

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0032.12
(921 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
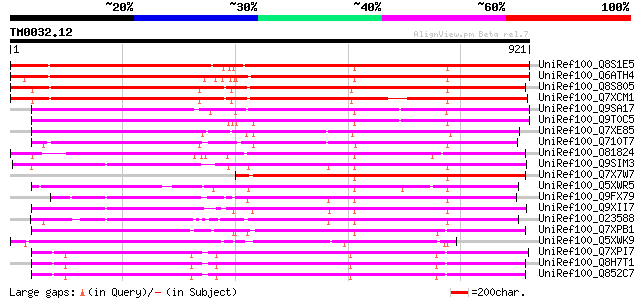
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8S1E5 Putative gag/pol polyprotein [Oryza sativa] 887 0.0
UniRef100_Q6ATH4 Putative polyprotein [Oryza sativa] 879 0.0
UniRef100_Q8S805 Putative copia-type polyprotein [Oryza sativa] 858 0.0
UniRef100_Q7XCM1 Putative gag-pol polyprotein [Oryza sativa] 817 0.0
UniRef100_Q9SA17 F28K20.17 protein [Arabidopsis thaliana] 663 0.0
UniRef100_Q9T0C5 Retrotransposon like protein [Arabidopsis thali... 620 e-176
UniRef100_Q7XE85 Putative pol polyprotein [Oryza sativa] 606 e-172
UniRef100_Q710T7 Gag-pol polyprotein [Populus deltoides] 567 e-160
UniRef100_O81824 Hypothetical protein AT4g27210 [Arabidopsis tha... 561 e-158
UniRef100_Q9SIM3 Putative retroelement pol polyprotein [Arabidop... 551 e-155
UniRef100_Q7X7W7 OSJNBa0061G20.2 protein [Oryza sativa] 528 e-148
UniRef100_Q5XWR5 Putative retroelement pol polyprotein-like [Sol... 520 e-145
UniRef100_Q9FX79 Putative retroelement polyprotein [Arabidopsis ... 518 e-145
UniRef100_Q9XII7 Putative retroelement pol polyprotein [Arabidop... 514 e-144
UniRef100_O23588 Retrotransposon like protein [Arabidopsis thali... 509 e-142
UniRef100_Q7XPB1 OSJNBb0026E15.10 protein [Oryza sativa] 493 e-137
UniRef100_Q5XWK9 Gag-pol polyprotein-like [Solanum tuberosum] 490 e-137
UniRef100_Q7XPI7 OSJNBb0004A17.2 protein [Oryza sativa] 488 e-136
UniRef100_Q8H7T1 Putative Zea mays retrotransposon Opie-2 [Oryza... 487 e-136
UniRef100_Q852C7 Putative gag-pol polyprotein [Oryza sativa] 487 e-136
>UniRef100_Q8S1E5 Putative gag/pol polyprotein [Oryza sativa]
Length = 1090
Score = 887 bits (2291), Expect = 0.0
Identities = 487/998 (48%), Positives = 621/998 (61%), Gaps = 83/998 (8%)
Query: 1 FGFSVSDFQTGMPLLRCNSLGDLYPV---TCSSHFAGL---ASGVWHNRLGHPTSSALNH 54
FGFSV D QT +LRCNS G+LY + T SS GL +S +WH RLGHP +A++
Sbjct: 80 FGFSVKDLQTRRVILRCNSRGELYTLPAATPSSAAHGLLATSSTLWHCRLGHPGPAAIHG 139
Query: 55 LRNNKLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSFDILHSDLWTSPILSTAGH 114
LRN I C+ +S +C +C LGKH RLPF +S + T F+++H D+WTSP++ST+G
Sbjct: 140 LRNIASISCNKIDTS-LCHACQLGKHTRLPFHNSSSRTSVPFELVHCDVWTSPVMSTSGF 198
Query: 115 RYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQFSANIKCLQCDNGREYDNDSF 174
+YY++VLDD + F W F + KS V+ + TQF +K Q DNGRE+ N +
Sbjct: 199 KYYLVVLDDFSHFCWTFLLRLKSDVHRHIVEFVEYVSTQFGLPLKSFQADNGREFVNTAI 258
Query: 175 RRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYL 234
+ + G R SCP+TS QNGK+ER +RTINN IRTLL +S+PPS+W AL ATYL
Sbjct: 259 TTFLASRGTQLRLSCPYTSPQNGKAERMLRTINNSIRTLLIQASMPPSYWAEALATATYL 318
Query: 235 LNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFFPSATINKLQPRSSPCVFLGYP 294
LN P ++ P Q L+ P +SHLRVFGCLC+P + T +KL PRS+ CVFLGYP
Sbjct: 319 LNRRPSSSIHQSLPFQLLHRTIPDFSHLRVFGCLCYPNLSATTPHKLSPRSTACVFLGYP 378
Query: 295 MNHRGYKCYDLSNRKLIISRHVIFDESRFPFADLPLESTSSYDCFTEDLPPSLIHHWQTT 354
+H+GY+C DLS ++IISRHV+FDES+FPFA P + SS+D + L P+ +
Sbjct: 379 TSHKGYRCLDLSTHRIIISRHVVFDESQFPFAATP-PAASSFDFLLQGLSPADAPSLEVE 437
Query: 355 STRPPDLSIPPSSPTDST-MPSPA----------PTSSPTSSAPL--------------- 388
RP L++ PS+ + +P P+ + +P++ APL
Sbjct: 438 QPRP--LTVAPSTEVEQPYLPLPSRRLSAGTVTVASEAPSAGAPLVGTSSADATPPGSAT 495
Query: 389 ----------------PLPPVPPT-----------PPTRTMTTHSMHGISKPKKPFSLSV 421
P+ VPP+ P +M T S G +P L+
Sbjct: 496 RASTIVSPFRHVYTRRPVTTVPPSSSTAVTNAVAAPQPHSMVTRSQSGSLRPVD--RLTY 553
Query: 422 SIDDPSISPLPCNPKQALSDPNWKFAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRH 481
+ + SP+P N AL+DPNW+ AM E+ L+ N TW LV RP N+ IF+H
Sbjct: 554 TATQAAASPVPANYHSALADPNWRAAMADEYKELVDNGTWRLVSRPPRANIATGKWIFKH 613
Query: 482 KKQSNGLFEHYKARLVGDGWSQIAGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDV 541
K S+G YKAR V G+SQ G+D DETF+ VVK ATIR VLSIA SR+WPIH LDV
Sbjct: 614 KFHSDGSLARYKARWVVRGYSQQHGIDYDETFSPVVKLATIRVVLSIAASRAWPIHQLDV 673
Query: 542 HNAFLHGDLHETVYMHQPLGFRDSQHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGF 601
NAFLHG L ETVY QP GF D PD VC +KSLYGLKQAPRAWYQRFA Y+ +GF
Sbjct: 674 KNAFLHGHLKETVYCQQPSGFVDPTAPDAVCLLQKSLYGLKQAPRAWYQRFATYIRQMGF 733
Query: 602 QHSSSDHS-----------YILLYVDDIILVASSHDLRKSFMALLAFEFAMKDLGPLSYF 650
S+SD S Y+LLYVDDIIL AS+ L + A L EFAM DLG L +F
Sbjct: 734 MPSASDTSLFVYKDGDRIAYLLLYVDDIILTASTTTLLQQLTARLHSEFAMTDLGDLHFF 793
Query: 651 LGIAVTRYVGGLFLSQSTYASEIIARAGMASCNPSATPVDTK*KLSSSSGTPCEDVTLYQ 710
LGI+V R GLFLSQ YA +++ RAGMA C+ ++TPVDT KLS++ G P D + Y+
Sbjct: 794 LGISVKRSPDGLFLSQRQYAVDLLQRAGMAECHSTSTPVDTHAKLSATDGLPVADPSAYR 853
Query: 711 SLAGALQYLTFTRPNISYVVQQVCLHMHAPHTEHMLAMFVAL*LTVFTCIPPLLRNLFLI 770
S+AGALQYLT TRP+++Y VQQVCL MH P H+ + L + L I
Sbjct: 854 SIAGALQYLTLTRPDLAYAVQQVCLFMHDPREPHLALVKRILRYVKGSLSIGLHIGSGPI 913
Query: 771 RMLT-------GGMSCTRRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVS 823
+ LT G +RRSTSGYCV+LGDNL+SWSSKRQ T+S SSAEAEYR VA+ V+
Sbjct: 914 QSLTAYSDADWAGCPNSRRSTSGYCVYLGDNLVSWSSKRQTTVSRSSAEAEYRAVAHAVA 973
Query: 824 ESCWIRNLLLELHFPLSQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQ 883
E CW+R LL ELH P++ T+V+CDNVS++Y++ NPVHH+RTKHIE+DIHFVREKVA GQ
Sbjct: 974 ECCWLRQLLQELHVPIASATIVYCDNVSAVYMTANPVHHRRTKHIEIDIHFVREKVALGQ 1033
Query: 884 ARILHVPSRHQIADIFTKGLPRVLFDDFRSSLSFGEPP 921
R+L+VPS HQ ADI TKGLP LF DFRSSL PP
Sbjct: 1034 VRVLYVPSSHQFADIMTKGLPVQLFTDFRSSLCVRAPP 1071
>UniRef100_Q6ATH4 Putative polyprotein [Oryza sativa]
Length = 1480
Score = 879 bits (2270), Expect = 0.0
Identities = 486/1033 (47%), Positives = 621/1033 (60%), Gaps = 116/1033 (11%)
Query: 1 FGFSVSDFQTGMPLLRCNSLGDLY------PVTCSSHFAGLASGVWHNRLGHPTSSALNH 54
FGFSV D T +LRCNS GDLY P + F +S +WH+RLGHP+ +A+
Sbjct: 445 FGFSVKDLPTRRVILRCNSRGDLYTLPTTVPAITAHSFLAKSSTLWHHRLGHPSPAAVQT 504
Query: 55 LRNNKLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSFDILHSDLWTSPILSTAGH 114
L ++ C S + +C +C LGKH RL FS S + T F+++H D+WTSP+LS +G
Sbjct: 505 LHKLAILSCTRSNNK-LCHACHLGKHTRLSFSKSSSSTSSPFELVHCDVWTSPVLSLSGF 563
Query: 115 RYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQFSANIKCLQCDNGREYDNDSF 174
+YY++VLDD T F W FP+ KS V+ +KTQFS I+C Q DNG ++ N +
Sbjct: 564 KYYLVVLDDFTHFCWTFPLRHKSDVHQHLLEFVAYVKTQFSLPIRCFQADNGTKFVNHAT 623
Query: 175 RRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYL 234
+ + G++ R SCP+TS QNGK+ER +RTIN IRTLL +S+PPS+W AL ATYL
Sbjct: 624 TSFFASRGIVLRLSCPYTSPQNGKAERVLRTINKSIRTLLIQASMPPSYWAEALATATYL 683
Query: 235 LNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFFPSATINKLQPRSSPCVFLGYP 294
LN P ++RN P Q L+++ P YS+L+VFGCLC+P + T +KL PRS+P VFLGY
Sbjct: 684 LNRRPSTSVRNSIPYQLLHNKLPDYSNLQVFGCLCYPNLSAMTSHKLSPRSAPYVFLGYS 743
Query: 295 MNHRGYKCYDLSNRKLIISRHVIFDESRFPFADLPLESTSSYDCFTEDL-----PPSLIH 349
+H+G++C D+S R+L ISRHV+FDE FPFA +P + SS+D + P S +
Sbjct: 744 ASHKGFRCLDISTRRLYISRHVVFDEKTFPFAAIP-QDASSFDFLLQGFSIAVAPSSEVE 802
Query: 350 HWQTTSTRP-PDLSIP---------------------------PSSPTDSTMPS------ 375
+ +S P P++ P SP D+ +P
Sbjct: 803 RPRFSSMTPSPEVEQPIPDDDTSGTELFQLLPGLRSSAAGRPLAGSPVDARLPGGCANDA 862
Query: 376 ----------------------PAPTSSPTSSAPLP-----LPPVPPTP-----PTR--- 400
PAP+ PT+S P L P P P R
Sbjct: 863 ANGSSSSNLSPVMDPPAASVVRPAPSEGPTTSLISPYRHTYLRRSQPAPTAIHRPIRASR 922
Query: 401 --------------TMTTHSMHGISKPKKPFSLSVSIDDPSISPLPCNPKQALSDPNWKF 446
TM T S G +P + F+ + + D +SP+P N + AL+DPNW+
Sbjct: 923 AFHSATDQQQQTGHTMVTRSQTGHLRPIQRFTYTATHD--VVSPVPSNYRSALADPNWRA 980
Query: 447 AMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVGDGWSQIAG 506
A E+ AL+ NNTW LVPRP NV+ IF+HK S+G +KAR V G+SQ G
Sbjct: 981 ATANEYKALVDNNTWRLVPRPPGANVVTGKWIFKHKFHSDGTLARHKARWVVRGYSQQHG 1040
Query: 507 VDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLGFRDSQ 566
+D DETF+ VVKPATI VLSIA SRSWPIH LDV NAFLHG+L ETVY QP GF D
Sbjct: 1041 IDYDETFSPVVKPATIHVVLSIAASRSWPIHQLDVKNAFLHGNLEETVYYQQPSGFVDPS 1100
Query: 567 HPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHS-----------YILLYV 615
P+ VC +KSLYGLKQAPRAWYQRFA Y+ +GF S+S+ S Y+LLYV
Sbjct: 1101 APNAVCLLQKSLYGLKQAPRAWYQRFATYIRQLGFTSSASNTSLFVYKDGDNIAYLLLYV 1160
Query: 616 DDIILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQSTYASEIIA 675
DDIIL ASS L A L EFAM DLG L +FLGI+VTR GLFLSQ Y +++
Sbjct: 1161 DDIILTASSATLLHHITARLHSEFAMTDLGDLHFFLGISVTRSSDGLFLSQRQYVVDLLK 1220
Query: 676 RAGMASCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPNISYVVQQVCL 735
RAGM+ C+ +ATPVD + KLS++ G P D T Y+SL GALQYLT TRP ++Y VQQVCL
Sbjct: 1221 RAGMSECHSTATPVDARSKLSATDGAPVSDPTEYRSLVGALQYLTLTRPELAYAVQQVCL 1280
Query: 736 HMHAPHTEHMLAMFVAL*LTVFTCIPPLLRNLFLIRMLT-------GGMSCTRRSTSGYC 788
MH P H+ + L T L + LT G + RSTSGYC
Sbjct: 1281 FMHDPREPHLALIKRILRYIKGTLHIGLHLGTTPVDSLTAYSDADWAGCPDSHRSTSGYC 1340
Query: 789 VFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQETLVHCD 848
