

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0034b.2
(494 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
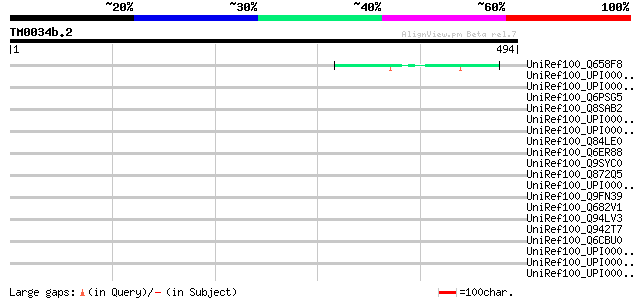
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q658F8 Hypothetical protein P0015E04.9 [Oryza sativa] 52 5e-05
UniRef100_UPI00004399A0 UPI00004399A0 UniRef100 entry 42 0.031
UniRef100_UPI000021E5EA UPI000021E5EA UniRef100 entry 42 0.031
UniRef100_Q6PSG5 Excretory/secretory protein Juv-p120 precursor ... 39 0.26
UniRef100_Q8SAB2 Putative gypsy-type retrotransposon RIRE2 prote... 39 0.34
UniRef100_UPI000049A2DE UPI000049A2DE UniRef100 entry 37 1.00
UniRef100_UPI00003642E5 UPI00003642E5 UniRef100 entry 37 1.7
UniRef100_Q84LE0 Phytocyanin protein, PUP2 [Arabidopsis thaliana] 36 2.2
UniRef100_Q6ER88 Translational activator protein-like [Oryza sat... 36 2.2
UniRef100_Q9SYC0 F11M15.4 protein [Arabidopsis thaliana] 36 2.2
UniRef100_Q872Q5 Hypothetical protein B13B3.200 [Neurospora crassa] 36 2.2
UniRef100_UPI00001CE674 UPI00001CE674 UniRef100 entry 36 2.9
UniRef100_Q9FN39 Similarity to phytocyanin/early nodulin-like pr... 36 2.9
UniRef100_Q682V1 Predicted GPI-anchored protein [Arabidopsis tha... 36 2.9
UniRef100_Q94LV3 Putative gypsy-type retrotransposon protein [Or... 36 2.9
UniRef100_Q942T7 P0560B06.7 protein [Oryza sativa] 35 3.8
UniRef100_Q6CBU0 Yarrowia lipolytica chromosome C of strain CLIB... 35 3.8
UniRef100_UPI00001CEB15 UPI00001CEB15 UniRef100 entry 35 5.0
UniRef100_UPI00002B2EF3 UPI00002B2EF3 UniRef100 entry 35 5.0
UniRef100_UPI0000278408 UPI0000278408 UniRef100 entry 35 5.0
>UniRef100_Q658F8 Hypothetical protein P0015E04.9 [Oryza sativa]
Length = 569
Score = 51.6 bits (122), Expect = 5e-05
Identities = 49/171 (28%), Positives = 66/171 (37%), Gaps = 24/171 (14%)
Query: 317 MFSELGIRLPFSPFVQTVLRDINAAPCQLHPNAWAFIRCFEILSAAVGIAPSP------- 369
M E +RLP PF VL AP QL PNAW + F L G+AP P
Sbjct: 92 MEEERLMRLPLQPFFAAVLAHFGLAPSQLTPNAWRLLAGFAALCRRAGVAPPPPPPLTVF 151
Query: 370 TSFFYLYDVDPKSIKNKGWISLKARAGRKCLHPHKSNAKSSFARKYFRVAVHPAYPEAFT 429
FF P + W S +AR S + S + V ++P + E F
Sbjct: 152 RHFFTAIGCPPGA-----WYSFRARG---------SASTGSLFTRLSNVTMNPWWKEQFI 197
Query: 430 LRDGTALF--PLYWTEKPNGITDPSEESLSANDKAFLNLLA-PLPILDCTT 477
L A + P+ W + P + + D A LLA ++D TT
Sbjct: 198 LVSSPAPWPCPVRWGKPRKHANFPPVLTTAEKDMATKLLLARGSSLIDLTT 248
>UniRef100_UPI00004399A0 UPI00004399A0 UniRef100 entry
