

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0023.5
(266 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
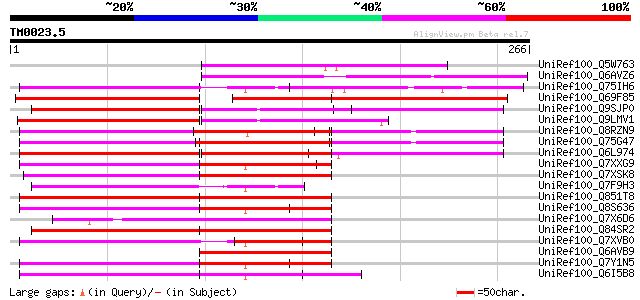
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q5W763 Putative polyprotein [Oryza sativa] 92 1e-17
UniRef100_Q6AVZ6 Putative polyprotein [Oryza sativa] 92 1e-17
UniRef100_Q75IH6 Putative polyprotein [Oryza sativa] 92 2e-17
UniRef100_Q69F85 Gag-pol polyprotein [Phaseolus vulgaris] 91 4e-17
UniRef100_Q9SJP0 Putative retroelement pol polyprotein [Arabidop... 81 3e-14
UniRef100_Q9LMV1 F5M15.26 [Arabidopsis thaliana] 80 7e-14
UniRef100_Q8RZN9 Putative polyprotein [Oryza sativa] 79 1e-13
UniRef100_Q75G47 Putative polyprotein [Oryza sativa] 79 2e-13
UniRef100_Q6L974 GAG-POL [Vitis vinifera] 79 2e-13
UniRef100_Q7XXG9 OSJNBb0016B03.14 protein [Oryza sativa] 78 2e-13
UniRef100_Q7XSK8 OSJNBa0059D20.5 protein [Oryza sativa] 78 3e-13
UniRef100_Q7F9H3 OSJNBa0032N05.3 protein [Oryza sativa] 77 4e-13
UniRef100_Q851T8 Putative polyprotein [Oryza sativa] 77 5e-13
UniRef100_Q8S636 Putative polyprotein [Oryza sativa] 76 1e-12
UniRef100_Q7X6D6 OSJNBa0093P23.3 protein [Oryza sativa] 76 1e-12
UniRef100_Q84SR2 Putative GAG-POL [Oryza sativa] 76 1e-12
UniRef100_Q7XVB0 OSJNBa0081G05.5 protein [Oryza sativa] 76 1e-12
UniRef100_Q6AVB9 Putative polyprotein [Oryza sativa] 76 1e-12
UniRef100_Q7Y1N5 Putative polyprotein [Oryza sativa] 75 1e-12
UniRef100_Q6I5B8 Putative polyprotein [Oryza sativa] 75 1e-12
>UniRef100_Q5W763 Putative polyprotein [Oryza sativa]
Length = 1551
Score = 92.4 bits (228), Expect = 1e-17
Identities = 54/147 (36%), Positives = 80/147 (53%), Gaps = 21/147 (14%)
Query: 99 PPEQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKIR 158
P ++ T+ WPFA+WG D++GLF A + VAVD F+KWIEA+ V TIT K R
Sbjct: 1390 PAQEQQTIPLSWPFAVWGLDMVGLFKKAIGGYTHLFVAVDKFSKWIEAKPVITITADKAR 1449
Query: 159 NF-----------------LWRRIT----STGETPYKLTYGSDAMLPVEVENQSWRVITM 197
+F LW T +TG++P+ L YG++AMLP EVE +S R
Sbjct: 1450 DFFINIVHRFGWVEQLSSVLWSLHTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFHNF 1509
Query: 198 NEEENNDNLIANLIMLPELQREAHLRN 224
+EE ++ + +L L E + A +++
Sbjct: 1510 SEERYGEDRVDDLNRLEEAREAALIQS 1536
>UniRef100_Q6AVZ6 Putative polyprotein [Oryza sativa]
Length = 1844
Score = 92.4 bits (228), Expect = 1e-17
Identities = 61/168 (36%), Positives = 90/168 (53%), Gaps = 13/168 (7%)
Query: 99 PPEQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKIR 158
P ++L T+ WPFA+WG D++G F A + VA+D F+KWIEA+ V TIT K R
Sbjct: 1648 PAQELQTIPLSWPFAVWGLDMVGPFKKAVGGYTHLFVAIDKFSKWIEAKPVVTITADKAR 1707
Query: 159 NFLWRRITSTGETPYKLTYGSDAMLPVEVENQSWRVITMNEEENNDNLIANLIMLPELQR 218
+F + + ++AMLP EV+ +S R EE ++ + +L L E+ R
Sbjct: 1708 DFF-----------INIVHRAEAMLPSEVKFESLRFHNFREERYEEDRVDDLHRLEEV-R 1755
Query: 219 EAHLRNEAGKGRVARKYASK-VDSRKMKARDLVLRQRTMTSRCNKLTP 265
EA L A + R+Y ++ V SR DLVLR+ T +KL+P
Sbjct: 1756 EAALIQSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRDRHKLSP 1803
Score = 35.4 bits (80), Expect = 1.5
Identities = 24/66 (36%), Positives = 35/66 (52%), Gaps = 10/66 (15%)
Query: 6 PTPEVPLILYLAVTSTAVSAMLLQEE------HKKYKMIYFVSHTLQGAEVRYQK----I 55
P + PL+LY++ TS VS +L+ E K + IYFVS L ++ RY + I
Sbjct: 1284 PHLQEPLLLYVSATSQVVSTVLVVEREEEGHIQKVQRPIYFVSEVLADSKTRYPQPRTSI 1343