VFLGDNL+SWSSKRQ T+S SSAEAEYR VA+ ++E CW+R LL ELH L+ T+V+CD
Sbjct: 1341 VFLGDNLVSWSSKRQTTVSRSSAEAEYRAVAHAIAECCWLRQLLSELHISLTSATVVYCD 1400
Query: 849 NVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADIFTKGLPRVLF 908
NVS++Y++ NPVHH+RTKHIE+DIHFVREKVA GQ R+LH+PS HQ ADI TKGLP LF
Sbjct: 1401 NVSAVYMAANPVHHRRTKHIEIDIHFVREKVALGQVRVLHLPSSHQFADIMTKGLPVQLF 1460
Query: 909 DDFRSSLSFGEPP 921
+FRSSL + P
Sbjct: 1461 TEFRSSLCVRDNP 1473
>UniRef100_Q8S805 Putative copia-type polyprotein [Oryza sativa]
Length = 1803
Score = 858 bits (2216), Expect = 0.0
Identities = 459/947 (48%), Positives = 595/947 (62%), Gaps = 40/947 (4%)
Query: 2 GFSVSDFQTGMPLLRCNSLGDLYPVTCSSHFAGLASGV-----WHNRLGHPTSSALNHLR 56
GFS+ D QT + LRC+S GDLYP+ S A A+ WH RLGHP S++L+ +
Sbjct: 457 GFSIKDLQTQVVKLRCDSPGDLYPLRLPSPHALSATSSPSVEHWHLRLGHPGSASLSKVL 516
Query: 57 NNKLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSFDILHSDLWTSPILSTAGHRY 116
+ C+ S C +C +G +VRLPF SS + TL F ++H+D+WTSPI S +G++Y
Sbjct: 517 GSFDFQCNKSAPHH-CSACHVGTNVRLPFHSSSSQTLFPFQLVHTDVWTSPIYSNSGYKY 575
Query: 117 YVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQFSANIKCLQCDNGREYDNDSFRR 176
YV+ LDD T ++W FP+ KS+V+ + TQF + LQ DNG+EYD+ + R
Sbjct: 576 YVVFLDDFTHYIWTFPVRNKSEVFHTVRSFFAYAHTQFGLPVLALQTDNGKEYDSYALRS 635
Query: 177 YCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLN 236
+G + R SCP++S QNGK+ER +RTIN+ +RT+L HS+ P SFW ALQ A +L+N
Sbjct: 636 LLSLHGAVLRLSCPYSSQQNGKAERILRTINDCVRTMLVHSAAPLSFWAEALQTAMHLIN 695
Query: 237 ILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFFPSATINKLQPRSSPCVFLGYPMN 296
P + + P Q L P+Y HLRVFGCLC+P + +KL PRS CVF+GYP +
Sbjct: 696 RRPCRATGSLKPYQLLLGAPPTYDHLRVFGCLCYPNTIATAPHKLSPRSLACVFIGYPAD 755
Query: 297 HRGYKCYDLSNRKLIISRHVIFDESRFPFADLPLESTSS------YDCFTEDLPPSLIHH 350
HRGY+CYD+ +R++ SRHV F E FPF D P S+ D LP H
Sbjct: 756 HRGYRCYDMVSRRVFTSRHVTFVEDVFPFRDAPSPRPSAPPPPDHGDDTIVLLPAPAQHV 815
Query: 351 WQTTSTRPPDLSIPPSSPTDSTMPSPAPTSSPTSSAPLPLP----PVPPTPPTRTMTTHS 406
T P + P SP ST S AP AP P P P +PP MTT +
Sbjct: 816 VTPVGTAPAHDAASPPSPASSTPSSAAPAH---DVAPPPSPETSSPASASPPRHAMTTRA 872
Query: 407 MHGISKPKKPFSLSVSIDDPSISPLPCNPKQALSDPNWKFAMQPEFNALIRNNTWELVPR 466
GISKP ++++ + ++SP P + + AL DPNW+ AMQ EF+AL+ N TW LVPR
Sbjct: 873 RAGISKPNPRYAMTAT---STLSPTPSSVRVALRDPNWRAAMQAEFDALLANRTWTLVPR 929
Query: 467 PCDVNVIRYM*IFRHKKQSNGLFEHYKARLVGDGWSQIAGVDCDETFNLVVKPATIRTVL 526
P +I +F+ K ++G + YKAR V G++Q GVD ETF+ VVKPATIRTVL
Sbjct: 930 PPGARIITGKWVFKTKLHADGSLDKYKARWVVRGFNQRPGVDFGETFSPVVKPATIRTVL 989
Query: 527 SIALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLGFRDSQHPDYVCRRKKSLYGLKQAPR 586
++ S+ WP H LDV NAFLHG L E V QP GF D+ P VC +SLYGL+QAPR
Sbjct: 990 TLISSKQWPAHQLDVSNAFLHGHLQERVLCQQPTGFEDAARPADVCLLSRSLYGLRQAPR 1049
Query: 587 AWYQRFADYVSSIGFQHS-----------SSDHSYILLYVDDIILVASSHDLRKSFMALL 635
AW++RFAD+ +S+GF S SD +Y+LLYVDD+IL ASS L + + L
Sbjct: 1050 AWFKRFADHATSLGFVQSRADPSLFVLRRGSDTAYLLLYVDDMILSASSSSLLQRIIDRL 1109
Query: 636 AFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQSTYASEIIARAGMASCNPSATPVDTK*KL 695
EF +KD+GPL YFLGI V R G LSQS YA++++ RAGMA+C ATP D K KL
Sbjct: 1110 QAEFKVKDMGPLKYFLGIEVQRTADGFVLSQSKYATDVLERAGMANCKAVATPADAKPKL 1169
Query: 696 SSSSGTPCEDVTLYQSLAGALQYLTFTRPNISYVVQQVCLHMHAPHTEHMLAMFVAL*LT 755
SS G +D + Y+S+AGALQYLT TRP+I+Y VQQVCLHMHAP H+ + L
Sbjct: 1170 SSDEGPLFQDSSWYRSIAGALQYLTLTRPDIAYAVQQVCLHMHAPREAHVTLLKRILRYI 1229
Query: 756 VFTCIPPLLRNLFLIRMLT-------GGMSCTRRSTSGYCVFLGDNLISWSSKRQPTLSH 808
T L LT G TRRSTSG+C+FLGD+LISWSSKRQ T+S
Sbjct: 1230 KGTAAFGLHLRASTSPTLTAFSDADWAGCPDTRRSTSGFCIFLGDSLISWSSKRQTTVSR 1289
Query: 809 SSAEAEYRGVANVVSESCWIRNLLLELHFPLSQETLVHCDNVSSIYLSGNPVHHQRTKHI 868
SSAEAEYRGVAN V+E W+R LL ELH + Q T+ +CDN+SS+Y+S NPVHH+RTKHI
Sbjct: 1290 SSAEAEYRGVANAVAECTWLRQLLGELHCRVPQATIAYCDNISSVYMSKNPVHHKRTKHI 1349
Query: 869 EMDIHFVREKVARGQARILHVPSRHQIADIFTKGLPRVLFDDFRSSL 915
E+DIHFVREKVA G+ R+L +PS HQ AD+FTKGLP +F++FR+SL
Sbjct: 1350 ELDIHFVREKVALGELRVLPIPSAHQFADVFTKGLPSSMFNEFRASL 1396
>UniRef100_Q7XCM1 Putative gag-pol polyprotein [Oryza sativa]
Length = 1417
Score = 817 bits (2110), Expect = 0.0
Identities = 445/951 (46%), Positives = 579/951 (60%), Gaps = 72/951 (7%)
Query: 2 GFSVSDFQTGMPLLRCNSLGDLYPVTCSSHFAGLASGV-----WHNRLGHPTSSALNHLR 56
GFS+ D QT + LRC+S GDLYP+ S A A+ WH RLGHP S++L+ +
Sbjct: 457 GFSIKDLQTQVVKLRCDSPGDLYPLRLPSPHALSATSSPSVEHWHLRLGHPGSASLSKVL 516
Query: 57 NNKLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSFDILHSDLWTSPILSTAGHRY 116
+ C+ S C +C +G +VRLPF SS + TL F ++H+D+WTSPI S +G++Y
Sbjct: 517 GSFDFQCNKSAPHH-CSACHVGTNVRLPFHSSSSQTLFPFQLVHTDVWTSPIYSNSGYKY 575
Query: 117 YVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQFSANIKCLQCDNGREYDNDSFRR 176
YV+ LDD T ++W FP+ KS+V+ + TQF + LQ DNG+EYD+ + R
Sbjct: 576 YVVFLDDFTHYIWTFPVRNKSEVFHTVRSFFAYAHTQFGLPVLALQTDNGKEYDSYALRS 635
Query: 177 YCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLN 236
+G + R SCP++S QNGK+ER +RTIN+ +RT+L HS+ P SFW ALQ AT+L+N
Sbjct: 636 LLSLHGAVLRLSCPYSSQQNGKAERILRTINDYVRTMLVHSAAPLSFWAEALQTATHLIN 695
Query: 237 ILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFFPSATINKLQPRSSPCVFLGYPMN 296
P + + +P Q L P+Y HLRVFGCLC+P + +KL PRS CVF+GYP +
Sbjct: 696 RRPCRATGSLTPYQLLLGAPPTYDHLRVFGCLCYPNTIATAPHKLSPRSLACVFIGYPAD 755
Query: 297 HRGYKCYDLSNRKLIISRHVIFDESRFPFADLPLESTSS------YDCFTEDLPPSLIHH 350
HRGY+CYD+ +R++ SRHV F E FPF D P S+ D LP H
Sbjct: 756 HRGYRCYDMVSRRVFTSRHVTFVEDVFPFRDAPSPRPSAPPPPDHGDDTIVLLPAPAQHV 815
Query: 351 WQTTSTRPPDLSIPPSSPTDSTMPSPAPTSSPTSSAPLPLP----PVPPTPPTRTMTTHS 406
T P + P SP ST S AP AP P P P +PP MTT +
Sbjct: 816 VTPVGTAPAHDAASPPSPASSTPSSAAPAH---DVAPPPSPETSSPASASPPRHAMTTRA 872
Query: 407 MHGISKPKKPFSLSVSIDDPSISPLPCNPKQALSDPNWKFAMQPEFNALIRNNTWELVPR 466
GISKP ++++ + ++SP P + + AL DPNW+ AMQ EF+AL+ N TW LVPR
Sbjct: 873 RAGISKPNPRYAMTAT---STLSPTPSSVRAALRDPNWRAAMQAEFDALLANRTWTLVPR 929
Query: 467 PCDVNVIRYM*IFRHKKQSNGLFEHYKARLVGDGWSQIAGVDCDETFNLVVKPATIRTVL 526
P +I +F+ K ++G + YKAR V G++Q GVD ETF+ VVKPATIRTVL
Sbjct: 930 PPGARIITGKWVFKTKLHADGSLDKYKARWVVRGFNQRPGVDFGETFSPVVKPATIRTVL 989
Query: 527 SIALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLGFRDSQHPDYVCRRKKSLYGLKQAPR 586
++ S+ WP H LDV NAFLHG L E V QP GF D+ P VC +SLYGL+QAPR
Sbjct: 990 TLISSKQWPAHQLDVSNAFLHGHLQERVLCQQPTGFEDAARPADVCLLSRSLYGLRQAPR 1049
Query: 587 AWYQRFADYVSSIGFQHS-----------SSDHSYILLYVDDIILVASSHDLRKSFMALL 635
AW++RFAD+ +S+GF S SD +Y+LLYVDD+IL ASS L + + L
Sbjct: 1050 AWFKRFADHATSLGFVQSRADPSLFVLRRGSDTAYLLLYVDDMILSASSSSLLQRIIDRL 1109
Query: 636 AFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQSTYASEIIARAGMASCNPSATPVDTK*KL 695
EF +KD+GPL YFLGI V R G LSQS YA+
Sbjct: 1110 QAEFKVKDMGPLKYFLGIEVQRTADGFVLSQSKYAT------------------------ 1145
Query: 696 SSSSGTPCEDVTLYQSLAGALQYLTFTRPNISYVVQQVCLHMHAPHTEHMLAMFVAL*LT 755
+D + Y+S+AGALQYLT TRP+I+Y VQQVCLHMHAP H+ + L
Sbjct: 1146 --------DDSSWYRSIAGALQYLTLTRPDIAYAVQQVCLHMHAPREAHVTLLKRILRYI 1197
Query: 756 VFTCIPPLLRNLFLIRMLT-------GGMSCTRRSTSGYCVFLGDNLISWSSKRQPTLSH 808
T L LT G TRRSTSG+C+FLGD+LISWSSKRQ T+S
Sbjct: 1198 KGTAAFGLHLRASTSPTLTAFSDADWAGCPDTRRSTSGFCIFLGDSLISWSSKRQTTVSR 1257
Query: 809 SSAEAEYRGVANVVSESCWIRNLLLELHFPLSQETLVHCDNVSSIYLSGNPVHHQRTKHI 868
SSAEAEYRGVAN V+E W+R LL ELH + Q T+ +CDN+SS+Y+S NPVHH+RTKHI
Sbjct: 1258 SSAEAEYRGVANAVAECTWLRQLLGELHCRVPQATIAYCDNISSVYMSKNPVHHKRTKHI 1317
Query: 869 EMDIHFVREKVARGQARILHVPSRHQIADIFTKGLPRVLFDDFRSSLSFGE 919
E+DIHFVREKVA G+ R+L +PS HQ AD+FTKGLP +F++FR+SL G+
Sbjct: 1318 ELDIHFVREKVALGELRVLPIPSAHQFADVFTKGLPSSMFNEFRASLCVGD 1368
>UniRef100_Q9SA17 F28K20.17 protein [Arabidopsis thaliana]
Length = 1415
Score = 663 bits (1710), Expect = 0.0
Identities = 367/923 (39%), Positives = 506/923 (54%), Gaps = 52/923 (5%)
Query: 39 VWHNRLGHPTSSALNHLRNNKLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSFDI 98
VWH+RLGH S AL HL+N+K I + SR+S VC+ C +GK RLPF S++ L D
Sbjct: 455 VWHHRLGHANSKALQHLQNSKAIQINKSRTSPVCEPCQMGKSSRLPFLISDSRVLHPLDR 514
Query: 99 LHSDLW-TSPILSTAGHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQFSAN 157
+H DLW SP++S G +YY + +DD++ + W +P+ KS+ +F + L++ Q +
Sbjct: 515 IHCDLWGPSPVVSNQGLKYYAIFVDDYSRYSWFYPLHNKSEFLSVFISFQKLVENQLNTK 574
Query: 158 IKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLLAHS 217
IK Q D G E+ ++ + + +G+ R SCP+T QNG +ERK R + + ++L HS
Sbjct: 575 IKVFQSDGGGEFVSNKLKTHLSEHGIHHRISCPYTPQQNGLAERKHRHLVELGLSMLFHS 634
Query: 218 SVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFFPSAT 277
P FW + A Y++N LP L+N SP + L+ P YS LRVFG C+P
Sbjct: 635 HTPQKFWVESFFTANYIINRLPSSVLKNLSPYEALFGEKPDYSSLRVFGSACYPCLRPLA 694
Query: 278 INKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFADLPLESTSSYD 337
NK PRS CVFLGY ++GY+C+ K+ ISR+VIF+ES PF + Y