Length = 274
Score = 42.4 bits (98), Expect = 0.031
Identities = 44/148 (29%), Positives = 65/148 (43%), Gaps = 20/148 (13%)
Query: 67 SISLFLRNRHFPFHN--RPIVRSNSP----LHATHLPKTIMTSNLPHASNTRHPS----- 115
SIS+F + H +H+ P + S P +H + P ++S P + HPS
Sbjct: 52 SISIFHPSIHIYYHSIHHPYLISIHPYLYSIHPSIHPS--ISSFHPSSIFNIHPSILPYL 109
Query: 116 -SFSPSPIKTLLPHSHPFLRPLFTSLIAYQTPSAHFRSGAVAYHPFRYPLFTSYSGESHC 174
SF PS I + P HP+L + S+ PS H + HP +P F Y H
Sbjct: 110 LSFHPSSIFNIHPSIHPYLYSIHPSIHPSIHPSIH-----PSIHPSIHPSFHIYYHSIHH 164
Query: 175 PYLCITHILIFSF-LSLHCLLKMASNPT 201
PYL H I F S+H + + +P+
Sbjct: 165 PYLISIHPSISIFHPSIHPSIHPSIHPS 192
>UniRef100_UPI000021E5EA UPI000021E5EA UniRef100 entry
Length = 426
Score = 42.4 bits (98), Expect = 0.031
Identities = 48/176 (27%), Positives = 70/176 (39%), Gaps = 21/176 (11%)
Query: 90 PLHATHLPKTIMTSNLPHASNTRHPSSFSPSPIKTLLPHSHPFLRPLFTSLIAYQTPSAH 149
P H T +P P S T H S PS + T P SH PL + A+++P +H
Sbjct: 80 PSHCTQVPLPHTAHRFP--SLTLHTGS-PPSSLHTGSPPSHCTQVPLPHT--AHRSPPSH 134
Query: 150 FRSGAVAYHPFRYPLFTSYSGE--SHCPYLCITH-ILIFSFLSLHCLLKMASNPTSPNHS 206
+ + + R P T Y+G SHC + + H + F LSLH + P+H
Sbjct: 135 YTQAPLPHTAHRVPSLTLYTGSPPSHCTQVPLPHTVHRFPSLSLH-------TGSPPSHC 187
Query: 207 S------SSSGSDPDATAAGGLRLEAPEIPPYRLKTYATLPLPVQIAPTHEAIDQP 256
+ ++ S P G L A +P L T + Q+ H A P
Sbjct: 188 TQVPLPHTAHRSPPSHCTQGPLPHTAHRVPSLTLYTGSPPSHCTQVPLPHTAHRSP 243
Score = 37.7 bits (86), Expect = 0.77
Identities = 43/157 (27%), Positives = 57/157 (35%), Gaps = 29/157 (18%)
Query: 90 PLHATHLPKTIMTSNLPHASNTRHPSSFSPSPIKTLLPHSHPFLRPLFTSLIAYQTPSAH 149
P H T P LP S T H S + LPH+ A+++P +H
Sbjct: 11 PSHYTQAPLPHTIHRLP--SLTLHTGSPPSHCTQVPLPHT------------AHRSPPSH 56
Query: 150 FRSGAVAYHPFRYPLFTSYSGE--SHCPYLCITHIL-IFSFLSLHCLLKMASNPTSPNHS 206
+ + + R P T Y+G SHC + + H F L+LH T S
Sbjct: 57 YTQAPLPHTAHRVPSLTLYTGSPPSHCTQVPLPHTAHRFPSLTLH---------TGSPPS 107
Query: 207 SSSSGSDPDATAAGGLRLEAPEIPPYRLKTYATLPLP 243
S +GS P L A PP Y PLP
Sbjct: 108 SLHTGSPPSHCTQVPLPHTAHRSPP---SHYTQAPLP 141
>UniRef100_Q6PSG5 Excretory/secretory protein Juv-p120 precursor [Litomosoides
sigmodontis]
Length = 719
Score = 39.3 bits (90), Expect = 0.26
Identities = 55/217 (25%), Positives = 82/217 (37%), Gaps = 17/217 (7%)
Query: 54 PPHVAAPRDASVTSISLFLRNRHFPFHNRPIVRSNSPLHATHLPKTIMTSNLPHA----S 109
P H PR +S ++ + F HFP+ + +P TH P I +S A +