Query: 56 EKAALA 61
+ ALA
Sbjct: 1344 KSQALA 1349
>UniRef100_Q75IH6 Putative polyprotein [Oryza sativa]
Length = 1048
Score = 91.7 bits (226), Expect = 2e-17
Identities = 66/202 (32%), Positives = 94/202 (45%), Gaps = 41/202 (20%)
Query: 98 APPEQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKI 157
AP L + WPFA WG DI+G F VA+ KF +VAV+YF++WIEAE + TIT+A +
Sbjct: 809 APQYDLQPIAPVWPFARWGLDIIGPFPVARNGYKFTIVAVEYFSRWIEAEPLGTITLAAV 868
Query: 158 RNFLWRR-----------ITSTGE-----------------------TPYKLTYGSDAML 183
+ F+W+ IT G+ +P+ YG +AM
Sbjct: 869 QKFVWKNIVCRFGVPKEFITDNGKQFDSDKFREMCEGLNLQIRPTKFSPFMPLYGDEAMT 928
Query: 184 PVEVENQSWRVITMNEEENNDNLIANLIMLPELQREA--HLRNEAGKGRVARKYASKVDS 241
P E+ S RVI EE + +L +L ++ EA H+ A A Y KV
Sbjct: 929 PTELGANSPRVIFSGGEEGRE---VSLELLEGVRVEALEHMHKYATSTSAA--YNKKVRP 983
Query: 242 RKMKARDLVLRQRTMTSRCNKL 263
R++ LVLR++ KL
Sbjct: 984 RELMPGQLVLRKKANPIEVGKL 1005
Score = 78.2 bits (191), Expect = 2e-13
Identities = 48/140 (34%), Positives = 79/140 (56%), Gaps = 16/140 (11%)
Query: 6 PTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKT 65
P P L+LYLA + AVSA L+QE K +YFVS LQGA+ RY ++EK A A++
Sbjct: 417 PPPGSELLLYLAASPVAVSAALVQEIESGQKPVYFVSEALQGAKTRYIEMEKLAYALVMA 476
Query: 66 ARRLRPYFQSFQIKVKTDILLRQVWQKPDLSGAPPEQLTTMTAPWPFAIWGADI--LGLF 123
+R+L+ YFQ+ ++ V + LL ++ + +++G + W A++ L
Sbjct: 477 SRKLKHYFQAHKVIVPSQYLLGEILRGKEVTGR-------------LSKWAAELSPFDLH 523
Query: 124 SVAKAQMKFIVVAVDYFTKW 143
VA++ +K V+A D+ +W
Sbjct: 524 FVARSAIKSQVLA-DFVAEW 542
>UniRef100_Q69F85 Gag-pol polyprotein [Phaseolus vulgaris]
Length = 1859
Score = 90.5 bits (223), Expect = 4e-17
Identities = 43/94 (45%), Positives = 66/94 (69%)
Query: 4 QKPTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAIL 63
QKP P+++YLAV++ AVS+ L+QE + + +YFVS L AE RYQ +EK A A++
Sbjct: 1190 QKPNAREPIVVYLAVSNEAVSSALVQEIKAEERPVYFVSRVLHDAETRYQMVEKVAFALV 1249
Query: 64 KTARRLRPYFQSFQIKVKTDILLRQVWQKPDLSG 97
TARR+R YFQ+ ++ V+T+ + ++ KPDL+G
Sbjct: 1250 ITARRMRMYFQNHKVIVRTNYPIMKILTKPDLAG 1283
Score = 75.1 bits (183), Expect = 2e-12
Identities = 39/90 (43%), Positives = 59/90 (65%)
Query: 166 TSTGETPYKLTYGSDAMLPVEVENQSWRVITMNEEENNDNLIANLIMLPELQREAHLRNE 225
++T ETPY LTYG++AM+PVE+ S R T++ + N ++L+ L ++ EL+ + +R E
Sbjct: 1718 STTQETPYSLTYGTEAMIPVEIGEPSLRRQTLDLDLNKESLLVGLDLINELRDKCKIREE 1777
Query: 226 AGKGRVARKYASKVDSRKMKARDLVLRQRT 255
A K R AR+Y SKV R + DLV R R+
Sbjct: 1778 ACKIRAARRYNSKVKPRSYQKGDLVWRMRS 1807
Score = 62.8 bits (151), Expect = 9e-09
Identities = 25/51 (49%), Positives = 37/51 (72%)
Query: 115 WGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKIRNFLWRRI 165
+G DI+G F+ K Q KF++V +DYFTKWIEAE + IT +++F+W+ I
Sbjct: 1581 YGMDIIGPFTPGKGQCKFLLVGIDYFTKWIEAEPLTAITARNVQSFVWKNI 1631
>UniRef100_Q9SJP0 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1787
Score = 80.9 bits (198), Expect = 3e-14
Identities = 39/86 (45%), Positives = 62/86 (71%)
Query: 12 LILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKTARRLRP 71
L LY+AV+++AVS +L++E+ + I++VS TL GAE+RY +EK A A++ +AR+LRP
Sbjct: 1059 LYLYIAVSTSAVSGVLVREDRGEQHPIFYVSKTLDGAELRYPTLEKLAFAVVISARKLRP 1118
Query: 72 YFQSFQIKVKTDILLRQVWQKPDLSG 97
YF+S+ ++V T+ LR + P SG
Sbjct: 1119 YFKSYTVEVLTNQPLRTILHSPSQSG 1144
Score = 66.2 bits (160), Expect = 8e-10