Sbjct: 695 QNKFDPRSLQCVFLGYNSQYKGYRCFYPPTGKVYISRNVIFNESELPFKE-------KYQ 747
Query: 338 CFTEDLPPSLIHHWQTTS--------------TRPPDLSIPPSSPTDSTMPSPAPTSSPT 383
L+ WQ ++P DL+ S + P PTS+
Sbjct: 748 SLVPQYSTPLLQAWQHNKISEISVPAAPVQLFSKPIDLNTYAGSQVTEQLTDPEPTSNNE 807
Query: 384 SSAPLPLPPVPPTPP-------TRTMTTHSMHGISKPKKPFSLSVSIDDPSISPLPCNPK 436
S P + MTT S GI KP ++L I + P
Sbjct: 808 GSDEEVNPVAEEIAANQEQVINSHAMTTRSKAGIQKPNTRYAL---ITSRMNTAEPKTLA 864
Query: 437 QALSDPNWKFAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARL 496
A+ P W A+ E N + +TW LVP D+N++ +F+ K +G + KARL
Sbjct: 865 SAMKHPGWNEAVHEEINRVHMLHTWSLVPPTDDMNILSSKWVFKTKLHPDGSIDKLKARL 924
Query: 497 VGDGWSQIAGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYM 556
V G+ Q GVD ETF+ VV+ ATIR VL ++ S+ WPI LDV NAFLHG+L E V+M
Sbjct: 925 VAKGFDQEEGVDYLETFSPVVRTATIRLVLDVSTSKGWPIKQLDVSNAFLHGELQEPVFM 984
Query: 557 HQPLGFRDSQHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHS------- 609
+QP GF D Q P +VCR K++YGLKQAPRAW+ F++++ GF S SD S
Sbjct: 985 YQPSGFIDPQKPTHVCRLTKAIYGLKQAPRAWFDTFSNFLLDYGFVCSKSDPSLFVCHQD 1044
Query: 610 ----YILLYVDDIILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLS 665
Y+LLYVDDI+L S L + + L F+MKDLGP YFLGI + Y GLFL
Sbjct: 1045 GKILYLLLYVDDILLTGSDQSLLEDLLQALKNRFSMKDLGPPRYFLGIQIEDYANGLFLH 1104
Query: 666 QSTYASEIIARAGMASCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPN 725
Q+ YA++I+ +AGM+ CNP TP+ +L + + + T ++SLAG LQYLT TRP+
Sbjct: 1105 QTAYATDILQQAGMSDCNPMPTPLPQ--QLDNLNSELFAEPTYFRSLAGKLQYLTITRPD 1162
Query: 726 ISYVVQQVCLHMHAPHTEH--MLAMFVAL*LTVFTCIPPLLRNLFLIRML-----TGGMS 778
I + V +C MH+P T +L + P+ RN L G
Sbjct: 1163 IQFAVNFICQRMHSPTTSDFGLLKRILRYIKGTIGMGLPIKRNSTLTLSAYSDSDHAGCK 1222
Query: 779 CTRRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFP 838
TRRST+G+C+ LG NLISWS+KRQPT+S+SS EAEYR + E WI LL +L P
Sbjct: 1223 NTRRSTTGFCILLGSNLISWSAKRQPTVSNSSTEAEYRALTYAAREITWISFLLRDLGIP 1282
Query: 839 LSQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADI 898
T V+CDN+S++YLS NP H R+KH + D H++RE+VA G H+ + Q+AD+
Sbjct: 1283 QYLPTQVYCDNLSAVYLSANPALHNRSKHFDTDYHYIREQVALGLIETQHISATFQLADV 1342
Query: 899 FTKGLPRVLFDDFRSSLSFGEPP 921
FTK LPR F D RS L P
Sbjct: 1343 FTKSLPRRAFVDLRSKLGVSGSP 1365
>UniRef100_Q9T0C5 Retrotransposon like protein [Arabidopsis thaliana]
Length = 1515
Score = 620 bits (1600), Expect = e-176
Identities = 364/972 (37%), Positives = 518/972 (52%), Gaps = 92/972 (9%)
Query: 39 VWHNRLGHPTSSALNHLRNNKLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSFDI 98
VWH RLGHP L HL K I + + SS +C++C +GK RLPF +SE ++ R +
Sbjct: 457 VWHQRLGHPNKEVLQHLIKTKAIVVNKT-SSNMCEACQMGKVCRLPFVASEFVSSRPLER 515
Query: 99 LHSDLW-TSPILSTAGHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQFSAN 157
+H DLW +P+ S G +YYV+ +D+++ F W +P+ KS + +F L++ Q+
Sbjct: 516 IHCDLWGPAPVTSAQGFQYYVIFIDNYSRFTWFYPLKLKSDFFSVFVLFQQLVENQYQHK 575
Query: 158 IKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLLAHS 217
I QCD G E+ + F + + G+ SCPHT QNG +ER+ R + + +L+ HS
Sbjct: 576 IAMFQCDGGGEFVSYKFVAHLASCGIKQLISCPHTPQQNGIAERRHRYLTELGLSLMFHS 635
Query: 218 SVPPSFWHHALQMATYLLNILPRKTLR-NDSPTQRLYHRDPSYSHLRVFGCLCFPFFPSA 276
VP W A + +L N+LP TL N SP + L+ P Y+ LRVFG C+P+
Sbjct: 636 KVPHKLWVEAFFTSNFLSNLLPSSTLSDNKSPYEMLHGTPPVYTALRVFGSACYPYLRPY 695
Query: 277 TINKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFADL--PLESTS 334
NK P+S CVFLGY ++GY+C K+ I RHV+FDE +FP++D+ ++ S
Sbjct: 696 AKNKFDPKSLLCVFLGYNNKYKGYRCLHPPTGKVYICRHVLFDERKFPYSDIYSQFQTIS 755
Query: 335 SYDCFT--EDLPPSLIHHWQTTSTRPPDLSIPPSSPTDSTMPSPAPTSSPTSSAP----- 387
FT + S +T ST D+ P ++ + S AP + T++AP
Sbjct: 756 GSPLFTAWQKGFSSTALSRETPSTNVEDIIFPSATVSSSVPTGCAPNIAETATAPDVDVA 815
Query: 388 ----LPLPPVP------PTPPTRT------MTTHSMHGISKPKKPFSLSVSI-DDPSISP 430
+ +PP P PT P + +T S IS P S++VS+ +D P
Sbjct: 816 AAHDMVVPPSPITSTSLPTQPEESTSDQNHYSTDSETAISSAMTPQSINVSLFEDSDFPP 875
Query: 431 L-------------------------------------------PCNPKQALSDPNWKFA 447
L P + K+AL D W A
Sbjct: 876 LQSVISSTTAAPETSHPMITRAKSGITKPNPKYALFSVKSNYPEPKSVKEALKDEGWTNA 935
Query: 448 MQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVGDGWSQIAGV 507
M E + +TW+LVP ++ +F+ K S+G + KARLV G+ Q GV
Sbjct: 936 MGEEMGTMHETDTWDLVPPEMVDRLLGCKWVFKTKLNSDGSLDRLKARLVARGYEQEEGV 995
Query: 508 DCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLGFRDSQH 567
D ET++ VV+ AT+R++L +A W + LDV NAFLH +L ETV+M QP GF D
Sbjct: 996 DYVETYSPVVRSATVRSILHVATINKWSLKQLDVKNAFLHDELKETVFMTQPPGFEDPSR 1055
Query: 568 PDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHS-----------YILLYVD 616
PDYVC+ KK++Y LKQAPRAW+ +F+ Y+ GF S SD S ++LLYVD
Sbjct: 1056 PDYVCKLKKAIYDLKQAPRAWFDKFSSYLLKYGFICSFSDPSLFVYLKGRDVMFLLLYVD 1115
Query: 617 DIILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQSTYASEIIAR 676
D+IL ++ L + + +L+ EF MKD+G L YFLGI + GLFLSQ Y S+++
Sbjct: 1116 DMILTGNNDVLLQQLLNILSTEFRMKDMGALHYFLGIQAHYHNDGLFLSQEKYTSDLLVN 1175
Query: 677 AGMASCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPNISYVVQQVCLH 736
AGM+ C+ TP+ L + P + T ++ LAG LQYLT TRP+I + V VC
Sbjct: 1176 AGMSDCSSMPTPLQL--DLLQGNNKPFPEPTYFRRLAGKLQYLTLTRPDIQFAVNFVCQK 1233
Query: 737 MHAPHTE--HMLAMFVAL*LTVFTCIPPLLRNL-FLIRMLT----GGMSCTRRSTSGYCV 789
MHAP H+L + T L N ++R + G TRRST G+C
Sbjct: 1234 MHAPTMSDFHLLKRILHYLKGTMTMGINLSSNTDSVLRCYSDSDWAGCKDTRRSTGGFCT 1293
Query: 790 FLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQETLVHCDN 849
FLG N+ISWS+KR PT+S SS EAEYR ++ SE WI LL E+ P Q ++CDN
Sbjct: 1294 FLGYNIISWSAKRHPTVSKSSTEAEYRTLSFAASEVSWIGFLLQEIGLPQQQIPEMYCDN 1353
Query: 850 VSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADIFTKGLPRVLFD 909
+S++YLS NP H R+KH ++D ++VRE+VA G + H+P+ Q+ADIFTK LP+ F
Sbjct: 1354 LSAVYLSANPALHSRSKHFQVDYYYVRERVALGALTVKHIPASQQLADIFTKSLPQAPFC 1413
Query: 910 DFRSSLSFGEPP 921
D R L PP
Sbjct: 1414 DLRFKLGVVLPP 1425
>UniRef100_Q7XE85 Putative pol polyprotein [Oryza sativa]
Length = 1688
Score = 606 bits (1563), Expect = e-172
Identities = 352/929 (37%), Positives = 501/929 (53%), Gaps = 68/929 (7%)
Query: 40 WHNRLGHPTSSALNHLRNNKLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSFDIL 99
WH+RLGH S L L N ++ P ++ VC C LGK V+LP+ SS + + R FD++
Sbjct: 325 WHHRLGHLCGSRLATLINQGVLGSVPVDTTFVCKGCKLGKQVQLPYPSSTSRSSRPFDLV 384
Query: 100 HSDLW-TSPILSTAGHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQFSANI 158
HSD+W SP S GH YYV+ +DD++ + W++ + +SQ+ I+ + A +I TQFS+ I
Sbjct: 385 HSDVWGKSPFPSKGGHNYYVIFVDDYSRYTWIYFMKHRSQLISIYQSFAQMIHTQFSSAI 444
Query: 159 KCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLLAHSS 218
+ + D+G EY +++FR + + G + + SCP +QNG +ERK R I RTLL S
Sbjct: 445 RIFRSDSGGEYMSNAFREFLVSQGTLPQLSCPGAHAQNGVAERKHRHIIETARTLLIASF 504
Query: 219 VPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFFPSATI 278
VP FW A+ A YL+N+ P +L+ SP + L+ P Y HLRVFGC C+
Sbjct: 505 VPAHFWAEAISTAVYLINMQPSSSLQGRSPGEVLFGSPPRYDHLRVFGCTCYVLLAPRER 564
Query: 279 NKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFADLPLESTSSYDC 338
KL +S CVFLGY + H+GY+CYD S R++ ISR V FDE++ F + +S +
Sbjct: 565 TKLTAQSVECVFLGYSLEHKGYRCYDPSARRIRISRDVTFDENKPFFYSSTNQPSSPENS 624
Query: 339 FT----------EDLPPSLIHHWQTTSTRPPDLSIPPSSPTDSTMPSPAPTSSPTSSAPL 388
+ E LP S I + S PP + P P PSP+P S P S P
Sbjct: 625 ISFLYLPPIPSPESLPSSPI--TPSPSPIPPSVPSPTYVPPPPPSPSPSPVSPPPSHIPA 682
Query: 389 PLPPVPPTPPTRTMTTHSMHGISKPKKPFSL----------SVSIDDPSISPL------- 431
P P P T T+ T H +PK P + S+DD S +P
Sbjct: 683 SSSP-PHVPSTITLDTFPFHYSRRPKIPNESQPSQPTLEDPTCSVDDSSPAPRYNLRARD 741
Query: 432 -----------------PCNPKQALSDPNWKFAMQPEFNALIRNNTWELVPRPCDVNVIR 474
P ++A+ P+WK AM E AL R NTW++VP P I
Sbjct: 742 ALRAPNRDDFVVGVVFEPSTYQEAIVLPHWKLAMSEELAALERTNTWDVVPLPSHAVPIT 801
Query: 475 YM*IFRHKKQSNGLFEHYKARLVGDGWSQIAGVDCDETFNLVVKPATIRTVLSIALSRSW 534
+++ K +S+G E YKARLV G+ Q G D DETF V T+RT++++A +RSW
Sbjct: 802 CKWVYKVKTKSDGQVERYKARLVARGFQQAHGRDYDETFAPVAHMTTVRTLIAVAATRSW 861
Query: 535 PIH*LDVHNAFLHGDLHETVYMHQPLGFRDSQHPDYVCRRKKSLYGLKQAPRAWYQRFAD 594
I +DV NAFLHGDLHE VYMH P G P +V R +++LYGLKQAPRAW+ RF+
Sbjct: 862 TISQMDVKNAFLHGDLHEEVYMHPPPGV--EAPPGHVFRLRRALYGLKQAPRAWFARFSS 919
Query: 595 YVSSIGFQHSSSD-----------HSYILLYVDDIILVASSHDLRKSFMALLAFEFAMKD 643
V + GF S D + +LLYVDD+++ + L+ +F M D
Sbjct: 920 VVLAAGFSPSDHDPALFIHTSSRGRTLLLLYVDDMLITGDDLEYIAFVKGKLSEQFMMSD 979
Query: 644 LGPLSYFLGIAVTRYVGGLFLSQSTYASEIIARAGMASCNPSATPVDTK*KLSSSSGTPC 703
LGPLSYFLGI VT V G +LSQ Y +++A++G+ + TP++ +L S+ GTP
Sbjct: 980 LGPLSYFLGIEVTSTVDGYYLSQHRYIEDLLAQSGLTDSRTTTTPMELHVRLRSTDGTPL 1039
Query: 704 EDVTLYQSLAGALQYLTFTRPNISYVVQQVCLHMHAP---HTEHMLAMFVAL*LTVFTCI 760
+D + Y+ L G+L YLT TRP+I+Y V + + AP H H+L + L T C+
Sbjct: 1040 DDPSRYRHLVGSLVYLTVTRPDIAYAVHILSQFVSAPISVHYGHLLRVLRYLRGTTTQCL 1099
Query: 761 PPLLRNLFLIRMLTGGMSCT----RRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYR 816
+ +R + + RRS +GYC+FLG +L++W SK+Q +S SS EAE R