Sbjct: 64 PTHFPYPRFSSTSTAAPFPPT-HFPYPRFSSTSTAAPFSPTHFPYPIFSSTSTAAPFPPT 122
Query: 110 NTRHPSSFSPSPIKTLLPHSHPFLRPLFTSLIAYQTPSAHF-------RSGAVAYHPFRY 162
+ +PS S S P P+ R TS A +P+ HF S A + P +
Sbjct: 123 HFPYPSFSSTSTAAPFPPTHFPYPRFSSTSTAAPFSPT-HFPYPIFSSTSTAAPFPPTHF 181
Query: 163 PLFTSYSGESHCPYLCITHILIFSFLSLHCLLKMASNPTSPNHSSSSSGSDPDATAAGGL 222
P + S+S S TH SF S + S T P + + + +
Sbjct: 182 P-YPSFSSTSTAAPFPPTHFPYPSFSSTS--TAVTSPRTFPYPRTFGTTPVFNHSVTFPT 238
Query: 223 RLEAPEIPPYRLKTYATLPLPVQIAPTHEAIDQPSIF 259
P IP + L T+ T P PT + P+ F
Sbjct: 239 AFPYPGIPTFPLLTFPT-AFPYPGIPTFPPVTFPTAF 274
>UniRef100_Q8SAB2 Putative gypsy-type retrotransposon RIRE2 protein [Sorghum bicolor]
Length = 447
Score = 38.9 bits (89), Expect = 0.34
Identities = 37/119 (31%), Positives = 52/119 (43%), Gaps = 7/119 (5%)
Query: 318 FSELGIRLPFSPFVQTVLRDINAAPCQLHPNAWAFIRCFEILSAA-VGIAPSPTSFFYLY 376
F + G +P PF+Q +L C LH N+ I F LS A VGI P F YL+
Sbjct: 66 FLKRGFGVPVHPFLQGLLLYYEIGICNLHRNSILLISTFIHLSEAYVGIEPHFDLFRYLF 125
Query: 377 DVDPKSI----KNKGWISLKARAGRK--CLHPHKSNAKSSFARKYFRVAVHPAYPEAFT 429
+ K K G + L R G K LH + + + + RK+ + A P + T
Sbjct: 126 CLRKKGAVGGSKVAGGVYLNLRDGMKNRYLHCPWNTSLTEWYRKWSTSGKNRAAPRSAT 184
>UniRef100_UPI000049A2DE UPI000049A2DE UniRef100 entry
Length = 492
Score = 37.4 bits (85), Expect = 1.00
Identities = 35/149 (23%), Positives = 66/149 (43%), Gaps = 7/149 (4%)
Query: 65 VTSISLFLRNRHFPFHNR--PIVRSNSPLHATHLPKTIMTSNLPHASNTRHPSSFSPSPI 122
V S S+ + + P H+ P+ S+ P+H++ +P + +S+ P S++ HP S P+
Sbjct: 235 VHSSSVPVHSSSHPVHSSSTPVQSSSHPVHSSSVP--VHSSSTPVQSSS-HPVHSSSHPV 291
Query: 123 KTLLPHSHPFLRPLFTSLIAYQTPSAHFRSGAVAYHPFRYPLFTSYS--GESHCPYLCIT 180
++ H P+ +S + + S S + H P+ +S S P +
Sbjct: 292 QSSSHPVHSSSVPVHSSSVPVHSSSVPVHSSSHPVHSSSVPVHSSSHPVHSSSTPVQSSS 351
Query: 181 HILIFSFLSLHCLLKMASNPTSPNHSSSS 209
H + S + + + +SP H SSS
Sbjct: 352 HPVHSSSVPVQSSSTPVHSSSSPKHGSSS 380
Score = 37.0 bits (84), Expect = 1.3
Identities = 35/154 (22%), Positives = 64/154 (40%), Gaps = 20/154 (12%)
Query: 70 LFLRNRHFPFHNR----PIVRSNSPLHATHLP-----KTIMTSNLPHASNTR------HP 114
+ +R RH P S+SP+H++ +P + +S++P S++ HP
Sbjct: 98 VIIRYRHIVIKEESSYIPTTSSSSPVHSSSVPVHSSSHPVRSSSIPIRSSSTPVKSSSHP 157
Query: 115 SSFSPSPIKTLLPHSHPFLRPLFTSLIAYQTPSAHFRSGAVAYHPFRYPLFTSYSGESHC 174
S +P+K+ H P+ +S Q+ S S + H +P+ + S
Sbjct: 158 VHSSSTPVKSSSVPIHSSSHPVHSSSTPVQSSSHPVHSSSHPVHSSSHPVHS-----SSV 212