Identities = 31/77 (40%), Positives = 45/77 (58%), Gaps = 1/77 (1%)
Query: 99 PPEQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKIR 158
P E L T P+PF W DI+ F ++ Q K+I+V DYFTKW EAE+ I +++
Sbjct: 1490 PSEVLQTAMPPYPFMRWAMDIIRPFPASR-QKKYILVMTDYFTKWFEAESYARIQSKEVQ 1548
Query: 159 NFLWRRITSTGETPYKL 175
NF+W+ I PY++
Sbjct: 1549 NFVWKNIICHHGLPYEI 1565
Score = 46.2 bits (108), Expect = 9e-04
Identities = 33/89 (37%), Positives = 48/89 (53%), Gaps = 2/89 (2%)
Query: 167 STGETPYKLTYGSDAMLPVEVENQS-WRVITMNEEENNDN-LIANLIMLPELQREAHLRN 224
ST TP+ LTYG +A+ P E + R + +N+ ND L+ NL L E + +A LR
Sbjct: 1643 STDRTPFSLTYGMEALAPCEAGLPTVRRSMLINDPTLNDQMLLDNLDTLEEQRDQALLRI 1702
Query: 225 EAGKGRVARKYASKVDSRKMKARDLVLRQ 253
+ + A+ Y KV +R DLVLR+
Sbjct: 1703 QNYQQAAAKFYNKKVKNRLFAEGDLVLRK 1731
>UniRef100_Q9LMV1 F5M15.26 [Arabidopsis thaliana]
Length = 1838
Score = 79.7 bits (195), Expect = 7e-14
Identities = 44/93 (47%), Positives = 60/93 (64%)
Query: 5 KPTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILK 64
KP L LY+AV+ AVS++L++E+ + + I++ S +L AE RY IEKAALA++
Sbjct: 1114 KPEVGETLYLYIAVSDHAVSSVLVREDRGEQRPIFYTSKSLVEAETRYPVIEKAALAVVT 1173
Query: 65 TARRLRPYFQSFQIKVKTDILLRQVWQKPDLSG 97
+AR+LRPYFQS I V TD LR P SG
Sbjct: 1174 SARKLRPYFQSHTIAVLTDQPLRVALHSPSQSG 1206
Score = 71.2 bits (173), Expect = 3e-11
Identities = 37/99 (37%), Positives = 55/99 (55%), Gaps = 4/99 (4%)
Query: 99 PPEQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKIR 158
P E L AP+PF W DI+G ++ Q +FI+V DYFTKW+EAE+ TI ++
Sbjct: 1541 PTELLRAGVAPYPFMRWAMDIVGPMPASR-QKRFILVMTDYFTKWVEAESYATIRANDVQ 1599
Query: 159 NFLWRRITSTGETPYK-LTYGSDAMLPVEVEN--QSWRV 194
NF+W+ I PY+ +T + + EN SW++
Sbjct: 1600 NFVWKFIICRHGLPYEIITDNGSQFISLSFENFCASWKI 1638
Score = 45.8 bits (107), Expect = 0.001
Identities = 30/90 (33%), Positives = 47/90 (51%), Gaps = 2/90 (2%)
Query: 166 TSTGETPYKLTYGSDAMLPVEVENQSWR--VITMNEEENNDNLIANLIMLPELQREAHLR 223
++T +TP+ YG +AM P EV S R ++ N E N+ ++ L L E++ A R
Sbjct: 1693 SATDQTPFAHAYGMEAMAPAEVGYSSLRRSMMVKNPELNDRMMLDRLDDLEEIRNAALCR 1752
Query: 224 NEAGKGRVARKYASKVDSRKMKARDLVLRQ 253
+ + A+ Y KV +R DLVLR+
Sbjct: 1753 IQNYQLAAAKHYNQKVHNRHFDVGDLVLRK 1782
>UniRef100_Q8RZN9 Putative polyprotein [Oryza sativa]
Length = 2001
Score = 79.0 bits (193), Expect = 1e-13
Identities = 32/71 (45%), Positives = 50/71 (70%)
Query: 95 LSGAPPEQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITV 154
++ APP++L + WPF WG DI+G AK ++F++VA++YF++WIEAEAV IT
Sbjct: 1705 MTKAPPKELQPIPPVWPFYRWGIDIVGPLPRAKGDLRFVIVAIEYFSRWIEAEAVARITS 1764
Query: 155 AKIRNFLWRRI 165
A ++ F+W+ I
Sbjct: 1765 AAVQKFVWKNI 1775
Score = 73.6 bits (179), Expect = 5e-12
Identities = 47/153 (30%), Positives = 77/153 (49%), Gaps = 16/153 (10%)
Query: 6 PTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKT 65
P PL++Y+A T VSA+L+QEE + +YFVS LQG + RY ++EK AI+
Sbjct: 1299 PPKGQPLLMYVAATPATVSAVLVQEEENRQVPVYFVSEALQGPKTRYSEVEKLIYAIVMA 1358
Query: 66 ARRLRPYFQSFQIKVKTDILLRQVWQKPDLSGAPPEQLTTMTAPWPFAIWGADIL--GLF 123
+R+LR YF S I + + + +V +++G A W ++L L
Sbjct: 1359 SRKLRHYFLSHDITIPSAYPIGEVLTNKEVAGR-------------IAKWAMELLPFDLK 1405
Query: 124 SVAKAQMKFIVVAVDYFTKWIEAEAVETITVAK 156
+++ +K V+A D+ +W E + V K
Sbjct: 1406 YISRTAIKSQVLA-DFVAEWTPNEVEQQEEVKK 1437
Score = 47.8 bits (112), Expect = 3e-04
