

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0033b.5
(88 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
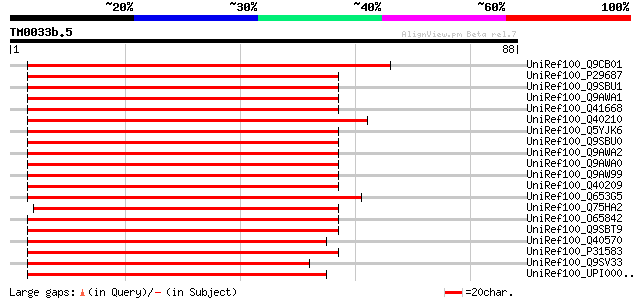
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9CB01 Ara6 [Arabidopsis thaliana] 76 2e-13
UniRef100_P29687 Ras-related protein Rab5 [Nicotiana tabacum] 72 3e-12
UniRef100_Q9SBU1 GTP binding protein [Cichorium intybus x Cichor... 72 3e-12
UniRef100_Q9AWA1 GTP binding protein [Cichorium intybus x Cichor... 72 3e-12
UniRef100_Q41668 Guanine nucleotide regulatory protein [Vicia faba] 72 4e-12
UniRef100_Q40210 RAB5B [Lotus japonicus] 72 4e-12
UniRef100_Q5YJK6 Ras-related protein [Hyacinthus orientalis] 71 6e-12
UniRef100_Q9SBU0 GTP binding protein [Cichorium intybus x Cichor... 70 1e-11
UniRef100_Q9AWA2 GTP binding protein [Cichorium intybus x Cichor... 70 1e-11
UniRef100_Q9AWA0 GTP binding protein [Cichorium intybus x Cichor... 70 1e-11
UniRef100_Q9AW99 GTP binding protein [Cichorium intybus x Cichor... 70 1e-11
UniRef100_Q40209 RAB5A [Lotus japonicus] 70 1e-11
UniRef100_Q653G5 Putative GTP binding protein [Oryza sativa] 70 2e-11
UniRef100_Q75HA2 Putative small GTP-binding protein [Oryza sativa] 70 2e-11
UniRef100_O65842 Small GTP-binding protein [Mesembryanthemum cry... 69 2e-11
UniRef100_Q9SBT9 GTP binding protein [Cichorium intybus x Cichor... 69 4e-11
UniRef100_Q40570 Ras-related GTP-binding protein [Nicotiana taba... 68 5e-11
UniRef100_P31583 Ras-related protein RHN1 [Nicotiana plumbaginif... 68 6e-11
UniRef100_Q9SV33 Small GTP-binding protein-like [Arabidopsis tha... 67 8e-11
UniRef100_UPI00002FAACD UPI00002FAACD UniRef100 entry 67 1e-10
>UniRef100_Q9CB01 Ara6 [Arabidopsis thaliana]
Length = 202
Score = 76.3 bits (186), Expect = 2e-13
Identities = 37/63 (58%), Positives = 47/63 (73%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGRGFESPG 63
+VMALV NK+DL KREV E+G + A++NGMF++ETSAKT +NIN LF EIG+ P
Sbjct: 140 IVMALVGNKADLHEKREVPTEDGMELAEKNGMFFIETSAKTADNINQLFEEIGKRLPRPA 199
Query: 64 RSS 66
SS
Sbjct: 200 PSS 202
>UniRef100_P29687 Ras-related protein Rab5 [Nicotiana tabacum]
Length = 200
Score = 72.4 bits (176), Expect = 3e-12
Identities = 32/54 (59%), Positives = 43/54 (79%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
+VMAL NK+DLE KR+V EE +A+ENG+F+METSAKT N+ND+F+EI +
Sbjct: 116 MVMALAGNKADLEDKRKVTAEEARLYAEENGLFFMETSAKTATNVNDIFYEIAK 169
>UniRef100_Q9SBU1 GTP binding protein [Cichorium intybus x Cichorium endivia]
Length = 200
Score = 72.0 bits (175), Expect = 3e-12
Identities = 32/54 (59%), Positives = 43/54 (79%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
+VMAL NK+DLE KR+V +EE +A+ENG+F+METSAKT N+ND+F EI +
Sbjct: 116 MVMALAGNKADLEDKRKVTVEEARVYAEENGLFFMETSAKTAANVNDVFHEIAK 169
>UniRef100_Q9AWA1 GTP binding protein [Cichorium intybus x Cichorium endivia]
Length = 196
Score = 72.0 bits (175), Expect = 3e-12
Identities = 33/54 (61%), Positives = 42/54 (77%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
+VMAL NK+DLE KREV EE +A+ENG+F+METSAKT N+ND+F EI +
Sbjct: 112 MVMALAGNKADLEDKREVTAEEARVYAEENGLFFMETSAKTAANVNDVFHEIAK 165
>UniRef100_Q41668 Guanine nucleotide regulatory protein [Vicia faba]
