

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0037a.10
(168 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
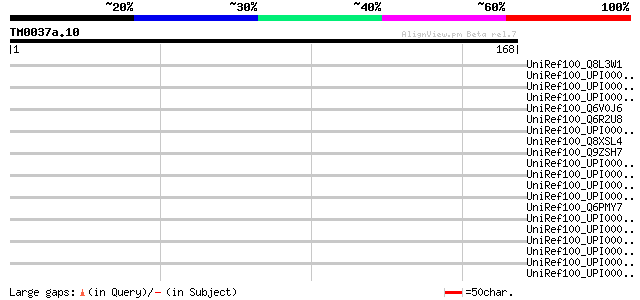
Sequences producing significant alignments: (bits) Value
UniRef100_Q8L3W1 Reduced vernalization response 1 [Arabidopsis t... 37 0.19
UniRef100_UPI00002D57DA UPI00002D57DA UniRef100 entry 35 0.73
UniRef100_UPI000042C26F UPI000042C26F UniRef100 entry 35 0.96
UniRef100_UPI00003417F8 UPI00003417F8 UniRef100 entry 35 0.96
UniRef100_Q6V0J6 Reduced vernalization response 1 [Brassica camp... 35 0.96
UniRef100_Q6R2U8 Reduced vernalization response 1 [Brassica camp... 34 1.3
UniRef100_UPI000046B7EE UPI000046B7EE UniRef100 entry 33 3.6
UniRef100_Q8XSL4 PROBABLE RHS-RELATED PROTEIN [Ralstonia solanac... 33 3.6
UniRef100_Q9ZSH7 T15B16.18 protein [Arabidopsis thaliana] 33 3.6
UniRef100_UPI00002F9FDF UPI00002F9FDF UniRef100 entry 32 4.8
UniRef100_UPI000021E16F UPI000021E16F UniRef100 entry 32 6.2
UniRef100_UPI00004530DC UPI00004530DC UniRef100 entry 32 6.2
UniRef100_UPI000042DD99 UPI000042DD99 UniRef100 entry 32 6.2
UniRef100_Q6PMY7 Polyprotein [Foot-and-mouth disease virus Asia 1] 32 6.2
UniRef100_UPI0000431DAB UPI0000431DAB UniRef100 entry 32 8.1
UniRef100_UPI0000431DA9 UPI0000431DA9 UniRef100 entry 32 8.1
UniRef100_UPI0000431DA8 UPI0000431DA8 UniRef100 entry 32 8.1
UniRef100_UPI0000431DA7 UPI0000431DA7 UniRef100 entry 32 8.1
UniRef100_UPI0000431DA6 UPI0000431DA6 UniRef100 entry 32 8.1
UniRef100_UPI0000431DA5 UPI0000431DA5 UniRef100 entry 32 8.1
>UniRef100_Q8L3W1 Reduced vernalization response 1 [Arabidopsis thaliana]
Length = 341
Score = 37.0 bits (84), Expect = 0.19
Identities = 27/99 (27%), Positives = 44/99 (44%), Gaps = 4/99 (4%)
Query: 4 VSRKFAVEFAHDLGLQWKLVDPFGSVHLVSYESNSEEPYLFEGLLEIRKFYSLQGMNWFL 63
V KF +F +L + L P G V V + + +G E YS++ +
Sbjct: 22 VPDKFVSKFKDELSVAVALTVPDGHVWRVGLRKADNKIWFQDGWQEFVDRYSIRIGYLLI 81
Query: 64 LKYQGCSNFDLFIYNDSIEEVCYLTRSLPPSITPTTHSH 102
+Y+G S F ++I+N S E+ Y + L S H+H
Sbjct: 82 FRYEGNSAFSVYIFNLSHSEINYHSTGLMDS----AHNH 116
>UniRef100_UPI00002D57DA UPI00002D57DA UniRef100 entry
Length = 516
