G S + K
Sbjct: 1814 VNWWNHVPHVAVNRLERPAYVFAAESVVIDE---KPWYYDIKNFLKTQEYPEGASKNDKK 1870
Query: 1214 KYRALNYTIIGNE---LFKKNVDGTLLKCLSEVDAFIAVSAAHGGLCGAHQAGAKMKWIL 1270
R L + N+ L+K+N D LL+C+ +A + + H G G H G M L
Sbjct: 1871 TLRRLAGSFYLNQDDVLYKRNFDMVLLRCMDRPEADMLMQEVHEGSFGTHAGGHAMAKKL 1930
Query: 1271 FRQGMYWPTIMKDCIEYAKGCQDCQKHSGIQHVPASELHSIIKSWPFRGWAIDLIGEIHP 1330
R G YW T+ DC +YA+ C CQ ++ HVP S L+ + WPF W ID+IG+I P
Sbjct: 1931 LRAGYYWMTMESDCFKYARKCHKCQIYADRVHVPPSPLNVMNSPWPFAMWGIDMIGKIEP 1990
Query: 1331 ALSRQHKYIIVAIDYFIKWVEAIPLQSVTQETVIEFIQNHIVYRFGLPESLTTDQGTMFV 1390
S H++I+VAIDYF KWVEA ++T++ V FI+ I+ R+G+PE + TD G+
Sbjct: 1991 TASNGHRFILVAIDYFTKWVEAASYANITKQVVTRFIKKEIICRYGVPERIITDNGSNLN 2050
Query: 1391 GRKVAAFAESWGIKLLTSIPYYAQANGQVEAANKILISLIKKHVGRKPKSWHESLSQVLW 1450
+ + + + I+ S PY + NG VEAANK + +++K V K WHE L L
Sbjct: 2051 NKMMKELCKDFKIEHHNSSPYRPKMNGAVEAANKNIKKIVRKMVVTY-KDWHEMLPFALH 2109
Query: 1451 AYRNSPREATGTTPFRLAYGQEAVLPAEVYLQSFRIQRQEDIPSEVYWNMMLDELVNLDE 1510
YR S R +TG TP+ L YG EAVLP EV + S R+ + + +EL ++E
Sbjct: 2110 GYRTSVRTSTGATPYSLVYGMEAVLPVEVEIPSLRVLLDVKLDEAEWIRTRFNELSLIEE 2169
Query: 1511 ERVLALDVLTRQKDRIAKAYNKKVKNRSFVTGDYVWKVILPTDKKDRAYGKWAPNWEGPF 1570
R+ + + R+ +A+++KV+ RS+ GD V K ILP +R GKW PN+EGP+
Sbjct: 2170 RRLAVVCHGQLYQRRMKRAFDQKVRPRSYQIGDLVLKRILPPGTDNR--GKWTPNYEGPY 2227
Query: 1571 KVKKVLSNNAYVIKELSGQRQFVTINDKYLKAY 1603
VKKV S A ++ + G+ +N +K Y
Sbjct: 2228 VVKKVFSGGALMLTTMDGEDFPSPVNSDVVKKY 2260
Score = 72.8 bits (177), Expect = 8e-11
Identities = 35/119 (29%), Positives = 68/119 (56%), Gaps = 1/119 (0%)
Query: 369 KSHLKPLFVWAKVEDKGVNKILVDGGAAINLMPRFMMKRLGKTETDLIPHDMVLSDYEGK 428
++H K L + + + ++ +LVD G+++N++P+ ++K++ L P D+++ ++
Sbjct: 823 RNHNKALHITMECKGAVLSHVLVDTGSSLNVLPKQILKKIDVEGFVLTPSDLIVRAFDRS 882
Query: 429 TSTSMGAIMLNITAGTVSRSTLFIVVPSKANYNLLLGREWIHGVGAVPSTLHQRIS-IW 486
+ G + L + G +F V+ + Y+ LLGR WIH GAV STLHQ++ +W
Sbjct: 883 KRSVCGEVTLPVKIGPEVFDIIFYVMDIQPAYSCLLGRPWIHAAGAVSSTLHQKLKYVW 941
>UniRef100_Q9LMV1 F5M15.26 [Arabidopsis thaliana]
Length = 1838
Score = 743 bits (1917), Expect = 0.0
Identities = 441/1252 (35%), Positives = 675/1252 (53%), Gaps = 48/1252 (3%)
Query: 371 HLKPLFVWAKVEDKGVNKILVDGGAAINLMPRFMMKRLGKTETDLIPHDMVLSDYEGKTS 430
H PL V + D V ++L+D G++++L+ + ++ + T+ + P L+ ++G
Sbjct: 614 HNDPLVVELIISDSRVTRVLIDTGSSVDLIFKDVLTAMNITDRQIKPVSKPLAGFDGDFV 673
Query: 431 TSMGAIMLNITAGTVSRSTLFIVVPSKANYNLLLGREWIHGVGAVPSTLHQRISIWKPDG 490
++G I L I G + F+V+ A YN++LG WIH + A+PST HQ + +G
Sbjct: 674 MTIGTIKLPIFVGGLIAWVKFVVIGKPAVYNVILGTPWIHQMQAIPSTYHQCVKFPTHNG 733
Query: 491 VVENVQADQSYYLAEAGYVDKKNFEKSPLVDATKMEAQDPLEKVDLGDGSKKRPTYISSL 550
+ + + +++E+S L E V++ + R + +
Sbjct: 734 IF-------TLRAPKEAKTPSRSYEESELCRT---------EMVNIDESDPTRCVGVGAE 777
Query: 551 IDPELRGRMIELLKEFKDCFA*DNDEMPGLSRDLVELQLPIKEDKKLVK*LPRRFHPDVL 610
I P +R +I LLK FA ++M G+ + +L + K VK R+ P+
Sbjct: 778 ISPSIRLELIALLKRNSKTFAWSIEDMKGIDPAITAHELNVDPTFKPVKQKRRKLGPERA 837
Query: 611 VKIKEEIERLLRCKFIRAAKYVEWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPI 670
+ EE+E+LL+ I KY EWLAN V V KKNGK RVC+D+ DLN A PKD Y +P
Sbjct: 838 RAVNEEVEKLLKAGQIIEVKYPEWLANPVVVKKKNGKWRVCVDYTDLNKACPKDSYPLPH 897
Query: 671 AEMMVDSAAGHEYLSLLDGYSGYNQIFIAKEDVSKTAFRCSGALGTYEWVVMPFGLKNAG 730
+ +V++ +G+ LS +D +SGYNQI + K+D KT+F GTY + VM FGLKNAG
Sbjct: 898 IDRLVEATSGNGLLSFMDAFSGYNQILMHKDDQEKTSFVTDR--GTYCYKVMSFGLKNAG 955
Query: 731 ATYQRVMNTIFHDFIETFMQVYIDDLVVKSPSRDEHLSHLRKSFERMRIHGLKMNPLKCA 790
ATYQR +N + D I ++VYIDD++VKS ++H+ HL K F+ + +G+K+NP KC
Sbjct: 956 ATYQRFVNKMLADQIGRTVEVYIDDMLVKSLKPEDHVEHLSKCFDVLNTYGMKLNPTKCT 1015
Query: 791 FGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKVNFLRRFIDNLSDK 850
FGV +G+FLG+VV K+GIE N + +AIL+ P + +++Q L G++ L RFI +DK
Sbjct: 1016 FGVTSGEFLGYVVTKRGIEANPKQIRAILELPSPRNAREVQRLTGRIAALNRFISRSTDK 1075
Query: 851 TKPFSSLLRLKREDIFRWEA*HQKAFDELKNYLAIPPVMIPPIKGKPMRLYISATDETIG 910
PF +L LKR F W+ ++AF++LK+YL+ PP+++ P G+ + LYI+ +D +
Sbjct: 1076 CLPFYNL--LKRRAQFDWDKDSEEAFEKLKDYLSTPPILVKPEVGETLYLYIAVSDHAVS 1133
Query: 911 SMLAQEDEDGIERAVFYLSRVLNDAKTRYTMIEKLCLCLYFSCTKLKYYIKPIDVMVFSH 970
S+L +ED G +R +FY S+ L +A+TRY +IEK L + S KL+ Y + + V +
Sbjct: 1134 SVLVREDR-GEQRPIFYTSKSLVEAETRYPVIEKAALAVVTSARKLRPYFQSHTIAVLTD 1192
Query: 971 FDIIKHMLSKPILHSRIGKWSLALTEYSLTYAPLKTIKGQAVADFLADHTLPE-EIAYVG 1029
++ L P R+ KW++ L+EY + + P +K Q +ADFL + L E A G
Sbjct: 1193 -QPLRVALHSPSQSGRMTKWAVELSEYDIDFRPRPAMKSQVLADFLIELPLQSAERAVSG 1251
Query: 1030 L--QPWKLFFDDSSHKEGSGIGMFIVSPQGIPTKFMFRIRESCSNNESEYEALVSGLEIL 1087
+ W L+ D SS GSGIG+ +VSP + FR+R +NN +EYE L++GL +
Sbjct: 1252 NRGEEWSLYVDGSSSARGSGIGIRLVSPTAEVLEQSFRLRFVATNNVAEYEVLIAGLRLA 1311
Query: 1088 LALGAKNVEVKGDSELVVKQLTKEYKCISKNLAKYYVKAMSLLANFDQAGVSYIPRVSNQ 1147
+ + DS+L+ QL+ EY+ ++ + Y + +F+ +S IPR N
Sbjct: 1312 AGMQITTIHAFTDSQLIAGQLSGEYEAKNEKMDAYLKIVQLMTKDFENFKLSKIPRGDNA 1371
Query: 1148 EANELAQIASGYMIDKQKLKELIRI----KEKLSPLDLEVMV--------IDNLTPNDWR 1195
A+ LA +A + L+ +I + K + D +V D P DWR
Sbjct: 1372 PADALAALA---LTSDSDLRRIIPVESIDKPSIDSTDAVEIVNTIRSSNAPDPADPTDWR 1428
Query: 1196 KPIVEYLQNPVGSTD----IKVKYRALNYTIIGNELFKKNVDGTLLKCLSEVDAFIAVSA 1251
I +YL + +D +++ +A YT++ L K + G +L CL + +
Sbjct: 1429 VEIRDYLSDGTLPSDKWTARRLRIKAAKYTLMKEHLLKVSAFGAMLNCLHGTEINEIMKE 1488
Query: 1252 AHGGLCGAHQAGAKMKWILFRQGMYWPTIMKDCIEYAKGCQDCQKHSGIQHVPASELHSI 1311
H G G H G + L + G YWPT++ DC + C+ CQ+H+ H P L +
Sbjct: 1489 THEGAAGNHSGGRALALKLKKLGFYWPTMISDCKTFTAKCEQCQRHAPTIHQPTELLRAG 1548
Query: 1312 IKSWPFRGWAIDLIGEIHPALSRQHKYIIVAIDYFIKWVEAIPLQSVTQETVIEFIQNHI 1371
+ +PF WA+D++G + PA SRQ ++I+V DYF KWVEA ++ V F+ I
Sbjct: 1549 VAPYPFMRWAMDIVGPM-PA-SRQKRFILVMTDYFTKWVEAESYATIRANDVQNFVWKFI 1606
Query: 1372 VYRFGLPESLTTDQGTMFVGRKVAAFAESWGIKLLTSIPYYAQANGQVEAANKILISLIK 1431
+ R GLP + TD G+ F+ F SW I+L S P Y Q NGQ EA NK ++S +K
Sbjct: 1607 ICRHGLPYEIITDNGSQFISLSFENFCASWKIRLNKSTPRYPQGNGQAEATNKTILSGLK 1666
Query: 1432 KHVGRKPKSWHESLSQVLWAYRNSPREATGTTPFRLAYGQEAVLPAEVYLQSFRIQRQED 1491
K + K +W + L VLW+YR +PR AT TPF AYG EA+ PAEV S R
Sbjct: 1667 KRLDEKKGAWADELDGVLWSYRTTPRSATDQTPFAHAYGMEAMAPAEVGYSSLRRSMMVK 1726
Query: 1492 IPSEVYWNMMLDELVNLDEERVLALDVLTRQKDRIAKAYNKKVKNRSFVTGDYVWKVILP 1551
P E+ MMLD L +L+E R AL + + AK YN+KV NR F GD V + +
Sbjct: 1727 NP-ELNDRMMLDRLDDLEEIRNAALCRIQNYQLAAAKHYNQKVHNRHFDVGDLVLRKVFE 1785
Query: 1552 TDKKDRAYGKWAPNWEGPFKVKKVLSNNAYVIKELSGQRQFVTINDKYLKAY 1603
+ A GK NWEG ++V K++ Y + +SG T N +LK Y
Sbjct: 1786 NTAEINA-GKLGANWEGSYQVSKIVRPGDYELLTMSGTAVPRTWNSMHLKRY 1836
>UniRef100_Q8LNE4 Putative retroelement [Oryza sativa]
Length = 1170
Score = 700 bits (1806), Expect = 0.0
Identities = 365/757 (48%), Positives = 503/757 (66%), Gaps = 33/757 (4%)
Query: 356 DDKAIFQRPSEHMKSHLKPLFVWAKVEDKGVNKILVDGGAAINLMPRFMMKRLGKTETDL 415
+D+AIF++P HLKPL++ V K ++K++VDGGAA+NLMP ++LG+ L
Sbjct: 431 EDEAIFEKPEGTENRHLKPLYINGYVNGKTMSKMMVDGGAAVNLMPYATFRKLGRNTEAL 490
Query: 416 IPHDMVLSDYEGKTSTSMGAIMLNITAGTVSRSTLFIVVPSKANYNLLLGREWIHGVGAV 475
I +MVL D+ G S + G + + +T G+ + T F V+ +Y+LLLGR+WIH +
Sbjct: 491 IKTNMVLKDFGGNPSETRGVLNVELTVGSKTIPTTFFVINGNGSYSLLLGRDWIHANCCI 550
Query: 476 PSTLHQRISIWKPDGVVENVQADQS-------YYLAEAGYVDKKN-FEKSPLVDATKMEA 527
