

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0030.7
(504 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
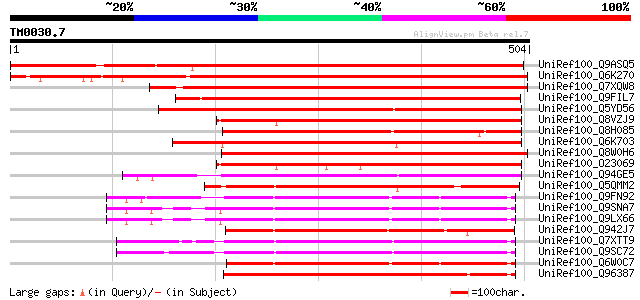
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9ASQ5 At2g11520/F14P14.15 [Arabidopsis thaliana] 560 e-158
UniRef100_Q6K270 Nodulation receptor kinase-like protein [Oryza ... 517 e-145
UniRef100_Q7XQW8 OSJNBa0014K14.7 protein [Oryza sativa] 478 e-133
UniRef100_Q9FIL7 Similarity to protein kinase [Arabidopsis thali... 345 2e-93
UniRef100_Q5YD56 Calcium/calmodulin-regulated receptor-like kina... 338 3e-91
UniRef100_Q8VZJ9 Hypothetical protein At4g00330 [Arabidopsis tha... 337 5e-91
UniRef100_Q8H085 Hypothetical protein OSJNBb0050N02.6 [Oryza sat... 328 2e-88
UniRef100_Q6K703 Receptor protein kinase-like [Oryza sativa] 328 3e-88
UniRef100_Q8W0H6 P0529E05.2 protein [Oryza sativa] 325 2e-87
UniRef100_O23069 A_IG005I10.8 protein [Arabidopsis thaliana] 325 2e-87
UniRef100_Q94GE5 Putative receptor kinase [Oryza sativa] 269 1e-70
UniRef100_Q5QMM2 Protein kinase-like [Oryza sativa] 246 1e-63
UniRef100_Q9FN92 Receptor-like protein kinase [Arabidopsis thali... 244 4e-63
UniRef100_Q9SNA7 Receptor-like protein kinase homolog [Arabidops... 244 5e-63
UniRef100_Q9LX66 Receptor protein kinase-like [Arabidopsis thali... 244 5e-63
UniRef100_Q942J7 Putative receptor protein kinase [Oryza sativa] 244 5e-63
UniRef100_Q7XTT9 OSJNBa0058K23.13 protein [Oryza sativa] 243 7e-63
UniRef100_Q9SC72 L1332.5 protein [Oryza sativa] 242 2e-62
UniRef100_Q6W0C7 Pto-like serine/threonine kinase [Capsicum chin... 242 2e-62
UniRef100_Q96387 Receptor-like protein kinase precursor [Cathara... 241 5e-62
>UniRef100_Q9ASQ5 At2g11520/F14P14.15 [Arabidopsis thaliana]
Length = 510
Score = 560 bits (1444), Expect = e-158
Identities = 290/504 (57%), Positives = 368/504 (72%), Gaps = 13/504 (2%)
Query: 1 MAITVLY-LVILLQLSTFLGSSVVLQSKECGTDWLAHSYSSHGQELFYINGNVVNQVSFC 59
+ IT L+ L++LLQ+ S+ + S C +D L ++ LF INGN V ++ FC
Sbjct: 11 LVITALFGLLMLLQIKETSASTSFVSSSVCKSDHLTYTKPYQQGSLFTINGNPVEKLRFC 70
Query: 60 EALQLYIANGCDVKDYFGSSGCAVGGSFVNFPSKAGRKLLQKDSSKESTSQGDSKALPTN 119
EAL+ + ANGC +D F C + S GR+ L++ + K+S +
Sbjct: 71 EALRFHKANGCIFEDSFSDDFCTIH-------SLLGRRFLEEKTVKDSKNSKPKTEYSHV 123
Query: 120 KVGIFAGGALLACCAVLCPCFYGKKRKATAHAVLAKDPISMDSATSFDASVSASVSG--- 176
KV I G LL CCA+ CPCF+ K+RKA +H VL K+ S+ +SF+ S S+
Sbjct: 124 KVSIAGSGFLLLCCALCCPCFH-KERKANSHEVLPKESNSVHQVSSFEMSPSSEKIPQSP 182
Query: 177 -KIPASPLRVPSSPSRFSMSPKLSRLHTLNLNLSQVVRATRNFSETLQIGQGGFGTVYKA 235
+ P SP RVP SPSR++MSP+ SRL LNL +SQ+ AT NF+++ QIG+GGFG V+K
Sbjct: 183 FRAPPSPSRVPQSPSRYAMSPRPSRLGPLNLTMSQINTATGNFADSHQIGEGGFGVVFKG 242
Query: 236 NLEDGLVVAVKRAKREHFESLRTEFSSEVELLAKIDHRNLVKLLGFIDKGNERILITEYV 295
L+DG VVA+KRAK+EHFE+LRTEF SEV+LL+KI HRNLVKLLG++DKG+ER++ITEYV
Sbjct: 243 VLDDGQVVAIKRAKKEHFENLRTEFKSEVDLLSKIGHRNLVKLLGYVDKGDERLIITEYV 302
Query: 296 PNGTLREHLDGLRGKILDFNQRLEIAIDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESM 355
NGTLR+HLDG RG L+FNQRLEI IDV HGLTYLH YAE+QIIHRD+KSSNILLT+SM
Sbjct: 303 RNGTLRDHLDGARGTKLNFNQRLEIVIDVCHGLTYLHSYAERQIIHRDIKSSNILLTDSM 362
Query: 356 RAKVADFGFARLGPVNGDQTHISTKVKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEIL 415
RAKVADFGFAR GP + +QTHI T+VKGTVGYLDPEYMKT+ LT KSDVYSFGILL+EIL
Sbjct: 363 RAKVADFGFARGGPTDSNQTHILTQVKGTVGYLDPEYMKTYHLTAKSDVYSFGILLVEIL 422
Query: 416 TGRRPVELKKAADERVTLRWAFRKYNEGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAP 475
TGRRPVE K+ DER+T+RWAF KYNEG V EL+DP E V+ +L KM L+FQCAAP
Sbjct: 423 TGRRPVEAKRLPDERITVRWAFDKYNEGRVFELVDPNARERVDEKILRKMFSLAFQCAAP 482
Query: 476 IRTDRPNMKSVGEQLWAIRADYLK 499
+ +RP+M++VG+QLWAIR+ YL+
Sbjct: 483 TKKERPDMEAVGKQLWAIRSSYLR 506
>UniRef100_Q6K270 Nodulation receptor kinase-like protein [Oryza sativa]
Length = 526
Score = 517 bits (1331), Expect = e-145
Identities = 281/532 (52%), Positives = 371/532 (68%), Gaps = 36/532 (6%)
Query: 1 MAITVLYLVILLQLSTFLGSSVVLQSKE-------CGTDWLAHSYSSHGQELFYINGNVV 53