Score = 35.0 bits (79), Expect = 0.73
Identities = 27/111 (24%), Positives = 45/111 (40%), Gaps = 20/111 (18%)
Query: 53 FYSLQGMNW----FLLKYQGC----------SNFDLFIYNDSIEEVCYLTRSLPPSITPT 98
F+ + G+NW F C NF+L I+ + + + + P S TP
Sbjct: 75 FFLILGVNWQSYVFHASVVNCLVTIATFILLRNFNLNIFYSFLYSIFFSILAYPSSGTPF 134
Query: 99 THSHTSFHGECTTGSKSVTEE------WIFYPCFFTTRLSPNQVSSSQLVM 143
H++F T S + + WIF+P FF QV +S +++
Sbjct: 135 VDHHSTFFSLLGTYSLILAIKNDKKHYWIFFPIFFIFAFLSKQVPASYIIL 185
>UniRef100_UPI000042C26F UPI000042C26F UniRef100 entry
Length = 945
Score = 34.7 bits (78), Expect = 0.96
Identities = 21/100 (21%), Positives = 48/100 (48%), Gaps = 4/100 (4%)
Query: 32 VSYESNSEEPYLFEGL--LEIRKFYSLQGMNW-FLLKYQGCSNFDLFIYNDSIEEVC-YL 87
+ Y ++ PY + +++ +FY L+ +NW F + + + F + +++VC Y
Sbjct: 291 IIYSCSASTPYSINNIFVMKLDRFYGLKWVNWIFQILFTVSTQLLGFSFAMIMKKVCVYP 350
Query: 88 TRSLPPSITPTTHSHTSFHGECTTGSKSVTEEWIFYPCFF 127
++++ P+I PT + + + + S+ W P F
Sbjct: 351 SKAIWPTILPTIALNRALMNQDNASNNSIVHGWKISPYSF 390
>UniRef100_UPI00003417F8 UPI00003417F8 UniRef100 entry
Length = 84
Score = 34.7 bits (78), Expect = 0.96
Identities = 18/59 (30%), Positives = 31/59 (52%)
Query: 1 MAIVSRKFAVEFAHDLGLQWKLVDPFGSVHLVSYESNSEEPYLFEGLLEIRKFYSLQGM 59
MA+ S+K A+ + +QWK V G +H+V+ + E+ + + GL Y + GM
Sbjct: 1 MAVESKKLAIYKKDERKIQWKTVVAIGMIHVVALFAFREDLFSYSGLAVGIFMYFIAGM 59
>UniRef100_Q6V0J6 Reduced vernalization response 1 [Brassica campestris]
Length = 329
Score = 34.7 bits (78), Expect = 0.96
Identities = 27/99 (27%), Positives = 43/99 (43%), Gaps = 2/99 (2%)
Query: 4 VSRKFAVEFAHDLGLQWKLVDPFGSVHLVSYES--NSEEPYLFEGLLEIRKFYSLQGMNW 61
V KF +F +L + L P G V V N+ + + +G E YS++
Sbjct: 22 VPDKFVSKFKDELSVAVALTVPDGHVWRVGLRKADNNNKIWFQDGWQEFVDRYSIRIGYL 81
Query: 62 FLLKYQGCSNFDLFIYNDSIEEVCYLTRSLPPSITPTTH 100
+ +Y+G S F + IYN E+ Y + L S + H
Sbjct: 82 LIFRYEGNSAFSVCIYNLPQSEINYHSTGLMDSASHNNH 120
>UniRef100_Q6R2U8 Reduced vernalization response 1 [Brassica campestris]
Length = 329
Score = 34.3 bits (77), Expect = 1.3
Identities = 27/99 (27%), Positives = 42/99 (42%), Gaps = 2/99 (2%)
Query: 4 VSRKFAVEFAHDLGLQWKLVDPFGSVHLVSYES--NSEEPYLFEGLLEIRKFYSLQGMNW 61