PST+HQ + W+ D E V AD YY G V+ N + K + D +
Sbjct: 551 PSTMHQCLIQWQGDKT-EIVPADSQLKMENPCYYFE--GVVEGSNVYAKDTVDDLDDKQG 607
Query: 528 QD-----PLEKVDLGDGSKKRPTYISSLIDPELRGRMIELLKEFKDCFA*DNDEMPGLSR 582
QD LE++D+G G + RPT+I + PE R ++IELLKEF+DCFA + EMPGLSR
Sbjct: 608 QDFMSADELEEIDIGPGDRPRPTFIRKNLSPEFRTKLIELLKEFRDCFAWEYYEMPGLSR 667
Query: 583 DLVELQLPIKEDKKLVK*LPRRFHPDVLVKIKEEIERLLRCKFIRAAKYVEWLANVVPVI 642
+VE +LPIK + + PRR ++L +K EI+RL + VPVI
Sbjct: 668 SIVEHRLPIKPGVRPCQQPPRRCKANMLEPVKAEIKRLY---------------DAVPVI 712
Query: 643 KKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAKED 702
KKNGK+RVCIDFRDLN ATPKDEY MP+A+ +VD+A+G++ LS +DG +GYNQIF+A+ED
Sbjct: 713 KKNGKVRVCIDFRDLNKATPKDEYPMPVADQLVDAASGNKILSFVDGNAGYNQIFMAEED 772
Query: 703 VSKTAFRCSGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDLVVKSPS 762
+ K AFRC A+G +EWVVM FGLK+AGATYQR MN I+HD I ++VYID++VVKS
Sbjct: 773 IHKMAFRCPSAIGLFEWVVMTFGLKSAGATYQRAMNYIYHDLIGWLVEVYIDNVVVKSKE 832
Query: 763 RDEHLSHLRKSFERMRIHGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTS 822
++H++ LRK FER R +GLKMNP KCAFGV AG FLGF+VH++GIE+ + AI
Sbjct: 833 VEDHIADLRKVFERTRKYGLKMNPTKCAFGVSAGQFLGFLVHERGIEVTQRSVNAIKKIQ 892
Query: 823 PPTSKKQLQSLLGKVNFLRRFIDNLSDKTKPFSSLLRLKREDIFRWEA*HQKAFDELKNY 882
PP +K +LQ ++GK+NF+RRFI NLS + +PF+ LLRLK F W A QKA D +K Y
Sbjct: 893 PPENKTELQEMIGKINFVRRFISNLSGRLEPFTPLLRLKANQQFTWGAEQQKALDNIKEY 952
Query: 883 LAIPPVMIPPIKGKPMRLYISATDETIGSMLAQEDEDGIERAVFYLSRVLNDAKTRYTMI 942
L+ PPV+IPP KG P RLY+SA +++IGS+L QE E G ER VFYLSR L DA+T+Y+ +
Sbjct: 953 LSSPPVLIPPQKGIPFRLYLSAGEKSIGSVLIQELE-GKERVVFYLSRRLLDAETQYSPV 1011
Query: 943 EKLCLCLYFSCTKLKYYIKPIDVMVFSHFDIIKHMLSKPILHSRIGKWSLALTEYSLTYA 1002
EKLCLCLYFSCT+L++Y+ + V D++++ML PIL R+GKW +LTE+ L Y
Sbjct: 1012 EKLCLCLYFSCTRLRHYLLSNECTVICKADVVRYMLLAPILKRRVGKWIFSLTEFDLRYE 1071
Query: 1003 PLKTIKGQAVADFLADHTLPEEIAYVGLQPWKLFFDDSSHKEGSGIGMFIVSPQGIPTKF 1062
K IKGQA+ADF+ +H + I V + PW LFFD S G GI + I+SP+ +F
Sbjct: 1072 SPKAIKGQAIADFIVEHR-DDSIGSVEIVPWTLFFDGSVCTHGCGISLVIISPRAACFEF 1130
Query: 1063 MFRIRESCSNNESEYEALVSGLEILLALGAKNVEVKG 1099
+ I+ +NN++EYEA++ GL++L + A ++ G
Sbjct: 1131 AYTIKPYATNNQAEYEAVLKGLQLLKEVEADTLKSWG 1167
>UniRef100_Q7XPQ7 OSJNBa0053K19.16 protein [Oryza sativa]
Length = 2010
Score = 684 bits (1764), Expect = 0.0
Identities = 434/1286 (33%), Positives = 687/1286 (52%), Gaps = 103/1286 (8%)
Query: 374 PLFVWAKVEDKGVNKILVDGGAAINLMPRFMMKRLGKTETDL-IPHDMVLSD-------Y 425
PL + V + + + L+DGG+A+N++ KT D+ IPH +
Sbjct: 770 PLVLDPVVRNVKLRRTLIDGGSALNIL-------FAKTLDDMQIPHSELKPSNAPFHGVI 822
Query: 426 EGKTSTSMGAIMLNITAGTV----SRSTLFIVVPSKANYNLLLGREWIHGVGAVPSTLHQ 481
G ++T +G I L +T GT + + F V + Y+ +LGR + AVP +
Sbjct: 823 PGLSATPLGQITLPVTFGTRENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYM 882
Query: 482 RISIWKPDGVVENVQADQSYYLAEAGYVDKKNFEKSP------------LVDATKMEAQD 529
+ + P GV+ ++++D + +A DK++ + + L AT E ++
Sbjct: 883 MMKMPGPRGVL-SLRSD----IKQAVTCDKESCDMAQTREMASAREDIRLAAATASEGEE 937
Query: 530 PLEKVDLGDGSKKRPTYISSLIDPELRGRMIELLKEFKDCFA*DNDEMPGLSRDLVELQL 589
P K+ S+ + I +DP + KD FA +MPG+ R+++E L
Sbjct: 938 PATKISKSGESEAKTKKIP--LDPSDPAKTANN----KDIFAWKPSDMPGIPREVIEHSL 991
Query: 590 PIKEDKKLVK*LPRRFHPDVLVKIKEEIERLLRCKFIRAAKYVEWLANVVPVIKKNGKMR 649
+KED K +K RRF D IKEE+ +LL FI+ + +WLAN V V KK G+ R
Sbjct: 992 HVKEDAKPIKQRLRRFAQDRKDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWR 1051
Query: 650 VCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAKEDVSKTAFR 709
+C+D+ DLN + PKD + +P + +VDS AG E LS LD YSGY+QI + + D KT+F
Sbjct: 1052 MCVDYTDLNKSCPKDPFGLPRIDQVVDSTAGRELLSFLDCYSGYHQIRLKESDCLKTSFI 1111
Query: 710 CSGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDLVVKSPSRDEHLSH 769
G Y +V MPFGLKNAGATYQR++ F I ++ Y+DD+VVK+ +D+ +S
Sbjct: 1112 TP--FGAYCYVTMPFGLKNAGATYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISD 1169
Query: 770 LRKSFERMRIHGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQ 829
L ++F +R +K+NP KC FGV +G +GF+V +GI+ N K AIL+ PP+++K
Sbjct: 1170 LEETFASIRAFRMKLNPEKCTFGVPSGKLVGFMVSHRGIQANPEKVTAILNMKPPSTQKD 1229
Query: 830 LQSLLGKVNFLRRFIDNLSDKTKPFSSLLRLKREDIFRWEA*HQKAFDELKNYLAIPPVM 889
+Q L G + L RF+ L ++ PF LL K+ D F+W QKAF++ K L PPV+
Sbjct: 1230 VQKLTGCMAALSRFVSRLGERGMPFFKLL--KKTDNFQWGPEAQKAFEDFKKLLTEPPVL 1287
Query: 890 IPPIKGKPMRLYISATDETIGSMLAQE-DEDG----IERAVFYLSRVLNDAKTRYTMIEK 944
P +P+ LY+SAT + + ++L E +E+G ++R ++++S VL D+KTRY ++K
Sbjct: 1288 ASPHPQEPLLLYVSATSQVVSTVLVVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQK 1347
Query: 945 LCLCLYFSCTKLKYYIKPIDVMVFSHFDIIKHMLSKPILHSRIGKWSLALTEYSLTYAPL 1004
L + + KL +Y + V V + F + +L + RI KW+L L +++ P
Sbjct: 1348 LLYGILITTRKLSHYFQGHSVTVVTSFPL-GDILHNREANGRIAKWALELMSLDISFKPR 1406
Query: 1005 KTIKGQAVADFLADHT-LPEEIAYVGLQPWKLFFDDSSHKEGSGIGMFIVSPQGIPTKFM 1063
+IK QA+ADF+A+ T E+ ++ W + FD S G+G G+ ++SP G ++
Sbjct: 1407 ISIKSQALADFVAEWTECQEDTPAENMEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYV 1466
Query: 1064 FRIRESCSNNESEYEALVSGLEILLALGAKNVEVKGDSELVVKQLTKEYKCISKNLAKYY 1123
I S S+N +EYEAL+ GL I ++LG K + V+GDS+LVV Q+ KE+ C+ N+ Y
Sbjct: 1467 LWIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYR 1526
Query: 1124 VKAMSLLANFDQAGVSYIPRVSNQEANELAQIASGYMIDKQKLKELIRIKEKLSPLDLEV 1183
+ L FD +S++ R +N+ A+ LA S K +++P D+ V
Sbjct: 1527 QEVRKLEDKFDGLELSHVLRHNNEAADRLANFGS---------------KREVAPSDVFV 1571
Query: 1184 MVIDNLT------------------PNDWRKPIVEYLQNPVGSTDI----KVKYRALNYT 1221
+ T DWR+P++ +L + D ++ R+ Y
Sbjct: 1572 EHLYTPTVPHKDTTQVAGTHDVAMVETDWREPLIRFLTSQELPQDKDEAERISRRSKLYV 1631
Query: 1222 IIGNELFKKNVDGTLLKCLSEVDAFIAVSAAHGGLCGAHQAGAKMKWILFRQGMYWPTIM 1281
+ EL+KK+ G L +C+S + + H G+CG H A + +RQG +WPT +
Sbjct: 1632 MHEAELYKKSPSGILQRCVSLEEGRQLLQDIHSGICGNHAAARTIVGKAYRQGFFWPTAV 1691
Query: 1282 KDCIEYAKGCQDCQKHSGIQHVPASELHSIIKSWPFRGWAIDLIGEIHPALSRQHKYIIV 1341
D + + C+ CQ + H+PA EL +I SWPF W +D++G A+ + ++ V
Sbjct: 1692 SDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGLDMVGPFKKAVG-GYTHLFV 1750
Query: 1342 AIDYFIKWVEAIPLQSVTQETVIEFIQNHIVYRFGLPESLTTDQGTMFVGRKVAAFAESW 1401
AID F KW+EA P+ ++T + +F N IV+RFG+P + TD GT F G F E +
Sbjct: 1751 AIDKFSKWIEAKPVVTITADNARDFFIN-IVHRFGVPNRIITDNGTQFTGGVFKDFCEDF 1809
Query: 1402 GIKLLTSIPYYAQANGQVEAANKILISLIKKHVGRKPK----SWHESLSQVLWAYRNSPR 1457
GIK+ + + +NGQVE AN +++ IK V + K W + L VLW+ R +P
Sbjct: 1810 GIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYAGKWVQQLPSVLWSLRTTPS 1869
Query: 1458 EATGTTPFRLAYGQEAVLPAEVYLQSFRIQRQEDIPSEVYWNMMLDELVNLDEERVLALD 1517
ATG +PF L YG EA+LP+EV +S R + + E Y +D+L L+E R AL
Sbjct: 1870 RATGQSPFFLVYGAEAMLPSEVEFESLRFR---NFREERYEEDRVDDLHRLEEVREAALI 1926
Query: 1518 VLTRQKDRIAKAYNKKVKNRSFVTGDYVWKVILPTDKKDRAYGKWAPNWEGPFKVKKVLS 1577
R + + +N+ V++R+F+ GD V + I T +DR K +P WEGPF + +V
Sbjct: 1927 RSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTT--RDR--HKLSPLWEGPFIISEVTR 1982
Query: 1578 NNAYVIKELSGQRQFVTINDKYLKAY 1603
+Y +K G + N ++L+ +
Sbjct: 1983 PGSYRLKREDGTLVDNSWNIEHLRRF 2008
>UniRef100_Q7X6L5 OSJNBb0093G06.3 protein [Oryza sativa]
Length = 1986
Score = 683 bits (1763), Expect = 0.0
Identities = 426/1255 (33%), Positives = 680/1255 (53%), Gaps = 67/1255 (5%)