Sbjct: 1100 FYAASSPLQLRAFSDSTWASDPIDRRSVTGYCIFLGTSLLTWKSKKQTAVSRSSTEAELR 1159
Query: 817 GVANVVSESCWIRNLLLELHFPLSQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVR 876
+A SE W+R LL + T + CDN +I ++ +P+ H+ TKHI +D F R
Sbjct: 1160 ALATTTSEIVWLRWLLADFGVSCDVPTPLLCDNTGAIQIANDPIKHELTKHIGVDASFTR 1219
Query: 877 EKVARGQARILHVPSRHQIADIFTKGLPR 905
+ + +VPS Q+AD FTK R
Sbjct: 1220 SHCQQSTIALHYVPSELQVADFFTKAQTR 1248
>UniRef100_Q710T7 Gag-pol polyprotein [Populus deltoides]
Length = 1382
Score = 567 bits (1462), Expect = e-160
Identities = 352/903 (38%), Positives = 483/903 (52%), Gaps = 62/903 (6%)
Query: 39 VWHNRLGHPTSSALNHLRNN----KLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLR 94
+WH+RLGH +SS L L + L CD S C C L K LPF+ S +++
Sbjct: 480 LWHSRLGHVSSSRLRFLASTGALGNLKTCDISD----CSGCKLAKFSALPFNRSTSVSSS 535
Query: 95 SFDILHSDLW-TSPILSTAGHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQ 153
FD++HSD+W SP+ + G RYYV +DDHT + W++ + +S+ ++I+ LIKTQ
Sbjct: 536 PFDLIHSDVWGPSPVSTKGGSRYYVSFIDDHTRYCWVYLMKHRSEFFEIYAAFRALIKTQ 595
Query: 154 FSANIKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTL 213
SA IKC +CD G EY ++ F + +G I + SC T QNG +ERK R I R+L
Sbjct: 596 HSAVIKCFRCDLGGEYTSNKFCQMLALDGTIHQTSCTDTPEQNGVAERKHRHIVETARSL 655
Query: 214 LAHSSVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFF 273
L + V FW A+ A L+N +P SP ++LY P YS RVFGC F
Sbjct: 656 LLSAFVLSEFWGEAVLTAVSLINTIPSSHSSGLSPFEKLYGHVPDYSSFRVFGCTYFVLH 715
Query: 274 PSATINKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFADLPLEST 333
P NKL RS+ CVFLGY +GY+C+D +KL +S HV+F E PF +P +
Sbjct: 716 PHVERNKLSSRSAICVFLGYGEGKKGYRCFDPITQKLYVSHHVVFLE-HIPFFSIPSTTH 774
Query: 334 S-------SYDCFTEDLPPSLIHHWQTTSTRPPDLSI-PPSSPTDSTMPSPAPTSSPTSS 385
S D F+ED T P SI +S T+ S P +S +S+
Sbjct: 775 SLTKSDLIHIDPFSED---------SGNDTSPYVRSICTHNSAGTGTLLSGTPEASFSST 825
Query: 386 APLPLPPVPPTPPTRTMTTHSMHGISKPKKPFSLSVSIDDPSISPL---------PCNPK 436
AP + PP +++ I K K + S S + P + K
Sbjct: 826 APQASSEIVDPPPRQSIR------IRKSTKLPDFAYSCYSSSFTSFLAYIHCLFEPSSYK 879
Query: 437 QALSDPNWKFAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARL 496
+A+ DP + AM E +AL + +TW+LVP P +V+ +++ K S+G E YKARL
Sbjct: 880 EAILDPLGQQAMDEELSALHKTDTWDLVPLPPGKSVVGCRWVYKIKTNSDGSIERYKARL 939
Query: 497 VGDGWSQIAGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYM 556
V G+SQ G+D +ETF + K TIRT++++A R W I LDV NAFL+GDL E VYM
Sbjct: 940 VAKGYSQQYGMDYEETFAPIAKMTTIRTLIAVASIRQWHISQLDVKNAFLNGDLQEEVYM 999
Query: 557 HQPLGFRDSQHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHSYIL---- 612
P G S YVC+ KK+LYGLKQAPRAW+++F+ +SS+GF SS D + +
Sbjct: 1000 APPPGI--SHDSGYVCKLKKALYGLKQAPRAWFEKFSIVISSLGFVSSSHDSALFIKCTD 1057
Query: 613 -------LYVDDIILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLS 665
LYVDD+I+ D LA F MKDLG L YFLGI V G LS
Sbjct: 1058 AGRIILSLYVDDMIITGDDIDGISVLKTELARRFEMKDLGYLRYFLGIEVAYSPRGYLLS 1117
Query: 666 QSTYASEIIARAGMASCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPN 725
QS Y + I+ RA + TP++ + SSS G P D TLY+++ G+L YLT T P+
Sbjct: 1118 QSKYVANILERARLTDNKTVDTPIEVNARYSSSDGLPLIDPTLYRTIVGSLVYLTITHPD 1177
Query: 726 ISYVVQQVCLHMHAPHTEHMLAMFVAL*L---TVFTCIPPLLRNLFLIRMLT----GGMS 778
I+Y V V + +P T H A+ L TVF + + +R + G
Sbjct: 1178 IAYAVHVVSQFVASPTTIHWAAVLRILRYLRGTVFQSLLLSSTSSLELRAYSDADHGSDP 1237
Query: 779 CTRRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFP 838
R+S +G+C+FLGD+LISW SK+Q +S SS EAEY +A+ E W R LL ++
Sbjct: 1238 TDRKSVTGFCIFLGDSLISWKSKKQSIVSQSSTEAEYCAMASTTKEIVWSRWLLADMGIS 1297
Query: 839 LSQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADI 898
S T ++CDN SSI ++ N V H+RTKHIE+D H R + G + VPS QIAD
Sbjct: 1298 FSHLTPMYCDNQSSIQIAHNSVFHERTKHIEIDCHLTRHHLKHGTIALPFVPSSLQIADF 1357
Query: 899 FTK 901
FTK
Sbjct: 1358 FTK 1360
>UniRef100_O81824 Hypothetical protein AT4g27210 [Arabidopsis thaliana]
Length = 1318
Score = 561 bits (1447), Expect = e-158
Identities = 355/996 (35%), Positives = 479/996 (47%), Gaps = 128/996 (12%)
Query: 2 GFSVSDFQTGMPLLRCNSLGDLYPVTCSSHFAGLASG--------VWHNRLGHPTSSALN 53
G V+D T LL ++ LY + F S VWH RLGHP L
Sbjct: 266 GVRVNDKATKKLLLMGSNRDGLYCLKDDKQFQAFFSTRQRSASDEVWHRRLGHPHPQILQ 325
Query: 54 HLRNNKLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSFDILHSDLW-TSPILSTA 112
L +H DLW + I S
Sbjct: 326 PLER-----------------------------------------VHCDLWGPTTITSVQ 344
Query: 113 GHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQFSANIKCLQCDNGREYDND 172
G RYY + +D ++ F W++P+ KS Y+IF L++ Q S I QCD G E+ +
Sbjct: 345 GFRYYAVFIDHYSRFSWIYPLKLKSDFYNIFLAFHKLVENQLSQKISVFQCDGGGEFVSH 404
Query: 173 SFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLLAHSSVPPSFWHHALQMAT 232
F ++ ++G+ + SCPHT QNG +ERK R + + ++L S VP FW A A
Sbjct: 405 KFLQHLQSHGIQQQLSCPHTPQQNGLAERKHRHLVELGLSMLFQSHVPHKFWVEAFFTAN 464
Query: 233 YLLNILPRKTLRND-SPTQRLYHRDPSYSHLRVFGCLCFPFFPSATINKLQPRSSPCVFL 291
+L+N+LP L+ SP ++LY + P Y+ LR FG CFP NK P S CVFL
Sbjct: 465 FLINLLPTSALKESISPYEKLYDKKPDYTSLRSFGSACFPTLRDYAENKFNPCSLKCVFL 524
Query: 292 GYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFA----------------------DLP 329
GY ++GY+C +L ISRHVIFDES +PF+ D P
Sbjct: 525 GYNEKYKGYRCLYPPTGRLYISRHVIFDESVYPFSHTYKHLHPQPRTPLLAAWLRSSDSP 584
Query: 330 LESTSSYDC-----FTE-DLPP----------------SLIHHWQTTSTRPPDLSIPPSS 367
STS+ FT D PP S+ H T+ + PD ++
Sbjct: 585 APSTSTSPSSRSPLFTSADFPPLPQRKTPLLPTLVPISSVSHASNITTQQSPDFDSERTT 644
Query: 368 PTDSTMPSPAPTSSPTSS--------APLPLPPVPPTPPTRTMTTHSMHGISKPKKPFSL 419
DS + SS S A + + + M T + GISKP +
Sbjct: 645 DFDSASIGDSSHSSQAGSDSEETIQQASVNVHQTHASTNVHPMVTRAKVGISKPNPRY-- 702
Query: 420 SVSIDDPSISPLPCNPKQALSDPNWKFAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IF 479
V + P P AL P W AM E TW LVP D++V+ +F
Sbjct: 703 -VFLSHKVSYPEPKTVTAALKHPGWTGAMTEEIGNCSETQTWSLVPYKSDMHVLGSKWVF 761
Query: 480 RHKKQSNGLFEHYKARLVGDGWSQIAGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*L 539
R K ++G KAR+V G+ Q G+D ET++ VV+ T+R VL +A + +W I +
Sbjct: 762 RTKLHADGTLNKLKARIVAKGFLQEEGIDYLETYSPVVRTPTVRLVLHLATALNWDIKQM 821
Query: 540 DVHNAFLHGDLHETVYMHQPLGFRDSQHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSI 599
DV NAFLHGDL ETVYM QP GF D PD+VC KS+YGLKQ+PRAW+ +F+ ++
Sbjct: 822 DVKNAFLHGDLKETVYMTQPAGFVDPSKPDHVCLLHKSIYGLKQSPRAWFDKFSTFLLEF 881
Query: 600 GFQHSSSDHS-----------YILLYVDDIILVASSHDLRKSFMALLAFEFAMKDLGPLS 648
GF S SD S +LLYVDD+++ +S S +A L EF M D+G L
Sbjct: 882 GFFCSKSDPSLFIYAHNNNLILLLLYVDDMVITGNSSQTLTSLLAALNKEFRMTDMGQLH 941
Query: 649 YFLGIAVTRYVGGLFLSQSTYASEIIARAGMASCNPSATPVDTK*KLSSSSGTPCEDVTL 708
YFLGI V R GLF+SQ YA +++ A M C P TP+ + D T
Sbjct: 942 YFLGIQVQRQQNGLFMSQQKYAEDLLIAASMEHCTPLPTPLPVQLDRVPHQEELFSDPTY 1001
Query: 709 YQSLAGALQYLTFTRPNISYVVQQVCLHMHAPHTE--HMLAMF-------VAL*LTVFTC 759
++S+AG LQYLT TRP+I + V VC MH P H+L + + ++
Sbjct: 1002 FRSIAGKLQYLTLTRPDIQFAVNFVCQKMHQPTISDFHLLKRILRYIKGTITMGISYSRD 1061
Query: 760 IPPLLRNLFLIRMLTGGMSCTRRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVA 819
P LL+ G TRRS G C F+G NL+SWSSK+ PT+S SS EAEY+ ++
Sbjct: 1062 SPTLLQ--AYSDSDWGNCKQTRRSVGGLCTFMGTNLVSWSSKKHPTVSRSSTEAEYKSLS 1119
Query: 820 NVVSESCWIRNLLLELHFPLSQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKV 879
+ SE W+ LL EL PL + CDN+S++YL+ NP H RTKH ++D HFVRE+V
Sbjct: 1120 DAASEILWLSTLLRELRIPLPDTPELFCDNLSAVYLTANPAFHARTKHFDIDFHFVRERV 1179
Query: 880 ARGQARILHVPSRHQIADIFTKGLPRVLFDDFRSSL 915
A + H+P QIADIFTK LP F R L
Sbjct: 1180 ALKALVVKHIPGSEQIADIFTKSLPYEAFIHLRGKL 1215
>UniRef100_Q9SIM3 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1461
Score = 551 bits (1420), Expect = e-155
Identities = 329/947 (34%), Positives = 498/947 (51%), Gaps = 59/947 (6%)
Query: 5 VSDFQTGMPLLRCNSLGDLYPVTCSSHFAGLAS----GVWHNRLGHPTSSALNHLRNNKL 60
+ D G+ L +G+LY + S + + VWH RLGHP+ S L+ L
Sbjct: 530 IQDLTKGLTLGEGKRIGNLYVLDTQSPAISVNAVVDVSVWHKRLGHPSFSRLDSLSEVLG 589
Query: 61 IYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSFDILHSDLWTSPILSTA-GHRYYVL 119
++ S C C L K +L F S+ I +F++LH D+W + T G++Y++
Sbjct: 590 TTRHKNKKSAYCHVCHLAKQKKLSFPSANNICNSTFELLHIDVWGPFSVETVEGYKYFLT 649
Query: 120 VLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQFSANIKCLQCDNGREYDNDSFRRYCH 179
++DDH+ W++ + KS V +F L++ Q+ +K ++ DN +E +F +
Sbjct: 650 IVDDHSRATWIYLLKSKSDVLTVFPAFIDLVENQYDTRVKSVRSDNAKEL---AFTEFYK 706
Query: 180 ANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILP 239
A G++ SCP T QN ERK + I N+ R L+ S++ +W + A +L+N P
Sbjct: 707 AKGIVSFHSCPETPEQNSVVERKHQHILNVARALMFQSNMSLPYWGDCVLTAVFLINRTP 766
Query: 240 RKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFFPSATINKLQPRSSPCVFLGYPMNHRG 299
L N +P + L + P YS L+ FGCLC+ S +K PRS CVFLGYP +G
Sbjct: 767 SALLSNKTPFEVLTGKLPDYSQLKTFGCLCYSSTSSKQRHKFLPRSRACVFLGYPFGFKG 826
Query: 300 YKCYDLSNRKLIISRHVIFDESRFPFADLPLESTSSYDCFTEDLPPSLIHHWQTTSTRPP 359