V KF F +L + L P G V V N+ + + +G E YS++
Sbjct: 22 VPDKFVSRFKDELSVAVALTVPDGHVWRVGLRKADNNNKIWFQDGWQEFVDRYSIRIGYL 81
Query: 62 FLLKYQGCSNFDLFIYNDSIEEVCYLTRSLPPSITPTTH 100
+ +Y+G S F + IYN E+ Y + L S + H
Sbjct: 82 LIFRYEGNSAFSVCIYNLPQSEINYHSTGLMDSASHNNH 120
>UniRef100_UPI000046B7EE UPI000046B7EE UniRef100 entry
Length = 502
Score = 32.7 bits (73), Expect = 3.6
Identities = 21/96 (21%), Positives = 41/96 (41%), Gaps = 6/96 (6%)
Query: 32 VSYESNSEEPYLFEGLLEIRKFYSLQGMNWFLLKYQGCSNFDLFIYNDSIEEVCYLTRSL 91
+ YE N ++ +FE + ++K Y ++ +N +L ++ C F YN + YL + +
Sbjct: 236 IYYEFNGKKYVVFENMYNLKKVYKIKYINLCILFHRYC----FFFYNYNPLIFSYLFKHI 291
Query: 92 PPSITPTTHSHTSFHGECTTGSKSVTEEWIFYPCFF 127
H C K T + ++ CF+
Sbjct: 292 YCMCIECVQKHLIIGNSCI--KKDKTPDKNYHDCFY 325
>UniRef100_Q8XSL4 PROBABLE RHS-RELATED PROTEIN [Ralstonia solanacearum]
Length = 881
Score = 32.7 bits (73), Expect = 3.6
Identities = 18/59 (30%), Positives = 25/59 (41%)
Query: 91 LPPSITPTTHSHTSFHGECTTGSKSVTEEWIFYPCFFTTRLSPNQVSSSQLVMNQFCYV 149
L S PT T+++G+ S T Y C PNQ S+SQL + Y+
Sbjct: 84 LQASSPPTASQVTTYYGDFACPSSVTTNGQAVYRCSHVQYTFPNQPSASQLAATRSAYI 142
>UniRef100_Q9ZSH7 T15B16.18 protein [Arabidopsis thaliana]
Length = 190
Score = 32.7 bits (73), Expect = 3.6
Identities = 19/65 (29%), Positives = 33/65 (50%)
Query: 22 LVDPFGSVHLVSYESNSEEPYLFEGLLEIRKFYSLQGMNWFLLKYQGCSNFDLFIYNDSI 81
LV P G + + EE + EG E + +S++ ++ L +Y+ S+F + I+N S
Sbjct: 64 LVTPAGYKRSIKLKRIGEEIWFHEGWSEFAEAHSIEEGHFLLFEYKKNSSFRVIIFNASA 123
Query: 82 EEVCY 86
E Y
Sbjct: 124 CETNY 128
>UniRef100_UPI00002F9FDF UPI00002F9FDF UniRef100 entry
Length = 891
Score = 32.3 bits (72), Expect = 4.8
Identities = 19/78 (24%), Positives = 33/78 (41%)
Query: 44 FEGLLEIRKFYSLQGMNWFLLKYQGCSNFDLFIYNDSIEEVCYLTRSLPPSITPTTHSHT 103
+ G LE+ K + QG N QG + + ++ + YL R + + +T
Sbjct: 28 YHGHLEVVKLLTSQGANVKCKDKQGYTPLHAAAVSGQLDVIKYLLRVVSEIDDSNAYGNT 87
Query: 104 SFHGECTTGSKSVTEEWI 121
+ H C TG +V E +
Sbjct: 88 ALHMACYTGQDTVANELV 105
>UniRef100_UPI000021E16F UPI000021E16F UniRef100 entry
Length = 1125
Score = 32.0 bits (71), Expect = 6.2
Identities = 24/95 (25%), Positives = 44/95 (46%), Gaps = 9/95 (9%)