Query: 374 PLFVWAKVEDKGVNKILVDGGAAINLMPRFMMKRLGKTETDLIPHDMVLSDY-EGKTSTS 432
PL + V + + + L+DGG+A+N++ + + ++L P + G ++T
Sbjct: 772 PLVLDPVVRNVKLRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATP 831
Query: 433 MGAIMLNITAGTV----SRSTLFIVVPSKANYNLLLGREWIHGVGAVPSTLHQRISIWKP 488
+G I L +T GT + + F V + Y+ +LGR + AVP + + + P
Sbjct: 832 LGQITLPVTFGTRENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKMPGP 891
Query: 489 DGVVENVQADQSYYLAEAGYVDKKNFEKSPLVDATKMEAQDPLEKVDLGDGSKKRPTYIS 548
GV+ ++++D + +A DK++ + A E E++ L ++
Sbjct: 892 RGVL-SLRSD----IKQAVTCDKESCDM-----AQTREMASAREEIRLAATTESA----- 936
Query: 549 SLIDPELRGRMIELLKEFKDCFA*DNDEMPGLSRDLVELQLPIKEDKKLVK*LPRRFHPD 608
+I L+ KD FA +MPG+ R+++E L +KED K +K RRF D
Sbjct: 937 ----------LITFLQNNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQD 986
Query: 609 VLVKIKEEIERLLRCKFIRAAKYVEWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHM 668
IKEE+ +LL FI+ + +WLAN V V KK G+ R+C+D+ DLN + PKD + +
Sbjct: 987 RKDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGL 1046
Query: 669 PIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAKEDVSKTAFRCSGALGTYEWVVMPFGLKN 728
P + +VDS AG E LS LD YSGY+QI + + D KT+F G Y +V MPFGLKN
Sbjct: 1047 PRIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITP--FGAYCYVTMPFGLKN 1104
Query: 729 AGATYQRVMNTIFHDFIETFMQVYIDDLVVKSPSRDEHLSHLRKSFERMRIHGLKMNPLK 788
AGATYQR++ F I ++ Y+DD+VVK+ +D+ +S L ++F +R +K+NP K
Sbjct: 1105 AGATYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEK 1164
Query: 789 CAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKVNFLRRFIDNLS 848
C FGV +G LGF+V +GI+ N K AIL+ PP+++K +Q L G + L RF+ L
Sbjct: 1165 CTFGVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLG 1224
Query: 849 DKTKPFSSLLRLKREDIFRWEA*HQKAFDELKNYLAIPPVMIPPIKGKPMRLYISATDET 908
++ PF L LK+ D F+W QKAF++ K L PPV+ P +P+ LY+SAT +
Sbjct: 1225 ERGMPFFKL--LKKTDNFQWGPEAQKAFEDFKQLLTKPPVLASPHPQEPLLLYVSATSQV 1282
Query: 909 IGSMLAQE-DEDG----IERAVFYLSRVLNDAKTRYTMIEKLCLCLYFSCTKLKYYIKPI 963
+ ++L E +E+G ++R ++++S VL D+KTRY ++KL + + KL +Y +
Sbjct: 1283 VSTVLVVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGH 1342
Query: 964 DVMVFSHFDIIKHMLSKPILHSRIGKWSLALTEYSLTYAPLKTIKGQAVADFLADHT-LP 1022
V V + F + +L + RI KW+L L +++ P +IK QA+ADF+A+ T
Sbjct: 1343 SVTVVTSFP-LGDVLHNREANGRIAKWALELMSLDISFKPRTSIKSQALADFVAEWTECQ 1401
Query: 1023 EEIAYVGLQPWKLFFDDSSHKEGSGIGMFIVSPQGIPTKFMFRIRESCSNNESEYEALVS 1082
E+ ++ W + FD S G+G G+ ++SP G ++ I S S+N +EYEAL+
Sbjct: 1402 EDTPVEKIEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLH 1461
Query: 1083 GLEILLALGAKNVEVKGDSELVVKQLTKEYKCISKNLAKYYVKAMSLLANFDQAGVSYIP 1142
GL I ++LG K + V+GDS+LVV Q+ KE+ CI N+ Y + L FD +S++
Sbjct: 1462 GLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCIDDNMMAYRQEVRKLEDKFDGLELSHVL 1521
Query: 1143 RVSNQEANELA------QIASGYMIDKQKLKELIRIKEKLSPLDLEVMVIDNLTPNDWRK 1196
R +N+ A+ LA + A + + + K+ D +V + DWR+
Sbjct: 1522 RHNNEAADRLANFGSKREAAPSDVFVEHLYTPTVPHKDTTQDADTHDVV---MVEADWRE 1578
Query: 1197 PIVEYLQNPVGSTD----IKVKYRALNYTIIGNELFKKNVDGTLLKCLSEVDAFIAVSAA 1252
P + +L + D ++ R+ Y + +EL+KK+ G L +C+S + +
Sbjct: 1579 PFIRFLSSQELPQDKDEAERISRRSKLYVMHESELYKKSPSGILQRCVSLEEGRQLLKDI 1638
Query: 1253 HGGLCGAHQAGAKMKWILFRQGMYWPTIMKDCIEYAKGCQDCQKHSGIQHVPASELHSII 1312
H G+CG H A + +RQG +WPT + D + + C+ CQ + H+PA EL +I
Sbjct: 1639 HSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIP 1698
Query: 1313 KSWPFRGWAIDLIGEIHPALSRQHKYIIVAIDYFIKWVEAIPLQSVTQETVIEFIQNHIV 1372
SWPF W +D++G A+ + ++ VAID F KW+EA P+ ++T + +F N IV
Sbjct: 1699 LSWPFAVWGLDMVGPFKKAVG-GYTHLFVAIDKFSKWIEAKPVVTITADNARDFFIN-IV 1756
Query: 1373 YRFGLPESLTTDQGTMFVGRKVAAFAESWGIKLLTSIPYYAQANGQVEAANKILISLIKK 1432
+RFG+P + TD GT F G F E +GIK+ + + +NGQVE AN +++ IK
Sbjct: 1757 HRFGVPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKA 1816
Query: 1433 HVGRKPK----SWHESLSQVLWAYRNSPREATGTTPFRLAYGQEAVLPAEVYLQSFRIQR 1488
V + K W + L VLW+ R +P ATG +PF L YG EA+LP+EV +S R +
Sbjct: 1817 RVFDRLKPYAGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFR- 1875
Query: 1489 QEDIPSEVYWNMMLDELVNLDEERVLALDVLTRQKDRIAKAYNKKVKNRSFVTGDYVWKV 1548
+ E Y +D+L L+E R AL R + + +N+ V++R+F+ GD V +
Sbjct: 1876 --NFREERYEEDRVDDLHRLEEAREAALIQSARYLQGLRRYHNRNVRSRAFLVGDLVLRK 1933
Query: 1549 ILPTDKKDRAYGKWAPNWEGPFKVKKVLSNNAYVIKELSGQRQFVTINDKYLKAY 1603
I T +DR K +P WEGPF + +V +Y +K G + N ++L+ +
Sbjct: 1934 IQTT--RDR--HKLSPLWEGPFIISEVTRPGSYRLKREDGTLVDNSWNIEHLRRF 1984
>UniRef100_Q8LMM4 Putative gag-pol [Oryza sativa]
Length = 2017
Score = 682 bits (1759), Expect = 0.0
Identities = 428/1267 (33%), Positives = 683/1267 (53%), Gaps = 65/1267 (5%)
Query: 374 PLFVWAKVEDKGVNKILVDGGAAINLMPRFMMKRLGKTETDLIPHDMVLSDY-EGKTSTS 432
PL + V + + + L+DGG+A+N++ + + ++L P + G ++T
Sbjct: 777 PLVLDPVVRNVKLRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATP 836
Query: 433 MGAIMLNITAGTV----SRSTLFIVVPSKANYNLLLGREWIHGVGAVPSTLHQRISIWKP 488
+G I L +T GT + + F V + Y+ +LGR + AVP + + + P
Sbjct: 837 LGQITLPVTFGTRENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKMPGP 896
Query: 489 DGVVENVQADQSYYLAEAGYVDKKNFEKSP------------LVDATKMEAQDPLEKVDL 536
GV+ ++++D + +A DK++ + + L AT E + P K+
Sbjct: 897 RGVL-SLRSD----IKQAVTCDKESCDMAQTREMASAREDIRLAAATASEGEVPATKISK 951
Query: 537 GDGSKKRPTYISSLIDPELRGRMIELLKEFKDCFA*DNDEMPGLSRDLVELQLPIKEDKK 596
S+ + I +DP + KD FA +MPG+ R+++E L +KED K
Sbjct: 952 SGESEAKTKKIP--LDPSDPTKTANN----KDIFAWKPSDMPGIPREVIEHSLHVKEDAK 1005
Query: 597 LVK*LPRRFHPDVLVKIKEEIERLLRCKFIRAAKYVEWLANVVPVIKKNGKMRVCIDFRD 656
+K RRF D IKEE+ +LL FI+ + +WLAN V V KK G+ R+C+D+ D
Sbjct: 1006 PIKQRLRRFAQDRKDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTD 1065
Query: 657 LNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAKEDVSKTAFRCSGALGT 716
LN + PKD + +P + +VDS AG E LS LD YSGY+QI + + D KT+F G
Sbjct: 1066 LNKSCPKDPFGLPRIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITP--FGA 1123
Query: 717 YEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDLVVKSPSRDEHLSHLRKSFER 776
Y +V MPFGLKNAGATYQR++ F I ++ Y+DD+VVK+ +D+ + L ++F
Sbjct: 1124 YCYVTMPFGLKNAGATYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLILDLEETFAS 1183
Query: 777 MRIHGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGK 836
+R +K+NP KC FGV +G LGF+V +GI+ N K AIL+ PP+++K +Q L G
Sbjct: 1184 IRAFRMKLNPEKCTFGVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGC 1243
Query: 837 VNFLRRFIDNLSDKTKPFSSLLRLKREDIFRWEA*HQKAFDELKNYLAIPPVMIPPIKGK 896
+ L RF+ L ++ PF L LK+ D F+W QKAF++ K L PPV+ P +
Sbjct: 1244 MAALSRFVSRLGERGMPFFKL--LKKTDDFQWGPEAQKAFEDFKKLLTEPPVLASPHPQE 1301
Query: 897 PMRLYISATDETIGSMLAQE-DEDG----IERAVFYLSRVLNDAKTRYTMIEKLCLCLYF 951
P+ LY+SAT + + ++L E +E+G ++R ++++S VL D+KTRY ++KL +
Sbjct: 1302 PLLLYVSATSQVVSTVLVVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILI 1361
Query: 952 SCTKLKYYIKPIDVMVFSHFDIIKHMLSKPILHSRIGKWSLALTEYSLTYAPLKTIKGQA 1011
+ KL +Y + V V + F + +L + RI KW+L L L++ P +IK QA
Sbjct: 1362 TTRKLSHYFQGHSVTVVTSFP-LGDILHNREANGRIAKWALELMSLDLSFKPRISIKSQA 1420
Query: 1012 VADFLADHT-LPEEIAYVGLQPWKLFFDDSSHKEGSGIGMFIVSPQGIPTKFMFRIRESC 1070
+ADF+A+ T E+ ++ W + FD S G+G G+ ++SP G ++ I S