YK DL + + ISR+V F E FP A +T++ D FT P
Sbjct: 827 YKLLDLESNVVHISRNVEFHEELFPLASSQQSATTASDVFT-----------------PM 869
Query: 360 DLSIPPSSPTDSTMPSPAPTSSPTSSAP----LPLPPVPPTPPTRTMTTHSMHGISKPKK 415
D P SS T P+P SP++ P + H IS
Sbjct: 870 D---PLSSGNSITSHLPSPQISPSTQISKRRITKFPAHLQDYHCYFVNKDDSHPISSSLS 926
Query: 416 PFSLSVS----IDDPSISPLPCNPKQALSDPNWKFAMQPEFNALIRNNTWELVPRPCDVN 471
+S S I++ S P+P + +A W A+ E A+ R +TWE+ P
Sbjct: 927 YSQISPSHMLYINNISKIPIPQSYHEAKDSKEWCGAIDQEIGAMERTDTWEITSLPPGKK 986
Query: 472 VIRYM*IFRHKKQSNGLFEHYKARLVGDGWSQIAGVDCDETFNLVVKPATIRTVLSIALS 531
+ +F K ++G E +KAR+V G++Q G+D ETF+ V K AT++ +L ++ S
Sbjct: 987 AVGCKWVFTVKFHADGSLERFKARIVAKGYTQKEGLDYTETFSPVAKMATVKLLLKVSAS 1046
Query: 532 RSWPIH*LDVHNAFLHGDLHETVYMHQPLGFRD----SQHPDYVCRRKKSLYGLKQAPRA 587
+ W ++ LD+ NAFL+GDL ET+YM P G+ D S P+ VCR KKS+YGLKQA R
Sbjct: 1047 KKWYLNQLDISNAFLNGDLEETIYMKLPDGYADIKGTSLPPNVVCRLKKSIYGLKQASRQ 1106
Query: 588 WYQRFADYVSSIGFQHSSSDHS-----------YILLYVDDIILVASSHDLRKSFMALLA 636
W+ +F++ + ++GF+ DH+ +L+YVDDI++ +++ +S L
Sbjct: 1107 WFLKFSNSLLALGFEKQHGDHTLFVRCIGSEFIVLLVYVDDIVIASTTEQAAQSLTEALK 1166
Query: 637 FEFAMKDLGPLSYFLGIAVTRYVGGLFLSQSTYASEIIARAGMASCNPSATPVDTK*KLS 696
F +++LGPL YFLG+ V R G+ LSQ YA E++ A M C PS+ P+ +LS
Sbjct: 1167 ASFKLRELGPLKYFLGLEVARTSEGISLSQRKYALELLTSADMLDCKPSSIPMTPNIRLS 1226
Query: 697 SSSGTPCEDVTLYQSLAGALQYLTFTRPNISYVVQQVCLHMHAPHTEHMLAMFVAL*LTV 756
+ G ED +Y+ L G L YLT TRP+I++ V ++C AP T H+ A++ L
Sbjct: 1227 KNDGLLLEDKEMYRRLVGKLMYLTITRPDITFAVNKLCQFSSAPRTAHLAAVYKVLQYIK 1286
Query: 757 FTCIPPLLRNL---FLIRMLT----GGMSCTRRSTSGYCVFLGDNLISWSSKRQPTLSHS 809
T L + ++ T G +RRST+G+ +F+G +LISW SK+QPT+S S
Sbjct: 1287 GTVGQGLFYSAEDDLTLKGYTDADWGTCPDSRRSTTGFTMFVGSSLISWRSKKQPTVSRS 1346
Query: 810 SAEAEYRGVANVVSESCWIRNLLLELHFPLSQETLVHCDNVSSIYLSGNPVHHQRTKHIE 869
SAEAEYR +A E W+ LLL L S +++ D+ +++Y++ NPV H+RTKHIE
Sbjct: 1347 SAEAEYRALALASCEMAWLSTLLLALRVH-SGVPILYSDSTAAVYIATNPVFHERTKHIE 1405
Query: 870 MDIHFVREKVARGQARILHVPSRHQIADIFTKGLPRVLFDDFRSSLS 916
+D H VREK+ GQ ++LHV ++ Q+ADI TK L F S +S
Sbjct: 1406 IDCHTVREKLDNGQLKLLHVKTKDQVADILTKPLFPYQFAHLLSKMS 1452
>UniRef100_Q7X7W7 OSJNBa0061G20.2 protein [Oryza sativa]
Length = 542
Score = 528 bits (1360), Expect = e-148
Identities = 279/532 (52%), Positives = 346/532 (64%), Gaps = 23/532 (4%)
Query: 402 MTTHSMHGISKPKKPFSLSVSIDDPSISPLPCNPKQALSDPNWKFAMQPEFNALIRNNTW 461
M T GI +P ++ + DD +P + L DP W AMQ E+ AL N TW
Sbjct: 1 MVTRWKAGIVQPNPRYAHLATADD-----VPTTVRAVLRDPAWFAAMQDEYRALQDNGTW 55
Query: 462 ELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVGDGWSQIAGVDCDETFNLVVKPAT 521
LVPRP +VI IF++K ++G E KAR V G++Q G+D D+TF+ VVKPAT
Sbjct: 56 ALVPRPRGAHVITGKWIFKNKFHADGTLERRKARWVARGFTQRPGLDFDKTFSPVVKPAT 115
Query: 522 IRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLGFRDSQHPDYVCRRKKSLYGL 581
IRTVL +A R WP+H LDV NAFLHGDL E Y HQP GF D PD VC +KSLYGL
Sbjct: 116 IRTVLHLAAMRDWPVHQLDVKNAFLHGDLTEHFYCHQPAGFVDPSQPDAVCLLRKSLYGL 175
Query: 582 KQAPRAWYQRFADYVSSIGFQHSSSDHS-----------YILLYVDDIILVASSHDLRKS 630
KQAPRAW+QRF ++ +GF S SD+S ++LLYVDDI+L ASS L +
Sbjct: 176 KQAPRAWFQRFGTHLHHLGFVSSKSDNSLFVLRRGTDEAHLLLYVDDIVLAASSQRLLQH 235
Query: 631 FMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQSTYASEIIARAGMASCNPSATPVD 690
+ L EFAMKDLGP+ +FLGI V R G FLSQ YA +++ RAG+A C P+ TP+D
Sbjct: 236 IINQLRVEFAMKDLGPVHFFLGIQVRRTADGFFLSQEQYAGDVLDRAGLADCKPAPTPID 295
Query: 691 TK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPNISYVVQQVCLHMHAPHTEHMLAMFV 750
TK K+SS++G P D T Y+S+ GALQYLT TRP++SY VQQVCLHMH+P H +
Sbjct: 296 TKAKVSSTTGQPYSDPTFYRSIVGALQYLTLTRPDLSYAVQQVCLHMHSPRDVHWTLVKR 355
Query: 751 AL*LTVFTCIPPLLRNLFLIRMLTG-------GMSCTRRSTSGYCVFLGDNLISWSSKRQ 803
L T L LT G TRRSTSG+CVF GD+ SWSSKRQ
Sbjct: 356 ILRYVRGTTHKGLQLRRSSTPSLTAYSDVDWAGCPDTRRSTSGFCVFFGDSSESWSSKRQ 415
Query: 804 PTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQETLVHCDNVSSIYLSGNPVHHQ 863
+S SSAEAEYRGVAN +E CW+R+LL ELH L + TLV+CDN+S++YLS NP+HH
Sbjct: 416 SVVSRSSAEAEYRGVANAAAECCWLRHLLGELHVKLDKATLVYCDNISAVYLSKNPLHHG 475
Query: 864 RTKHIEMDIHFVREKVARGQARILHVPSRHQIADIFTKGLPRVLFDDFRSSL 915
R KH+E+D+HFVREKVA G R+ H+P+R Q+ADI TK LP LF+DFRSSL
Sbjct: 476 RAKHVELDVHFVREKVAVGDIRVAHIPTRQQLADIMTKRLPTALFEDFRSSL 527
>UniRef100_Q5XWR5 Putative retroelement pol polyprotein-like [Solanum tuberosum]
Length = 1476
Score = 520 bits (1338), Expect = e-145
Identities = 323/910 (35%), Positives = 460/910 (50%), Gaps = 69/910 (7%)
Query: 39 VWHNRLGHPTSSALNHLRNNKLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSFDI 98
+WH RLGH S L R K+ CD C L + VRLPF S++ + FD+
Sbjct: 552 MWHKRLGHIPMSVL---RKIKMFDSPQKLVLPSCDVCPLARQVRLPFPISQSRSENCFDL 608
Query: 99 LHSDLWTSPILSTAGH-RYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQFSAN 157
+H D+W +T RY++ V+DDH+ + W+F + KS V + +I TQF
Sbjct: 609 IHLDVWGPYKAATHNKMRYFLTVVDDHSRWTWIFLMHLKSDVSTVLQNFILMIDTQFGQK 668
Query: 158 IKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLLAHS 217
IK + DNG E+ N ++G++ + SCPHT QNG ER+ + I R L
Sbjct: 669 IKIFRSDNGTEFFNAQCDGLFKSHGIVHQSSCPHTPQQNGVVERRHKHILETARALRFQG 728
Query: 218 SVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFFPSAT 277
+P FW + A +++N +P L N SP + +Y R P S++RV GCLC
Sbjct: 729 HLPIRFWGECVLSAVHIINRIPSSVLHNKSPFELMYKRSPDLSYMRVIGCLCH------- 781
Query: 278 INKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFADLPLESTSSYD 337
L +GYK YDL ++ +SR ++F+E+ FPF L
Sbjct: 782 ----------ATNLVNTSTQKGYKLYDLEHQHFFVSRDMVFNEAVFPFQSPALADPHDTP 831
Query: 338 CFTEDLPPSLIHHWQTTSTRPP--------DLSIPPSSPTDSTMPSPAPTSSPTSSAPLP 389
F PP H + +P ++ PPS+ +D + P + P
Sbjct: 832 VFLAS-PPCSSHTEDADAVQPAIITSEEIIPVASPPSAVSDDHLHPPPERRRSYRTGKPP 890
Query: 390 LPPVPPTPPTRTMTTHSMHGISKPKKPFSLSVS----IDDPSISPLPCNPKQALSDPNWK 445
+ + + + H ++ IS LS + I S+ P QA +D W
Sbjct: 891 IWQKDFITTSTSRSNHCLYPISDNIDYSCLSSTYQCYIASSSVETEPQFYYQAANDCRWV 950
Query: 446 FAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVGDGWSQIA 505
AM+ E AL N TWE+V P I +++ K +++G E +KARLV G++Q
Sbjct: 951 HAMKEEIQALEDNKTWEVVSLPKGKKAIGCKWVYKIKYKASGEIERFKARLVAKGYNQKE 1010
Query: 506 GVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLGFR-D 564
G+D ETF+ VVK T+RTVL++A+S+ W I +DV+NAFL GDL E VYM P GF+ D
Sbjct: 1011 GLDYQETFSPVVKMVTLRTVLTLAVSKGWDIQQMDVYNAFLQGDLIEEVYMQLPQGFQYD 1070
Query: 565 SQHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHSY-----------ILL 613
VCR KSLYGLKQA R W + + + GFQ S D+S +L+
Sbjct: 1071 KTGDPKVCRLLKSLYGLKQASRQWNVKLTTALLAAGFQQSHLDYSLMLKRTADGIVIVLI 1130
Query: 614 YVDDIILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQSTYASEI 673
YVDD+++ SS L +L F +KDLG L YFLG+ R G+ + Q YA E+
Sbjct: 1131 YVDDLLITGSSLQLIDDAKQVLKANFKIKDLGTLRYFLGMEFARNASGMLMHQRKYALEL 1190
Query: 674 IARAGMASCNPSATPVDTK*KL----------SSSSGTPCEDVTLYQSLAGALQYLTFTR 723
I+ G+ PS TPV+ KL SS + + D T YQ L G L YLT TR
Sbjct: 1191 ISDLGLGGSKPSVTPVELHLKLTTREFDLHVGSSGADSLLADPTEYQRLVGRLLYLTITR 1250
Query: 724 PNISYVVQQVCLHMHAPHTEHMLAMFVAL*LTVFTCIPPLLRNLFLIRMLT--------- 774
P+IS+ VQ + MHAP HM A A+ + + P L ++
Sbjct: 1251 PDISFAVQHLSQFMHAPKVSHMEA---AIRVVKYVKQAPGLGLYMAVQTADTLQAYCDAD 1307
Query: 775 -GGMSCTRRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLL 833
G TR+S +GY + G L+SW SK+QPT+S SSAEAEYR +A+ V+E W+ L
Sbjct: 1308 WGSCINTRKSITGYMIQFGSALLSWKSKKQPTISRSSAEAEYRSLASTVAELVWLTGLFK 1367
Query: 834 ELHFPLSQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRH 893
EL PLS ++CD+ ++I ++ NPV H+RTKHI++D HF+REKV G I ++P++
Sbjct: 1368 ELDMPLSLPVSLYCDSKAAIQIAANPVFHERTKHIDIDCHFIREKVQAGLVMIHYLPTQE 1427
Query: 894 QIADIFTKGL 903
Q ADI TKGL
Sbjct: 1428 QPADILTKGL 1437
>UniRef100_Q9FX79 Putative retroelement polyprotein [Arabidopsis thaliana]
Length = 1413
Score = 518 bits (1335), Expect = e-145
Identities = 297/856 (34%), Positives = 453/856 (52%), Gaps = 47/856 (5%)
Query: 72 CDSCVLGKHVRLPFSSSETITLRSFDILHSDLW---TSPILSTAGHRYYVLVLDDHTDFL 128
CD C K +L + S I L FD+LH D+W + P + G+ Y++ ++DDHT
Sbjct: 566 CDICQRAKQKKLTYPSRHNICLAPFDLLHIDVWGPFSEP--TQEGYHYFLTIVDDHTRVT 623
Query: 129 WMFPISKKSQVYDIFTTLATLIKTQFSANIKCLQCDNGREYDNDSFRRYCHANGLIFRFS 188
W++ + KS V IF T+++TQ+ +K ++ DN E F G++ S
Sbjct: 624 WVYLMKYKSDVLTIFPDFITMVETQYDTKVKAVRSDNAPEL---KFEELYRRKGIVAYHS 680
Query: 189 CPHTSSQNGKSERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLRNDSP 248
CP T QN ERK + I N+ R LL S +P S+W + A +++N P + N +
Sbjct: 681 CPETPEQNSVVERKHQHILNVARALLFQSQIPLSYWGDCILTAVFIINRTPSPVISNKTL 740
Query: 249 TQRLYHRDPSYSHLRVFGCLCFPFFPSATINKLQPRSSPCVFLGYPMNHRGYKCYDLSNR 308
+ L + P Y+HL+ FGCLC+ +K + R+ C FLGYP ++GYK DL +
Sbjct: 741 FEMLTKKVPDYTHLKSFGCLCYASTSPKQRHKFEDRARTCAFLGYPSGYKGYKLLDLESH 800
Query: 309 KLIISRHVIFDESRFPFADLPLESTSSYDCFTEDLPPSLIHHWQTTSTRPPDLSIPPSSP 368
+ ISR+V+F E FPF P E+ S F H + + P +P
Sbjct: 801 TIFISRNVVFYEDLFPFKTKPAENEESSVFFP--------HIYVDRNDSHPSQPLPVQET 852