Query: 73 DLFIYNDSIEEVCYLTRSLPPSITPTTHSHTSFHGECTTGSKSVTEEWIF---YPCFFTT 129
DL ++ D+I +C++ R I + + G +G +S+T F F T
Sbjct: 327 DLVLFEDAIHHICHINR-----ILESPRGNALLVGVGGSGKQSLTRLAAFISSMDVFQIT 381
Query: 130 RLSPNQVSSSQLVMNQFCYVLMTNESVYGIFVSLN 164
Q+ +++ N+ Y+LM N S++ + VS N
Sbjct: 382 LRKGYQIPDFKVMENKATYILMGNNSLF-VIVSTN 415
>UniRef100_UPI00004530DC UPI00004530DC UniRef100 entry
Length = 1491
Score = 32.0 bits (71), Expect = 6.2
Identities = 25/103 (24%), Positives = 47/103 (45%), Gaps = 8/103 (7%)
Query: 43 LFEGLLEIRKFYSLQGMNWFLLKYQGCSNFDLFIYNDSIEEVCYLTRSLPPSITPTTHSH 102
+F + EI + YSL+ +NW + C ++N E+ YL S P P ++H
Sbjct: 978 VFNSIQEILEKYSLENINWVITGQ--CLRLLHKLHNTF--ELEYLDLSNDPERPPNHNNH 1033
Query: 103 T----SFHGECTTGSKSVTEEWIFYPCFFTTRLSPNQVSSSQL 141
T +++ + + + PC+ T R + + SS++L
Sbjct: 1034 TFSFLNYNKKAERFYRQIRNNHQSLPCWMTLRSNTRESSSNRL 1076
>UniRef100_UPI000042DD99 UPI000042DD99 UniRef100 entry
Length = 1491
Score = 32.0 bits (71), Expect = 6.2
Identities = 25/103 (24%), Positives = 47/103 (45%), Gaps = 8/103 (7%)
Query: 43 LFEGLLEIRKFYSLQGMNWFLLKYQGCSNFDLFIYNDSIEEVCYLTRSLPPSITPTTHSH 102
+F + EI + YSL+ +NW + C ++N E+ YL S P P ++H
Sbjct: 978 VFNSIQEILEKYSLENINWVITGQ--CLRLLHKLHNTF--ELEYLDLSNDPERPPNHNNH 1033
Query: 103 T----SFHGECTTGSKSVTEEWIFYPCFFTTRLSPNQVSSSQL 141
T +++ + + + PC+ T R + + SS++L
Sbjct: 1034 TFSFFNYNKKAERFYRQIRNNHQSLPCWMTLRSNTRESSSNRL 1076
>UniRef100_Q6PMY7 Polyprotein [Foot-and-mouth disease virus Asia 1]
Length = 2329
Score = 32.0 bits (71), Expect = 6.2
Identities = 27/115 (23%), Positives = 45/115 (38%), Gaps = 5/115 (4%)
Query: 56 LQGMNWFLLKYQGCSNFDLFIYNDSIEEVCYLTRSLPPSITPTT---HSHTSFHGECTTG 112
L G+ + +Y G N + + Y+ +PP +TP T H+ H E TG
Sbjct: 594 LAGLAQYYTQYSGTMNVHFMFTGPTDAKARYMVAYVPPGMTPPTDPEHAAHCIHSEWDTG 653
Query: 113 SKSVTEEWIFYPCFFTTRLSPNQVSSSQLVMNQFCYVLMTNESVYG--IFVSLNA 165
S I Y + + V+ + V C +T+ G + VS++A
Sbjct: 654 LNSKFTFSIPYLSAADYAYTASDVAETTSVQGWVCIYQITHGKAEGDALVVSVSA 708
>UniRef100_UPI0000431DAB UPI0000431DAB UniRef100 entry
Length = 1127
Score = 31.6 bits (70), Expect = 8.1
Identities = 30/130 (23%), Positives = 57/130 (43%), Gaps = 9/130 (6%)