Sbjct: 1421 LADFVAEWTECQEDTTVKKMEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSA 1480
Query: 1071 SNNESEYEALVSGLEILLALGAKNVEVKGDSELVVKQLTKEYKCISKNLAKYYVKAMSLL 1130
S+N +EYEAL+ GL I ++LG K + V+GDS+LVV Q+ KE+ C+ N+ Y + L
Sbjct: 1481 SHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMKAYRQEVRKLE 1540
Query: 1131 ANFDQAGVSYIPRVSNQEANELA------QIASGYMIDKQKLKELIRIKEKLSPLDLEVM 1184
FD +S++ R N+ A+ LA ++A + + + K+ + +
Sbjct: 1541 DKFDGLELSHVLRHDNEAADRLANFGSKREVAPSDVFVEHLYTPTVPHKDTTQAAGIHDV 1600
Query: 1185 VIDNLTPNDWRKPIVEYLQNPVGSTD----IKVKYRALNYTIIGNELFKKNVDGTLLKCL 1240
+ DWR+P++ +L + D ++ R+ Y + EL+KK+ G L +C+
Sbjct: 1601 A---MVETDWREPLIRFLTSQELPQDKDEAERISRRSKLYVMHEAELYKKSPSGILQRCV 1657
Query: 1241 SEVDAFIAVSAAHGGLCGAHQAGAKMKWILFRQGMYWPTIMKDCIEYAKGCQDCQKHSGI 1300
S + + H G+CG H A + +RQG +WPT + D + + C+ CQ +
Sbjct: 1658 SLEEGRQLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQ 1717
Query: 1301 QHVPASELHSIIKSWPFRGWAIDLIGEIHPALSRQHKYIIVAIDYFIKWVEAIPLQSVTQ 1360
H+PA EL +I SWPF W +D++G A+ + ++ VAID F KW+EA P+ ++T
Sbjct: 1718 IHLPAQELQTIPLSWPFAVWGLDMVGPFKKAVG-GYTHLFVAIDKFSKWIEAKPVVTITA 1776
Query: 1361 ETVIEFIQNHIVYRFGLPESLTTDQGTMFVGRKVAAFAESWGIKLLTSIPYYAQANGQVE 1420
+ +F N IV+RFG+P + TD GT F G F E +GIK+ + + +NGQVE
Sbjct: 1777 DNARDFFIN-IVHRFGVPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVE 1835
Query: 1421 AANKILISLIKKHVGRKPK----SWHESLSQVLWAYRNSPREATGTTPFRLAYGQEAVLP 1476
AN +++ IK V + K W + L VLW+ R +P ATG +PF L YG EA+LP
Sbjct: 1836 RANGMILQGIKARVFDRLKPYAGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLP 1895
Query: 1477 AEVYLQSFRIQRQEDIPSEVYWNMMLDELVNLDEERVLALDVLTRQKDRIAKAYNKKVKN 1536
+EV +S R + + E Y +D+L L+E R AL R + + +N+ V++
Sbjct: 1896 SEVEFESLRFR---NFREERYEEDRVDDLHRLEEVREAALIRSARYLQGLRRYHNRNVRS 1952
Query: 1537 RSFVTGDYVWKVILPTDKKDRAYGKWAPNWEGPFKVKKVLSNNAYVIKELSGQRQFVTIN 1596
R+F+ GD V + I T +DR K +P WEGPF + +V +Y +K G + N
Sbjct: 1953 RAFLVGDLVLRKIQTT--RDR--HKLSPLWEGPFIISEVTRPGSYRLKREDGTLVDNSWN 2008
Query: 1597 DKYLKAY 1603
+YL+ +
Sbjct: 2009 IEYLRRF 2015
>UniRef100_Q7X8E7 OSJNBa0042F21.5 protein [Oryza sativa]
Length = 1950
Score = 681 bits (1756), Expect = 0.0
Identities = 428/1267 (33%), Positives = 683/1267 (53%), Gaps = 65/1267 (5%)
Query: 374 PLFVWAKVEDKGVNKILVDGGAAINLMPRFMMKRLGKTETDLIPHDMVLSDY-EGKTSTS 432
PL + V + + + L+DGG+A+N++ + + ++L P + G ++T
Sbjct: 710 PLVLDPVVRNVKLRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATP 769
Query: 433 MGAIMLNITAGTV----SRSTLFIVVPSKANYNLLLGREWIHGVGAVPSTLHQRISIWKP 488
+G I L +T GT + + F V + Y+ +LGR + AVP + + + P
Sbjct: 770 LGQITLPVTFGTRENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKMPGP 829
Query: 489 DGVVENVQADQSYYLAEAGYVDKKNFEKSP------------LVDATKMEAQDPLEKVDL 536
GV+ ++++D + +A DK++ + + L AT E + P K
Sbjct: 830 RGVL-SLRSD----IKQAVTCDKESCDMAQTREMASAREDIRLAAATASEGEIPATKTSK 884
Query: 537 GDGSKKRPTYISSLIDPELRGRMIELLKEFKDCFA*DNDEMPGLSRDLVELQLPIKEDKK 596
S+ + I +DP + KD FA +MPG+ R+++E L +KED K
Sbjct: 885 SGESEAKTKKIP--LDPSDPTKTANN----KDIFAWKPSDMPGIPREVIEHSLHVKEDAK 938
Query: 597 LVK*LPRRFHPDVLVKIKEEIERLLRCKFIRAAKYVEWLANVVPVIKKNGKMRVCIDFRD 656
+K RRF D IKEE+ +LL FI+ + +WLAN V V KK G+ R+C+D+ D
Sbjct: 939 PIKQRLRRFAQDRKDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTD 998
Query: 657 LNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAKEDVSKTAFRCSGALGT 716
LN + PKD + +P + +VDS AG E LS LD YSGY+QI + + D KT+F G
Sbjct: 999 LNKSCPKDPFGLPRIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITP--FGA 1056
Query: 717 YEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDLVVKSPSRDEHLSHLRKSFER 776
Y +V MPFGLKNAGATYQR++ F I ++ Y+DD+VVK+ +D+ +S L ++F
Sbjct: 1057 YCYVTMPFGLKNAGATYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFAS 1116
Query: 777 MRIHGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGK 836
+R +K+NP KC FGV +G LGF+V +GI+ N K AIL+ PP+++K +Q L G
Sbjct: 1117 IRAFRMKLNPEKCTFGVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGC 1176
Query: 837 VNFLRRFIDNLSDKTKPFSSLLRLKREDIFRWEA*HQKAFDELKNYLAIPPVMIPPIKGK 896
+ L RF+ L ++ PF L LK+ D F+W QKAF++ K L PP++ P +
Sbjct: 1177 MAALSRFVSRLGERGMPFFKL--LKKTDNFQWGPEAQKAFEDFKKLLTEPPILASPHPQE 1234
Query: 897 PMRLYISATDETIGSMLAQE-DEDG----IERAVFYLSRVLNDAKTRYTMIEKLCLCLYF 951
P+ LY+SAT + + ++L E +E+G ++R ++++S VL D+KTRY ++KL +
Sbjct: 1235 PLLLYVSATSQVVSTVLVVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILI 1294
Query: 952 SCTKLKYYIKPIDVMVFSHFDIIKHMLSKPILHSRIGKWSLALTEYSLTYAPLKTIKGQA 1011
+ KL +Y + V V + F + +L + RI KW+L L +++ P +IK QA
Sbjct: 1295 TTRKLSHYFQGHSVTVVTSFP-LGDILHNREANGRIAKWALELMSLDISFKPRISIKSQA 1353
Query: 1012 VADFLADHT-LPEEIAYVGLQPWKLFFDDSSHKEGSGIGMFIVSPQGIPTKFMFRIRESC 1070
+ADF+A+ T E+ ++ W + FD S G+G G+ ++SP G ++ I S
Sbjct: 1354 LADFVAEWTECQEDTPAENMEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSA 1413
Query: 1071 SNNESEYEALVSGLEILLALGAKNVEVKGDSELVVKQLTKEYKCISKNLAKYYVKAMSLL 1130
S+N +EYEAL+ GL I ++LG K + V+GDS+LVV Q+ KE+ C+ N+ Y + L
Sbjct: 1414 SHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLE 1473
Query: 1131 ANFDQAGVSYIPRVSNQEANELA------QIASGYMIDKQKLKELIRIKEKLSPLDLEVM 1184
FD +S++ R +N+ A+ LA ++A + + + K+ D +
Sbjct: 1474 DKFDGLELSHVLRHNNEAADRLANFGSKREMAPSDVFVEHLYTPTVPHKDTTQDADTHDV 1533
Query: 1185 VIDNLTPNDWRKPIVEYLQNPVGSTD----IKVKYRALNYTIIGNELFKKNVDGTLLKCL 1240
L DWR+P + +L + D ++ R+ Y + EL+KK+ G L +C+
Sbjct: 1534 A---LVEADWREPFIRFLTSQELPQDKDEAERISRRSKLYAMHEAELYKKSPSGILQRCV 1590
Query: 1241 SEVDAFIAVSAAHGGLCGAHQAGAKMKWILFRQGMYWPTIMKDCIEYAKGCQDCQKHSGI 1300
S + + H G+CG H A + +RQG +WPT + D + + C+ CQ +
Sbjct: 1591 SLEEGRQLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQ 1650
Query: 1301 QHVPASELHSIIKSWPFRGWAIDLIGEIHPALSRQHKYIIVAIDYFIKWVEAIPLQSVTQ 1360
H+PA EL +I SWPF W +D++G A+ + ++ VAID F KW+EA P+ ++T
Sbjct: 1651 IHLPAQELQTIPLSWPFAVWGLDMVGPFKKAVG-GYTHLFVAIDKFSKWIEAKPVVTITA 1709
Query: 1361 ETVIEFIQNHIVYRFGLPESLTTDQGTMFVGRKVAAFAESWGIKLLTSIPYYAQANGQVE 1420
+ +F N IV+RFG+P + TD GT F G F E +GIK+ + + +NGQVE
Sbjct: 1710 DNARDFFIN-IVHRFGVPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVE 1768
Query: 1421 AANKILISLIKKHVGRKPK----SWHESLSQVLWAYRNSPREATGTTPFRLAYGQEAVLP 1476
AN +++ IK V + K W + L VLW+ R +P ATG +PF L YG EA+LP
Sbjct: 1769 RANGMILQGIKARVFDRLKPYAGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLP 1828
Query: 1477 AEVYLQSFRIQRQEDIPSEVYWNMMLDELVNLDEERVLALDVLTRQKDRIAKAYNKKVKN 1536
+EV +S R + + E Y +D+L L+E R AL R + + +N+ V++
Sbjct: 1829 SEVEFESLRFR---NFREERYEEDRVDDLHRLEEAREAALIRSARYLQGLRRYHNRNVRS 1885
Query: 1537 RSFVTGDYVWKVILPTDKKDRAYGKWAPNWEGPFKVKKVLSNNAYVIKELSGQRQFVTIN 1596
R+F+ GD V + I T +DR K +P WEGPF + +V +Y +K G + N
Sbjct: 1886 RAFLVGDLVLRKIQTT--RDR--HKLSPLWEGPFIISEVTRPGSYRLKREDGTLVDNSWN 1941
Query: 1597 DKYLKAY 1603
++L+ +
Sbjct: 1942 IEHLRRF 1948
>UniRef100_Q8LMM6 Putative gag-pol [Oryza sativa]
Length = 1986
Score = 676 bits (1745), Expect = 0.0
Identities = 428/1264 (33%), Positives = 679/1264 (52%), Gaps = 85/1264 (6%)
Query: 374 PLFVWAKVEDKGVNKILVDGGAAINLMPRFMMKRLGKTETDL-IPHDMVLSD-------Y 425
PL + V + + + L+DGG+A+N++ KT D+ IP + S