Query: 369 TDSTMPSPAPTSSPTSSAPLPLPPVPPTPPTRTMTTHSMHGISKPKKPFSLS----VSID 424
+ S +P+ S S P L ++T+ + H IS+ SLS + I+
Sbjct: 853 SASNVPAEKQNSR-VSRPPAYLKDYH----CNSVTSSTDHPISEVLSYSSLSDPYMIFIN 907
Query: 425 DPSISPLPCNPKQALSDPNWKFAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQ 484
+ P P QA W AM E AL N TW + P + +++ K
Sbjct: 908 AVNKIPEPHTYAQARQIKEWCDAMGMEITALEDNGTWVVCSLPVGKKAVGCKWVYKIKLN 967
Query: 485 SNGLFEHYKARLVGDGWSQIAGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNA 544
++G E YKARLV G++Q G+D +TF+ V K T++ ++++A ++ W + LD+ NA
Sbjct: 968 ADGSLERYKARLVAKGYTQTEGLDYVDTFSPVAKLTTVKLLIAVAAAKGWSLSQLDISNA 1027
Query: 545 FLHGDLHETVYMHQPLGFR----DSQHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIG 600
FL+G L E +YM P G+ DS P+ VCR KKSLYGLKQA R WY +F++ + ++G
Sbjct: 1028 FLNGSLDEEIYMTLPPGYSPRQGDSFPPNAVCRLKKSLYGLKQASRQWYLKFSESLKALG 1087
Query: 601 FQHSSSDHSY-----------ILLYVDDIILVASSHDLRKSFMALLAFEFAMKDLGPLSY 649
F SS DH+ +L+YVDDII+ +S + L ++DLG L Y
Sbjct: 1088 FTQSSGDHTLFTRKSKNSYMAVLVYVDDIIIASSCDRETELLRDALQRSSKLRDLGTLRY 1147
Query: 650 FLGIAVTRYVGGLFLSQSTYASEIIARAGMASCNPSATPVDTK*KLSSSSGTPCEDVTLY 709
FLG+ + R G+ + Q Y E++A G+ C S+ P++ KLS G +D Y
Sbjct: 1148 FLGLEIARNTDGISICQRKYTLELLAETGLLGCKSSSVPMEPNQKLSQEDGELIDDAEHY 1207
Query: 710 QSLAGALQYLTFTRPNISYVVQQVCLHMHAPHTEHMLAMFVAL*LTVFTCIPPLLRNLFL 769
+ L G L YLTFTRP+I+Y V ++C AP H+ A++ + T L + +
Sbjct: 1208 RKLVGKLMYLTFTRPDITYAVHRLCQFTSAPRVPHLKAVYKIIYYLKGTVGQGLFYSANV 1267
Query: 770 IRMLTG-------GMSCTRRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVV 822
L+G S +R+ T+GYC+FLG +L++W SK+Q +S SSAEAEY+ ++ V
Sbjct: 1268 DLKLSGFADSDFSSCSDSRKLTTGYCMFLGTSLVAWKSKKQEVISMSSAEAEYKAMSMAV 1327
Query: 823 SESCWIRNLLLELHFPLSQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARG 882
E W+R LL +L +S+ ++++CDN ++I+++ NPV H+RTKHIE D H +REK+ G
Sbjct: 1328 REMMWLRFLLEDLWIDVSEASVLYCDNTAAIHIANNPVFHERTKHIERDYHHIREKIILG 1387
Query: 883 QARILHVPSRHQIADI 898
R LHV + +Q+ADI
Sbjct: 1388 LIRTLHVRTENQLADI 1403
>UniRef100_Q9XII7 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1454
Score = 514 bits (1325), Expect = e-144
Identities = 312/920 (33%), Positives = 468/920 (49%), Gaps = 73/920 (7%)
Query: 39 VWHNRLGHPTSSALNHLRNNKLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSFDI 98
+WH RLGH + L+ + ++ ++ S C C L K +L F +S + FD+
Sbjct: 557 MWHRRLGHASLQRLDAISDSLGTTRHKNKGSDFCHVCHLAKQRKLSFPTSNKVCKEIFDL 616
Query: 99 LHSDLWTSPILSTA-GHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQFSAN 157
LH D+W + T G++Y++ ++DDH+ WM+ + KS+V +F ++ Q+
Sbjct: 617 LHIDVWGPFSVETVEGYKYFLTIVDDHSRATWMYLLKTKSEVLTVFPAFIQQVENQYKVK 676
Query: 158 IKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLLAHS 217
+K ++ DN E F + G++ SCP T QN ERK + I N+ R L+ S
Sbjct: 677 VKAVRSDNAPEL---KFTSFYAEKGIVSFHSCPETPEQNSVVERKHQHILNVARALMFQS 733
Query: 218 SVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFFPSAT 277
VP S W + A +L+N P + L N +P + L P Y LR FGCLC+
Sbjct: 734 QVPLSLWGDCVLTAVFLINRTPSQLLMNKTPYEILTGTAPVYEQLRTFGCLCYSSTSPKQ 793
Query: 278 INKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFADLPLESTSSYD 337
+K QPRS C+FLGYP ++GYK DL + + ISR+V F E FP A P S SS
Sbjct: 794 RHKFQPRSRACLFLGYPSGYKGYKLMDLESNTVFISRNVQFHEEVFPLAKNP-GSESSLK 852
Query: 338 CFTEDLPPSLIHHWQTTSTRPPDLSIPPSSPTDSTMPSPAPTSSPTSSAPLPLPPVPP-- 395
FT +P S S T SP SS P + +PP
Sbjct: 853 LFTPMVPVS-------------------SGIISDTTHSP-------SSLPSQISDLPPQI 886
Query: 396 ------TPPTRTMTTHSMHGISKPKKPFSLSVSIDDPSIS-----------PLPCNPKQA 438
PP H S K P S ++S S S P+P N +A
Sbjct: 887 SSQRVRKPPAHLNDYHCNTMQSDHKYPISSTISYSKISPSHMCYINNITKIPIPTNYAEA 946
Query: 439 LSDPNWKFAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVG 498
W A+ E A+ + NTWE+ P + +F K ++G E YKARLV
Sbjct: 947 QDTKEWCEAVDAEIGAMEKTNTWEITTLPKGKKAVGCKWVFTLKFLADGNLERYKARLVA 1006
Query: 499 DGWSQIAGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQ 558
G++Q G+D +TF+ V K TI+ +L ++ S+ W + LDV NAFL+G+L E ++M
Sbjct: 1007 KGYTQKEGLDYTDTFSPVAKMTTIKLLLKVSASKKWFLKQLDVSNAFLNGELEEEIFMKI 1066
Query: 559 PLGFRDSQ----HPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHS----- 609
P G+ + + + V R K+S+YGLKQA R W+++F+ + S+GF+ + DH+
Sbjct: 1067 PEGYAERKGIVLPSNVVLRLKRSIYGLKQASRQWFKKFSSSLLSLGFKKTHGDHTLFLKM 1126
Query: 610 ------YILLYVDDIILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLF 663
+L+YVDDI++ ++S L F ++DLG L YFLG+ V R G+
Sbjct: 1127 YDGEFVIVLVYVDDIVIASTSEAAAAQLTEELDQRFKLRDLGDLKYFLGLEVARTTAGIS 1186
Query: 664 LSQSTYASEIIARAGMASCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTR 723
+ Q YA E++ GM +C P + P+ K+ G ED+ Y+ + G L YLT TR
Sbjct: 1187 ICQRKYALELLQSTGMLACKPVSVPMIPNLKMRKDDGDLIEDIEQYRRIVGKLMYLTITR 1246
Query: 724 PNISYVVQQVCLHMHAPHTEHMLAMFVAL*LTVFTCIPPLLRNLFLIRMLTG-----GMS 778
P+I++ V ++C AP T H+ A + L T L + L G S
Sbjct: 1247 PDITFAVNKLCQFSSAPRTTHLTAAYRVLQYIKGTVGQGLFYSASSDLTLKGFADSDWAS 1306
Query: 779 C--TRRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELH 836
C +RRST+ + +F+GD+LISW SK+Q T+S SSAEAEYR +A E W+ LL+ L
Sbjct: 1307 CQDSRRSTTSFTMFVGDSLISWRSKKQHTVSRSSAEAEYRALALATCEMVWLFTLLVSLQ 1366
Query: 837 FPLSQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIA 896
+++ D+ ++IY++ NPV H+RTKHI++D H VRE++ G+ ++LHV + Q+A
Sbjct: 1367 -ASPPVPILYSDSTAAIYIATNPVFHERTKHIKLDCHTVRERLDNGELKLLHVRTEDQVA 1425
Query: 897 DIFTKGLPRVLFDDFRSSLS 916
DI TK L F+ +S +S
Sbjct: 1426 DILTKPLFPYQFEHLKSKMS 1445
>UniRef100_O23588 Retrotransposon like protein [Arabidopsis thaliana]
Length = 1433
Score = 509 bits (1310), Expect = e-142
Identities = 307/894 (34%), Positives = 454/894 (50%), Gaps = 60/894 (6%)
Query: 37 SGVWHNRLGHPTSSALNHLRNN-----KLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETI 91
S WH RLGHP S ++ L + K I + S VC C L K L F S + +
Sbjct: 550 SVTWHKRLGHPAYSKIDLLSDVLNLKVKKINKEHSPVCHVCHVCHLSKQKHLSFQSRQNM 609
Query: 92 TLRSFDILHSDLWTSPILSTAGHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIK 151
+FD++H D W + T D W++ + KS V +F ++
Sbjct: 610 CSAAFDLVHIDTWGPFSVPT-------------NDATWIYLLKNKSDVLHVFPAFINMVH 656
Query: 152 TQFSANIKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIR 211
TQ+ +K ++ DN E F A+G++ SCP T QN ERK + I N+ R
Sbjct: 657 TQYQTKLKSVRSDNAHEL---KFTDLFAAHGIVAYHSCPETPEQNSVVERKHQHILNVAR 713
Query: 212 TLLAHSSVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFP 271
LL S++P FW + A +L+N LP L N SP ++L + P+Y L+ FGCLC+
Sbjct: 714 ALLFQSNIPLEFWGDCVLTAVFLINRLPTPVLNNKSPYEKLKNIPPAYESLKTFGCLCYS 773
Query: 272 FFPSATINKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFADLPLE 331
+K +PR+ CVFLGYP+ ++GYK D+ + ISRHVIF E FPF +
Sbjct: 774 STSPKQRHKFEPRARACVFLGYPLGYKGYKLLDIETHAVSISRHVIFHEDIFPF----IS 829
Query: 332 STSSYDCFTEDLPPSLIHHWQTTSTRPPDLSIPPSSPTDSTMPSPAPTSSPTSSAPLPLP 391
ST D +D P L R DL + +S D T P +SS PL
Sbjct: 830 STIKDD--IKDFFPLL-----QFPARTDDLPLEQTSIID-THPHQDVSSSKALVPFDPLS 881
Query: 392 PVPPTPPTRTMTTHSMHGISKPKKPFSLSVSIDDPSISPLPCNPKQALSDPNWKFAMQPE 451
PP H + ++P F I++ + + +P +A W AM+ E
Sbjct: 882 KRQKKPPKHLQDFHCYNNTTEPFHAF-----INNITNAVIPQRYSEAKDFKAWCDAMKEE 936
Query: 452 FNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVGDGWSQIAGVDCDE 511
A++R NTW +V P + I +F K ++G E YKARLV G++Q G+D +E
Sbjct: 937 IGAMVRTNTWSVVSLPPNKKAIGCKWVFTIKHNADGSIERYKARLVAKGYTQEEGLDYEE 996
Query: 512 TFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLGFRD----SQH 567
TF+ V K ++R +L +A W +H LD+ NAFL+GDL E +YM P G+ D +
Sbjct: 997 TFSPVAKLTSVRMMLLLAAKMKWSVHQLDISNAFLNGDLDEEIYMKIPPGYADLVGEALP 1056
Query: 568 PDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHSY-----------ILLYVD 616
P +CR KS+YGLKQA R WY + ++ + +GFQ S++DH+ +L+YVD
Sbjct: 1057 PHAICRLHKSIYGLKQASRQWYLKLSNTLKGMGFQKSNADHTLFIKYANGVLMGVLVYVD 1116
Query: 617 DIILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQSTYASEIIAR 676
DI++V++S D F A L F ++DLG YFLGI + R G+ + Q Y E+++
Sbjct: 1117 DIMIVSNSDDAVAQFTAELKSYFKLRDLGAAKYFLGIEIARSEKGISICQRKYILELLST 1176
Query: 677 AGMASCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPNISYVVQQVCLH 736
G PS+ P+D KL+ G P D T Y+ L G L YL TRP+I+Y V +C
Sbjct: 1177 TGFLGSKPSSIPLDPSVKLNKEDGVPLTDSTSYRKLVGKLMYLQITRPDIAYAVNTLCQF 1236
Query: 737 MHAPHTEHMLAMFVAL*LTVFTCIPPLLRNL---FLIRMLT----GGMSCTRRSTSGYCV 789
HAP + H+ A+ L T L + F +R T G + +RR + YC+
Sbjct: 1237 SHAPTSVHLSAVHKVLRYLKGTVGQGLFYSADDKFDLRGYTDSDFGSCTDSRRCVAAYCM 1296
Query: 790 FLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQETLVHCDN 849
F+GD L+SW SK+Q T+S S+AEAE+R ++ E W+ L + P ++CDN
Sbjct: 1297 FIGDYLVSWKSKKQDTVSMSTAEAEFRAMSQGTKEMIWLSRLFDDFKVPFIPPAYLYCDN 1356
Query: 850 VSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADIFTKGL 903
+++++ N V H+RTK +E+D + RE V G + + V + Q+AD TK +
Sbjct: 1357 TAALHIVNNSVFHERTKFVELDCYKTREAVESGFLKTMFVETGEQVADPLTKAI 1410