MA L+ +LL L + SS+ L S CG D +A +S G +NG +V
Sbjct: 1 MASFPLFPALLLLLCS---SSLALDSVSEPTLSWTCGDDQVAILDTSDGGRNLSVNGELV 57
Query: 54 -NQVSFCEALQLYIANGC-----DVKDYFGS--SGCAVGGSFVNFPSKAGRKLLQKDSSK 105
++V C+ L+ Y + C + + G+ C G N RKLL++ S
Sbjct: 58 QDRVLGCQKLRSYYVSRCLRCGQQSEAWRGAWKHYCREGSESSN-AQNIPRKLLRQPSMN 116
Query: 106 EST--------------SQGDSKALPTNKVGIFAGGALLACCAVLCPCFYGKKRKATAHA 151
++ +Q D+ +L + G +L CC ++ PCF+ +K++ + H
Sbjct: 117 DAKIEDDPCKNMGIHGHNQDDNDSLEGQDHLLAVPGVILLCCGLMIPCFHAEKKEVSRHN 176
Query: 152 VLAKDPISMDSATSFDASVSASVSGKIPASPLRVPSSPSRFSMSPKLSRLHTLNLNLSQV 211
+ +++S S D S S S K+P +P R+P SPSRF+ SP+++R+ ++NL + Q+
Sbjct: 177 TTSIQRNAVESIASLDVSTS---SEKVPPTPHRIPPSPSRFAQSPQIARVGSVNLTVQQI 233
Query: 212 VRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVELLAKID 271
+RAT+NFS + ++G+GGFGTVY+A L DG VVAVKRAK++ F R EFS+EVELLAKID
Sbjct: 234 LRATQNFSPSFKLGEGGFGTVYRAVLPDGQVVAVKRAKKDQFAGPRDEFSNEVELLAKID 293
Query: 272 HRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAHGLTYL 331
HRNLV+LLGF DKG+ERI+ITEYVPNGTLREHLDG G+ LDFNQRLEIAIDVAH LTYL
Sbjct: 294 HRNLVRLLGFTDKGHERIIITEYVPNGTLREHLDGQYGRTLDFNQRLEIAIDVAHALTYL 353
Query: 332 HLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGYLDPE 391
HLYAEK IIHRDVKSSNILLTES RAKV+DFGFAR GP + ++THISTKVKGT GYLDPE
Sbjct: 354 HLYAEKTIIHRDVKSSNILLTESYRAKVSDFGFARSGPSDTEKTHISTKVKGTAGYLDPE 413
Query: 392 YMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGSVVELMDP 451
Y++T+QLTPKSDV+SFGILL+EIL+ RRPVELK+AA+ER+T+RW F+K+NEG+ E++DP
Sbjct: 414 YLRTYQLTPKSDVFSFGILLVEILSARRPVELKRAAEERITIRWTFKKFNEGNRREILDP 473
Query: 452 LMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLWAIRADYLKSARR 503
L+E+ V+ +VL ++L+L+FQCAAP R DRP MK VGEQLW IR +Y KS RR
Sbjct: 474 LLEDPVDDEVLERLLNLAFQCAAPTREDRPTMKEVGEQLWEIRKEYGKSVRR 525
>UniRef100_Q7XQW8 OSJNBa0014K14.7 protein [Oryza sativa]
Length = 367
Score = 478 bits (1229), Expect = e-133
Identities = 233/368 (63%), Positives = 288/368 (77%), Gaps = 6/368 (1%)
Query: 136 LCPCFYGKKRKATAHAVLAKDPISMDSATSFDASVSASVSGKIPASPLRVPSSPSRFSMS 195
+CPCF +++ + VL +D S++S S S+S ++P SPLRVP+SPSRFS+S
Sbjct: 1 MCPCFGSRRKDGSEDPVLGRDGNSLNS------SELRSMSDRVPPSPLRVPASPSRFSLS 54
Query: 196 PKLSRLHTLNLNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFES 255
SR LNL+L QV++ T NF+ L IG+G FG VY+A L DG +VA+KRAK EHF S
Sbjct: 55 SSPSRNEPLNLSLEQVIKLTHNFAPDLMIGEGYFGKVYRAQLRDGHIVAIKRAKMEHFAS 114
Query: 256 LRTEFSSEVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFN 315
LR EFS+E+ LL KI+HRNLV+LLG+IDK NERI+ITEYVPNGTLREHLDG RG +L FN
Sbjct: 115 LRAEFSNEIALLKKIEHRNLVQLLGYIDKRNERIVITEYVPNGTLREHLDGQRGLVLSFN 174
Query: 316 QRLEIAIDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQT 375
QRLEIAIDVAHGLTYLHLYAEK IIHRDVKSSNILL E RAKVADFGFAR GP DQ+
Sbjct: 175 QRLEIAIDVAHGLTYLHLYAEKPIIHRDVKSSNILLNEGFRAKVADFGFARTGPTEPDQS 234
Query: 376 HISTKVKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRW 435
I T V+GT GY+DPEY++T+ LT KSDV+S+G+LLLEIL+GRRP+E+++AA ER+T+RW
Sbjct: 235 QIQTDVRGTAGYVDPEYLRTNHLTVKSDVFSYGVLLLEILSGRRPIEVRRAARERITVRW 294
Query: 436 AFRKYNEGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLWAIRA 495
AF KYN G V E++DP++ E+VN D+L K+ D++FQC AP R DRP MK V E+LW IR
Sbjct: 295 AFEKYNRGDVKEILDPMLTESVNEDILNKIFDVAFQCVAPTRADRPTMKEVAERLWKIRR 354
Query: 496 DYLKSARR 503
DY K+ RR
Sbjct: 355 DYAKTQRR 362
>UniRef100_Q9FIL7 Similarity to protein kinase [Arabidopsis thaliana]
Length = 470
Score = 345 bits (884), Expect = 2e-93
Identities = 176/337 (52%), Positives = 237/337 (70%), Gaps = 3/337 (0%)
Query: 162 SATSFDASVSASVSGKIPASPLRVPSSPSRFSMSPKLSRLHTLNLNLSQVVRATRNFSET 221
S +SF S VSG+ S R SP + S S K + + ++ RAT NFS
Sbjct: 92 SPSSFGRSTERKVSGQYRFSGSRF-QSPGKDSSSSKSWHQGPVIFSFGELQRATANFSSV 150
Query: 222 LQIGQGGFGTVYKANLEDGLVVAVKRAKREHF-ESLRTEFSSEVELLAKIDHRNLVKLLG 280
QIG+GGFGTV+K L+DG +VA+KRA++ ++ +S EF +E+ L+KI+H NLVKL G
Sbjct: 151 HQIGEGGFGTVFKGKLDDGTIVAIKRARKNNYGKSWLLEFKNEIYTLSKIEHMNLVKLYG 210
Query: 281 FIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAHGLTYLHLYAEKQII 340
F++ G+E++++ EYV NG LREHLDGLRG L+ +RLEIAIDVAH LTYLH Y + II
Sbjct: 211 FLEHGDEKVIVVEYVANGNLREHLDGLRGNRLEMAERLEIAIDVAHALTYLHTYTDSPII 270
Query: 341 HRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGYLDPEYMKTHQLTP 400
HRD+K+SNIL+T +RAKVADFGFARL + THIST+VKG+ GY+DP+Y++T QLT
Sbjct: 271 HRDIKASNILITNKLRAKVADFGFARLVSEDLGATHISTQVKGSAGYVDPDYLRTFQLTD 330
Query: 401 KSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGSVVELMDPLMEEAVNA- 459
KSDVYSFG+LL+EILTGRRP+ELK+ +R+T++WA R+ + V +MDP ++ A
Sbjct: 331 KSDVYSFGVLLVEILTGRRPIELKRPRKDRLTVKWALRRLKDDEAVLIMDPFLKRNRAAI 390
Query: 460 DVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLWAIRAD 496
+V KML L+ +C P R RP MK + E+LWAIR +
Sbjct: 391 EVAEKMLRLASECVTPTRATRPAMKGIAEKLWAIRRE 427
>UniRef100_Q5YD56 Calcium/calmodulin-regulated receptor-like kinase [Medicago sativa]
Length = 456
Score = 338 bits (866), Expect = 3e-91
Identities = 177/355 (49%), Positives = 235/355 (65%), Gaps = 3/355 (0%)
Query: 145 RKATAHAVLAKDPISMDSATSFDASVSASVS-GKIPASPLRVPSSPSRFSMSPKLSRLHT 203
R+ TA + D S S S +S G + + S S S S +L T
Sbjct: 55 RRKTASKINGSDDRKNTSKPRLMLSSSTDLSSGSSNKNSSKWRFSSSYASSSTTSEQLGT 114
Query: 204 LNLNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSE 263
N ++ ++T FS QIG+GGFGTVY+ L DG +VAVKRAK+E +S EF +E
Sbjct: 115 GNFTFEEIYKSTAKFSSDNQIGEGGFGTVYRGKLNDGTIVAVKRAKKEALQSHLYEFKNE 174
Query: 264 VELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAID 323
+ L+KI+H NLV+L G+++ G+E+++I EYV NG L EHLDG+RG L+ +RL+IAID
Sbjct: 175 IYTLSKIEHLNLVRLYGYLEHGDEKLIIVEYVGNGNLTEHLDGIRGDGLEIGERLDIAID 234
Query: 324 VAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKG 383
+AH +TYLH+Y + IIHRD+K+SNIL++E++RAKVADFGFARL G THIST+VKG
Sbjct: 235 IAHAITYLHMYTDNPIIHRDIKASNILISENLRAKVADFGFARLSEDPG-ATHISTQVKG 293
Query: 384 TVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEG 443
T GY+DPEY++T+QLT KSDVYSFG+LL+E++TGR PVE KK DERVT+RWA + G
Sbjct: 294 TAGYMDPEYLRTYQLTEKSDVYSFGVLLVEMMTGRHPVEPKKKIDERVTIRWAMKMLKNG 353
Query: 444 SVVELMDP-LMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLWAIRADY 497
V MDP L + V+ K+ L+FQC AP RP MK+ E LW IR D+
Sbjct: 354 DAVFAMDPRLRRSPASIKVVKKVFKLAFQCLAPSIHSRPAMKNCAEVLWGIRKDF 408
>UniRef100_Q8VZJ9 Hypothetical protein At4g00330 [Arabidopsis thaliana]
Length = 411
Score = 337 bits (864), Expect = 5e-91
Identities = 167/301 (55%), Positives = 216/301 (71%), Gaps = 6/301 (1%)
Query: 202 HTLNLNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLR---T 258
HT ++ AT+NFS + +IGQGGFGTVYK L DG AVKRAK+ + +
Sbjct: 104 HT-RFTFDEIYDATKNFSPSFRIGQGGFGTVYKVKLRDGKTFAVKRAKKSMHDDRQGADA 162
Query: 259 EFSSEVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRL 318
EF SE++ LA++ H +LVK GF+ +E+IL+ EYV NGTLR+HLD GK LD RL
Sbjct: 163 EFMSEIQTLAQVTHLSLVKYYGFVVHNDEKILVVEYVANGTLRDHLDCKEGKTLDMATRL 222
Query: 319 EIAIDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGP-VNGDQTHI 377
+IA DVAH +TYLH+Y + IIHRD+KSSNILLTE+ RAKVADFGFARL P + TH+
Sbjct: 223 DIATDVAHAITYLHMYTQPPIIHRDIKSSNILLTENYRAKVADFGFARLAPDTDSGATHV 282
Query: 378 STKVKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAF 437
ST+VKGT GYLDPEY+ T+QLT KSDVYSFG+LL+E+LTGRRP+EL + ER+T+RWA
Sbjct: 283 STQVKGTAGYLDPEYLTTYQLTEKSDVYSFGVLLVELLTGRRPIELSRGQKERITIRWAI 342
Query: 438 RKYNEGSVVELMDPLMEE-AVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLWAIRAD 496
+K+ G + ++DP +E+ + N L K+L+++FQC AP R RP+MK E LW IR D
Sbjct: 343 KKFTSGDTISVLDPKLEQNSANNLALEKVLEMAFQCLAPHRRSRPSMKKCSEILWGIRKD 402
Query: 497 Y 497
Y
Sbjct: 403 Y 403
>UniRef100_Q8H085 Hypothetical protein OSJNBb0050N02.6 [Oryza sativa]
Length = 430
Score = 328 bits (842), Expect = 2e-88
Identities = 160/294 (54%), Positives = 223/294 (75%), Gaps = 6/294 (2%)
Query: 207 NLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFES-LRTEFSSEVE 265
+L Q+ +AT+NFS L+IGQGG GTVYK L DG ++AVKRAK+ ++ + EF +E+E
Sbjct: 129 SLPQIQKATKNFSPNLKIGQGGSGTVYKGQLNDGTLIAVKRAKKNVYDKHMGREFRNEIE 188
Query: 266 LLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVA 325