Sbjct: 772 PLVLDPVVRNVKLRRTLIDGGSALNIL-------FAKTLDDMQIPRSELKSSNAPFHGVI 824
Query: 426 EGKTSTSMGAIMLNITAGTV----SRSTLFIVVPSKANYNLLLGREWIHGVGAVPSTLHQ 481
G ++T +G I L +T GT + + F V + Y+ +LG + AVP +
Sbjct: 825 PGLSATPLGQITLPVTFGTRENFRTENISFEVADFETAYHAILGHPALAKFMAVPHYTYM 884
Query: 482 RISIWKPDGVVENVQADQSYYLAEAGYVDKKNFEKSPLVDATKMEAQDPLEKVDLGDGSK 541
+ + P GV+ +++D + +A DK++ + A E E++ L ++
Sbjct: 885 MMKMPGPRGVL-TLRSD----IKQAVTCDKESCDM-----AQTREIASAREEIRLAATTE 934
Query: 542 KRPTYISSLIDPELRGRMIELLKEFKDCFA*DNDEMPGLSRDLVELQLPIKEDKKLVK*L 601
+I L+ KD FA +MPG+ R+++E L +KED K +K
Sbjct: 935 SA---------------LITFLQNNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQR 979
Query: 602 PRRFHPDVLVKIKEEIERLLRCKFIRAAKYVEWLANVVPVIKKNGKMRVCIDFRDLNAAT 661
RRF D IKEE+ +LL FI+ + +WLAN V V KK G+ R+C+D+ DLN +
Sbjct: 980 LRRFAQDRKDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSC 1039
Query: 662 PKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAKEDVSKTAFRCSGALGTYEWVV 721
PKD + +P + +VDS AG E LS LD YSGY+QI + + D KT+F G Y +V
Sbjct: 1040 PKDPFGLPRIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITP--FGAYCYVT 1097
Query: 722 MPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDLVVKSPSRDEHLSHLRKSFERMRIHG 781
MPFGLKNAGATYQR++ F I ++ Y+DD+VVK+ +D+ +S L ++F +R
Sbjct: 1098 MPFGLKNAGATYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFR 1157
Query: 782 LKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKVNFLR 841
+K+NP KC FGV +G LGF+V +GI+ N K AIL+ PP+++K +Q L G + L
Sbjct: 1158 MKLNPEKCTFGVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALS 1217
Query: 842 RFIDNLSDKTKPFSSLLRLKREDIFRWEA*HQKAFDELKNYLAIPPVMIPPIKGKPMRLY 901
RF+ L ++ PF L LK+ D F+W QKAF++ K L PPV+ P +P+ LY
Sbjct: 1218 RFVSRLGERGMPFFKL--LKKTDNFQWGPEAQKAFEDFKQLLTKPPVLASPHPQEPLLLY 1275
Query: 902 ISATDETIGSMLAQE-DEDG----IERAVFYLSRVLNDAKTRYTMIEKLCLCLYFSCTKL 956
+SAT + + ++L E +E+G ++R ++++S VL D+K RY ++KL + + KL
Sbjct: 1276 VSATSQVVSTVLVVEREEEGHVQKVQRPIYFVSEVLADSKARYPQVQKLLYGILITTRKL 1335
Query: 957 KYYIKPIDVMVFSHFDIIKHMLSKPILHSRIGKWSLALTEYSLTYAPLKTIKGQAVADFL 1016
+Y + V V + F + +L + RI KW+L L +++ P +IK QA+ADF+
Sbjct: 1336 SHYFQGHSVTVVTSFP-LGDVLHNREANGRIAKWALELMSLDISFKPRTSIKSQALADFV 1394
Query: 1017 ADHT-LPEEIAYVGLQPWKLFFDDSSHKEGSGIGMFIVSPQGIPTKFMFRIRESCSNNES 1075
A+ T E+ ++ W + FD S G+G G+ ++SP G ++ I S S+N +
Sbjct: 1395 AEWTECQEDTPVEKIEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVA 1454
Query: 1076 EYEALVSGLEILLALGAKNVEVKGDSELVVKQLTKEYKCISKNLAKYYVKAMSLLANFDQ 1135
EYEAL+ GL I ++LG K + V+GDS+LVV Q+ KE+ CI N+ Y + L FD
Sbjct: 1455 EYEALLHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCIDDNMMAYRQEVRKLEDKFDG 1514
Query: 1136 AGVSYIPRVSNQEANELAQIASGYMIDKQKLKELIRIKEKLSPL--------DLEVMVID 1187
+S++ R +N+ A+ LA S ++ + ++ +P D + I
Sbjct: 1515 LELSHVLRHNNEAADRLANFGS----KRETAPSDVFVEHLYTPTVPHKDTTQDADTRNI- 1569
Query: 1188 NLTPNDWRKPIVEYLQNPVGSTD----IKVKYRALNYTIIGNELFKKNVDGTLLKCLSEV 1243
+ DWR+P + +L + D ++ R+ Y + +EL+KK+ G L +C+S
Sbjct: 1570 AMVEADWREPFIRFLTSQELPQDKDEAERISRRSKLYVLHESELYKKSPSGILQRCVSLE 1629
Query: 1244 DAFIAVSAAHGGLCGAHQAGAKMKWILFRQGMYWPTIMKDCIEYAKGCQDCQKHSGIQHV 1303
+ + H G+CG H A + +RQG +WPT + D + + C+ CQ + H+
Sbjct: 1630 EGRQLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHL 1689
Query: 1304 PASELHSIIKSWPFRGWAIDLIGEIHPALSRQHKYIIVAIDYFIKWVEAIPLQSVTQETV 1363
PA EL +I SWPF W +D++G A+ + ++ VAID F KW+EA P+ ++T +
Sbjct: 1690 PAQELQTIPLSWPFAVWGLDMVGPFKKAVG-GYTHLFVAIDKFSKWIEAKPVVTITADNA 1748
Query: 1364 IEFIQNHIVYRFGLPESLTTDQGTMFVGRKVAAFAESWGIKLLTSIPYYAQANGQVEAAN 1423
+F N IV+RFG+P + TD GT F G F E +GIK+ + + +NGQVE AN
Sbjct: 1749 RDFFIN-IVHRFGVPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERAN 1807
Query: 1424 KILISLIKKHVGRKPK----SWHESLSQVLWAYRNSPREATGTTPFRLAYGQEAVLPAEV 1479
+++ IK V + K W + L VLW+ R +P ATG +PF L YG EA+LP+EV
Sbjct: 1808 GMILQGIKARVFDRLKPYAGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEV 1867
Query: 1480 YLQSFRIQRQEDIPSEVYWNMMLDELVNLDEERVLALDVLTRQKDRIAKAYNKKVKNRSF 1539
+S R + + E Y +D+L L+E R AL R + + +N+ V++R+F
Sbjct: 1868 EFESLRFR---NFREERYEEDRVDDLHRLEEVREAALIQSARYLQGLRRYHNRNVRSRAF 1924
Query: 1540 VTGDYVWKVILPTDKKDRAYGKWAPNWEGPFKVKKVLSNNAYVIKELSGQRQFVTINDKY 1599
+ GD V + I T +DR K +P WEGPF + +V +Y +K G + N ++
Sbjct: 1925 LVGDLVLRKIQTT--RDR--HKLSPLWEGPFIISEVTRPGSYRLKREDGTLVDNSWNIEH 1980
Query: 1600 LKAY 1603
L+ +
Sbjct: 1981 LRRF 1984
>UniRef100_Q8S7A3 Putative retroelement [Oryza sativa]
Length = 2017
Score = 676 bits (1745), Expect = 0.0
Identities = 426/1252 (34%), Positives = 673/1252 (53%), Gaps = 65/1252 (5%)
Query: 374 PLFVWAKVEDKGVNKILVDGGAAINLMPRFMMKRLGKTETDLIPHDMVLSDY-EGKTSTS 432
PL + V + + + L+DGG+A+N++ + + ++L P + G ++T
Sbjct: 777 PLVLDPVVRNVKLRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATP 836
Query: 433 MGAIMLNITAGTV----SRSTLFIVVPSKANYNLLLGREWIHGVGAVPSTLHQRISIWKP 488
+G I L +T GT + + F V + Y+ +LGR + AVP + + + P
Sbjct: 837 LGQITLPVTFGTRENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKLPGP 896
Query: 489 DGVVENVQADQSYYLAEAGYVDKKNFEKSP------------LVDATKMEAQDPLEKVDL 536
GV+ ++++D + +A DK++ + + L AT E + P K+
Sbjct: 897 RGVL-SLRSD----IKQAVTCDKESCDMAQTREMASAREDIRLAAATASEGEVPATKISK 951
Query: 537 GDGSKKRPTYISSLIDPELRGRMIELLKEFKDCFA*DNDEMPGLSRDLVELQLPIKEDKK 596
S+ + I +DP + KD FA +MPG+ R+++E L +KED K
Sbjct: 952 SGESEAKTKKIP--LDPSDSTKTANN----KDIFAWKPSDMPGIPREVIEHSLHVKEDAK 1005
Query: 597 LVK*LPRRFHPDVLVKIKEEIERLLRCKFIRAAKYVEWLANVVPVIKKNGKMRVCIDFRD 656
+K RRF D IKEE+ +LL FI+ + +WLAN V V KK G+ R+C+D+ D
Sbjct: 1006 PIKQRLRRFAQDRKDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTD 1065
Query: 657 LNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAKEDVSKTAFRCSGALGT 716
LN PKD + +P + +VDS AG E LS LD YSGY+QI + + D KT+F G
Sbjct: 1066 LNKCCPKDPFGLPRIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITP--FGA 1123
Query: 717 YEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDLVVKSPSRDEHLSHLRKSFER 776
Y +V MPFGLKNAGATYQR++ F I ++ Y+DD+VVK+ +D+ +S L ++F
Sbjct: 1124 YCYVTMPFGLKNAGATYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFAS 1183
Query: 777 MRIHGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGK 836
+R +K+NP KC FGV +G LGF+V +GI+ N K AIL+ PP+++K +Q L G
Sbjct: 1184 IRAFRMKLNPEKCTFGVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGC 1243
Query: 837 VNFLRRFIDNLSDKTKPFSSLLRLKREDIFRWEA*HQKAFDELKNYLAIPPVMIPPIKGK 896
+ L RF+ L ++ PF LL K+ D F W QKAF++ K L PPV+ P +
Sbjct: 1244 MAALSRFVSRLGERGMPFFKLL--KKTDDFHWGPEAQKAFEDFKKLLTEPPVLASPHPQE 1301
Query: 897 PMRLYISATDETIGSMLAQE-DEDG----IERAVFYLSRVLNDAKTRYTMIEKLCLCLYF 951
P+ LY+SAT + + ++L E +EDG ++R ++++S VL D+KTRY ++KL +
Sbjct: 1302 PLLLYVSATSQVVSTVLVVEREEDGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILI 1361
Query: 952 SCTKLKYYIKPIDVMVFSHFDIIKHMLSKPILHSRIGKWSLALTEYSLTYAPLKTIKGQA 1011
+ KL +Y + V V + F + +L + RI KW+L L +++ P +IK QA
Sbjct: 1362 TTRKLSHYFQGHLVTVVTSFPL-GDILHNREANGRIAKWALELMSLDISFKPRISIKSQA 1420
Query: 1012 VADFLADHT-LPEEIAYVGLQPWKLFFDDSSHKEGSGIGMFIVSPQGIPTKFMFRIRESC 1070
+ADF+A+ T E+ ++ W + FD S G+G G+ ++SP G ++ I S
Sbjct: 1421 LADFVAEWTECQEDTPAEKVEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSA 1480
