

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0022.15
(313 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
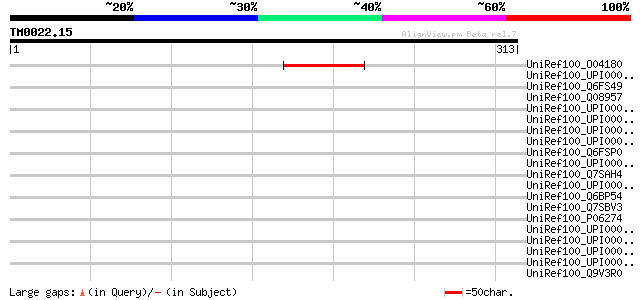
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O04180 Pge1 protein [Lotus japonicus] 60 1e-07
UniRef100_UPI000023D13A UPI000023D13A UniRef100 entry 43 0.012
UniRef100_Q6FS49 Candida glabrata strain CBS138 chromosome H com... 43 0.012
UniRef100_Q08957 S.cerevisiae chromosome XVI reading frame ORF Y... 40 0.062
UniRef100_UPI0000306A5A UPI0000306A5A UniRef100 entry 40 0.081
UniRef100_UPI000042F4A0 UPI000042F4A0 UniRef100 entry 40 0.11
UniRef100_UPI000042D64F UPI000042D64F UniRef100 entry 40 0.11
UniRef100_UPI0000235728 UPI0000235728 UniRef100 entry 39 0.18
UniRef100_Q6FSP0 Similarities with sp|Q08957 Saccharomyces cerev... 39 0.18
UniRef100_UPI000023ED11 UPI000023ED11 UniRef100 entry 38 0.31
UniRef100_Q7SAH4 Predicted protein [Neurospora crassa] 38 0.31
UniRef100_UPI000021AD4C UPI000021AD4C UniRef100 entry 37 0.52
UniRef100_Q6BP54 Similar to CA1250|IPF9626 Candida albicans IPF9... 37 0.52
UniRef100_Q7SBV3 Hypothetical protein [Neurospora crassa] 37 0.89
UniRef100_P06274 DNA-directed RNA polymerase beta'' chain [March... 35 2.0
UniRef100_UPI00002350D1 UPI00002350D1 UniRef100 entry 35 2.6
UniRef100_UPI00002C71B7 UPI00002C71B7 UniRef100 entry 35 3.4
UniRef100_UPI00003C212D UPI00003C212D UniRef100 entry 34 4.4
UniRef100_UPI00003607EB UPI00003607EB UniRef100 entry 33 7.6
UniRef100_Q9V3R0 CG7631-PA [Drosophila melanogaster] 33 7.6
>UniRef100_O04180 Pge1 protein [Lotus japonicus]
Length = 210
Score = 59.7 bits (143), Expect = 1e-07
Identities = 23/50 (46%), Positives = 37/50 (74%)
Query: 170 KVFASRDEILEWARNLGKQHGFIIVITRSDNGGLKRKTFMILGCERCDKY 219
+ FAS ++++WAR +GK++G+++++ RSD G KRK + LGCER KY
Sbjct: 155 EAFASHTDLIDWARCVGKENGYVVIVIRSDYGSAKRKPLITLGCERGGKY 204
>UniRef100_UPI000023D13A UPI000023D13A UniRef100 entry
Length = 207
Score = 42.7 bits (99), Expect = 0.012
Identities = 27/96 (28%), Positives = 48/96 (49%), Gaps = 5/96 (5%)
Query: 171 VFASRDEILEWARNLGKQHGFIIVITRSDNGGLKRKTFMILGCERCDKYIPYKEVLKHQS 230
+F + D+++ + + K G+ IV R+ N + T L C+R + Y K ++
Sbjct: 16 IFRTFDDLMASVQRVAKDQGYGIVKLRASNYRDGKPTRYDLVCDRGG--VKYNSTAKKRN 73
Query: 231 TGTKKCYCPFRLRA-RGTKVG--WKVSVMNGYHIHE 263
T+K CPFR +A ++G W+ ++ G H HE
Sbjct: 74 PSTRKIDCPFRAKAVCEVQLGNQWRFAIQEGRHNHE 109
>UniRef100_Q6FS49 Candida glabrata strain CBS138 chromosome H complete sequence
[Candida glabrata]
Length = 535
Score = 42.7 bits (99), Expect = 0.012
Identities = 40/146 (27%), Positives = 60/146 (40%), Gaps = 22/146 (15%)