Identities = 31/89 (34%), Positives = 48/89 (53%), Gaps = 2/89 (2%)
Query: 165 ITSTGETPYKLTYGSDAMLPVEVENQSWRVITMNEEENNDNLIANLIMLPELQREAHLRN 224
+ STG TP++L YG +AM P EV S R+I ++E + L ML E++ EA +
Sbjct: 1862 VRSTGMTPFRLVYGDEAMTPSEVGAHSPRMIFDQKDEEGREI--TLEMLDEIRVEALEKM 1919
Query: 225 EAGKGRVARKYASKVDSRKMKARDLVLRQ 253
+ Y KV +R ++ DLVL++
Sbjct: 1920 ASYTEGTKSYYNQKVKTRPIEEGDLVLKK 1948
>UniRef100_Q75G47 Putative polyprotein [Oryza sativa]
Length = 1459
Score = 78.6 bits (192), Expect = 2e-13
Identities = 33/70 (47%), Positives = 49/70 (69%)
Query: 96 SGAPPEQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVA 155
S APP++L + WPF WG DI+G AK ++F++VA++YF++WIEAEAV IT A
Sbjct: 1164 SKAPPKELQPIPPVWPFYRWGIDIVGPLPRAKGDLRFVIVAIEYFSRWIEAEAVARITSA 1223
Query: 156 KIRNFLWRRI 165
++ F+W+ I
Sbjct: 1224 AMQKFVWKNI 1233
Score = 71.6 bits (174), Expect = 2e-11
Identities = 36/92 (39%), Positives = 55/92 (59%)
Query: 6 PTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKT 65
P PL+LY+A T VSA+L+QE+ K +YFVS LQG + RY ++EK AI+
Sbjct: 756 PQKGEPLLLYVAATPVTVSAVLVQEQGNKQMPVYFVSEALQGPKTRYIEVEKMIYAIVMA 815
Query: 66 ARRLRPYFQSFQIKVKTDILLRQVWQKPDLSG 97
+R+LR YF S I + + + +V +++G
Sbjct: 816 SRKLRHYFLSHDITIPSTYPIGEVLSNKEIAG 847
Score = 52.8 bits (125), Expect = 9e-06
Identities = 33/89 (37%), Positives = 50/89 (56%), Gaps = 2/89 (2%)
Query: 165 ITSTGETPYKLTYGSDAMLPVEVENQSWRVITMNEEENNDNLIANLIMLPELQREAHLRN 224
+ +TG TP++L YG + M P EV S RVI E+E + +L ML E++ EA +
Sbjct: 1320 VWATGMTPFRLVYGDEPMTPSEVGVNSPRVIFYQEDEQGRKV--SLEMLDEIRVEALQKM 1377
Query: 225 EAGKGRVARKYASKVDSRKMKARDLVLRQ 253
EA ++Y KV R ++ DLVL++
Sbjct: 1378 EAYTEGTRKQYNKKVRPRNIEEGDLVLKK 1406
>UniRef100_Q6L974 GAG-POL [Vitis vinifera]
Length = 1027
Score = 78.6 bits (192), Expect = 2e-13
Identities = 42/93 (45%), Positives = 59/93 (63%), Gaps = 1/93 (1%)
Query: 6 PTPEVPLILYLAVTSTAVSAMLLQ-EEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILK 64
P P+ L +YLAV+ A+SA+L + K+ K IY+VS L E RY K+E +LA+
Sbjct: 312 PIPKEKLYMYLAVSEWAISAVLFRCPSPKEQKPIYYVSRALADVETRYSKMELISLALRS 371
Query: 65 TARRLRPYFQSFQIKVKTDILLRQVWQKPDLSG 97
A++LRPYFQ+ + V TD LR + KPDL+G
Sbjct: 372 AAQKLRPYFQAHPVIVLTDQPLRNILHKPDLTG 404
Score = 70.5 bits (171), Expect = 4e-11
Identities = 30/67 (44%), Positives = 42/67 (61%)
Query: 99 PPEQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKIR 158
P L +++ PWPFA WG DI+ A AQ KF++VA DYF+KW+EAEA + +
Sbjct: 731 PSTTLKSISGPWPFAQWGMDIVRPLPTAPAQKKFLLVATDYFSKWVEAEAYASTKDKDVT 790
Query: 159 NFLWRRI 165
F+W+ I
Sbjct: 791 KFVWKNI 797
Score = 48.9 bits (115), Expect = 1e-04
Identities = 32/104 (30%), Positives = 50/104 (47%), Gaps = 4/104 (3%)
Query: 154 VAKIRNFLWRRITS----TGETPYKLTYGSDAMLPVEVENQSWRVITMNEEENNDNLIAN 209
V ++ LW T+ TG TP+ L YG DA++P+E+ + + + N L N
Sbjct: 868 VEELPGVLWAYRTTPGRPTGNTPFALAYGMDAVIPIEIGLPTIWTNAAKQSDANMQLGRN 927
Query: 210 LIMLPELQREAHLRNEAGKGRVARKYASKVDSRKMKARDLVLRQ 253
L E++ A +R + R + Y KV R +K LVLR+
Sbjct: 928 LDWTDEVRESASIRMADYQQRASAHYNRKVRPRSLKNGTLVLRK 971
>UniRef100_Q7XXG9 OSJNBb0016B03.14 protein [Oryza sativa]
Length = 1323
Score = 78.2 bits (191), Expect = 2e-13
Identities = 51/154 (33%), Positives = 83/154 (53%), Gaps = 16/154 (10%)
Query: 6 PTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKT 65
