

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0036b.1
(122 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
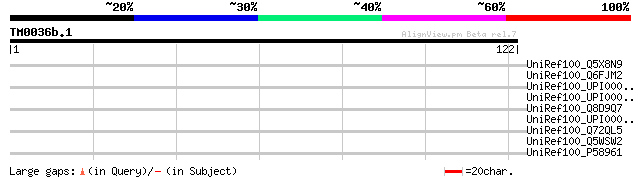
UniRef100_Q5X8N9 Hypothetical protein [Legionella pneumophila st... 31 5.6
UniRef100_Q6FJM2 Similar to sp|P36070 Saccharomyces cerevisiae Y... 31 5.6
UniRef100_UPI000042E14E UPI000042E14E UniRef100 entry 31 7.4
UniRef100_UPI000042DD1D UPI000042DD1D UniRef100 entry 31 7.4
UniRef100_Q8D9Q7 Hypothetical protein [Vibrio vulnificus] 31 7.4
UniRef100_UPI0000077E87 UPI0000077E87 UniRef100 entry 30 9.6
UniRef100_Q72QL5 Response regulator [Leptospira interrogans] 30 9.6
UniRef100_Q5WSW2 Hypothetical protein [Legionella pneumophila st... 30 9.6
UniRef100_P58961 Putative gustatory receptor 47b [Drosophila mel... 30 9.6
>UniRef100_Q5X8N9 Hypothetical protein [Legionella pneumophila str. Paris]
Length = 269
Score = 31.2 bits (69), Expect = 5.6
Identities = 13/39 (33%), Positives = 23/39 (58%)
Query: 7 WYEPNDSLRSDRFIVSLKDHSSTCNFSELLEILSRHVVA 45
W PN+ + D ++S+ D TC+ + LE LS+ ++A
Sbjct: 223 WTTPNEVVEQDLEMISIIDEVRTCDLKQSLENLSQPLIA 261
>UniRef100_Q6FJM2 Similar to sp|P36070 Saccharomyces cerevisiae YKL125w RRN3 Cter
[Candida glabrata]
Length = 547
Score = 31.2 bits (69), Expect = 5.6
Identities = 16/39 (41%), Positives = 24/39 (61%)
Query: 25 DHSSTCNFSELLEILSRHVVAGIFYKGLNIVLCILILQK 63
D ST NF+ LLE+LS ++ KGL ++ CI+ +K
Sbjct: 18 DRISTKNFTILLEVLSNNINKVDSSKGLPLIQCIINFEK 56
>UniRef100_UPI000042E14E UPI000042E14E UniRef100 entry
Length = 943
Score = 30.8 bits (68), Expect = 7.4
Identities = 16/56 (28%), Positives = 27/56 (47%), Gaps = 1/56 (1%)
Query: 47 IFYKGLNIVLCILILQKDKLIRTHTCTLLHQLMGKSKVFDANCYSYPPPLFRSLKI 102
+F ++ +CIL+ + H+ L H + SK F Y+Y P F+S+ I
Sbjct: 889 LFAMWFSLTVCILVFMEGTSAMLHSLRL-HWVEAMSKFFQGEGYAYEPFTFKSIDI 943
>UniRef100_UPI000042DD1D UPI000042DD1D UniRef100 entry
Length = 943
Score = 30.8 bits (68), Expect = 7.4
Identities = 16/56 (28%), Positives = 27/56 (47%), Gaps = 1/56 (1%)
Query: 47 IFYKGLNIVLCILILQKDKLIRTHTCTLLHQLMGKSKVFDANCYSYPPPLFRSLKI 102
+F ++ +CIL+ + H+ L H + SK F Y+Y P F+S+ I
Sbjct: 889 LFAMWFSLTVCILVFMEGTSAMLHSLRL-HWVEAMSKFFQGEGYAYEPFTFKSIDI 943
>UniRef100_Q8D9Q7 Hypothetical protein [Vibrio vulnificus]
Length = 207
Score = 30.8 bits (68), Expect = 7.4
Identities = 23/88 (26%), Positives = 40/88 (45%), Gaps = 20/88 (22%)
Query: 1 MSDELDWYEPNDSLRSDRFIVSLKDH----------------SSTCNFSELLEILSRHVV 44
M D++E S+ +R I+S++DH SS N S + E+LSRH+
Sbjct: 108 MGKAQDYFEGVKSIAVERGILSIRDHIEKVANIRNTILYASDSSLPNVSNVEEMLSRHIG 167