Query: 37 NSEEPYLFEGLLEIRKFYSLQGMNWFLLKYQGCSNFDLFIYNDSIEEVCYLTRSLPPSIT 96
N +EP +E LL+ YSL + + S + ++ND++E + + R+L
Sbjct: 113 NEDEPRFYEDLLDYEAVYSLFLEIYEDYNERNVSKLHMVLFNDALEHLTRIHRAL----- 167
Query: 97 PTTHSHTSFHGECTTGSKSVTEEWIFYPCFFTTRLS----PNQVSSSQLVMNQFCYVLMT 152
H G +G KSV + + F T ++S N+ + + + N + V +
Sbjct: 168 RMHKGHVLVIGIGGSGKKSVIKLAAYAAGFRTFQISLSRGYNEPAFREDMKNLYNMVGVD 227
Query: 153 NESVYGIFVS 162
N+ + +F S
Sbjct: 228 NKKIVFLFTS 237
>UniRef100_UPI0000431DA9 UPI0000431DA9 UniRef100 entry
Length = 3158
Score = 31.6 bits (70), Expect = 8.1
Identities = 30/130 (23%), Positives = 57/130 (43%), Gaps = 9/130 (6%)
Query: 37 NSEEPYLFEGLLEIRKFYSLQGMNWFLLKYQGCSNFDLFIYNDSIEEVCYLTRSLPPSIT 96
N +EP +E LL+ YSL + + S + ++ND++E + + R+L
Sbjct: 1537 NEDEPRFYEDLLDYEAVYSLFLEIYEDYNERNVSKLHMVLFNDALEHLTRIHRAL----- 1591
Query: 97 PTTHSHTSFHGECTTGSKSVTEEWIFYPCFFTTRLS----PNQVSSSQLVMNQFCYVLMT 152
H G +G KSV + + F T ++S N+ + + + N + V +
Sbjct: 1592 RMHKGHVLVIGIGGSGKKSVIKLAAYAAGFRTFQISLSRGYNEPAFREDMKNLYNMVGVD 1651
Query: 153 NESVYGIFVS 162
N+ + +F S
Sbjct: 1652 NKKIVFLFTS 1661
>UniRef100_UPI0000431DA8 UPI0000431DA8 UniRef100 entry
Length = 2876
Score = 31.6 bits (70), Expect = 8.1
Identities = 30/130 (23%), Positives = 57/130 (43%), Gaps = 9/130 (6%)
Query: 37 NSEEPYLFEGLLEIRKFYSLQGMNWFLLKYQGCSNFDLFIYNDSIEEVCYLTRSLPPSIT 96
N +EP +E LL+ YSL + + S + ++ND++E + + R+L
Sbjct: 1205 NEDEPRFYEDLLDYEAVYSLFLEIYEDYNERNVSKLHMVLFNDALEHLTRIHRAL----- 1259
Query: 97 PTTHSHTSFHGECTTGSKSVTEEWIFYPCFFTTRLS----PNQVSSSQLVMNQFCYVLMT 152
H G +G KSV + + F T ++S N+ + + + N + V +
Sbjct: 1260 RMHKGHVLVIGIGGSGKKSVIKLAAYAAGFRTFQISLSRGYNEPAFREDMKNLYNMVGVD 1319
Query: 153 NESVYGIFVS 162
N+ + +F S
Sbjct: 1320 NKKIVFLFTS 1329
>UniRef100_UPI0000431DA7 UPI0000431DA7 UniRef100 entry
Length = 1931
Score = 31.6 bits (70), Expect = 8.1
Identities = 30/130 (23%), Positives = 57/130 (43%), Gaps = 9/130 (6%)
Query: 37 NSEEPYLFEGLLEIRKFYSLQGMNWFLLKYQGCSNFDLFIYNDSIEEVCYLTRSLPPSIT 96
N +EP +E LL+ YSL + + S + ++ND++E + + R+L
Sbjct: 317 NEDEPRFYEDLLDYEAVYSLFLEIYEDYNERNVSKLHMVLFNDALEHLTRIHRAL----- 371
Query: 97 PTTHSHTSFHGECTTGSKSVTEEWIFYPCFFTTRLS----PNQVSSSQLVMNQFCYVLMT 152