Query: 160 VDYTQSFTTDKV--FASRDEILEWARNLGKQHGFIIVITRSDN-----------GGLKRK 206
+D Q D V F ++EI W + + G IVI RSD+ G K +
Sbjct: 1 MDSNQLIHLDSVPNFKDKNEIKPWLQKIFYPQGIEIVIERSDSLKVVFKCKAAKRGRKSQ 60
Query: 207 TFMILGCERCDKYIPYKEVLKHQSTGTKKC-----YCPFRLRARGT--KVGWKVSVMNGY 259
+ + + + +K +S G K+C +CPFR+RA + + W V V+N
Sbjct: 61 EELTEEERLRQEALAREHEMKRKSMGNKRCVSRYNHCPFRVRATYSLKRKKWSVVVLNNT 120
Query: 260 HIHERSETLLGHPYVGRPFTNKRLAD 285
H H L Y + F K AD
Sbjct: 121 HTHPLKFNPLSEEY--KKFKEKLRAD 144
>UniRef100_Q08957 S.cerevisiae chromosome XVI reading frame ORF YPL202c
[Saccharomyces cerevisiae]
Length = 416
Score = 40.4 bits (93), Expect = 0.062
Identities = 35/122 (28%), Positives = 50/122 (40%), Gaps = 12/122 (9%)
Query: 146 KLVRAEDVQDEPLAVDYTQSFTTDKV--FASRDEILEWARNLGKQHGFIIVITRSDNGGL 203
KL + D ++ D + D V F R EI W + + G IVI RSD+ +
Sbjct: 23 KLTASPDNLASMMSKDQNKLIHLDPVPSFEDRHEIKPWLQKIFYPQGIDIVIERSDSSKV 82
Query: 204 KRKTFMILGCERCDKYIPYKEVLKHQSTGTKKCYCPFRLRARGT--KVGWKVSVMNGYHI 261
K C + K + + ++ CPFR+RA + W V VMN H
Sbjct: 83 TFK------CRSVRSKVGLNP--KSKGSSSRSHACPFRIRAAYSVRLQKWNVVVMNNIHS 134
Query: 262 HE 263
HE
Sbjct: 135 HE 136
>UniRef100_UPI0000306A5A UPI0000306A5A UniRef100 entry
Length = 262
Score = 40.0 bits (92), Expect = 0.081
Identities = 42/174 (24%), Positives = 77/174 (44%), Gaps = 17/174 (9%)
Query: 143 LTGKLVRAEDVQDEPLAVDYTQSFTTDKVFASRDEILEWARNLGKQHGFIIVITRSD--- 199
L G L+ A D+ D V Y SF + +F + + ++ K+ FI+V + S
Sbjct: 2 LKGNLIAALDIHDN---VGYVDSFFPNCIFFKHENLF-FSYLKKKKIDFIVVCSPSYLHF 57
Query: 200 ---NGGLKRKTFMILGCERCDKYIPYKEVLKHQSTGTKKCYCPFRLRA--RGTKVGWKVS 254
L +I+ Y YK++ + + KKCYC F+LR + K+ +++
Sbjct: 58 KHIRSSLLTGKNVIVEKPPLLSYHEYKKINRLEKLSKKKCYCIFQLRLDDKLKKLKKEIN 117
Query: 255 VMNG-YHIHERSETLLGHPYVGRPFTNKRLADGPRPVRLANLMVGCVKSIVWVF 307
+ N Y++ + T G Y +K+L+ G L N+ + ++W+F
Sbjct: 118 LKNKFYNVEIKYYTYRGDWYFKSWKNDKKLSGG----LLVNIAIHFFDILIWMF 167
>UniRef100_UPI000042F4A0 UPI000042F4A0 UniRef100 entry
Length = 573
Score = 39.7 bits (91), Expect = 0.11
Identities = 34/120 (28%), Positives = 49/120 (40%), Gaps = 22/120 (18%)
Query: 169 DKVFASRDEILEWARNLGKQHGFIIVITRSDNGGLKRKTFMILGCERCDKYIPYK----E 224
++VF SRDE+ E+ + +GF +VI S+ K + CE +Y K +
Sbjct: 208 EQVFNSRDELNEFIAEFARDNGFGVVIAHSN------KKAIYYTCELGGRYRHKKNKKID 261
Query: 225 VLKHQSTG----------TKKCYCPFRLRARGTKV--GWKVSVMNGYHIHERSETLLGHP 272
V K G TKK CPF + A K W + H H + + L HP
Sbjct: 262 VTKQIDVGDGYMLDPDTKTKKLKCPFAMTASYKKSANAWTLRTTCNEHNHPQLDPLSNHP 321
>UniRef100_UPI000042D64F UPI000042D64F UniRef100 entry
Length = 573
Score = 39.7 bits (91), Expect = 0.11
Identities = 34/120 (28%), Positives = 49/120 (40%), Gaps = 22/120 (18%)