L I+H NLV+ G+++ G E+++I EYVPNG LREHLD + GKIL+F+ RL+I+IDVA
Sbjct: 189 TLQCIEHLNLVRFHGYLEFGGEQLIIVEYVPNGNLREHLDCVNGKILEFSLRLDISIDVA 248
Query: 326 HGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTV 385
H +TYLH Y++ +IHRD+KSSNILLT + RAKVADFGFA+L P D +H+ST+VKGT
Sbjct: 249 HAVTYLHTYSDHPVIHRDIKSSNILLTNNCRAKVADFGFAKLAPT--DASHVSTQVKGTA 306
Query: 386 GYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGSV 445
GYLDPEY++T+QL KSDVYSFG+LL+E++TGRRP+E ++A ERVT +WA K+ EG+
Sbjct: 307 GYLDPEYLRTYQLNEKSDVYSFGVLLVELITGRRPIEPRRAIVERVTAKWAMEKFVEGNA 366
Query: 446 VELMDPLME--EAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLWAIRADY 497
++ +DP +E +A+N V K +L+ QC A + +RP+M+ E LW+IR D+
Sbjct: 367 IQTLDPNLEATDAINLAV-EKTYELALQCLATTKRNRPSMRRCAEILWSIRKDF 419
>UniRef100_Q6K703 Receptor protein kinase-like [Oryza sativa]
Length = 472
Score = 328 bits (840), Expect = 3e-88
Identities = 173/360 (48%), Positives = 234/360 (64%), Gaps = 21/360 (5%)
Query: 159 SMDSATSFDASVSASVSGKIPASPLRVPSSPSRFSMSPKLSRLHTLN------------- 205
S +ATS S + S + P ++ P P S K S N
Sbjct: 74 SSAAATSASGSNAKRRSRREPNDLIKKPPLPGPGSDQGKASMRGLYNSSRGRGIATQFQS 133
Query: 206 --LNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAK-REHFESLRTEFSS 262
++ +++RAT NFS L++GQGGFG VY+ L DG +VAVKRAK R+ + EF S
Sbjct: 134 SVFSMEEILRATNNFSPALKVGQGGFGAVYRGVLPDGTLVAVKRAKLRDQNPHVDVEFRS 193
Query: 263 EVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAI 322
EV+ +A+I+H++LV+ G+++ G ER+++ E+VPNGTLREHLD G+ LD RLEIAI
Sbjct: 194 EVKAMARIEHQSLVRFYGYLECGQERVIVVEFVPNGTLREHLDRCNGRFLDMGARLEIAI 253
Query: 323 DVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQ----THIS 378
DVAH +TYLH+YA+ IIHRD+KSSN+LLT S+RAKV DFGFARLG TH++
Sbjct: 254 DVAHAVTYLHMYADHPIIHRDIKSSNVLLTPSLRAKVGDFGFARLGVGEAGAADGVTHVT 313
Query: 379 TKVKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFR 438
T+VKGT GYLDPEY+KT QLT +SDVYSFG+LLLEI +GRRP+E ++ ER+T RWA R
Sbjct: 314 TQVKGTAGYLDPEYLKTCQLTDRSDVYSFGVLLLEIASGRRPIEARREMRERLTARWAMR 373
Query: 439 KYNEGSVVELMDP-LMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLWAIRADY 497
K EG+ +++DP L A +++L+F+C AP+R +RP+M LWA+R Y
Sbjct: 374 KLAEGAAADVLDPHLPRTPATARAAEMVMELAFRCLAPVRQERPSMGECCRALWAVRKTY 433
>UniRef100_Q8W0H6 P0529E05.2 protein [Oryza sativa]
Length = 333
Score = 325 bits (833), Expect = 2e-87
Identities = 160/300 (53%), Positives = 215/300 (71%), Gaps = 2/300 (0%)
Query: 206 LNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFES-LRTEFSSEV 264
L + ++ AT NFSE +IG G FGTVYK L DG ++AVKRA + ++ L EF SE+
Sbjct: 6 LLVQEICMATSNFSEQNRIGLGNFGTVYKGKLRDGSIIAVKRATKNMYDRHLSEEFRSEI 65
Query: 265 ELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDV 324
+ L+K++H NLVK LG+++ +ER+++ EYV NG+LREHLDGLRG+ L+F+QRL IAID+
Sbjct: 66 QTLSKVEHLNLVKFLGYLEHEDERLILVEYVNNGSLREHLDGLRGEPLEFSQRLNIAIDI 125
Query: 325 AHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGT 384
H ++YLH Y + IIHRD+KSSNILLT+ +RAKVADFGFARL P N + TH+ST VKGT
Sbjct: 126 VHAVSYLHGYTDHPIIHRDIKSSNILLTDQLRAKVADFGFARLAPDNTEATHVSTMVKGT 185
Query: 385 VGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGS 444
GY+DPEYM+T+QLT +SDVYSFG+LL+E+LTGRRP+E + +R+T +WA RK +G
Sbjct: 186 AGYVDPEYMRTNQLTDRSDVYSFGVLLVELLTGRRPIERGRGRHQRLTTQWALRKCRDGD 245
Query: 445 VVELMDPLMEEAVNADVLM-KMLDLSFQCAAPIRTDRPNMKSVGEQLWAIRADYLKSARR 503
V MD M M K++ L+ +C AP R RP M+ E LW+IR D+ +R
Sbjct: 246 AVVAMDARMRRTSAVVAAMEKVMALAAECTAPDRAARPAMRRCAEVLWSIRRDFQHEQQR 305
>UniRef100_O23069 A_IG005I10.8 protein [Arabidopsis thaliana]
Length = 341
Score = 325 bits (833), Expect = 2e-87
Identities = 168/317 (52%), Positives = 216/317 (67%), Gaps = 22/317 (6%)
Query: 202 HTLNLNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLR---T 258
HT ++ AT+NFS + +IGQGGFGTVYK L DG AVKRAK+ + +
Sbjct: 18 HT-RFTFDEIYDATKNFSPSFRIGQGGFGTVYKVKLRDGKTFAVKRAKKSMHDDRQGADA 76
Query: 259 EFSSEVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDG---------LRG 309
EF SE++ LA++ H +LVK GF+ +E+IL+ EYV NGTLR+HLD G
Sbjct: 77 EFMSEIQTLAQVTHLSLVKYYGFVVHNDEKILVVEYVANGTLRDHLDCEYTSYYYLLFEG 136