PTP L+LYLA + AVSA L+QE K +YF+S LQGA+ RY ++EK A A++
Sbjct: 662 PTPGSELLLYLAASPVAVSAALVQETKFGQKPVYFISEALQGAKTRYIEMEKLAYALVMA 721
Query: 66 ARRLRPYFQSFQIKVKTDILLRQVWQKPDLSGAPPEQLTTMTAPWPFAIWGADI--LGLF 123
+R+L+ YFQ+ ++ V L ++ + +++G + W A++ L
Sbjct: 722 SRKLKHYFQAHKVIVPPQYPLGEILRGKEVTGR-------------LSKWAAELSPFDLH 768
Query: 124 SVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKI 157
VA+ +K V+A D+ +W A ET V ++
Sbjct: 769 FVARTAIKSQVLA-DFVAEWTLVFAPETELVEQL 801
Score = 71.2 bits (173), Expect = 3e-11
Identities = 32/68 (47%), Positives = 44/68 (64%)
Query: 98 APPEQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKI 157
AP L + W FA WG DI+G F VA+ KF +VAV+YF++WIEAE + IT A +
Sbjct: 1058 APQYDLQPIAPVWLFASWGLDIIGPFPVARNGYKFTIVAVEYFSRWIEAEPLGAITSATV 1117
Query: 158 RNFLWRRI 165
+ F+W+ I
Sbjct: 1118 QKFVWKNI 1125
Score = 35.8 bits (81), Expect = 1.2
Identities = 35/109 (32%), Positives = 49/109 (44%), Gaps = 11/109 (10%)
Query: 161 LWR-RITSTGET---PYKLTYGSDAMLPVEVENQSWRVITMNEEENNDNLIANLIMLPEL 216
LW R T T T P+ L YG +AM P E+ S RV+ EE + +L +L +
Sbjct: 1204 LWALRTTPTRPTKFSPFMLFYGDEAMTPAELGANSPRVMFSRREEGRE---VSLELLEGV 1260
Query: 217 QREA--HLRNEAGKGRVARKYASKVDSRKMKARDLVLRQRTMTSRCNKL 263
+ EA H+R A + Y KV ++ LVLR + KL
Sbjct: 1261 RVEALEHMRKYATS--TSATYNKKVRPTELMPGHLVLRNKANPIAVGKL 1307
>UniRef100_Q7XSK8 OSJNBa0059D20.5 protein [Oryza sativa]
Length = 832
Score = 77.8 bits (190), Expect = 3e-13
Identities = 34/68 (50%), Positives = 47/68 (69%)
Query: 98 APPEQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKI 157
AP +L + WPFAIWG DI+G F VA+ KF +VAV+YF++WIEAE + IT A +
Sbjct: 631 APQYELQPIAPIWPFAIWGLDIIGPFPVARNGYKFAIVAVEYFSRWIEAEPLGAITSAAV 690
Query: 158 RNFLWRRI 165
+ F+W+ I
Sbjct: 691 QKFVWKNI 698
Score = 62.8 bits (151), Expect = 9e-09
Identities = 33/90 (36%), Positives = 54/90 (59%)
Query: 8 PEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKTAR 67
P L+LY A AVSA L+QE K +YFVS LQGA+ RY ++ K A A++ +R
Sbjct: 281 PGSELLLYPAALPVAVSAALVQETESGQKPVYFVSEALQGAKTRYIEMGKLAYALVMASR 340
Query: 68 RLRPYFQSFQIKVKTDILLRQVWQKPDLSG 97
+L+ Y Q+ ++ V + L ++ + +++G
Sbjct: 341 KLKHYLQAHKVIVSSQYPLGKILRGKEVTG 370
>UniRef100_Q7F9H3 OSJNBa0032N05.3 protein [Oryza sativa]
Length = 707
Score = 77.4 bits (189), Expect = 4e-13
Identities = 50/142 (35%), Positives = 80/142 (56%), Gaps = 16/142 (11%)
Query: 12 LILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKTARRLRP 71
L+LYLA + AVSA L+QE K +YF+S LQGA++RY ++EK A++ +R+L+
Sbjct: 373 LLLYLAASPVAVSAALVQETESGQKPVYFISEALQGAKLRYIEMEKFTYALVMASRKLKH 432
Query: 72 YFQSFQIKVKTDILLRQVWQKPDLSGAPPEQLTTMTAPWPFAIWGADI--LGLFSVAKAQ 129
YFQ+ ++ V + L +V + +++G W + W A++ L VA+
Sbjct: 433 YFQAHKVTVPSQYPLGEVLRGKEITG------------W-LSKWVAELSPFDLHFVARTA 479
Query: 130 MKFIVVAVDYFTKWIEAEAVET 151
+K V+A D+ KW A A ET
Sbjct: 480 IKSQVLA-DFVAKWTPAFASET 500
>UniRef100_Q851T8 Putative polyprotein [Oryza sativa]
Length = 658
Score = 77.0 bits (188), Expect = 5e-13
Identities = 33/68 (48%), Positives = 47/68 (68%)
Query: 98 APPEQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKI 157
AP L + WPFA WG DI+G FSVA+ KF++VA++YF++WIEAE + IT A +
Sbjct: 366 APHYNLQPIAPIWPFARWGLDIIGPFSVARNGYKFVIVAIEYFSRWIEAEPLGAITSAAV 425
Query: 158 RNFLWRRI 165
+ F+W+ I
Sbjct: 426 QKFVWKNI 433