Length = 200
Score = 71.6 bits (174), Expect = 4e-12
Identities = 32/54 (59%), Positives = 43/54 (79%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
+VMAL NKSDLE KR+V EE +A+ENG+F+METSAK+ N+ND+F+EI +
Sbjct: 116 MVMALTGNKSDLEDKRKVTAEEARVYAEENGLFFMETSAKSAANVNDVFYEIAK 169
>UniRef100_Q40210 RAB5B [Lotus japonicus]
Length = 200
Score = 71.6 bits (174), Expect = 4e-12
Identities = 33/59 (55%), Positives = 46/59 (77%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGRGFESP 62
+VMALV NK+DL KREV +++G +A++NGMF++ETSAKT +NIN+LF EI + P
Sbjct: 139 IVMALVGNKADLLEKREVAVQDGIDYAEKNGMFFIETSAKTADNINELFEEIAKRLPRP 197
>UniRef100_Q5YJK6 Ras-related protein [Hyacinthus orientalis]
Length = 96
Score = 71.2 bits (173), Expect = 6e-12
Identities = 33/54 (61%), Positives = 42/54 (77%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
+V ALV NKSDLE KR+V EE +A ENG+F++ETSAKT N+ND+F+EI R
Sbjct: 13 MVTALVGNKSDLEEKRKVAAEEAHAYANENGLFFLETSAKTAINVNDVFYEIAR 66
>UniRef100_Q9SBU0 GTP binding protein [Cichorium intybus x Cichorium endivia]
Length = 200
Score = 70.5 bits (171), Expect = 1e-11
Identities = 32/54 (59%), Positives = 42/54 (77%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
+VMAL NK+DLE KR+V EE +A+ENG+F+METSAKT N+ND+F EI +
Sbjct: 116 MVMALAGNKADLEDKRKVTAEEARVYAEENGLFFMETSAKTAANVNDVFHEIAK 169
>UniRef100_Q9AWA2 GTP binding protein [Cichorium intybus x Cichorium endivia]
Length = 196
Score = 70.5 bits (171), Expect = 1e-11
Identities = 32/54 (59%), Positives = 42/54 (77%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
+VMAL NK+DLE KR+V EE +A+ENG+F+METSAKT N+ND+F EI +
Sbjct: 112 MVMALAGNKADLEDKRKVTAEEARVYAEENGLFFMETSAKTAANVNDVFHEIAK 165
>UniRef100_Q9AWA0 GTP binding protein [Cichorium intybus x Cichorium endivia]
Length = 200
Score = 70.5 bits (171), Expect = 1e-11
Identities = 32/54 (59%), Positives = 42/54 (77%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
+VMAL NK+DLE KR+V EE +A+ENG+F+METSAKT N+ND+F EI +
Sbjct: 116 MVMALAGNKADLEDKRKVTAEEARVYAEENGLFFMETSAKTAANVNDVFHEIAK 169
>UniRef100_Q9AW99 GTP binding protein [Cichorium intybus x Cichorium endivia]
Length = 200
Score = 70.5 bits (171), Expect = 1e-11
Identities = 32/54 (59%), Positives = 42/54 (77%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
+VMAL NK+DLE KR+V EE +A+ENG+F+METSAKT N+ND+F EI +
Sbjct: 116 MVMALAGNKADLEDKRKVTAEEARVYAEENGLFFMETSAKTAANVNDVFHEIAK 169
>UniRef100_Q40209 RAB5A [Lotus japonicus]
Length = 200
Score = 70.5 bits (171), Expect = 1e-11
Identities = 31/54 (57%), Positives = 42/54 (77%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
+VMA NKSDLE KR+V +E +A+ENG+F+METSAKT N+ND+F+EI +
Sbjct: 116 MVMAFAGNKSDLEDKRKVTADEARVYAEENGLFFMETSAKTAANVNDVFYEIAK 169
>UniRef100_Q653G5 Putative GTP binding protein [Oryza sativa]
Length = 226
Score = 69.7 bits (169), Expect = 2e-11
Identities = 30/58 (51%), Positives = 46/58 (78%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGRGFES 61
LVMALV NK DLE KR+V +E ++A+ NG+F++ETSAKT +N+ +LF+E+G+ + +
Sbjct: 97 LVMALVGNKVDLEEKRQVGTQEAMEYAERNGLFFIETSAKTSQNVTELFYELGKHYRN 154
>UniRef100_Q75HA2 Putative small GTP-binding protein [Oryza sativa]
Length = 203