Query: 310 KILDFNQRLEIAIDVAHGLTYLHLYAEKQI-------IHRDVKSSNILLTESMRAKVADF 362
K LD RL+IA DVAH +TYLH+Y K I IHRD+KSSNILLTE+ RAKVADF
Sbjct: 137 KTLDMATRLDIATDVAHAITYLHMYTRKSIVYIEPPIIHRDIKSSNILLTENYRAKVADF 196
Query: 363 GFARLGP-VNGDQTHISTKVKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPV 421
GFARL P + TH+ST+VKGT GYLDPEY+ T+QLT KSDVYSFG+LL+E+LTGRRP+
Sbjct: 197 GFARLAPDTDSGATHVSTQVKGTAGYLDPEYLTTYQLTEKSDVYSFGVLLVELLTGRRPI 256
Query: 422 ELKKAADERVTLRWAFRKYNEGSVVELMDPLMEE-AVNADVLMKMLDLSFQCAAPIRTDR 480
EL + ER+T+RWA +K+ G + ++DP +E+ + N L K+L+++FQC AP R R
Sbjct: 257 ELSRGQKERITIRWAIKKFTSGDTISVLDPKLEQNSANNLALEKVLEMAFQCLAPHRRSR 316
Query: 481 PNMKSVGEQLWAIRADY 497
P+MK E LW IR DY
Sbjct: 317 PSMKKCSEILWGIRKDY 333
>UniRef100_Q94GE5 Putative receptor kinase [Oryza sativa]
Length = 444
Score = 269 bits (688), Expect = 1e-70
Identities = 145/396 (36%), Positives = 234/396 (58%), Gaps = 31/396 (7%)
Query: 110 QGDSKALPTNKVG---IFAGGALLACCAVLC---PCFYGKKRKATAHAVLAKDPISMDS- 162
Q S ++P++++ + + G+ A L CF +K + +++ IS S
Sbjct: 43 QASSYSVPSSEISRSSVRSSGSFRAAAQSLAGVFSCFVPRKSRNEDELEISRTTISQGSR 102
Query: 163 ATSFDASVSASVSGKIPASPLRVPSSPSRFSMSPKLSRLHTLNLNLSQVVRATRNFSETL 222
+T + S+ + +G S L ++++ +AT NFS+
Sbjct: 103 STGYQVSIDPAGTGYPQEST----------------------ELTVAEIFKATSNFSDKN 140
Query: 223 QIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVELLAKIDHRNLVKLLGFI 282
I QG + ++Y+ L DG +A+K A++ + + E E+E+L KIDH+NLV+ LGF
Sbjct: 141 IIKQGSYSSIYRGKLRDGSEIAIKCARKLNSQYASAELRRELEILQKIDHKNLVRFLGFF 200
Query: 283 DKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAHGLTYLHLYAEKQIIHR 342
++ +E + + EYV NG+LREHLD G L+ QRL IAIDVAH +TYLH + E++IIHR
Sbjct: 201 EREDESLTVVEYVSNGSLREHLDESCGNGLELAQRLNIAIDVAHAITYLHEFKEQRIIHR 260
Query: 343 DVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGYLDPEYMKTHQLTPKS 402
+V+SSN+LLT+++ AK+A G AR+ ++ T+ K GY+DPEY+ T++LT KS
Sbjct: 261 NVRSSNVLLTDTLTAKLAGVGLARMAGGESSESE-DTQGKSAAGYVDPEYLSTYELTDKS 319
Query: 403 DVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGSVVELMDP-LMEEAVNADV 461
DVYSFG+LL+E++TGR P+E ++ D R T +WA +++ G VV MDP + +
Sbjct: 320 DVYSFGVLLVELVTGRPPIERRRDLDPRPTTKWALQRFRGGEVVVAMDPRIRRSPASVAT 379
Query: 462 LMKMLDLSFQCAAPIRTDRPNMKSVGEQLWAIRADY 497
+ K+++L+ QC AP R +RP+M+ E LW++R +Y
Sbjct: 380 VEKVMELAEQCVAPARKERPSMRRCTEALWSVRREY 415
>UniRef100_Q5QMM2 Protein kinase-like [Oryza sativa]
Length = 478
Score = 246 bits (628), Expect = 1e-63
Identities = 135/310 (43%), Positives = 196/310 (62%), Gaps = 14/310 (4%)
Query: 190 SRFSMSPKLSRLHTLNLNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAK 249
SR+S+ R T + ++ AT +F+++ Q+GQGG+G VYK NL DG VA+KRA
Sbjct: 118 SRYSVKVDGVRCFTFD----EMAAATNDFTDSAQVGQGGYGKVYKGNLTDGTAVAIKRAH 173
Query: 250 REHFESLRTEFSSEVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRG 309
+ + EF +E+ELL+++ HRNLV L+G+ D+ +E++L+ E++PNGTLR+HL
Sbjct: 174 EGSLQGSK-EFCTEIELLSRLHHRNLVSLVGYCDEEDEQMLVYEFMPNGTLRDHLSAKSR 232
Query: 310 KILDFNQRLEIAIDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGP 369
+ L+F+QR+ IA+ A G+ YLH A+ I HRDVK+SNILL AKVADFG +RL P
Sbjct: 233 RPLNFSQRIHIALGAAKGILYLHTEADPPIFHRDVKASNILLDSKFVAKVADFGLSRLAP 292
Query: 370 V-NGDQT---HISTKVKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKK 425
V + D T HIST VKGT GYLDPEY TH+LT KSDVYS G++LLE+LTG +P++ K
Sbjct: 293 VPDVDGTMPAHISTVVKGTPGYLDPEYFLTHKLTDKSDVYSLGVVLLELLTGMKPIQHGK 352
Query: 426 AADERVTLRWAFRKYNEGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKS 485
V Y G + ++D + + + + + ++ L+ +C RP+M
Sbjct: 353 NIVREVN-----TAYQSGEIAGVIDERISSSSSPECVARLASLAVKCCKDETDARPSMAD 407
Query: 486 VGEQLWAIRA 495
V +L AIR+
Sbjct: 408 VVRELDAIRS 417
>UniRef100_Q9FN92 Receptor-like protein kinase [Arabidopsis thaliana]
Length = 829
Score = 244 bits (623), Expect = 4e-63
Identities = 161/406 (39%), Positives = 220/406 (53%), Gaps = 37/406 (9%)
Query: 95 GRKLLQKDSSKESTSQGD-----SKALPTNKVGIFAG---GALLACCAVLCPCFYGKKRK 146
G ++++ ++SK S G S + VG+ G G+LLA VL F K++