Score = 75.9 bits (185), Expect = 1e-12
Identities = 39/92 (42%), Positives = 60/92 (64%)
Query: 6 PTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKT 65
PTP L+LYLA + AVSA L+QE K +YFVS LQGA+ RY ++EK A A++
Sbjct: 70 PTPGSELLLYLAASPVAVSAALVQETESGQKPVYFVSEALQGAKTRYIEMEKLAYALVMA 129
Query: 66 ARRLRPYFQSFQIKVKTDILLRQVWQKPDLSG 97
+R+L+ YFQ+ ++ V + L ++ + ++SG
Sbjct: 130 SRKLKHYFQAHKVIVPSLYPLGEILRGKEISG 161
Score = 34.3 bits (77), Expect = 3.4
Identities = 31/112 (27%), Positives = 50/112 (43%), Gaps = 7/112 (6%)
Query: 156 KIRNFLWR-RITSTGET---PYKLTYGSDAMLPVEVENQSWRVITMNEEENNDNLIANLI 211
++ + LW R T T T P+ L YG +AM P E+ S RV+ EE ++ +L
Sbjct: 507 ELLSVLWALRTTPTRPTKFSPFMLLYGDEAMTPAELGANSPRVMFSGGEEGHE---VSLE 563
Query: 212 MLPELQREAHLRNEAGKGRVARKYASKVDSRKMKARDLVLRQRTMTSRCNKL 263
+L ++ EA + Y KV ++ L+LR++ KL
Sbjct: 564 LLEGVRVEALEHMHKYATSTSATYNKKVRPTELMPGHLILRKKANLVAVGKL 615
>UniRef100_Q8S636 Putative polyprotein [Oryza sativa]
Length = 1722
Score = 75.9 bits (185), Expect = 1e-12
Identities = 33/68 (48%), Positives = 46/68 (67%)
Query: 98 APPEQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKI 157
AP L + WPFA WG DI+G F VA+ KF++VAV+YF++WIEAE + IT A +
Sbjct: 1430 APQYDLQPIAPIWPFARWGLDIIGPFPVARNGYKFVIVAVEYFSRWIEAEPLGAITSAAV 1489
Query: 158 RNFLWRRI 165
+ F+W+ I
Sbjct: 1490 QKFVWKNI 1497
Score = 74.3 bits (181), Expect = 3e-12
Identities = 48/140 (34%), Positives = 76/140 (54%), Gaps = 16/140 (11%)
Query: 6 PTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKT 65
P P L+LYLA + AVSA L+QE K +YFVS LQGA+ RY ++EK A A++
Sbjct: 1059 PPPGSELLLYLAASPVAVSAALVQETKFGQKPVYFVSEALQGAKTRYIEMEKLAYALVMA 1118
Query: 66 ARRLRPYFQSFQIKVKTDILLRQVWQKPDLSGAPPEQLTTMTAPWPFAIWGADI--LGLF 123
+R+L+ YFQ+ ++ V + L ++ ++SG + W A++ L
Sbjct: 1119 SRKLKHYFQAHKVIVPSQYPLGEILLGKEVSGR-------------LSKWAAELSPFDLH 1165
Query: 124 SVAKAQMKFIVVAVDYFTKW 143
VA+ +K V+A D+ +W
Sbjct: 1166 FVARTAIKSQVLA-DFVAEW 1184
Score = 33.9 bits (76), Expect = 4.4
Identities = 26/93 (27%), Positives = 41/93 (43%), Gaps = 3/93 (3%)
Query: 171 TPYKLTYGSDAMLPVEVENQSWRVITMNEEENNDNLIANLIMLPELQREAHLRNEAGKGR 230
+P+ L YG +AM P E+ S RV+ EE + +L +L ++ EA
Sbjct: 1590 SPFMLLYGDEAMTPAELGTNSPRVMFSGGEEGRE---VSLELLEGVRVEALEHMHKYATS 1646
Query: 231 VARKYASKVDSRKMKARDLVLRQRTMTSRCNKL 263
+ Y KV ++ LVLR++ KL
Sbjct: 1647 TSATYNKKVRPTELMPGHLVLRKKANPIAVGKL 1679
>UniRef100_Q7X6D6 OSJNBa0093P23.3 protein [Oryza sativa]
Length = 598
Score = 75.9 bits (185), Expect = 1e-12
Identities = 53/147 (36%), Positives = 72/147 (48%), Gaps = 8/147 (5%)
Query: 23 VSAMLLQEEHKKYKMIY---FVSHTLQGAEVRYQKIEKAALAILKTARRLRPYFQSFQIK 79
V A LQ KYKM+ QG +Q AA + R Y+ +
Sbjct: 231 VEAKRLQLSATKYKMVSGEEMAKEIHQGLCGAHQ----AARTVASKVFRQGVYWPTVLKV 286
Query: 80 VKTDILLRQVWQKPDLSGAPPEQLTTMTAP-WPFAIWGADILGLFSVAKAQMKFIVVAVD 138
I + Q+ D S A P+ AP WPFA WG DI+G F VA+ KF +VAV+
Sbjct: 287 CVEQIKKCESCQRYDRSQAAPQYDLQPIAPIWPFARWGLDIIGPFPVARNGYKFTIVAVE 346
Query: 139 YFTKWIEAEAVETITVAKIRNFLWRRI 165
YF++WIEAE + IT A ++ F+W+ I
Sbjct: 347 YFSRWIEAEPLGAITSAVVQKFVWKNI 373
Score = 33.5 bits (75), Expect = 5.8
Identities = 32/112 (28%), Positives = 48/112 (42%), Gaps = 7/112 (6%)
Query: 156 KIRNFLWR-RITSTGET---PYKLTYGSDAMLPVEVENQSWRVITMNEEENNDNLIANLI 211
++ + LW R T T T P+ L YG AM P E+ S RV+ EE + +L