Score = 69.7 bits (169), Expect = 2e-11
Identities = 31/53 (58%), Positives = 42/53 (78%)
Query: 5 VMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
V+AL NK+DL R+V+IEE +AQENG+F+METSAKT N+ND+F+EI +
Sbjct: 118 VVALAGNKADLLETRQVQIEEAKTYAQENGLFFMETSAKTATNVNDIFYEIAK 170
>UniRef100_O65842 Small GTP-binding protein [Mesembryanthemum crystallinum]
Length = 201
Score = 69.3 bits (168), Expect = 2e-11
Identities = 30/54 (55%), Positives = 45/54 (82%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
++MALV NK+DL+ +REV ++G ++A++NGMF++ETSAKT +NIN LF EI +
Sbjct: 140 IIMALVGNKADLQERREVPAQDGIEYAEKNGMFFIETSAKTADNINQLFEEIAK 193
>UniRef100_Q9SBT9 GTP binding protein [Cichorium intybus x Cichorium endivia]
Length = 126
Score = 68.6 bits (166), Expect = 4e-11
Identities = 31/54 (57%), Positives = 42/54 (77%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
+VMAL NK+DLE KR+V EE +A+ENG+F+METSAKT ++ND+F EI +
Sbjct: 42 MVMALAGNKADLEDKRKVTAEEARVYAEENGLFFMETSAKTAADVNDVFHEIAK 95
>UniRef100_Q40570 Ras-related GTP-binding protein [Nicotiana tabacum]
Length = 200
Score = 68.2 bits (165), Expect = 5e-11
Identities = 31/52 (59%), Positives = 40/52 (76%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEI 55
+VMAL NK+DL R+V EE +AQENG+F+METSAKT N+ND+F+EI
Sbjct: 116 MVMALAGNKADLLDARKVAAEEAQTYAQENGLFFMETSAKTASNVNDIFYEI 167
>UniRef100_P31583 Ras-related protein RHN1 [Nicotiana plumbaginifolia]
Length = 200
Score = 67.8 bits (164), Expect = 6e-11
Identities = 30/54 (55%), Positives = 42/54 (77%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
+VMAL NK+DLE +R+V EE +A+ENG+F+METSAKT N+N +F+EI +
Sbjct: 116 MVMALAGNKADLEDRRKVTAEEARLYAEENGLFFMETSAKTAVNVNAIFYEIAK 169
>UniRef100_Q9SV33 Small GTP-binding protein-like [Arabidopsis thaliana]
Length = 224
Score = 67.4 bits (163), Expect = 8e-11
Identities = 31/49 (63%), Positives = 40/49 (81%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLF 52
+VMALV NK+DL KREV E+G + A++NGMF++ETSAKT +NIN LF
Sbjct: 138 IVMALVGNKADLHEKREVPTEDGMELAEKNGMFFIETSAKTADNINQLF 186
>UniRef100_UPI00002FAACD UPI00002FAACD UniRef100 entry
Length = 180
Score = 67.0 bits (162), Expect = 1e-10
Identities = 31/52 (59%), Positives = 39/52 (74%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEI 55
L+MAL NK D+E KREV+ EE +A ENG+F+METSAKT N+ +LF EI
Sbjct: 98 LIMALAGNKCDMEEKREVQREEAEAYASENGLFFMETSAKTAVNVTELFHEI 149
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.311 0.129 0.361
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 150,340,195
Number of Sequences: 2790947
Number of extensions: 5232708
Number of successful extensions: 11458
Number of sequences better than 10.0: 1763
Number of HSP's better than 10.0 without gapping: 1547
Number of HSP's successfully gapped in prelim test: 216
Number of HSP's that attempted gapping in prelim test: 9832
Number of HSP's gapped (non-prelim): 1766
length of query: 88
length of database: 848,049,833
effective HSP length: 64
effective length of query: 24
effective length of database: 669,429,225
effective search space: 16066301400
effective search space used: 16066301400
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0033b.5