Query: 1071 SNNESEYEALVSGLEILLALGAKNVEVKGDSELVVKQLTKEYKCISKNLAKYYVKAMSLL 1130
S+N +EYEAL+ GL I ++LG K + V+GDS+LVV Q+ KE+ C+ N+ Y + L
Sbjct: 1481 SHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLE 1540
Query: 1131 ANFDQAGVSYIPRVSNQEANELA------QIASGYMIDKQKLKELIRIKEKLSPLDLEVM 1184
FD +S++ R +N+ A+ LA ++A + + + K+ +
Sbjct: 1541 DKFDGLELSHVLRHNNKAADRLANFGSKREVAPSDVFVEHLYAPTVPHKDTTQVAGTHDV 1600
Query: 1185 VIDNLTPNDWRKPIVEYLQNPVGSTDI----KVKYRALNYTIIGNELFKKNVDGTLLKCL 1240
+ DWR+P++ +L + D ++ R+ Y + EL+KK+ G L +C+
Sbjct: 1601 A---MVEADWREPLIRFLTSQELPQDKDEAERISRRSRLYVMHEAELYKKSPSGILQRCV 1657
Query: 1241 SEVDAFIAVSAAHGGLCGAHQAGAKMKWILFRQGMYWPTIMKDCIEYAKGCQDCQKHSGI 1300
S + + H G+CG H A + +RQG +WPT + D + + C+ CQ +
Sbjct: 1658 SLEEGRQLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQ 1717
Query: 1301 QHVPASELHSIIKSWPFRGWAIDLIGEIHPALSRQHKYIIVAIDYFIKWVEAIPLQSVTQ 1360
H+PA EL +I SWPF W +D++G A + ++ VAID F KW+EA P+ ++T
Sbjct: 1718 IHLPAQELQTIPLSWPFAVWGLDMVGPFKKAFG-GYTHLFVAIDKFSKWIEAKPVVTITA 1776
Query: 1361 ETVIEFIQNHIVYRFGLPESLTTDQGTMFVGRKVAAFAESWGIKLLTSIPYYAQANGQVE 1420
+ +F N IV+RFG+P + TD G F G F E +GIK+ + + +NGQVE
Sbjct: 1777 DNARDFFIN-IVHRFGVPNRIITDNGRQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVE 1835
Query: 1421 AANKILISLIKKHVGRKPK----SWHESLSQVLWAYRNSPREATGTTPFRLAYGQEAVLP 1476
AN +++ IK V + K W E L VLW+ R +P ATG +PF L YG EA+LP
Sbjct: 1836 RANGMILQGIKARVFDRLKPYAGKWVEQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLP 1895
Query: 1477 AEVYLQSFRIQRQEDIPSEVYWNMMLDELVNLDEERVLALDVLTRQKDRIAKAYNKKVKN 1536
+EV +S R + + E Y +D+L L+E R AL R + + +N+ V++
Sbjct: 1896 SEVEFESLRFR---NFREERYEEDRVDDLHRLEEVREAALIQSARYLQGLRRYHNRNVRS 1952
Query: 1537 RSFVTGDYVWKVILPTDKKDRAYGKWAPNWEGPFKVKKVLSNNAYVIKELSG 1588
R+F+ GD V + I T +DR K +P WEGPF + +V +Y +K G
Sbjct: 1953 RAFLVGDLVLRKIQTT--RDR--HKLSPLWEGPFIISEVTRPGSYRLKREDG 2000
>UniRef100_Q75KG5 Putative polyprotein [Oryza sativa]
Length = 1991
Score = 676 bits (1743), Expect = 0.0
Identities = 427/1248 (34%), Positives = 672/1248 (53%), Gaps = 83/1248 (6%)
Query: 374 PLFVWAKVEDKGVNKILVDGGAAINLMPRFMMKRLGKTETDL-IPHDMVLSD-------Y 425
PL + V + + + L+DGG+A+N++ KT D+ IP + S
Sbjct: 777 PLVLDPVVRNVKLRRTLIDGGSALNIL-------FAKTLDDMQIPRSELKSSNAPFHGVI 829
Query: 426 EGKTSTSMGAIMLNITAGTV----SRSTLFIVVPSKANYNLLLGREWIHGVGAVPSTLHQ 481
G ++T +G I L +T GT + + F V + Y+ +LGR + AVP +
Sbjct: 830 PGLSATPLGQITLPVTFGTRENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYM 889
Query: 482 RISIWKPDGVVENVQADQSYYLAEAGYVDKKNFEKSPLVDATKMEAQDPLEKVDLGDGSK 541
+ + P GV+ +++D + +A DK++ + A E E++ L ++
Sbjct: 890 MMKMPGPRGVL-TLRSD----IKQAVTCDKESCDM-----AQTREMASAREEIRLAATTE 939
Query: 542 KRPTYISSLIDPELRGRMIELLKEFKDCFA*DNDEMPGLSRDLVELQLPIKEDKKLVK*L 601
+I L+ KD FA +MPG+ R+++E L +KED K +K
Sbjct: 940 SA---------------LITFLQNNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQR 984
Query: 602 PRRFHPDVLVKIKEEIERLLRCKFIRAAKYVEWLANVVPVIKKNGKMRVCIDFRDLNAAT 661
RRF D IKEE+ +LL FI+ + +WLAN V V KK G+ R+C+D+ DLN +
Sbjct: 985 LRRFAQDRKDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSC 1044
Query: 662 PKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAKEDVSKTAFRCSGALGTYEWVV 721
PKD + +P + +VDS AG E LS LD YSGY+QI + + D KT+F G Y +V
Sbjct: 1045 PKDPFGLPRIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITP--FGAYCYVT 1102
Query: 722 MPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDLVVKSPSRDEHLSHLRKSFERMRIHG 781
MPFGLKNAGATYQR++ F I ++ Y+DD+VVK+ +D+ + L ++F +R
Sbjct: 1103 MPFGLKNAGATYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLILDLEETFASIRAFR 1162
Query: 782 LKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKVNFLR 841
+K+NP KC FGV +G LGF+V +GI+ N K AIL+ PP+++K +Q L G + L
Sbjct: 1163 MKLNPEKCTFGVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALS 1222
Query: 842 RFIDNLSDKTKPFSSLLRLKREDIFRWEA*HQKAFDELKNYLAIPPVMIPPIKGKPMRLY 901
RF+ L ++ PF L LK+ D F+W QKAF++ K L PPV+ P +P+ LY
Sbjct: 1223 RFVSRLGERGMPFFKL--LKKTDNFQWGPEAQKAFEDFKQLLTKPPVLASPHPQEPLLLY 1280
Query: 902 ISATDETIGSMLAQE-DEDG----IERAVFYLSRVLNDAKTRYTMIEKLCLCLYFSCTKL 956
+SAT + + ++L E +E+G ++R ++++S VL D+KTRY ++KL + + KL
Sbjct: 1281 VSATSQVVSTVLVVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKL 1340
Query: 957 KYYIKPIDVMVFSHFDIIKHMLSKPILHSRIGKWSLALTEYSLTYAPLKTIKGQAVADFL 1016
+Y + V V + F + +L + RI KW+L L +++ P +IK QA+ADF+
Sbjct: 1341 SHYFQGHSVTVVTSFP-LGDVLHNREANGRIAKWALELMSLDISFKPRTSIKSQALADFV 1399
Query: 1017 ADHT-LPEEIAYVGLQPWKLFFDDSSHKEGSGIGMFIVSPQGIPTKFMFRIRESCSNNES 1075
A+ T E+ ++ W + FD S G+G G+ ++SP G ++ I S S+N +
Sbjct: 1400 AEWTECQEDTPVEKIEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVA 1459
Query: 1076 EYEALVSGLEILLALGAKNVEVKGDSELVVKQLTKEYKCISKNLAKYYVKAMSLLANFDQ 1135
EYEAL+ GL I ++LG K + V+GDS+LVV Q+ KE+ CI N+ Y + L FD
Sbjct: 1460 EYEALLHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCIDDNMMAYRQEVRKLEDKFDG 1519
Query: 1136 AGVSYIPRVSNQEANELAQIASGYMIDKQKLKELIRIKEKLSPL---DLEVMVIDN---- 1188
+S++ R +N+ A+ LA S ++ + ++ SP V D
Sbjct: 1520 LELSHVLRHNNEAADRLANFGS----KREAAPSDVFVEHLYSPTVPHKDATQVADTHDIA 1575
Query: 1189 LTPNDWRKPIVEYLQN---PVGSTDI-KVKYRALNYTIIGNELFKKNVDGTLLKCLSEVD 1244
+ DWR+P++ +L + P + ++ R+ Y + EL+KK+ G L +C+S +
Sbjct: 1576 MVEADWREPLIRFLTSQELPQYKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEE 1635
Query: 1245 AFIAVSAAHGGLCGAHQAGAKMKWILFRQGMYWPTIMKDCIEYAKGCQDCQKHSGIQHVP 1304
+ H G+CG H A + +RQG +WPT + D + + C+ CQ + H+P
Sbjct: 1636 GRQLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLP 1695
Query: 1305 ASELHSIIKSWPFRGWAIDLIGEIHPALSRQHKYIIVAIDYFIKWVEAIPLQSVTQETVI 1364
A EL +I SWPF W +D++G A+ + ++ +AID F KW+EA P+ ++T +
Sbjct: 1696 AQELQTIPLSWPFAVWGLDMVGPFKKAVG-GYTHLFMAIDKFSKWIEAKPVVTITADNAR 1754
Query: 1365 EFIQNHIVYRFGLPESLTTDQGTMFVGRKVAAFAESWGIKLLTSIPYYAQANGQVEAANK 1424
+F N IV+RFG+P + TD GT F G F E +GIK+ + + +NGQVE AN
Sbjct: 1755 DFFIN-IVHRFGVPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANG 1813
Query: 1425 ILISLIKKHVGRKPK----SWHESLSQVLWAYRNSPREATGTTPFRLAYGQEAVLPAEVY 1480
+++ IK V + K W L VLW+ R +P ATG +PF L YG EA+LP+EV
Sbjct: 1814 MILQRIKARVFDRLKPYAGKWVSQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVE 1873
Query: 1481 LQSFRIQRQEDIPSEVYWNMMLDELVNLDEERVLALDVLTRQKDRIAKAYNKKVKNRSFV 1540
+S R + + E Y +D+L L+E R AL R + + +N+ V++R+F+
Sbjct: 1874 FESLRFR---NFREERYEEDRVDDLHRLEEVREAALIQSARYLQGLRRYHNRNVRSRAFL 1930
Query: 1541 TGDYVWKVILPTDKKDRAYGKWAPNWEGPFKVKKVLSNNAYVIKELSG 1588
GD V + I T +DR K +P WEGPF + +V +Y +K G
Sbjct: 1931 VGDLVLRKIQTT--RDR--HKLSPLWEGPFIISEVTRPGSYRLKREDG 1974
>UniRef100_Q75IS9 Putative polyprotein [Oryza sativa]
Length = 1756
Score = 675 bits (1741), Expect = 0.0
Identities = 428/1279 (33%), Positives = 683/1279 (52%), Gaps = 89/1279 (6%)
Query: 374 PLFVWAKVEDKGVNKILVDGGAAINLMPRFMMKRLGKTETDLIPHDMVLSDY-EGKTSTS 432
PL + V + + + L+DGG+A+N++ + + ++L P + G ++T
Sbjct: 516 PLVLDPVVRNVKLRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATP 575
Query: 433 MGAIMLNITAGTV----SRSTLFIVVPSKANYNLLLGREWIHGVGAVPSTLHQRISIWKP 488
+G I L +T GT + + F V + Y+ +LGR + AVP + + + P