Query: 175 PYLCITHILIFSFLSLHCLLKMASNPTSPNHSSS 208
P +H + S + +H + + P HSSS
Sbjct: 213 PVHSSSHPVHSSSVPVHSSSHPVHSSSVPVHSSS 246
>UniRef100_UPI00003642E5 UPI00003642E5 UniRef100 entry
Length = 256
Score = 36.6 bits (83), Expect = 1.7
Identities = 46/181 (25%), Positives = 74/181 (40%), Gaps = 23/181 (12%)
Query: 44 PYFKTLLNLYPPHVAAPRDASVTSISLFLRNRHFPFHNRPIVRSNSPLHATHLPKTIMTS 103
P+ TL +L PH + S +L L + P H+ + S++ H P T+ S
Sbjct: 11 PHTLTLSHLSHPHTLSHPHTLSHSHTLTLSHSLTPSHSHTLSHSHTHSHTLSHPHTLSHS 70
Query: 104 NL-----PHA---SNT-----RHPSSFSPSPIKTLLPHSHPFLRPLFTSLIAYQTPSAHF 150
+ PH S+T HP + +PS T P SH L P + +++ +H
Sbjct: 71 HTHSHTHPHTLTLSHTLTHSHTHPHTHTPSHTLTPSPLSHS-LTPSHSHTLSHSHTHSHT 129
Query: 151 RSGAVAYHPFRYPLFTSYSGESHCPYLCITHILIFSFLSLHCLLKMASNPT-SPNHSSSS 209
H +P S++ SH P +H L + H L + P+ +P H+ S
Sbjct: 130 HP-----HTLSHPHTHSHT-HSHTPSHTHSHTLTHTL--THTLTHSSHTPSHTPAHTLSH 181
Query: 210 S 210
S
Sbjct: 182 S 182
>UniRef100_Q84LE0 Phytocyanin protein, PUP2 [Arabidopsis thaliana]
Length = 370
Score = 36.2 bits (82), Expect = 2.2
Identities = 29/92 (31%), Positives = 41/92 (44%), Gaps = 3/92 (3%)
Query: 59 APRDASVTSISLFLRNRHFPFHNRPIVRSNSPLHA-THLPKTIMTSNLPHASN--TRHPS 115
+P+ S S S H P H+ S+SP HA +H P + + HA + H
Sbjct: 210 SPKSPSPVSHSPSHSPAHTPSHSPAHTPSHSPAHAPSHSPAHAPSHSPAHAPSHSPAHSX 269
Query: 116 SFSPSPIKTLLPHSHPFLRPLFTSLIAYQTPS 147
S SP+ K+ P S P P S + Q+PS
Sbjct: 270 SHSPATPKSPSPSSSPAQSPATPSPMTPQSPS 301
>UniRef100_Q6ER88 Translational activator protein-like [Oryza sativa]
Length = 920
Score = 36.2 bits (82), Expect = 2.2
Identities = 34/133 (25%), Positives = 53/133 (39%), Gaps = 11/133 (8%)
Query: 64 SVTSISLFLRNRHFPFHNRPIVRSNSPLHATHLPKTIMTSNLPHASNTRHPSS---FSPS 120
S+ +++ L N P RS P A + P T+ ++P SN HP + P+
Sbjct: 508 SLRTVAAILPNNGLPEFE---ARSAEPSEAANAPPTV---SVPAGSNATHPVTPVPQPPA 561
Query: 121 PIKTLLPHSHPFLRPLFTSLIAYQTPSAHFRSGAVAYHPFRYPLFTSYSGESHCPYLCIT 180
P L P + F SL+ + SA + +A YP+ + + Y I
Sbjct: 562 PAADQLQDMIPMQKEAFISLLKAISTSAASATATIAIFVVSYPI--KWPDKVSVNYQMIL 619
Query: 181 HILIFSFLSLHCL 193
L+F F L L
Sbjct: 620 EALLFVFFPLALL 632
>UniRef100_Q9SYC0 F11M15.4 protein [Arabidopsis thaliana]
Length = 663
Score = 36.2 bits (82), Expect = 2.2
Identities = 35/141 (24%), Positives = 54/141 (37%), Gaps = 12/141 (8%)
Query: 313 AYEYMFSELGIRLPFSPFVQTVLRDINAAPCQLHPNAWAFIRCFEILSAAVGIAPSPTSF 372
A+E FSE G+ P + ++ + A QL PN + C L+ G + + F