H G +G KSV + + F T ++S N+ + + + N + V +
Sbjct: 372 RMHKGHVLVIGIGGSGKKSVIKLAAYAAGFRTFQISLSRGYNEPAFREDMKNLYNMVGVD 431
Query: 153 NESVYGIFVS 162
N+ + +F S
Sbjct: 432 NKKIVFLFTS 441
>UniRef100_UPI0000431DA6 UPI0000431DA6 UniRef100 entry
Length = 2896
Score = 31.6 bits (70), Expect = 8.1
Identities = 30/130 (23%), Positives = 57/130 (43%), Gaps = 9/130 (6%)
Query: 37 NSEEPYLFEGLLEIRKFYSLQGMNWFLLKYQGCSNFDLFIYNDSIEEVCYLTRSLPPSIT 96
N +EP +E LL+ YSL + + S + ++ND++E + + R+L
Sbjct: 1209 NEDEPRFYEDLLDYEAVYSLFLEIYEDYNERNVSKLHMVLFNDALEHLTRIHRAL----- 1263
Query: 97 PTTHSHTSFHGECTTGSKSVTEEWIFYPCFFTTRLS----PNQVSSSQLVMNQFCYVLMT 152
H G +G KSV + + F T ++S N+ + + + N + V +
Sbjct: 1264 RMHKGHVLVIGIGGSGKKSVIKLAAYAAGFRTFQISLSRGYNEPAFREDMKNLYNMVGVD 1323
Query: 153 NESVYGIFVS 162
N+ + +F S
Sbjct: 1324 NKKIVFLFTS 1333
>UniRef100_UPI0000431DA5 UPI0000431DA5 UniRef100 entry
Length = 3334
Score = 31.6 bits (70), Expect = 8.1
Identities = 30/130 (23%), Positives = 57/130 (43%), Gaps = 9/130 (6%)
Query: 37 NSEEPYLFEGLLEIRKFYSLQGMNWFLLKYQGCSNFDLFIYNDSIEEVCYLTRSLPPSIT 96
N +EP +E LL+ YSL + + S + ++ND++E + + R+L
Sbjct: 1561 NEDEPRFYEDLLDYEAVYSLFLEIYEDYNERNVSKLHMVLFNDALEHLTRIHRAL----- 1615
Query: 97 PTTHSHTSFHGECTTGSKSVTEEWIFYPCFFTTRLS----PNQVSSSQLVMNQFCYVLMT 152
H G +G KSV + + F T ++S N+ + + + N + V +
Sbjct: 1616 RMHKGHVLVIGIGGSGKKSVIKLAAYAAGFRTFQISLSRGYNEPAFREDMKNLYNMVGVD 1675
Query: 153 NESVYGIFVS 162
N+ + +F S
Sbjct: 1676 NKKIVFLFTS 1685
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.322 0.136 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 279,969,012
Number of Sequences: 2790947
Number of extensions: 10980232
Number of successful extensions: 24587
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 16
Number of HSP's that attempted gapping in prelim test: 24580
Number of HSP's gapped (non-prelim): 21
length of query: 168
length of database: 848,049,833
effective HSP length: 118
effective length of query: 50
effective length of database: 518,718,087
effective search space: 25935904350
effective search space used: 25935904350
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0037a.10