>UniRef100_Q7XPB1 OSJNBb0026E15.10 protein [Oryza sativa]
Length = 1449
Score = 493 bits (1268), Expect = e-137
Identities = 314/932 (33%), Positives = 471/932 (49%), Gaps = 77/932 (8%)
Query: 40 WHNRLGHPTSSALNHLRNNKLIYCDP--SRSSTVCDSCVLGKHVRLPF-SSSETITLRSF 96
WH R GH AL L +++ P + + VCD C+LGK R F + S+
Sbjct: 528 WHARYGHLNFPALRKLAQQEMVRGLPLLQQVTQVCDGCLLGKQRRAAFPTQSKYRADEHL 587
Query: 97 DILHSDLWTSPI--LSTAGHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQF 154
+++H DL PI + AG+RY++L++DD + ++W+ I K + + + +
Sbjct: 588 ELVHGDL-CGPIEPATPAGNRYFLLLVDDMSRYMWLTMIRSKDEAANAIKHFQARAEVES 646
Query: 155 SANIKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLL 214
++ L+ D G E+ + F YC G+ + + P++ QNG ER+ +TI R+++
Sbjct: 647 GRKLRALRMDRGSEFTSIEFGEYCANLGVGRQLTAPYSPQQNGVVERRNQTIVATARSMM 706
Query: 215 AHSSVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFFP 274
VP FW A+ A +LLN P K+L N +P + Y + P+ LR FGC+
Sbjct: 707 KAKGVPGRFWGEAMSTAVFLLNRSPTKSLDNQTPYEAWYGQWPAVHFLRTFGCVGHVKIT 766
Query: 275 SATINKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFADLPLESTS 334
+ KL RS+P V LGY + Y+ YD + ++ +SR V+FDE D+ +
Sbjct: 767 KPGLKKLDDRSAPMVLLGYEQGSKAYRLYDPVSERVHVSRDVVFDE------DIAWDWGP 820
Query: 335 SYDCFTEDLPPSLIHHWQTTSTRPPDLSIPPSSPTDSTMPSPAPTSSPTSSAPLPLPPVP 394
L P + TT+ + P SSPT + PSPAP S+PT+ AP P PP P
Sbjct: 821 VTPDGAPQLEPFTVEQVVTTTIG----TAPASSPTPPSPPSPAP-SAPTTPAP-PSPPSP 874
Query: 395 P-----TPPT---------------RTMTTHSMHGISKPK--KPFSLS------VSIDDP 426
TPPT R ++ G + P P L VS D+P
Sbjct: 875 EAVEFVTPPTQDSILDADADDDVVPRYRLVDNLLGNASPPGHAPRVLEQLELHVVSADEP 934
Query: 427 SISPLPCNPKQALSDPNWKFAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSN 486
+ + +A +DP+W+ AMQ E NA++ N+TW L P I +++ K+
Sbjct: 935 A------SLAEAEADPSWRGAMQDELNAIVDNDTWSLTDLPHGHRAIGLKWVYKLKRDEQ 988
Query: 487 GLFEHYKARLVGDGWSQIAGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFL 546
G YKARLV G+ Q GVD DE F LV + ++R +L++A + W +H +DV +AFL
Sbjct: 989 GAIVRYKARLVAKGYVQRQGVDFDEVFALVARLESVRLLLAVAAHQGWQVHHMDVKSAFL 1048
Query: 547 HGDLHETVYMHQPLGFRDSQHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSS 606
+G+L E VY+ QP GF D H + V R K+LYGL+QAPRAW + + S+GF SSS
Sbjct: 1049 NGELLEEVYVSQPPGFVDDNHKNKVYRLHKALYGLRQAPRAWNAKLDSSLLSLGFHRSSS 1108
Query: 607 DHS-----------YILLYVDDIILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAV 655
+H + +YVDD+I+ D +SF + F M DLG L Y+LGI V
Sbjct: 1109 EHGVYTRTRGGRRLMVGVYVDDLIITGDHDDEIRSFKGEMMKLFKMSDLGALRYYLGIEV 1168
Query: 656 TRYVGGLFLSQSTYASEIIARAGMASCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGA 715
T+ G+ L Q+ YA +I+ +AG+ CNP TP++ + KL S P D TLY+SL G+
Sbjct: 1169 TQDSDGITLGQAAYAGKILEKAGLKDCNPCQTPMEVRLKLRKGSDFPLVDATLYRSLVGS 1228
Query: 716 LQYLTFTRPNISYVVQQVCLHMHAPHTEHMLAMFVAL*LTVFT------------CIPPL 763
L+YL TRP++++ V V M +P +H+ A+ L T C P+
Sbjct: 1229 LRYLVNTRPDLAFSVGYVSRFMESPREDHLAAVRRILRYVAGTRCWGIRFGPGARCALPM 1288
Query: 764 LRNLFLIRMLTGGMSCTRRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVS 823
L + G R+STSG F+ ++W S +Q ++ SS EAEY A
Sbjct: 1289 LVGYSDSDL--AGDPDERKSTSGQIFFINGGPVTWQSSKQKVVALSSCEAEYIAAAAATC 1346
Query: 824 ESCWIRNLLLELHFPLSQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQ 883
+ W+ LL E+ L+ DN S+I L NPVHH R+KHI++ H++RE +
Sbjct: 1347 QGVWLARLLAEVLGDEITAPLLKVDNQSTISLIKNPVHHDRSKHIDVKYHYIRECAEKKL 1406
Query: 884 ARILHVPSRHQIADIFTKGLPRVLFDDFRSSL 915
++ V + Q+ DIFTK L R F + RS +
Sbjct: 1407 IEMMFVGTAEQLGDIFTKSLGRTRFQELRSKI 1438
>UniRef100_Q5XWK9 Gag-pol polyprotein-like [Solanum tuberosum]
Length = 1212
Score = 490 bits (1262), Expect = e-137
Identities = 308/829 (37%), Positives = 440/829 (52%), Gaps = 62/829 (7%)
Query: 2 GFSVSDFQTGMPLLRCNSLGDLYPV----------TCSSHFAGLASGVWHNRLGHPTSSA 51
G V D +G + + +G L+P+ C+S + VWH RLGHP S
Sbjct: 401 GCLVQDQVSGTIIAKGPKVGRLFPIHFSIPPVLSFACTS--TASKTEVWHKRLGHPNSVV 458
Query: 52 LNHLRNNKLIYCDP--SRSSTVCDSCVLGKHVRLPFSSSETITLRSFDILHSDLW-TSPI 108
L+H+ N+ L+ S +S C +C LGK LPF + + + FD++HSD+W SPI
Sbjct: 459 LSHISNSGLLGNKNKFSVASIDCSTCKLGKSKTLPFPNFGSRATKCFDVIHSDVWGISPI 518
Query: 109 LSTAGHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQFSANIKCLQCDNGRE 168
+S A +Y++ +DD++ F W++ + KS+V+ +F T I+TQFS IK L+ D+G E
Sbjct: 519 ISHAHFKYFMTFIDDYSRFTWVYFLRSKSEVFSMFKTFLAYIETQFSTCIKLLRSDSGGE 578
Query: 169 YDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLLAHSSVPPSFWHHAL 228
Y + F+++ G++ + SCP+T QNG +ERK R + ++ RTLL SSVP +W AL
Sbjct: 579 YMSYEFKKFLLDKGIVSQHSCPYTPQQNGVAERKNRHLLDVTRTLLIESSVPSKYWVEAL 638
Query: 229 QMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFFPSATINKLQPRSSPC 288
A YL+N LP K L +SP RLYH++P+YS FGC+CF P + NKL +S+ C
Sbjct: 639 STAVYLINRLPSKVLNLESPYFRLYHQNPNYSDFHTFGCVCFVHLPPSQCNKLSVQSTKC 698
Query: 289 VFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFADL-PLESTSSYDCFTEDLPPSL 347
F+GY + +G+ CYD + K ISR+V+F E+++ F + L S S EDL S
Sbjct: 699 AFMGYSTSQKGFICYDPCSHKFRISRNVVFFENQYFFPTIVDLSSVSPLLPTFEDLSSSF 758
Query: 348 --IHHWQTTSTRPPDLSIPPSSPTDSTMPSPAPTSSPTSSAPLPLPPVPPTPPTRTMTTH 405
R P L P + P T P S SS PL P + TR T
Sbjct: 759 KRFKPGFVYERRRPTLPYPNTDPPPETAPQ---LESENSSRSGPLEPTRRS--TRVSRTP 813
Query: 406 SMHGISKPKKPFSLSVSIDDPSISPLPCNPKQALSDPNWKFAMQPEFNALIRNNTWELVP 465
+ +G S S +P QA W+ AM+ E AL N+TW++V
Sbjct: 814 NWYGFSSTLSNIS------------VPSCYSQASKHECWQKAMEEELLALKENDTWDIVS 861
Query: 466 RPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVGDGWSQIAGVDCDETFNLVVKPATIRTV 525
P +V I ++ K S+G + YKARLV G Q GVD +ETF V K T+RT+
Sbjct: 862 CPSNVRPIGCKWVYSIKLHSDGTLDRYKARLVVLGNRQEYGVDYEETFAPVAKMTTVRTI 921
Query: 526 LSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLGFRDSQHPDYVCRRKKSLYGLKQAP 585
++IA S++W ++ DV NAFLHGDL E +YM P S D VC+ K+SLYGLKQAP
Sbjct: 922 IAIAASQNWSLYQKDVKNAFLHGDLKEDIYMKPPPDLFSSPTSD-VCKLKRSLYGLKQAP 980
Query: 586 RAWYQRF-----------ADYVSSIGFQHSSSDHSYILLYVDDIILVASSHDLRKSFMAL 634
RAW+ +F + Y SS+ + +S+ +L+YVDDII+ + L
Sbjct: 981 RAWFDKFRSTLLQFSFELSKYDSSLFLRKTSTSCVLLLVYVDDIIITGTDSSLITCLQQQ 1040
Query: 635 LAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQSTYASEIIARAGMASCNPSATPVDTK*K 694
L F MKDLG L+YFLG+ V G+FL+Q Y ++I+ AG+ + TP++ K
Sbjct: 1041 LKDSFHMKDLGTLTYFLGLEVHNVASGVFLNQHKYTQDLISLAGLQVSSSVDTPLEMNVK 1100
Query: 695 LSSSSGTPCEDVTLYQSLAGALQYLTFTRPNISYVVQQVCLHMHAPHTEHMLAMFVAL*L 754
G D T+++ L G+L YLT TRP+IS+ VQQV M AP H++A+ +
Sbjct: 1101 YRREEGDLLPDPTIFRQLVGSLNYLTITRPDISFAVQQVSQFMQAPRHLHLVAVCHIIRY 1160
Query: 755 TVFTCIPPLLRNLFL-----IRMLT------GGMSCTRRSTSGYCVFLG 792
+ T R LF IR+ G TRRS SG+C+FLG
Sbjct: 1161 LLGTS----TRGLFFPSGSPIRLNAFSDSDWAGCPDTRRSVSGWCMFLG 1205
>UniRef100_Q7XPI7 OSJNBb0004A17.2 protein [Oryza sativa]
Length = 1877
Score = 488 bits (1256), Expect = e-136
Identities = 315/921 (34%), Positives = 477/921 (51%), Gaps = 55/921 (5%)
Query: 40 WHNRLGHPTSSALNHLRNNKLIYCDPSRS---STVCDSCVLGKHVRLPFSSSETITLRSF 96
WH RLGH L+ + LI P VC C +H ++ SS + +T+
Sbjct: 968 WHRRLGHIGFDHLSRISGMDLIRGLPKLKVPKDLVCAPC---RHGKMTSSSHKPVTMVMT 1024
Query: 97 D----ILHSDLWTSPILSTAGHRYYVLVL-DDHTDFLWMFPISKKSQVYDIFTTLATLIK 151
D +LH D + + G ++YVLV+ DD + + W++ + K + + F +LA +
Sbjct: 1025 DGPGQLLHMDTVGPARVQSVGGKWYVLVVVDDFSRYSWVYFLESKEETFGFFQSLARSLA 1084
Query: 152 TQFSANIKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIR 211
+F ++ ++ DNG E+ N +F +C ++G+ +FS P+ QNG ERK RT+ M R
Sbjct: 1085 LEFPGALRAIRSDNGSEFKNSAFESFCDSSGVEHQFSSPYVPQQNGVVERKNRTLVEMAR 1144
Query: 212 TLLAHSSVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFP 271
T+L + P FW A+ A ++ N + +T+ + +P + + R P SHLRVFGC CF
Sbjct: 1145 TMLDEFTTPRKFWTEAISAACFISNRVFLRTILHKTPYELRFGRRPKVSHLRVFGCKCF- 1203
Query: 272 FFPSATINKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDES----RFPFAD 327
S ++K + RS +FLGY + R Y+ Y LS K++ + V FDE+ R +
Sbjct: 1204 VLKSGNLDKFESRSLDGIFLGYATHSRAYRVYVLSTNKIVETCEVTFDEASPGARPEISG 1263
Query: 328 LPLESTSSYDCFTEDLPPSLIHHWQTTSTRPPDLSIPP----SSPTDSTMPSPAPTSSPT 383
+P ES + +D S PP S PP SP+ ++ APT+S +
Sbjct: 1264 VPDESIFVDEDSDDD---------DDDSIPPPLDSTPPVQETGSPSTTSPSGDAPTTSSS 1314
Query: 384 SSAPLPLPPVPPTPPTRTMTTH---SMHGISKPKKPFSLSVSIDDPSI--SPLPCNPKQA 438
++ + PT P H SM G + + S + + + S P N A
Sbjct: 1315 AAEEIDGGTSGPTAPRHIQNRHPPDSMIGGLGERVTRNRSYELVNSAFVASFEPKNVCHA 1374
Query: 439 LSDPNWKFAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVG 498
LSD NW AM E RN W LV P NVI +F++K +G KARLV
Sbjct: 1375 LSDENWVNAMHEELENFERNKVWSLVEPPLGFNVIGTKWVFKNKLGEDGSIVRNKARLVA 1434
Query: 499 DGWSQIAGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQ 558
G++Q+ G+D +ETF V + IR +L+ A S+ + + +DV +AFL+G + E VY+ Q
Sbjct: 1435 QGFTQVEGLDFEETFAPVARLEAIRILLAFAASKGFKLFQMDVKSAFLNGVIEEEVYVKQ 1494
Query: 559 PLGFRDSQHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQ-----------HSSSD 607
P GF + + P++V + +K+LYGLKQAPRAWY+R ++ GF+ HS D
Sbjct: 1495 PPGFENPKFPNHVFKLEKALYGLKQAPRAWYERLKTFLLQNGFEMGAVDKTLFTLHSGID 1554