Sbjct: 576 LGQITLPVTFGTRENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKMPGP 635
Query: 489 DGVVENVQADQSYYLAEAGYVDKKNFEKSP------------LVDATKMEAQDPLEKVDL 536
V+ ++++D + +A DK++ + + L AT E + P K+
Sbjct: 636 RRVL-SLRSD----IKQAVTCDKESCDMAQTREMASAREDIRLAVATASEGEVPATKISK 690
Query: 537 GDGSKKRPTYISSLIDPELRGRMIELLKEFKDCFA*DNDEMPGLSRDLVELQLPIKEDKK 596
S+ + I +DP + KD FA +MPG+ R+++E L +KED K
Sbjct: 691 SGESEAKTKKIP--LDPSDSTKTANN----KDIFAWKPSDMPGIPREVIEHSLHVKEDAK 744
Query: 597 LVK*LPRRFHPDVLVKIKEEIERLLRCKFIRAAKYVEWLANVVPVIKKNGKMRVCIDFRD 656
+K RRF D IKEE+ +LL FI+ + +WLAN V V KK G+ R+C+D+ D
Sbjct: 745 PIKQRLRRFVQDRKDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTD 804
Query: 657 LNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAKEDVSKTAFRCSGALGT 716
LN PKD + +P + +VDS AG E LS LD YSGY+QI + + D KT+F +G
Sbjct: 805 LNKCCPKDPFGLPRIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITP--IGA 862
Query: 717 YEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDLVVKSPSRDEHLSHLRKSFER 776
Y +V MPFGLKNAGATYQR++ F I ++ Y+DD+V+K+ +D+ +S L ++F
Sbjct: 863 YCYVTMPFGLKNAGATYQRMIQRCFSTQIGRNVEAYVDDVVIKTKQKDDLISDLEETFAS 922
Query: 777 MRIHGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGK 836
+R +K+NP KC FGV +G LGF+V +GI+ N K AIL+ PP+++K +Q L G
Sbjct: 923 IRAFRMKLNPEKCTFGVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGC 982
Query: 837 VNFLRRFIDNLSDKTKPFSSLLRLKREDIFRWEA*HQKAFDELKNYLAIPPVMIPPIKGK 896
+ L RF+ L ++ PF LL K+ D F+W QKAF++ K +L PPV+ P +
Sbjct: 983 MAALSRFVSRLGERGMPFFKLL--KKTDDFQWGPEAQKAFEDFKKFLTEPPVLASPHPQE 1040
Query: 897 PMRLYISATDETIGSMLAQE-DEDG----IERAVFYLSRVLNDAKTRYTMIEKLCLCLYF 951
P+ LY+SAT + + ++L E +E+G ++R ++++S VL D+KTRY ++KL +
Sbjct: 1041 PLLLYVSATSQVVSTVLVVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILI 1100
Query: 952 SCTKLKYYIKPIDVMVFSHFDIIKHMLSKPILHSRIGKWSLALTEYSLTYAPLKTIKGQA 1011
+ KL +Y + V V + F + +L + RI KW+L L +++ P +IK QA
Sbjct: 1101 TTRKLSHYFQGHSVTVVTSFPL-GDILHNREANGRIAKWALELMSLDISFKPRISIKSQA 1159
Query: 1012 VADFLADHT-LPEEIAYVGLQPWKLFFDDSSHKEGSGIGMFIVSPQGIPTKFMFRIRESC 1070
+ADF+A+ T E+ ++ W + FD S G+G G+ ++SP G ++ I S
Sbjct: 1160 LADFVAEWTECQEDTPAEKMEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSA 1219
Query: 1071 SNNESEYEALVSGLEILLALGAKNVEVKGDSELVVKQLTKEYKCISKNLAKYYVKAMSLL 1130
S+N +EYEAL+ GL I ++LG K + V+GDS+LVV Q+ KE+ C+ N+ Y + L
Sbjct: 1220 SHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLE 1279
Query: 1131 ANFDQAGVSYIPRVSNQEANELAQIASGYMIDKQKLKELIRIKEKLSPLDLEVMVIDN-- 1188
FD +S++ R +N+ A+ LA S K +++P D+ V +
Sbjct: 1280 DKFDGLELSHVLRHNNEAADRLANFGS---------------KREVAPSDVFVEHLYTPT 1324
Query: 1189 ----------------LTPNDWRKPIVEYLQNPVGSTDI----KVKYRALNYTIIGNELF 1228
L DWR+P + +L + D ++ R+ Y + EL+
Sbjct: 1325 VPHKDTTQIAGTHDVALVEADWREPFIRFLTSQELPQDKDEAERISRRSKLYVMHEAELY 1384
Query: 1229 KKNVDGTLLKCLSEVDAFIAVSAAHGGLCGAHQAGAKMKWILFRQGMYWPTIMKDCIEYA 1288
KK+ G L +C+S + + H G+CG H A + +RQG +WPT + D +
Sbjct: 1385 KKSPSGILQRCVSLEEGRQLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIV 1444
Query: 1289 KGCQDCQKHSGIQHVPASELHSIIKSWPFRGWAIDLIGEIHPALSRQHKYIIVAIDYFIK 1348
+ C+ CQ + H+PA EL +I SWPF W +D++G A+ + ++ VAID F K
Sbjct: 1445 RTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGLDMVGPFKKAVG-GYTHLFVAIDKFSK 1503
Query: 1349 WVEAIPLQSVTQETVIEFIQNHIVYRFGLPESLTTDQGTMFVGRKVAAFAESWGIKLLTS 1408
W+EA P+ ++T + +F N IV+RFG+P + TD G F G F E +GIK+ +
Sbjct: 1504 WIEAKPVVTITADNARDFFIN-IVHRFGVPNRIITDNGRQFTGGVFKDFCEDFGIKICYA 1562
Query: 1409 IPYYAQANGQVEAANKILISLIKKHVGRKPK----SWHESLSQVLWAYRNSPREATGTTP 1464
+ +NGQVE AN +++ IK V + K W + L VLW+ R +P ATG +P
Sbjct: 1563 SVAHPMSNGQVERANGMILQGIKARVFDRLKPYAGKWVKQLPSVLWSLRTTPSRATGQSP 1622
Query: 1465 FRLAYGQEAVLPAEVYLQSFRIQRQEDIPSEVYWNMMLDELVNLDEERVLALDVLTRQKD 1524
F L YG EA+LP+EV +S R + + E Y +D+L L+E R AL R
Sbjct: 1623 FFLVYGAEAMLPSEVEFESLRFR---NFREERYEEDRVDDLHRLEEVREAALIQSARYLQ 1679
Query: 1525 RIAKAYNKKVKNRSFVTGDYVWKVILPTDKKDRAYGKWAPNWEGPFKVKKVLSNNAYVIK 1584
+ + +N+ V++R+F+ GD V + I T +DR K +P WEGPF + +V +Y +K
Sbjct: 1680 GLRRYHNRNVRSRAFLVGDLVLRKIQTT--RDR--HKLSPLWEGPFIISEVTRPGSYRLK 1735
Query: 1585 ELSGQRQFVTINDKYLKAY 1603
G + N ++L+ +
Sbjct: 1736 REDGTLVDNSWNIEHLRRF 1754
>UniRef100_Q8W3B5 Putative gag-pol [Oryza sativa]
Length = 2026
Score = 675 bits (1741), Expect = 0.0
Identities = 432/1281 (33%), Positives = 681/1281 (52%), Gaps = 93/1281 (7%)
Query: 374 PLFVWAKVEDKGVNKILVDGGAAINLMPRFMMKRLGKTETDLIPHDMVLSDY-EGKTSTS 432
PL + V + + + L+DGG+A+N++ + + ++L P + G ++T
Sbjct: 777 PLVLDPVVRNVKLRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATP 836
Query: 433 MGAIMLNITAGTV----SRSTLFIVVPSKANYNLLLGREWIHGVGAVPSTLHQRISIWKP 488
+G I L +T GT + + F V + Y+ +LGR + AVP + + + P
Sbjct: 837 LGQITLPVTFGTRENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKMPGP 896
Query: 489 DGVVENVQADQSYYLAEAGYVDKKNFEKSP------------LVDATKMEAQDPLEKVDL 536
GV+ ++++D + +A DK++ + + L AT E + P K
Sbjct: 897 RGVL-SLRSD----IKQAVTCDKESCDMAQTREMASAREDIRLAAATASEGEVPATKTSK 951
Query: 537 GDGSKKRPTYISSLIDPELRGRMIELLKEFKDCFA*DNDEMPGLSRDLVELQLPIKEDKK 596
S+ + I +DP + KD FA +MPG+ R+++E L +KED K
Sbjct: 952 SGESEAKTKKIP--LDPSNPTKTANN----KDIFAWKPSDMPGIPREVIEHSLHVKEDAK 1005
Query: 597 LVK*LPRRFHPDVLVKIKEEIERLLRCKFIRAAKYVEWLANVVPVIKKNGKMRVCIDFRD 656
+K RRF D IKEE+ +LL FI+ + +WLAN V V KK G+ R+C+D+ D
Sbjct: 1006 PIKQRLRRFAQDRKDAIKEELTKLLAAGFIKEVHHPDWLANPVLVRKKTGQWRMCVDYTD 1065
Query: 657 LNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAKEDVSKTAFRCSGALGT 716
LN + PKD + +P + +VDS AG E LS LD YSGY+QI + + D KT+F G
Sbjct: 1066 LNKSCPKDPFGLPRIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITP--FGA 1123
Query: 717 YEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDLVVKSPSRDEHLSHLRKSFER 776
Y +V MPFGLKNAGATYQR++ F I ++ Y+DD+VVK+ +D+ +S L ++F
Sbjct: 1124 YCYVTMPFGLKNAGATYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFAS 1183
Query: 777 MRIHGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGK 836
+R +K+NP KC FGV +G LGF+V +GI+ N K AIL+ PP+++K +Q L G
Sbjct: 1184 IRAFRMKLNPEKCTFGVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGC 1243
Query: 837 VNFLRRFIDNLSDKTKPFSSLLRLKREDIFRWEA*HQKAFDELKNYLAIPPVMIPPIKGK 896
+ L RF+ L ++ PF LL K+ D F+W QKAF++ K L PPV+ P +
Sbjct: 1244 MAALSRFVSRLGERGMPFFKLL--KKTDDFQWGPEAQKAFEDFKKLLTEPPVLASPHPQE 1301
Query: 897 PMRLYISATDETIGSMLAQE-DEDG----IERAVFYLSRVLNDAKTRYTMIEKLCLCLYF 951
P+ LY+SAT + + ++L E +E+G ++R ++++S VL D+KTRY ++KL +
Sbjct: 1302 PLLLYVSATSQVVSTVLVVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILI 1361
Query: 952 SCTKLKYYIKPIDVMVFSHFDIIKHMLSKPILHSRIGKWSLALTEYSLTYAPLKTIKGQA 1011
+ KL +Y + V V + F + +L + RI KW+L L +++ P +IK QA
Sbjct: 1362 TTRKLSHYFQGHSVTVVTSFPL-GDILHNREANGRIAKWALELMSLDISFKPRISIKSQA 1420
Query: 1012 VADFLADHT-LPEEIAYVGLQPWKLFFDDSSHKEGSGIGMFIVSPQGIPTKFMFRIRESC 1070
+ADF+A+ T E+ ++ W + FD S G+G G+ ++SP G ++ I S
Sbjct: 1421 LADFVAEWTECQEDTPAEKMEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSA 1480
Query: 1071 SNNESEYEALVSGLEILLALGAKNVEVKGDSELVVKQLTKEYKCISKNLAKYYVKAMSLL 1130
S+N +EYEAL+ GL I ++LG K + V+GDS+LVV Q+ KE+ C+ N+ Y + L
Sbjct: 1481 SHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLE 1540