Query: 169 DKVFASRDEILEWARNLGKQHGFIIVITRSDNGGLKRKTFMILGCERCDKYIPYK----E 224
++VF SRDE+ E+ + +GF +VI S+ K + CE +Y K +
Sbjct: 208 EQVFNSRDELNEFIAEFARDNGFGVVIAHSN------KKAIYYTCELGGRYRHKKNKKID 261
Query: 225 VLKHQSTG----------TKKCYCPFRLRARGTKV--GWKVSVMNGYHIHERSETLLGHP 272
V K G TKK CPF + A K W + H H + + L HP
Sbjct: 262 VTKQIDVGDGYMLDPDTKTKKLKCPFAMTASYKKSANAWTLRTTCNEHNHPQLDPLSNHP 321
>UniRef100_UPI0000235728 UPI0000235728 UniRef100 entry
Length = 339
Score = 38.9 bits (89), Expect = 0.18
Identities = 29/98 (29%), Positives = 45/98 (45%), Gaps = 8/98 (8%)
Query: 172 FASRDEILEWARNLGKQHGFIIVITRSDNGGLK---RKTFMILGCERCDKYIPY----KE 224
+ + +L + K HG+ +V+ S K R + L C+R Y P +E
Sbjct: 33 YPDKTSLLASVQAHAKAHGYNVVVKSSSTPTEKKPGRTAKVWLRCDRGGHYRPRNGLTEE 92
Query: 225 VLKHQSTGTKKCYCPFRLRARGTKVGWKVSVMNGYHIH 262
K + T ++ CPF L A GT W ++V+NG H H
Sbjct: 93 TRKRRRT-SRLMDCPFMLVAAGTPGIWTLTVLNGTHNH 129
>UniRef100_Q6FSP0 Similarities with sp|Q08957 Saccharomyces cerevisiae YPL202c
[Candida glabrata]
Length = 437
Score = 38.9 bits (89), Expect = 0.18
Identities = 34/117 (29%), Positives = 55/117 (46%), Gaps = 13/117 (11%)
Query: 152 DVQDEPLAVDYTQSFTTDKV--FASRDEILEWARNLGKQHGFIIVITRSDNGGLKRKTFM 209
D + L V+ Q D V F + ++ W + + G IVI RSD KT +
Sbjct: 20 DNLENALMVNENQLIHIDPVPDFQEKIDVKVWLQEMFFPMGIDIVIERSD------KTKI 73
Query: 210 ILGCERCD--KYIPYKEVLKHQSTGTKKCYCPFRLRAR-GTKV-GWKVSVMNGYHIH 262
I C+ D + P K++ + + K+ CPFR+R T++ WK+ ++N H H
Sbjct: 74 IFKCKPSDYRSHEP-KDLRQKEPVDDKRHICPFRVRCTFSTQLKKWKIVIINNSHSH 129
>UniRef100_UPI000023ED11 UPI000023ED11 UniRef100 entry
Length = 362
Score = 38.1 bits (87), Expect = 0.31
Identities = 28/100 (28%), Positives = 45/100 (45%), Gaps = 7/100 (7%)
Query: 171 VFASRDEILEWARNLGKQHGFIIVITRSDNGGLKRKTFMILGCERCDKYI-----PYKEV 225
+F S +E+ + + ++ G+ IV RS G + RCD PY+ +
Sbjct: 15 IFPSYEELFKDLSDRMEKDGYKIVKARSHRGKVGGADIPGNDIVRCDLVCDRGGRPYRCM 74
Query: 226 LKHQSTGTKKCYCPFRLRA--RGTKVGWKVSVMNGYHIHE 263
T TKK CP++ +A R T GW +++ H HE
Sbjct: 75 ATKHKTTTKKTDCPWKAKAVHRKTMGGWVLTITCDQHNHE 114
>UniRef100_Q7SAH4 Predicted protein [Neurospora crassa]
Length = 569
Score = 38.1 bits (87), Expect = 0.31
Identities = 36/121 (29%), Positives = 57/121 (46%), Gaps = 19/121 (15%)
Query: 173 ASRDEILEWARNLGKQHGFIIVITRSDNG--GLKRKTFMILGCERCDKYIPYKEV---LK 227
A +LE+ ++G + +VI RS GLK+ F+ C+R K P K V ++
Sbjct: 316 AIHKHVLEYCTSVG----YAVVIGRSKKTVPGLKKVLFV---CDRAGK--PPKRVSPEMR 366
Query: 228 HQSTGTKKCYCPFRLRARGTKVGWKV----SVMNGYHIHERSETLLGHPYVGRPFTNKRL 283
+ T ++KC CPF A + W + ++ H H SE+ L HP R +K +
Sbjct: 367 KRKTTSRKCDCPFGFFAIEQRTQWTIRYRPDAIHLQHNHGPSESPLLHP-AARKLDSKMV 425