Sbjct: 44 AFEIFFSECGLFFPLPDILIDLMHEFGIALPQLCPNVIRTVLCLLTLAEEDGFRLALSDF 103
Query: 373 FYLYDVDPKSIKNKGWISLKARAGRKCLH--PHKSNAKSSFARKYFRVAVHPAYPEAFTL 430
LY V K + L R G + P K + + YF V+ T
Sbjct: 104 LQLYAV--KKSRTNNTFFLSPRKGFRVFEDFPEKD---EHWRKSYFFFPVN-----ELTY 153
Query: 431 RDGTALFPLYWTEKPNGITDP 451
+ + LF WT + +T P
Sbjct: 154 EEKSDLFVSDWTTRAVKVTIP 174
>UniRef100_Q872Q5 Hypothetical protein B13B3.200 [Neurospora crassa]
Length = 550
Score = 36.2 bits (82), Expect = 2.2
Identities = 32/118 (27%), Positives = 45/118 (38%), Gaps = 12/118 (10%)
Query: 97 PKTIMTSNLPHASNTRHPSSFSPSPIKTLLPHSHPFLRPLFTSLIAYQTPSAHFRSGAVA 156
P T +S+ P +T P S S + T P +P + + S H S +
Sbjct: 89 PSTSGSSSHPRPPSTSKPPSISSTTTTTSTSDLKPNPKPKAFLRLLHIPHSPHRSSSLLI 148
Query: 157 YHPFRYPLFTSYSGESHCPYLCITHILIFSFLSLHCLLKMASNPTSPNHSSSSSGSDP 214
P LF + + H YL TH F L+ H NHSS+S+ S P
Sbjct: 149 PPPLLLRLFCLLNLDPHALYLHHTHTPGFHVLNCH------------NHSSTSTSSSP 194
>UniRef100_UPI00001CE674 UPI00001CE674 UniRef100 entry
Length = 1024
Score = 35.8 bits (81), Expect = 2.9
Identities = 31/99 (31%), Positives = 44/99 (44%), Gaps = 15/99 (15%)
Query: 44 PYFKTLLNLYPPHVAAPRDASVTSISLFLRNR-HFPFHNRPIVR---------SNSPLHA 93
P + L + PP +A R ++ IS + R R H PFH P VR + SP A
Sbjct: 554 PVDRPKLFITPPEGSARR-RTIHGISSYKREREHIPFHVSPGVRRTLSPSNLKARSPAPA 612
Query: 94 THLPKTIMTSNLPHASNTRHP-SSFSPSPIKTLLPHSHP 131
P + ++PH T P SS +P P++ S P
Sbjct: 613 RLWPP---SKSMPHLPGTPRPASSLTPGPVRAAASSSSP 648
>UniRef100_Q9FN39 Similarity to phytocyanin/early nodulin-like protein [Arabidopsis
thaliana]
Length = 370
Score = 35.8 bits (81), Expect = 2.9
Identities = 29/92 (31%), Positives = 41/92 (44%), Gaps = 3/92 (3%)
Query: 59 APRDASVTSISLFLRNRHFPFHNRPIVRSNSPLHA-THLPKTIMTSNLPHA--SNTRHPS 115
+P+ S S S H P H+ S+SP HA +H P + + HA + H
Sbjct: 210 SPKSPSPVSHSPSHSPAHTPSHSPAHTPSHSPAHAPSHSPAHAPSHSPAHAPSHSPAHSP 269
Query: 116 SFSPSPIKTLLPHSHPFLRPLFTSLIAYQTPS 147
S SP+ K+ P S P P S + Q+PS
Sbjct: 270 SHSPATPKSPSPSSSPAQSPATPSPMTPQSPS 301
>UniRef100_Q682V1 Predicted GPI-anchored protein [Arabidopsis thaliana]
Length = 370
Score = 35.8 bits (81), Expect = 2.9
Identities = 29/92 (31%), Positives = 41/92 (44%), Gaps = 3/92 (3%)
Query: 59 APRDASVTSISLFLRNRHFPFHNRPIVRSNSPLHA-THLPKTIMTSNLPHA--SNTRHPS 115
+P+ S S S H P H+ S+SP HA +H P + + HA + H
Sbjct: 210 SPKSPSPVSHSPSHSPAHTPSHSPAHTPSHSPAHAPSHSPAHAPSHSPAHAPSHSPAHSP 269