Query: 1131 ANFDQAGVSYIPRVSNQEANELAQIASGYMIDKQKLKELIRIKEKLSPLDLEVMVIDNLT 1190
FD +S++ R +N+ A+ LA S K +++P D V V T
Sbjct: 1541 DKFDGLELSHVLRHNNEAADRLANFGS---------------KREVAPSD--VFVEHLYT 1583
Query: 1191 PN--------------------DWRKPIVEYLQNPVGSTDI----KVKYRALNYTIIGNE 1226
P DWR+P++ +L + D ++ R+ Y + E
Sbjct: 1584 PTVPHKDTTQVAGTHDVAMVEADWREPLIRFLTSQELPQDKDEAERISRRSKLYVMHEAE 1643
Query: 1227 LFKKNVDGTLLKCLSEVDAFIAVSAAHGGLCGAHQAGAKMKWILFRQGMYWPTIMKDCIE 1286
L+KK+ G L +C+S + + H G+CG H A + +RQG +WPT + D +
Sbjct: 1644 LYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADK 1703
Query: 1287 YAKGCQDCQKHSGIQHVPASELHSIIKSWPFRGWAIDLIGEIHPALSRQHKYIIVAIDYF 1346
+ C+ CQ + H+PA EL +I SWPF W +D++G A+ + ++ VAID F
Sbjct: 1704 IVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGLDMVGPFKKAVG-GYTHLFVAIDKF 1762
Query: 1347 IKWVEAIPLQSVTQETVIEFIQNHIVYRFGLPESLTTDQGTMFVGRKVAAFAESWGIKLL 1406
KW+EA P+ ++T + +F N IV+RFG+P + TD G F G E +GIK+
Sbjct: 1763 SKWIEAKPVVTITADNAPDFFIN-IVHRFGVPNRIITDNGRQFTGGVFKDCCEDFGIKIC 1821
Query: 1407 TSIPYYAQANGQVEAANKILISLIKKHVGRKPK----SWHESLSQVLWAYRNSPREATGT 1462
+ + +NGQVE AN +++ IK V + K W E L VLW+ R +P ATG
Sbjct: 1822 YASVAHPMSNGQVERANGMILQGIKARVFDRLKPYAGKWVEQLPSVLWSLRTTPSRATGQ 1881
Query: 1463 TPFRLAYGQEAVLPAEVYLQSFRIQRQEDIPSEVYWNMMLDELVNLDEERVLALDVLTRQ 1522
+PF L YG EA+LP+EV +S R + + E Y +D+L L+E R AL R
Sbjct: 1882 SPFFLVYGAEAMLPSEVEFESLRFR---NFREERYEEDRVDDLHRLEEVREAALIQSARY 1938
Query: 1523 KDRIAKAYNKKVKNRSFVTGDYVWKVILPTDKKDRAYGKWAPNWEGPFKVKKVLSNNAYV 1582
+ + +N+ V++R+F+ GD V + I T +DR K +P WEGPF + V +Y
Sbjct: 1939 LQGLRRYHNRNVRSRAFLVGDLVLRKIQTT--RDR--HKLSPLWEGPFIISAVTRPGSYR 1994
Query: 1583 IKELSGQRQFVTINDKYLKAY 1603
+K G + N ++L+ +
Sbjct: 1995 LKREDGTLVDNSWNIEHLRRF 2015
>UniRef100_Q9ZNW4 Polyprotein [Sorghum bicolor]
Length = 1877
Score = 674 bits (1740), Expect = 0.0
Identities = 421/1268 (33%), Positives = 670/1268 (52%), Gaps = 61/1268 (4%)
Query: 371 HLKPLFVWAKVEDKGVNKILVDGGAAINLMPRFMMKRLGKTETDLIPHDMVLSDYEGKTS 430
H L + V GV++ILVD G++ +++ ++ + + L P + L + GK
Sbjct: 634 HTDALVITTNVAGWGVSRILVDSGSSADIIFAGTFDQMKLSRSQLQPSESPLIGFGGKQI 693
Query: 431 TSMGAIMLNITAGTVSRS----TLFIVVPSKANYNLLLGREWIHGVGAVPSTLHQRISIW 486
++G + L ++ GT + F VV YN + GR +++ A + + I
Sbjct: 694 HALGKVALPVSFGTTENARTEYVTFDVVDLHYPYNAIFGRGFLNKFNAAAHMGYLCMKIP 753
Query: 487 KPDGVVENVQADQSYYLAEAGYVDK---KNFEKSPLVDATKMEAQDPLE----KVDLGDG 539
GV+ V Q EA ++K K+F VD+T+ EA P + K +L D
Sbjct: 754 ALHGVI-TVHGSQK----EARNIEKAIYKSFRNINSVDSTQHEASQPPDMPRGKTNLADQ 808
Query: 540 SKKR-----------PTYISSLIDPELRGRMIELLKEFKDCFA*DNDEMPGLSRDLVELQ 588
+ + IS+ + E ++++L++ +D FA ++ G+SRD++E
Sbjct: 809 EETKCIPLQEAVPDKKVTISATLSREEELELLDVLQKNQDIFAWSAADLQGVSRDIIEHS 868
Query: 589 LPIKEDKKLVK*LPRRFHPDVLVKIKEEIERLLRCKFIRAAKYVEWLANVVPVIKKNGKM 648
L I + K R+ + + K E++RLL K IR Y EWLANVV V KKNGKM
Sbjct: 869 LDIDPRMRPKKQRQRKMSEERTLAAKAEVQRLLDAKVIREVIYPEWLANVVLVPKKNGKM 928
Query: 649 RVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAKEDVSKTAF 708
R+CIDF DLN A KD + +P + VD AAG + SLLD +SGY+QI++ KED K +F
Sbjct: 929 RMCIDFTDLNKACVKDSFPLPRIDTSVDKAAGCQRFSLLDCFSGYHQIWLKKEDEGKASF 988
Query: 709 RCSGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDLVVKSPSRDEHLS 768
+ GTY + MP GLKNAGAT+ R+M + ++ + Y+DD+VV S +++H+
Sbjct: 989 --TTPFGTYCYTRMPEGLKNAGATFSRMMGKVLGSQLQRNIIAYVDDVVVMSKRKEDHIK 1046
Query: 769 HLRKSFERMRIHGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKK 828
L+++F +R GLK+NP KC FGV G LG+++ +GI N +K KAI+ + P++KK
Sbjct: 1047 DLQETFVNLRSAGLKLNPEKCVFGVSKGKMLGYIISSEGIRANPDKTKAIMSMAEPSNKK 1106
Query: 829 QLQSLLGKVNFLRRFIDNLSDKTKPFSSLLRLKREDIFRWEA*HQKAFDELKNYLAIPPV 888
++Q L G++ L RFI ++++ PF +L RE W +AF +LKNY+A +
Sbjct: 1107 EVQRLTGRIAALNRFISRSAERSLPFFKVL---REGKTEWGPEQSEAFRQLKNYIATNLL 1163
Query: 889 MIPPIKGKPMRLYISATDETIGSMLAQEDED---GIERAVFYLSRVLNDAKTRYTMIEKL 945
+ P P+ LY++A++ + +L E D ++R V+Y+S L+ AK YT IEK+
Sbjct: 1164 VTVPEPDTPLLLYVAASEHAVSGVLVHETSDTKGTVQRPVYYVSEALSGAKLNYTEIEKI 1223
Query: 946 CLCLYFSCTKLKYYIKPIDVMVFSHFDIIKHMLSKPILHSRIGKWSLALTEYSLTYAPLK 1005
+ + KLK+Y + ++ V + + +L RIGKW+ L+++ +TY P
Sbjct: 1224 AYAVLCASRKLKHYFQSHEIKVPTS-QPLGDILRNKEASGRIGKWAAELSQFDITYVPRT 1282
Query: 1006 TIKGQAVADFLADHTLPEEIAYVGL-QPWKLFFDDSSHKEGSGIGMFIVSPQGIPTKFMF 1064
+IK QA+ADF+AD T + + QPW L+ D + + G+G +++P G+ KF
Sbjct: 1283 SIKSQALADFMADWTPSNKNEEKVIDQPWTLYTDGAWGQAGAGAAAVLIAPSGLKLKFAI 1342
Query: 1065 RIRESCSNNESEYEALVSGLEILLALGAKNVEVKGDSELVVKQLTKEYKCISKNLAKYYV 1124
R+ +NN +EYE L+ GL A GAK + +K DS++V Q+ KEY + LA+Y
Sbjct: 1343 RLEFKATNNIAEYEGLILGLNKAKASGAKTLVIKTDSQVVAGQVEKEYLAHNPELARYLA 1402
Query: 1125 KAMSLLANFDQAGVSYIPRVSNQEANELAQIASGYM-IDKQKLKELIR--IKEKLSPLDL 1181
L F + YIPR N EA+ELA+ A+ + + +++ E L
Sbjct: 1403 VVRGLERRFKGFTLQYIPRAENYEADELAKAAANNTPLPEGTFHQIVTTPATETLPKAFR 1462
Query: 1182 EVMVIDNLTPNDWRKPIVEYLQNPV----GSTDIKVKYRALNYTIIGNELFKKNVDGTLL 1237
V++ ++ DWR+ I + L+ +T +++ RA NYT I L+KK V LL
Sbjct: 1463 SVLLTES---EDWRQAIADCLKGKTTVDDEATAKRMEARARNYTSIDGILYKKGVVQPLL 1519
Query: 1238 KCLSEVDAFIAVSAAHGGLCGAHQAGAKMKWILFRQGMYWPTIMKDCIEYAKGCQDCQKH 1297
KC+S+ + + H G+CG+H + RQG YWPT ++D E K C+ CQ
Sbjct: 1520 KCISQSEGRELLREIHSGMCGSHIGPRALSAKALRQGFYWPTHIRDAEEIVKTCKACQTF 1579
Query: 1298 SGIQHVPASELHSIIKSWPFRGWAIDLIGEIHPALSRQHKYIIVAIDYFIKWVEAIPLQS 1357
S IQ P++ I SWP + W +DL+G + P +K+ +VAI+YF +W+EA PL +
Sbjct: 1580 SPIQSGPSALTQLIPASWPLQRWGMDLVGPM-PTAQGGNKFAVVAIEYFTRWIEAKPLTT 1638
Query: 1358 VTQETVIEFIQNHIVYRFGLPESLTTDQGTMFVGRKVAAFAESWGIKLLTSIPYYAQANG 1417
+T ET+ +F +IV RFG+P LT D G F F G K+ + Y+ ++NG
Sbjct: 1639 ITSETIRKFFWQNIVCRFGVPRLLTVDNGKQFDSDNFKEFCHLIGTKIAFASVYHPESNG 1698
Query: 1418 QVEAANKILISLIKKH-VGRKPKSWHESLSQVLWAYRNSPREATGTTPFRLAYGQEAVLP 1476
VE AN+ + S I K + + W E L +V+W++ + ATG TPF+L YG+EA+LP
Sbjct: 1699 AVERANRTIFSAISKTLLNLRKGKWVEELPRVVWSHNTTVSRATGFTPFKLLYGEEAMLP 1758
Query: 1477 AEVYLQSFRIQRQEDIPSEVYWNMMLDELVNLDEERVLALDVLTRQKDRIAKAYNKKVKN 1536
E+ QS R +Q+ E Y L+ + R+ A++ +TR + ++KV
Sbjct: 1759 EEIKHQSLRSMKQQLAEDEEYCKETLESI------RLEAVENITRYQQETKNWRDRKVVR 1812
Query: 1537 RSFVTGDYVWKVILPTDKKDRA-YGKWAPNWEGPFKVKKVLSNNAYVIKELSGQRQFVTI 1595
+ GD V + K D GK P WEGP+ + + ++ +K+L G+ T
Sbjct: 1813 KDIQNGDLVLR-----KKGDHPNAGKLQPKWEGPYTAIQAGRSGSFYLKDLEGRTSTHTW 1867
Query: 1596 NDKYLKAY 1603
N L+ +
Sbjct: 1868 NVDNLRRF 1875
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.325 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,574,234,679
Number of Sequences: 2790947
Number of extensions: 108011583
Number of successful extensions: 414534
Number of sequences better than 10.0: 28168
Number of HSP's better than 10.0 without gapping: 2998
Number of HSP's successfully gapped in prelim test: 25170
Number of HSP's that attempted gapping in prelim test: 374245
Number of HSP's gapped (non-prelim): 33676
length of query: 1610
length of database: 848,049,833
effective HSP length: 141
effective length of query: 1469
effective length of database: 454,526,306
effective search space: 667699143514
effective search space used: 667699143514
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 82 (36.2 bits)
Lotus: description of TM0041.7