Query: 608 HSYILLYVDDIILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQS 667
+ +YVDDII SSH L F +++ EF M +G L++FLG+ + + G+F+ Q+
Sbjct: 1555 FLLVQIYVDDIIFGGSSHALVAQFSDVMSREFEMSMMGELTFFLGLQIKQTKEGIFVHQT 1614
Query: 668 TYASEIIARAGMASCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPNIS 727
Y+ E++ + MA C P ATP+ T L D Y+S+ G+L YLT +RP+I
Sbjct: 1615 KYSKELLKKFDMADCKPIATPMATTSSLGPDEDGEEVDQREYRSMIGSLLYLTASRPDIH 1674
Query: 728 YVVQQVCLHMHAPHTEHMLA---MFVAL*LTV-----FTCIPPLLRNLFLIRMLTGGMSC 779
+ V +P T H A MF + T+ ++C L F G
Sbjct: 1675 FSVCLCARFQASPRTSHRQAVKRMFRYIKSTLEYGIWYSCSSALSVRAFSDADF-AGCKI 1733
Query: 780 TRRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPL 839
R+STSG C FLG +L+SWSS++Q +++ S+AEAEY A+ S+ W+ + L +
Sbjct: 1734 DRKSTSGTCHFLGTSLVSWSSRKQSSVAQSTAEAEYVAAASACSQVLWMISTLKDYGLSF 1793
Query: 840 SQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADIF 899
S L+ CDN S+I ++ NPV H RTKHIE+ HF+R+ V +G + V S Q+ADIF
Sbjct: 1794 SGVPLL-CDNTSAINIAKNPVQHSRTKHIEIRYHFLRDNVEKGTIVLEFVESEKQLADIF 1852
Query: 900 TKGLPRVLFDDFRSSLSFGEP 920
TK L R F+ RS L P
Sbjct: 1853 TKPLDRSRFEFLRSELGVIHP 1873
>UniRef100_Q8H7T1 Putative Zea mays retrotransposon Opie-2 [Oryza sativa]
Length = 2145
Score = 487 bits (1254), Expect = e-136
Identities = 314/916 (34%), Positives = 476/916 (51%), Gaps = 55/916 (6%)
Query: 40 WHNRLGHPTSSALNHLRNNKLIYCDPSRS---STVCDSCVLGKHVRLPFSSSETITLRSF 96
WH RLGH L+ + LI P VC C +H ++ SS + +T+
Sbjct: 886 WHRRLGHIGFDHLSRISGMDLIRGLPKLKVPKDLVCAPC---RHGKMTSSSHKPVTMVMT 942
Query: 97 D----ILHSDLWTSPILSTAGHRYYVLVL-DDHTDFLWMFPISKKSQVYDIFTTLATLIK 151
D +LH D + + G ++YVLV+ DD + + W++ + K + + F +LA +
Sbjct: 943 DGPGQLLHMDTVGPARVQSVGGKWYVLVVVDDFSRYSWVYFLESKEETFGFFQSLARSLA 1002
Query: 152 TQFSANIKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIR 211
+F ++ ++ DNG E+ N +F +C ++G+ +FS P+ QNG ERK RT+ M R
Sbjct: 1003 LEFPGALRAIRSDNGSEFKNSAFESFCDSSGVEHQFSSPYVPQQNGVVERKNRTLVEMAR 1062
Query: 212 TLLAHSSVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFP 271
T+L + P FW A+ A ++ N + +T+ + +P + + R P SHLRVFGC CF
Sbjct: 1063 TMLDEFTTPRKFWTEAISAACFISNRVFLRTILHKTPYELRFGRRPKVSHLRVFGCKCF- 1121
Query: 272 FFPSATINKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDES----RFPFAD 327
S ++K + RS +FLGY + R Y+ Y LS K++ + V FDE+ R +
Sbjct: 1122 VLKSGNLDKFESRSLDGIFLGYATHSRAYRVYVLSTNKIVETCEVTFDEASPGARPEISG 1181
Query: 328 LPLESTSSYDCFTEDLPPSLIHHWQTTSTRPPDLSIPP----SSPTDSTMPSPAPTSSPT 383
+P ES + +D S PP S PP SP+ ++ APT+S +
Sbjct: 1182 VPDESIFVDEDSDDD---------DDDSIPPPLDSTPPVQETGSPSTTSPSGDAPTTSSS 1232
Query: 384 SSAPLPLPPVPPTPPTRTMTTH---SMHGISKPKKPFSLSVSIDDPSI--SPLPCNPKQA 438
++ + PT P H SM G + + S + + + S P N A
Sbjct: 1233 AAEEIDGGTSGPTAPRHIQNRHPPDSMIGGLGERVTRNRSYELVNSAFVASFEPKNVCHA 1292
Query: 439 LSDPNWKFAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVG 498
LSD NW AM E RN W LV P NVI +F++K +G KARLV
Sbjct: 1293 LSDENWVNAMHEELENFERNKVWSLVEPPLGFNVIGTKWVFKNKLGEDGSIVRNKARLVA 1352
Query: 499 DGWSQIAGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQ 558
G++Q+ G+D +ETF V + IR +L+ A S+ + + +DV +AFL+G + E VY+ Q
Sbjct: 1353 QGFTQVEGLDFEETFAPVARLEAIRILLAFAASKGFKLFQMDVKSAFLNGVIEEEVYVKQ 1412
Query: 559 PLGFRDSQHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQ-----------HSSSD 607
P GF + + P++V + +K+LYGLKQAPRAWY+R ++ GF+ HS D
Sbjct: 1413 PPGFENPKFPNHVFKLEKALYGLKQAPRAWYERLKTFLLQNGFEMGAVDKTLFTLHSGID 1472
Query: 608 HSYILLYVDDIILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQS 667
+ +YVDDII SSH L F +++ EF M +G L++FLG+ + + G+F+ Q+
Sbjct: 1473 FLLVQIYVDDIIFGGSSHALVAQFSDVMSREFEMSMMGELTFFLGLQIKQTKEGIFVHQT 1532
Query: 668 TYASEIIARAGMASCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPNIS 727
Y+ E++ + MA C P ATP+ T L D Y+S+ G+L YLT +RP+I
Sbjct: 1533 KYSKELLKKFDMADCKPIATPMATTSSLGPDEDGEEVDQREYRSMIGSLLYLTASRPDIH 1592
Query: 728 YVVQQVCLHMHAPHTEHMLA---MFVAL*LTV-----FTCIPPLLRNLFLIRMLTGGMSC 779
+ V +P T H A MF + T+ ++C L F G
Sbjct: 1593 FSVCLCARFQASPRTSHRQAVKRMFRYIKSTLEYGIWYSCSSALSVRAFSDADF-AGCKI 1651
Query: 780 TRRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPL 839
R+STSG C FLG +L+SWSS++Q +++ S+AEAEY A+ S+ W+ + L +
Sbjct: 1652 DRKSTSGTCHFLGTSLVSWSSRKQSSVAQSTAEAEYVAAASACSQVLWMISTLKDYGLSF 1711
Query: 840 SQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADIF 899
S L+ CDN S+I ++ NPV H RTKHIE+ HF+R+ V +G + V S Q+ADIF
Sbjct: 1712 SGVPLL-CDNTSAINIAKNPVQHSRTKHIEIRYHFLRDNVEKGTIVLEFVESEKQLADIF 1770
Query: 900 TKGLPRVLFDDFRSSL 915
TK L R F+ RS L
Sbjct: 1771 TKPLDRSRFEFLRSEL 1786
>UniRef100_Q852C7 Putative gag-pol polyprotein [Oryza sativa]
Length = 1969
Score = 487 bits (1253), Expect = e-136
Identities = 313/916 (34%), Positives = 477/916 (51%), Gaps = 55/916 (6%)
Query: 40 WHNRLGHPTSSALNHLRNNKLIYCDPS---RSSTVCDSCVLGKHVRLPFSSSETITLRSF 96
WH RLGH L+ + LI P + VC C +H ++ SS + +T+
Sbjct: 886 WHRRLGHIGFDHLSRISGMDLIRGLPKLKVQKDLVCAPC---RHGKMTSSSHKPVTMVMT 942
Query: 97 D----ILHSDLWTSPILSTAGHRYYVLVL-DDHTDFLWMFPISKKSQVYDIFTTLATLIK 151
D +LH D + + G ++YVLV+ DD + + W++ + K + + F +LA +
Sbjct: 943 DGPGQLLHMDTVGPARVQSVGGKWYVLVVVDDFSRYSWVYFLESKEETFGFFQSLARSLA 1002
Query: 152 TQFSANIKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIR 211
+F ++ ++ DNG E+ N +F +C ++G+ +FS P+ QNG ERK RT+ M R
Sbjct: 1003 LEFPGALRAIRSDNGSEFKNSAFESFCDSSGVEHQFSSPYVPQQNGVVERKNRTLVEMAR 1062
Query: 212 TLLAHSSVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFP 271
T+L + P FW A+ A ++ N + +T+ + +P + + R P SHLRVFGC CF
Sbjct: 1063 TMLDEFTTPRKFWTEAISAACFISNRVFLRTILHKTPYELRFGRRPKVSHLRVFGCKCF- 1121
Query: 272 FFPSATINKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDES----RFPFAD 327
S ++K + RS +FLGY + R Y+ Y LS K++ + V FDE+ R +
Sbjct: 1122 VLKSGNLDKFESRSLDGIFLGYATHSRAYRVYVLSTNKIVETCEVTFDEASPGARPEISG 1181
Query: 328 LPLESTSSYDCFTEDLPPSLIHHWQTTSTRPPDLSIPP----SSPTDSTMPSPAPTSSPT 383
+P ES + +D S PP S PP SP+ ++ APT+S +
Sbjct: 1182 VPDESIFVDEDSDDD---------DDDSIPPPLDSTPPVQETGSPSTTSPSGDAPTTSSS 1232
Query: 384 SSAPLPLPPVPPTPPTRTMTTH---SMHGISKPKKPFSLSVSIDDPSI--SPLPCNPKQA 438
++ + PT P H SM G + + S + + + S P N A
Sbjct: 1233 AAEEIDGGTSGPTAPRHIQNRHPPDSMIGGLGERVTRNRSYELVNSAFVASFEPKNVCHA 1292
Query: 439 LSDPNWKFAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVG 498
LSD NW AM E RN W LV P NVI +F++K +G KARLV
Sbjct: 1293 LSDENWVNAMHEELENFERNKVWSLVEPPLGFNVIGTKWVFKNKLGEDGSIVRNKARLVA 1352
Query: 499 DGWSQIAGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQ 558
G++Q+ G+D +ETF V + IR +L+ A S+ + + +DV +AFL+G + E VY+ Q
Sbjct: 1353 QGFTQVEGLDFEETFAPVARLEAIRILLAFAASKGFKLFQMDVKSAFLNGVIEEEVYVKQ 1412
Query: 559 PLGFRDSQHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQ-----------HSSSD 607
P GF + + P++V + +K+LYGLKQAPRAWY+R ++ GF+ HS D
Sbjct: 1413 PPGFENPKFPNHVFKLEKALYGLKQAPRAWYERLKTFLLQNGFEMGAVDKTLFTLHSGID 1472
Query: 608 HSYILLYVDDIILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQS 667
+ +YVDDII SSH L F +++ EF M +G L++FLG+ + + G+F+ Q+
Sbjct: 1473 FLLVQIYVDDIIFGGSSHALVAQFSDVMSREFEMSMMGELTFFLGLQIKQTKEGIFVHQT 1532
Query: 668 TYASEIIARAGMASCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPNIS 727
Y+ E++ + MA C P ATP+ T L D Y+S+ G+L YLT +RP+I
Sbjct: 1533 KYSKELLKKFDMADCKPIATPMATTSSLGPDEDGEEVDQREYRSMIGSLLYLTASRPDIH 1592
Query: 728 YVVQQVCLHMHAPHTEHMLA---MFVAL*LTV-----FTCIPPLLRNLFLIRMLTGGMSC 779
+ V +P T H A +F + T+ ++C L F G
Sbjct: 1593 FSVCLCARFQASPRTSHRQAVKRIFRYIKSTLEYGIWYSCSSALSVRAFSDADF-AGCKI 1651
Query: 780 TRRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPL 839
R+STSG C FLG +L+SWSS++Q +++ S+AEAEY A+ S+ W+ + L +
Sbjct: 1652 DRKSTSGTCHFLGTSLVSWSSRKQSSVAQSTAEAEYVAAASACSQVLWMISTLKDYGLSF 1711
Query: 840 SQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADIF 899
S L+ CDN S+I ++ NPV H RTKHIE+ HF+R+ V +G + V S Q+ADIF
Sbjct: 1712 SGVPLL-CDNTSAINIAKNPVQHSRTKHIEIRYHFLRDNVEKGTIVLEFVESEKQLADIF 1770
Query: 900 TKGLPRVLFDDFRSSL 915
TK L R F+ RS L
Sbjct: 1771 TKPLDRSRFEFLRSEL 1786
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.325 0.138 0.439
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,570,526,666
Number of Sequences: 2790947
Number of extensions: 68805269
Number of successful extensions: 564270
Number of sequences better than 10.0: 5204
Number of HSP's better than 10.0 without gapping: 2474
Number of HSP's successfully gapped in prelim test: 3058
Number of HSP's that attempted gapping in prelim test: 463325
Number of HSP's gapped (non-prelim): 36414
length of query: 921
length of database: 848,049,833
effective HSP length: 137
effective length of query: 784
effective length of database: 465,690,094
effective search space: 365101033696
effective search space used: 365101033696
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 80 (35.4 bits)
Lotus: description of TM0032.12