Query: 116 SFSPSPIKTLLPHSHPFLRPLFTSLIAYQTPS 147
S SP+ K+ P S P P S + Q+PS
Sbjct: 270 SHSPATPKSPSPSSSPAQSPATPSPMTPQSPS 301
>UniRef100_Q94LV3 Putative gypsy-type retrotransposon protein [Oryza sativa]
Length = 742
Score = 35.8 bits (81), Expect = 2.9
Identities = 32/128 (25%), Positives = 58/128 (45%), Gaps = 5/128 (3%)
Query: 322 GIRLPFSPFVQTVLRDINAAPCQLHPNAWAFIRCF-EILSAAVGIAPSPTSFFYLYDVD- 379
G P S F+ VL + L+PN+ AF+ F + A +G+ P F + Y++
Sbjct: 27 GFLPPPSEFLLLVLNFYGLSLLHLNPNSIAFLSIFVHLCEAYIGVEPFLDLFRFYYELRW 86
Query: 380 PKSIKNKGWISLKARAGRKCLH-PHK-SNAKSSFARKYFRVAVHPAYPEAFTLRDGTALF 437
+S + G + + R G K + P + +++S + ++F + + + P D
Sbjct: 87 MESNRVSGCVGFRLRDGMKSRYIPFQCPSSRSKWRNRWFYLEIKDSNPVFVVPEDQPDKI 146
Query: 438 PLYWTEKP 445
P WT KP
Sbjct: 147 PA-WTAKP 153
>UniRef100_Q942T7 P0560B06.7 protein [Oryza sativa]
Length = 917
Score = 35.4 bits (80), Expect = 3.8
Identities = 29/89 (32%), Positives = 38/89 (42%), Gaps = 9/89 (10%)
Query: 295 RPWLHKPDGDVKRSHFFFAYEYMFSELGIRLPFSPFVQTVLRDINAAPCQLHPNAWAFIR 354
RP H P RS FF F+ G+ LPFS F VL + L PNA +
Sbjct: 47 RPAPHYPG----RSVFFLP----FAMAGLVLPFSSFFMDVLEFYDLQMAHLTPNAVMTLA 98
Query: 355 CF-EILSAAVGIAPSPTSFFYLYDVDPKS 382
F + +G+ PS F + + V P S
Sbjct: 99 IFAHLCEMFIGVRPSLRLFRWFFTVQPVS 127
>UniRef100_Q6CBU0 Yarrowia lipolytica chromosome C of strain CLIB99 of Yarrowia
lipolytica [Yarrowia lipolytica]
Length = 812
Score = 35.4 bits (80), Expect = 3.8
Identities = 55/213 (25%), Positives = 91/213 (41%), Gaps = 22/213 (10%)
Query: 85 VRSNSPLHATHLPKTIMTSNLPHASNTRHPSSFSPSPIKTLLPHSHPFLRPLFTSLIAYQ 144
V S+SP+ ++ +S+ P +S++ PSS SPS + S P +S +
Sbjct: 388 VSSSSPVSSSS-----PSSSNPGSSSSSSPSSSSPSSSSSSPSSSSSSSSPSSSSSSSSS 442
Query: 145 TPSAHFRSGAVAYHPFRYPLFTSYSGESHCPYLCITHILIFSFLSLHCLLKMASNPTSPN 204
+PS+ S + + +S S S P + S S +S+ TS +
Sbjct: 443 SPSSSSSSSSPS---------SSSSFSSSSPSSSSSSSSSSSSSSSPSASSSSSSSTSLS 493
Query: 205 HSSSSSGSDPDATAAGGLRLEAPEIPPY---RLKTYATLPLPVQIAPTHEAIDQPSIFQN 261
SSSSS S ++++ L L + P+ R+ + P Q E I Q + N
Sbjct: 494 SSSSSSSS---SSSSAPLPLVTQQGMPWGLSRISHQSPEMAPYQDPMVGEYIHQQFVDPN 550
Query: 262 SDLIVQFVNSLGGLSNDDEFSAKLRIWICSPGD 294
++V V+S G N D F+ K IW+ + D
Sbjct: 551 PKVVVYVVDS-GVNINHDNFATK-PIWLANYAD 581
>UniRef100_UPI00001CEB15 UPI00001CEB15 UniRef100 entry
Length = 798
Score = 35.0 bits (79), Expect = 5.0
Identities = 44/189 (23%), Positives = 70/189 (36%), Gaps = 36/189 (19%)