Sbjct: 373 GLEIMKMNNSKSQLSIGTFLPSGSSSTTKKNVGMIIGLTIGSLLAL-VVLGGFFVLYKKR 431
Query: 147 ATAHAVLAKDPISMDSATSFDASVSASVSGKIPASPLRVPSSPSRFSMSPKLSRLHTLNL 206
+K I + S + +S +++ S R+P
Sbjct: 432 GRDQDGNSKTWIPLSSNGTTSSSNGTTLASIASNSSYRIP-------------------- 471
Query: 207 NLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVEL 266
L V AT +F E IG GGFG VYK L DG VAVKRA + + L EF +E+E+
Sbjct: 472 -LVAVKEATNSFDENRAIGVGGFGKVYKGELHDGTKVAVKRANPKSQQGL-AEFRTEIEM 529
Query: 267 LAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAH 326
L++ HR+LV L+G+ D+ NE IL+ EY+ NGTL+ HL G L + QRLEI I A
Sbjct: 530 LSQFRHRHLVSLIGYCDENNEMILVYEYMENGTLKSHLYGSGLLSLSWKQRLEICIGSAR 589
Query: 327 GLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVG 386
GL YLH K +IHRDVKS+NILL E++ AKVADFG ++ GP DQTH+ST VKG+ G
Sbjct: 590 GLHYLHTGDAKPVIHRDVKSANILLDENLMAKVADFGLSKTGP-EIDQTHVSTAVKGSFG 648
Query: 387 YLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTL-RWAFRKYNEGSV 445
YLDPEY + QLT KSDVYSFG+++ E+L RPV E V L WA + +G +
Sbjct: 649 YLDPEYFRRQQLTEKSDVYSFGVVMFEVLCA-RPVIDPTLTREMVNLAEWAMKWQKKGQL 707
Query: 446 VELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLW 491
++DP + + D L K + +C A DRP+M G+ LW
Sbjct: 708 EHIIDPSLRGKIRPDSLRKFGETGEKCLADYGVDRPSM---GDVLW 750
>UniRef100_Q9SNA7 Receptor-like protein kinase homolog [Arabidopsis thaliana]
Length = 512
Score = 244 bits (622), Expect = 5e-63
Identities = 158/409 (38%), Positives = 222/409 (53%), Gaps = 41/409 (10%)
Query: 95 GRKLLQKDSSKESTSQGD----SKALPTNKVGIFAGGALLACCAVL----CPCFYGKKRK 146
G ++++ ++SK S G S + + +G+ G A+ + AV+ C Y K+++
Sbjct: 56 GLEIMKMNNSKGQLSTGTFVPGSSSSSKSNLGLIVGSAIGSLLAVVFLGSCFVLYKKRKR 115
Query: 147 ATAHAVLAKDPISMDSATSFDASVSASVSGKIPASPLRVPSSPSRFSMSPKLSRLHT--- 203
S T S++ + G S++S L+ + T
Sbjct: 116 GQ----------DGHSKTWMPFSINGTSMG-------------SKYSNGTTLTSITTNAN 152
Query: 204 LNLNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSE 263
+ + V AT NF E+ IG GGFG VYK L DG VAVKR + + L EF +E
Sbjct: 153 YRIPFAAVKDATNNFDESRNIGVGGFGKVYKGELNDGTKVAVKRGNPKSQQGL-AEFRTE 211
Query: 264 VELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAID 323
+E+L++ HR+LV L+G+ D+ NE ILI EY+ NGT++ HL G L + QRLEI I
Sbjct: 212 IEMLSQFRHRHLVSLIGYCDENNEMILIYEYMENGTVKSHLYGSGLPSLTWKQRLEICIG 271
Query: 324 VAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKG 383
A GL YLH K +IHRDVKS+NILL E+ AKVADFG ++ GP DQTH+ST VKG
Sbjct: 272 AARGLHYLHTGDSKPVIHRDVKSANILLDENFMAKVADFGLSKTGP-ELDQTHVSTAVKG 330
Query: 384 TVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTL-RWAFRKYNE 442
+ GYLDPEY + QLT KSDVYSFG++L E+L RPV E V L WA + +
Sbjct: 331 SFGYLDPEYFRRQQLTDKSDVYSFGVVLFEVLCA-RPVIDPTLPREMVNLAEWAMKWQKK 389
Query: 443 GSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLW 491
G + +++D + + D L K + +C A DRP+M G+ LW
Sbjct: 390 GQLDQIIDQSLRGNIRPDSLRKFAETGEKCLADYGVDRPSM---GDVLW 435
>UniRef100_Q9LX66 Receptor protein kinase-like [Arabidopsis thaliana]
Length = 830
Score = 244 bits (622), Expect = 5e-63
Identities = 158/409 (38%), Positives = 222/409 (53%), Gaps = 41/409 (10%)
Query: 95 GRKLLQKDSSKESTSQGD----SKALPTNKVGIFAGGALLACCAVL----CPCFYGKKRK 146
G ++++ ++SK S G S + + +G+ G A+ + AV+ C Y K+++
Sbjct: 374 GLEIMKMNNSKGQLSTGTFVPGSSSSSKSNLGLIVGSAIGSLLAVVFLGSCFVLYKKRKR 433
Query: 147 ATAHAVLAKDPISMDSATSFDASVSASVSGKIPASPLRVPSSPSRFSMSPKLSRLHT--- 203
S T S++ + G S++S L+ + T
Sbjct: 434 GQ----------DGHSKTWMPFSINGTSMG-------------SKYSNGTTLTSITTNAN 470
Query: 204 LNLNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSE 263
+ + V AT NF E+ IG GGFG VYK L DG VAVKR + + L EF +E
Sbjct: 471 YRIPFAAVKDATNNFDESRNIGVGGFGKVYKGELNDGTKVAVKRGNPKSQQGL-AEFRTE 529
Query: 264 VELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAID 323
+E+L++ HR+LV L+G+ D+ NE ILI EY+ NGT++ HL G L + QRLEI I
Sbjct: 530 IEMLSQFRHRHLVSLIGYCDENNEMILIYEYMENGTVKSHLYGSGLPSLTWKQRLEICIG 589
Query: 324 VAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKG 383
A GL YLH K +IHRDVKS+NILL E+ AKVADFG ++ GP DQTH+ST VKG
Sbjct: 590 AARGLHYLHTGDSKPVIHRDVKSANILLDENFMAKVADFGLSKTGP-ELDQTHVSTAVKG 648