Sbjct: 447 ELLSVLWALRTTPTRPTKFSPFMLLYGDKAMTPAELGANSPRVMFSGGEEGRE---VSLE 503
Query: 212 MLPELQREAHLRNEAGKGRVARKYASKVDSRKMKARDLVLRQRTMTSRCNKL 263
+L ++ EA + Y KV ++ LVLR++ KL
Sbjct: 504 LLEGVRVEALKHMHKYATSTSATYNKKVRPTELMPGHLVLRKKANPVAVGKL 555
>UniRef100_Q84SR2 Putative GAG-POL [Oryza sativa]
Length = 562
Score = 75.9 bits (185), Expect = 1e-12
Identities = 33/68 (48%), Positives = 46/68 (67%)
Query: 98 APPEQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKI 157
AP L + WPFA WG DI+G F VA+ KF++VAV+YF++WIEAE + IT A +
Sbjct: 405 APQYNLQPIAPIWPFARWGLDIIGPFPVARNGYKFVIVAVEYFSRWIEAEPLGAITSAAV 464
Query: 158 RNFLWRRI 165
+ F+W+ I
Sbjct: 465 QKFVWKNI 472
Score = 69.7 bits (169), Expect = 7e-11
Identities = 35/86 (40%), Positives = 57/86 (65%)
Query: 12 LILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKTARRLRP 71
L+LYLA + AVSA L+QE K +YFVS LQGA+ RY ++EK A A++ +R+L+
Sbjct: 171 LLLYLAASPVAVSAALVQETEFGQKPVYFVSEALQGAKTRYIEMEKLAYALVMASRKLKH 230
Query: 72 YFQSFQIKVKTDILLRQVWQKPDLSG 97
YFQ+ ++ + T L ++ + +++G
Sbjct: 231 YFQAHKVIMPTQYPLGEILRGKEVTG 256
>UniRef100_Q7XVB0 OSJNBa0081G05.5 protein [Oryza sativa]
Length = 926
Score = 75.9 bits (185), Expect = 1e-12
Identities = 49/147 (33%), Positives = 79/147 (53%), Gaps = 16/147 (10%)
Query: 6 PTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKT 65
P P L+LYLA + AVSA L+QE K +YFVS LQGA+ RY ++EK A A++
Sbjct: 294 PAPGSELLLYLAASPVAVSAALVQETEFGQKPVYFVSEALQGAKTRYIEMEKLAYALVMA 353
Query: 66 ARRLRPYFQSFQIKVKTDILLRQVWQKPDLSGAPPEQLTTMTAPWPFAIWGADI--LGLF 123
+R+L+ YFQ+ ++ V + L ++ + +++G + W A++ L
Sbjct: 354 SRKLKHYFQAHKVIVPSQYPLGEILRGKEVTGR-------------LSKWAAELSTFDLH 400
Query: 124 SVAKAQMKFIVVAVDYFTKWIEAEAVE 150
VA+ +K V+A D+ +W A E
Sbjct: 401 FVARTAIKSQVLA-DFVAEWTPVFAPE 426
Score = 57.8 bits (138), Expect = 3e-07
Identities = 25/50 (50%), Positives = 36/50 (72%)
Query: 116 GADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKIRNFLWRRI 165
G DI+G F VA+ KF +VAV+YF++WIEAE + IT A ++ F+W+ I
Sbjct: 641 GLDIIGPFPVARNGYKFAIVAVEYFSRWIEAEPLGAITSAAVQKFVWKNI 690
Score = 35.0 bits (79), Expect = 2.0
Identities = 35/111 (31%), Positives = 51/111 (45%), Gaps = 11/111 (9%)
Query: 159 NFLWR-RITSTGET---PYKLTYGSDAMLPVEVENQSWRVITMNEEENNDNLIANLIMLP 214
+ LW R T T T P+ L YG +AM P E+ S RV+ EE + +L +L
Sbjct: 767 SILWALRTTPTRPTKFSPFILLYGDEAMTPAELWANSPRVMFSGGEEGRE---VSLELLE 823
Query: 215 ELQREA--HLRNEAGKGRVARKYASKVDSRKMKARDLVLRQRTMTSRCNKL 263
++ EA H+R A + Y KV ++ LVLR++ KL
Sbjct: 824 GVRVEALEHMRKYATS--TSATYNKKVRPTELMPGHLVLRKKANPIAVGKL 872
>UniRef100_Q6AVB9 Putative polyprotein [Oryza sativa]
Length = 986
Score = 75.9 bits (185), Expect = 1e-12
Identities = 33/68 (48%), Positives = 46/68 (67%)
Query: 98 APPEQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKI 157
AP L + WPFA WG DI+G F VA+ KF++VAV+YF++WIEAE + IT A +
Sbjct: 694 APQYDLQPIAPIWPFARWGLDIIGPFPVARNGYKFVIVAVEYFSRWIEAEPLGAITSAAV 753
Query: 158 RNFLWRRI 165
+ F+W+ I
Sbjct: 754 QKFVWKNI 761
Score = 45.4 bits (106), Expect = 0.001
Identities = 23/44 (52%), Positives = 28/44 (63%)
Query: 6 PTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAE 49
P P L+LYLA + AVSA L+QE K +YFVS LQGA+
Sbjct: 303 PPPGSELLLYLAASPVAVSAALVQETESGQKPVYFVSEALQGAK 346
Score = 37.0 bits (84), Expect = 0.52