Query: 53 YPPHVAAPRDASVTSISLFLRNRHFPFHN--RPIVRSNSPLHATHLPKTIMTSNLPHASN 110
+PP ++APR +S+ R +P R S + ATHLP N
Sbjct: 429 WPPPLSAPRGPYHSSVVSATRPMVISATRPTQPSARKTSVISATHLPL-----------N 477
Query: 111 TRHPSSFSPSPIKTLLP-HSHPFLRPLFTSLIAYQTPSAHFRSGAVAYHPFRYP------ 163
HP + +P+ +LP H P ++ + L P H A HP + P
Sbjct: 478 PVHPPALAPTTPPAVLPEHQIPKIKASYPDL-----PFGHKPGITSATHPAQPPPHQPPI 532
Query: 164 LFTSYSG---ESHCPYLCITHILIFSFLSLHCLLKMASNPTSPNHSSSSSGSDPDATAAG 220
+ T Y P TH + L + + P ++S +G P
Sbjct: 533 ISTKYPQVFPPQQAPMSPDTHTITN--------LPLIPSHLDPGDTTSQAGHHPLLPDVP 584
Query: 221 GLRLEAPEI 229
G+R +AP++
Sbjct: 585 GIRTQAPQV 593
>UniRef100_UPI00002B2EF3 UPI00002B2EF3 UniRef100 entry
Length = 208
Score = 35.0 bits (79), Expect = 5.0
Identities = 31/94 (32%), Positives = 40/94 (41%), Gaps = 5/94 (5%)
Query: 83 PIVRSNSPLHATHLPKTIMTSNLPHASNTRHPSS-----FSPSPIKTLLPHSHPFLRPLF 137
P R +P T P T+ S P AS+T HP S +PSP +L PF R
Sbjct: 67 PQPRPLTPKQLTLQPATLQPSPNPSASSTPHPFSPYPLNRAPSPQNSLPFSQQPFNRAQT 126
Query: 138 TSLIAYQTPSAHFRSGAVAYHPFRYPLFTSYSGE 171
+TPSA S A + YP ++ S E
Sbjct: 127 PQPAPPRTPSAPTPSTAPPHPKTAYPSASNPSTE 160
>UniRef100_UPI0000278408 UPI0000278408 UniRef100 entry
Length = 172
Score = 35.0 bits (79), Expect = 5.0
Identities = 27/81 (33%), Positives = 36/81 (44%), Gaps = 5/81 (6%)
Query: 368 SPTSFFYLYDVDPKSIKNKGWISLKARAGRKCLHPHKSNAKSSFARKYFRVAVHPAYPEA 427
S TS YL D+ K G +G K + K + + F + YF + YP A
Sbjct: 83 SNTSLSYLRDIVGFIEKRYG--QADEISGVKVIDDKKISIEIDFPKAYFLAKL--TYPTA 138
Query: 428 FTL-RDGTALFPLYWTEKPNG 447
F + +D L P WT KPNG
Sbjct: 139 FVIDKDQVELNPRNWTRKPNG 159
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.322 0.138 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 895,596,508
Number of Sequences: 2790947
Number of extensions: 39601291
Number of successful extensions: 93142
Number of sequences better than 10.0: 35
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 30
Number of HSP's that attempted gapping in prelim test: 93068
Number of HSP's gapped (non-prelim): 80
length of query: 494
length of database: 848,049,833
effective HSP length: 131
effective length of query: 363
effective length of database: 482,435,776
effective search space: 175124186688
effective search space used: 175124186688
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 77 (34.3 bits)
Lotus: description of TM0034b.2