Query: 384 TVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTL-RWAFRKYNE 442
+ GYLDPEY + QLT KSDVYSFG++L E+L RPV E V L WA + +
Sbjct: 649 SFGYLDPEYFRRQQLTDKSDVYSFGVVLFEVLCA-RPVIDPTLPREMVNLAEWAMKWQKK 707
Query: 443 GSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLW 491
G + +++D + + D L K + +C A DRP+M G+ LW
Sbjct: 708 GQLDQIIDQSLRGNIRPDSLRKFAETGEKCLADYGVDRPSM---GDVLW 753
>UniRef100_Q942J7 Putative receptor protein kinase [Oryza sativa]
Length = 856
Score = 244 bits (622), Expect = 5e-63
Identities = 128/283 (45%), Positives = 189/283 (66%), Gaps = 6/283 (2%)
Query: 210 QVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVELLAK 269
++ R T NFSET +IG GG+G VYK L +G + A+KRA++ + EF +E+ELL++
Sbjct: 516 ELKRCTNNFSETQEIGSGGYGKVYKGMLANGQMAAIKRAQQGSMQGA-AEFKNEIELLSR 574
Query: 270 IDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAHGLT 329
+ H+NLV L+GF + E++L+ EY+PNGTLRE+L G G LD+ +RL+IA+ A GL
Sbjct: 575 VHHKNLVSLVGFCYEQGEQMLVYEYIPNGTLRENLKGKGGMHLDWKKRLQIAVGSAKGLA 634
Query: 330 YLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGYLD 389
YLH A+ IIHRD+KS+NILL ES+ AKVADFG ++L + + H+ST+VKGT+GYLD
Sbjct: 635 YLHELADPPIIHRDIKSTNILLDESLNAKVADFGLSKL-VSDTKKGHVSTQVKGTLGYLD 693
Query: 390 PEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEG--SVVE 447
PEY T QL+ KSDVYSFG+++LE++T R+P+E K +R A +Y++ +
Sbjct: 694 PEYYMTQQLSEKSDVYSFGVVMLELITSRQPIE--KGTYIVREIRTAIDQYDQEYYGLKS 751
Query: 448 LMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQL 490
L+DP + ++ + + L+ +C DRP M V ++L
Sbjct: 752 LIDPTIRDSAKMVGFRRFVQLAMECVEESAADRPTMNDVVKEL 794
>UniRef100_Q7XTT9 OSJNBa0058K23.13 protein [Oryza sativa]
Length = 844
Score = 243 bits (621), Expect = 7e-63
Identities = 150/390 (38%), Positives = 224/390 (56%), Gaps = 20/390 (5%)
Query: 104 SKESTSQGDSKALPTNKVGIFAGGALLACCAVLCP-CFYGKKRKATA-HAVLAKDPISMD 161
++ S+ ++ + +VGI + + VL C+ +KRKA A P+ +
Sbjct: 414 NQRGISKDRNRKILWEEVGIGSASFVTLTSVVLFAWCYVRRKRKADEKEAPPGWHPLVLH 473
Query: 162 SATSFDASVSASVSGKIPASPLRVPSSPSRFSMSPKLSRLHTLNLNLSQVVRATRNFSET 221
A ++ A +GK SPL SS M + S +S++ AT+NF E
Sbjct: 474 EAMK--STTDARAAGK---SPLTRNSSSIGHRMGRRFS--------ISEIRAATKNFDEA 520
Query: 222 LQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVELLAKIDHRNLVKLLGF 281
L IG GGFG VYK +++G VA+KRA + L+ EF +E+E+L+K+ HR+LV ++G+
Sbjct: 521 LLIGTGGFGKVYKGEVDEGTTVAIKRANPLCGQGLK-EFETEIEMLSKLRHRHLVAMIGY 579
Query: 282 IDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAHGLTYLHLYAEKQIIH 341
++ E IL+ EY+ GTLR HL G L + QR++ I A GL YLH A++ IIH
Sbjct: 580 CEEQKEMILVYEYMAKGTLRSHLYGSDLPPLTWKQRVDACIGAARGLHYLHTGADRGIIH 639
Query: 342 RDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGYLDPEYMKTHQLTPK 401
RDVK++NILL E+ AK+ADFG ++ GP DQTH+ST VKG+ GYLDPEY + QLT K
Sbjct: 640 RDVKTTNILLDENFVAKIADFGLSKTGPTL-DQTHVSTAVKGSFGYLDPEYFRRQQLTQK 698
Query: 402 SDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGSVVELMDPLMEEAVNADV 461
SDVYSFG++L E+ GR ++ D+ WA R + S+ ++DP ++ +++
Sbjct: 699 SDVYSFGVVLFEVACGRPVIDPTLPKDQINLAEWAMRWQRQRSLDAIVDPRLDGDFSSES 758
Query: 462 LMKMLDLSFQCAAPIRTDRPNMKSVGEQLW 491
L K +++ +C A RP+M GE LW
Sbjct: 759 LKKFGEIAEKCLADDGRSRPSM---GEVLW 785
>UniRef100_Q9SC72 L1332.5 protein [Oryza sativa]
Length = 844
Score = 242 bits (617), Expect = 2e-62
Identities = 147/389 (37%), Positives = 220/389 (55%), Gaps = 18/389 (4%)
Query: 104 SKESTSQGDSKALPTNKVGIFAGGALLACCAVLCP-CFYGKKRKATAHAVLAKDPISMDS 162
++ S+ ++ + +VGI + + VL C+ +KRKA + P
Sbjct: 414 NQRGISKDRNRKILWEEVGIGSASFVTLTSVVLFAWCYIRRKRKADEK----EPPPGWHP 469
Query: 163 ATSFDASVSASVSGKIPASPLRVPSSPSRFSMSPKLSRLHTLNLNLSQVVRATRNFSETL 222
+A S + + SPL SS M + S +S++ AT+NF E L
Sbjct: 470 LVLHEAMKSTTDARAAGKSPLTRNSSSIGHRMGRRFS--------ISEIRAATKNFDEAL 521
Query: 223 QIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVELLAKIDHRNLVKLLGFI 282
IG GGFG VYK +++G VA+KRA + L+ EF +E+E+L+K+ HR+LV ++G+
Sbjct: 522 LIGTGGFGKVYKGEVDEGTTVAIKRANPLCGQGLK-EFETEIEMLSKLRHRHLVAMIGYC 580