Identities = 29/99 (29%), Positives = 44/99 (44%), Gaps = 3/99 (3%)
Query: 165 ITSTGETPYKLTYGSDAMLPVEVENQSWRVITMNEEENNDNLIANLIMLPELQREAHLRN 224
I T +P+ L YG DAM+P E+ S RV+ EE + +L +L ++ EA
Sbjct: 848 IRPTKFSPFMLLYGDDAMIPAELGANSPRVMFSGGEEGRE---VSLELLEAVRVEALEHM 904
Query: 225 EAGKGRVARKYASKVDSRKMKARDLVLRQRTMTSRCNKL 263
+ Y KV ++ LVLR++ KL
Sbjct: 905 HKYATSTSATYNKKVRPTELMPGHLVLRKKANPLAVGKL 943
>UniRef100_Q7Y1N5 Putative polyprotein [Oryza sativa]
Length = 1862
Score = 75.5 bits (184), Expect = 1e-12
Identities = 46/140 (32%), Positives = 78/140 (54%), Gaps = 16/140 (11%)
Query: 6 PTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKT 65
P P L+LYLA + A+SA L+QE K +YFVS LQGA+ RY ++EK A A++
Sbjct: 1153 PPPGSELLLYLAASPVAISAALVQETESGQKPVYFVSEALQGAKTRYIEMEKLAYALVMA 1212
Query: 66 ARRLRPYFQSFQIKVKTDILLRQVWQKPDLSGAPPEQLTTMTAPWPFAIWGADI--LGLF 123
+R+L+ YFQ+ ++ V + L ++ + +++G + W A++ L+
Sbjct: 1213 SRKLKHYFQANKVIVPSQYPLGEILRGKEVTGR-------------LSKWAAELSPFDLY 1259
Query: 124 SVAKAQMKFIVVAVDYFTKW 143
VA + +K V+A D+ +W
Sbjct: 1260 FVAHSAIKSQVLA-DFVAEW 1278
Score = 74.3 bits (181), Expect = 3e-12
Identities = 33/68 (48%), Positives = 45/68 (65%)
Query: 98 APPEQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKI 157
AP L + WPFA WG DI+G F VA+ KF +VAV+YF++WIEAE + IT A +
Sbjct: 1571 APQYDLQPIAPIWPFARWGLDIIGPFPVARNGYKFAIVAVEYFSRWIEAEPLGAITSAAV 1630
Query: 158 RNFLWRRI 165
+ F+W+ I
Sbjct: 1631 QKFVWKNI 1638
>UniRef100_Q6I5B8 Putative polyprotein [Oryza sativa]
Length = 1136
Score = 75.5 bits (184), Expect = 1e-12
Identities = 35/83 (42%), Positives = 49/83 (58%)
Query: 98 APPEQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKI 157
AP L + WPFA WG DI+G F VA+ KF +VAV+YF++WIEAE + IT A +
Sbjct: 844 APQYDLQPIAPIWPFARWGLDIIGPFPVARNGYKFTIVAVEYFSRWIEAEPLGAITSAAV 903
Query: 158 RNFLWRRITSTGETPYKLTYGSD 180
+ F+W+ I P + +D
Sbjct: 904 QKFVWKNIVCRFGVPKEFITNND 926
Score = 72.4 bits (176), Expect = 1e-11
Identities = 47/147 (31%), Positives = 80/147 (53%), Gaps = 16/147 (10%)
Query: 6 PTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKT 65
P P L+LYLA + AVSA L+QE K +YFVS LQGA+ RY ++EK A A++
Sbjct: 426 PPPGSELLLYLAASPVAVSAALVQETEFGQKPVYFVSEALQGAKTRYIEMEKLAYALVMA 485
Query: 66 ARRLRPYFQSFQIKVKTDILLRQVWQKPDLSGAPPEQLTTMTAPWPFAIWGADI--LGLF 123
+ +L+ YFQ+ ++ V + L ++ + +++G + W A++ L
Sbjct: 486 SCQLKHYFQAHKVIVPSQYPLGEILRGKEVTGR-------------LSKWAAELSPFDLH 532
Query: 124 SVAKAQMKFIVVAVDYFTKWIEAEAVE 150
VA++ +K V+A D+ +W A++
Sbjct: 533 FVARSAIKSQVLA-DFVAEWTPVLALD 558
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.132 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 412,705,541
Number of Sequences: 2790947
Number of extensions: 15598954
Number of successful extensions: 40697
Number of sequences better than 10.0: 830
Number of HSP's better than 10.0 without gapping: 697
Number of HSP's successfully gapped in prelim test: 134
Number of HSP's that attempted gapping in prelim test: 38755
Number of HSP's gapped (non-prelim): 1829
length of query: 266
length of database: 848,049,833
effective HSP length: 125
effective length of query: 141
effective length of database: 499,181,458
effective search space: 70384585578
effective search space used: 70384585578
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0023.5