Query: 45 AGIFYKGLNIVLCILILQKDKLIRTHTC 72
A + + ++ +L++Q K C
Sbjct: 168 AALIIQ----MIYLLVVQNKKQNLVEQC 191
>UniRef100_UPI0000077E87 UPI0000077E87 UniRef100 entry
Length = 344
Score = 30.4 bits (67), Expect = 9.6
Identities = 25/92 (27%), Positives = 40/92 (43%), Gaps = 7/92 (7%)
Query: 8 YEPNDSLRSDRFIVSLKDHSSTCNF--SELLEILSRHVVAGIFYKGLNI-----VLCILI 60
Y+ +DS I L + S C+F +EL + AGIFY + V+C +
Sbjct: 166 YQADDSSEVHERIAYLFEMSKRCSFLLAELNGVFGFAAAAGIFYDFTIMTCFVYVICQKL 225
Query: 61 LQKDKLIRTHTCTLLHQLMGKSKVFDANCYSY 92
L+++ + LLH + KV + Y Y
Sbjct: 226 LEREPWDPEYVYMLLHVAIHTYKVVITSTYGY 257
>UniRef100_Q72QL5 Response regulator [Leptospira interrogans]
Length = 330
Score = 30.4 bits (67), Expect = 9.6
Identities = 12/40 (30%), Positives = 24/40 (60%)
Query: 4 ELDWYEPNDSLRSDRFIVSLKDHSSTCNFSELLEILSRHV 43
E D++E + + FI+ ++DH + + E+L+ L RH+
Sbjct: 229 ESDFFEIGYGIAENTFIIVIRDHFGSLDKKEILKRLDRHI 268
>UniRef100_Q5WSW2 Hypothetical protein [Legionella pneumophila str. Lens]
Length = 460
Score = 30.4 bits (67), Expect = 9.6
Identities = 22/70 (31%), Positives = 35/70 (49%), Gaps = 2/70 (2%)
Query: 48 FYKGLNIVLCILILQKDKLIRTHTCTLLHQLMGKSKVFDANCYSYPPPLFRSLKIGELSK 107
+Y+G + I+ K+KL ++ LHQL+ K+ + + P L L+I E
Sbjct: 332 YYRGKMVSAYEFIIAKNKLYQS--LRPLHQLLEKTDIVLTPALAQLPLLIGHLRIDEEFN 389
Query: 108 MPLFKKKKFS 117
L KKK+FS
Sbjct: 390 KYLQKKKEFS 399
>UniRef100_P58961 Putative gustatory receptor 47b [Drosophila melanogaster]
Length = 414
Score = 30.4 bits (67), Expect = 9.6
Identities = 25/92 (27%), Positives = 40/92 (43%), Gaps = 7/92 (7%)
Query: 8 YEPNDSLRSDRFIVSLKDHSSTCNF--SELLEILSRHVVAGIFYKGLNI-----VLCILI 60
Y+ +DS I L + S C+F +EL + AGIFY + V+C +
Sbjct: 218 YQADDSSEVHERIAYLFEMSKRCSFLLAELNGVFGFAAAAGIFYDFTIMTCFVYVICQKL 277
Query: 61 LQKDKLIRTHTCTLLHQLMGKSKVFDANCYSY 92
L+++ + LLH + KV + Y Y
Sbjct: 278 LEREPWDPEYVYMLLHVAIHTYKVVITSTYGY 309
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.324 0.140 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 200,470,494
Number of Sequences: 2790947
Number of extensions: 6868611
Number of successful extensions: 18107
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 18104
Number of HSP's gapped (non-prelim): 9
length of query: 122
length of database: 848,049,833
effective HSP length: 98
effective length of query: 24
effective length of database: 574,537,027
effective search space: 13788888648
effective search space used: 13788888648
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0036b.1