Query: 283 DKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAHGLTYLHLYAEKQIIHR 342
++ E IL+ EY+ GTLR HL G L + QR++ I A GL YLH A++ IIHR
Sbjct: 581 EEQKEMILVYEYMAKGTLRSHLYGSDLPPLTWKQRVDACIGAARGLHYLHTGADRGIIHR 640
Query: 343 DVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGYLDPEYMKTHQLTPKS 402
DVK++NILL E+ AK+ADFG ++ GP DQTH+ST VKG+ GYLDPEY + QLT KS
Sbjct: 641 DVKTTNILLDENFVAKIADFGLSKTGPTL-DQTHVSTAVKGSFGYLDPEYFRRQQLTQKS 699
Query: 403 DVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGSVVELMDPLMEEAVNADVL 462
DVYSFG++L E+ GR ++ D+ WA R + S+ ++DP ++ +++ L
Sbjct: 700 DVYSFGVVLFEVACGRPVIDPTLPKDQINLAEWAMRWQRQRSLDAIVDPRLDGDFSSESL 759
Query: 463 MKMLDLSFQCAAPIRTDRPNMKSVGEQLW 491
K +++ +C A RP+M GE LW
Sbjct: 760 KKFGEIAEKCLADDGRSRPSM---GEVLW 785
>UniRef100_Q6W0C7 Pto-like serine/threonine kinase [Capsicum chinense]
Length = 359
Score = 242 bits (617), Expect = 2e-62
Identities = 136/282 (48%), Positives = 179/282 (63%), Gaps = 7/282 (2%)
Query: 211 VVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVELLAKI 270
++ AT NF E+L IG GGFG VYK L DG +AVKR + + L EF +E+E+L++
Sbjct: 11 LLEATSNFDESLVIGIGGFGKVYKGVLYDGTKLAVKRGNPKSQQGL-AEFRTEIEMLSQF 69
Query: 271 DHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAHGLTY 330
HR+LV L+G+ D+ NE IL+ EY+ NGTL+ HL G + + QRLEI I A GL Y
Sbjct: 70 RHRHLVSLMGYCDEKNEMILVYEYMENGTLKSHLYGSDLPSMSWKQRLEICIGSARGLHY 129
Query: 331 LHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGYLDP 390
LH K +IHRDVKS+NILL ES AKVADFG ++ GP DQTH+ST VKG+ GYLDP
Sbjct: 130 LHTGYAKAVIHRDVKSANILLDESFMAKVADFGLSKTGP-ELDQTHVSTAVKGSFGYLDP 188
Query: 391 EYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTL-RWAFRKYNEGSVVELM 449
EY + QLT KSDVYSFG++L E+L RPV E V L WA + +G + +++
Sbjct: 189 EYFRRQQLTEKSDVYSFGVVLFEVLCA-RPVIDPSLPREMVNLAEWAMKWQKKGQLEQII 247
Query: 450 DPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLW 491
DP + + D L K + + +C A DRP+M G+ LW
Sbjct: 248 DPTLVGKIRPDSLRKFGETAEKCLADFGVDRPSM---GDVLW 286
>UniRef100_Q96387 Receptor-like protein kinase precursor [Catharanthus roseus]
Length = 803
Score = 241 bits (614), Expect = 5e-62
Identities = 136/285 (47%), Positives = 184/285 (63%), Gaps = 6/285 (2%)
Query: 208 LSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRA-KREHFESLRTEFSSEVEL 266
L+ V AT NFSE IG GGFG VYK +DG VAVKR + +EF +EVEL
Sbjct: 461 LAVVQEATDNFSENRVIGIGGFGKVYKGVFKDGTKVAVKRGISCSSSKQGLSEFRTEVEL 520
Query: 267 LAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAH 326
L++ HR+LV L+G+ D+ NE I+I E++ NGTLR+HL G L++ +R+EI I A
Sbjct: 521 LSQFRHRHLVSLIGYCDEKNEMIIIYEFMENGTLRDHLYGSDKPKLNWRKRVEICIGSAK 580
Query: 327 GLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVG 386
GL YLH K+IIHRDVKS+NILL E++ AKVADFG ++ GP + DQTH+ST VKG+ G
Sbjct: 581 GLHYLHTGTMKRIIHRDVKSANILLDENLMAKVADFGVSKTGPDHFDQTHVSTAVKGSFG 640
Query: 387 YLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGSVV 446
YLDPEY+ +LT KSDVYSFG+++LEILTGR ++ K + + WA + +G
Sbjct: 641 YLDPEYLTMQKLTEKSDVYSFGVVMLEILTGRPVIDPSKPREMVNLVEWAMKCSRKGE-- 698
Query: 447 ELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLW 491
E++D + V + L+K + + +C A DRP M G+ LW
Sbjct: 699 EIVDSDIVNEVRPESLIKFQETAEKCLAERGVDRPTM---GDVLW 740
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.135 0.387
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 782,836,031
Number of Sequences: 2790947
Number of extensions: 31673731
Number of successful extensions: 135241
Number of sequences better than 10.0: 18747
Number of HSP's better than 10.0 without gapping: 9642
Number of HSP's successfully gapped in prelim test: 9106
Number of HSP's that attempted gapping in prelim test: 94966
Number of HSP's gapped (non-prelim): 21217
length of query: 504
length of database: 848,049,833
effective HSP length: 132
effective length of query: 372
effective length of database: 479,644,829
effective search space: 178427876388
effective search space used: 178427876388
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 77 (34.3 bits)
Lotus: description of TM0030.7