Query: 284 A 284
A
Sbjct: 426 A 426
>UniRef100_UPI000021AD4C UPI000021AD4C UniRef100 entry
Length = 466
Score = 37.4 bits (85), Expect = 0.52
Identities = 24/96 (25%), Positives = 44/96 (45%), Gaps = 5/96 (5%)
Query: 171 VFASRDEILEWARNLGKQHGFIIVITRSDNGGLKRKTFMILGCERCDKYIPYKEVLKHQS 230
++ S +++L + K+ G+ +V R+ N + T L C+R + Y K ++
Sbjct: 248 IYRSFEDLLSAVQQFSKEQGYGVVKLRASNYRDGKPTRYDLVCDRGG--VKYSSTAKKRN 305
Query: 231 TGTKKCYCPFRLRA---RGTKVGWKVSVMNGYHIHE 263
T+K CP+R +A W+ +V H HE
Sbjct: 306 PSTRKVDCPWRAKAVCEVQLSNQWRFAVQEARHNHE 341
>UniRef100_Q6BP54 Similar to CA1250|IPF9626 Candida albicans IPF9626 unknown function
[Debaryomyces hansenii]
Length = 481
Score = 37.4 bits (85), Expect = 0.52
Identities = 29/115 (25%), Positives = 47/115 (40%), Gaps = 11/115 (9%)
Query: 169 DKVFASRDEILEWARNLGKQHGFIIVITRSDN---------GGLKRKTFMILGCERCDKY 219
++V +RD++ E+ + + +GF +VI S+ GG R+ G E
Sbjct: 159 EQVLHNRDDLNEFIQEFARDNGFGVVIAHSNKKAIYYTCELGGRYRQKKSKKGMEDARHL 218
Query: 220 IPYKEVLKHQSTGTKKCYCPFRLRARGTKVG--WKVSVMNGYHIHERSETLLGHP 272
+ +T TKK CPF + A K W + H H + + L HP
Sbjct: 219 EVDNGYILDPNTKTKKLRCPFSMTATYKKSTGMWTLKTTCNEHNHPQLDPLSNHP 273
>UniRef100_Q7SBV3 Hypothetical protein [Neurospora crassa]
Length = 704
Score = 36.6 bits (83), Expect = 0.89
Identities = 24/96 (25%), Positives = 46/96 (47%), Gaps = 5/96 (5%)
Query: 171 VFASRDEILEWARNLGKQHGFIIVITRSDNGGLKRKTFMILGCERCDKYIPYKEVLKHQS 230
++ + +++L + + K G+ +V R+ N + T L C+R + Y K ++
Sbjct: 460 IYRTFEDLLASVQKVAKDQGYGVVKLRASNYREGKPTRYDLVCDRGG--VKYSSTAKKRN 517
Query: 231 TGTKKCYCPFRLRA-RGTKVG--WKVSVMNGYHIHE 263
T+K CP+R +A +G W+ +V H HE
Sbjct: 518 PSTRKVDCPWRAKAVCEVNLGNQWRFAVQEARHNHE 553
>UniRef100_P06274 DNA-directed RNA polymerase beta'' chain [Marchantia polymorpha]
Length = 1386
Score = 35.4 bits (80), Expect = 2.0
Identities = 25/76 (32%), Positives = 39/76 (50%), Gaps = 6/76 (7%)
Query: 14 QSETQHSILTKVTRSSIYSEQLSEQRSSKKRR------KSSSPLLLLKLTGDGKMRENEN 67
+S+ ++IL K+ S Y+ L E KK+ K+ + L LK+ +G ++ NE
Sbjct: 535 KSQLTNNILNKINNSKNYNFILQEYNIKKKKNFYFLKNKNLTCPLFLKIKKNGVLKNNEI 594
Query: 68 FAILIFPAPSSVNSKI 83
FAIL P+ NS I
Sbjct: 595 FAILDDPSYKVKNSGI 610
>UniRef100_UPI00002350D1 UPI00002350D1 UniRef100 entry
Length = 404
Score = 35.0 bits (79), Expect = 2.6
Identities = 24/110 (21%), Positives = 46/110 (41%), Gaps = 5/110 (4%)
Query: 169 DKVFASRDEILEWARNLGKQHGFIIVITRSDNGGLKRKTFMILGCERCDKYIPYKEVLKH 228
D + + E ++ + + HG+ ++ R++ G + + L C++ Y +
Sbjct: 172 DATYRTETEAVQAIKTFAQDHGYAVITKRTNKGREGKIEAIYLTCDQGQVY--QSTAKER 229
Query: 229 QSTGTKKCYCPFRLRARGTKVG--WKVSVMNGYHIHERSETLLGHPYVGR 276
Q +++ CPF +R K W V V + H H S HP + R
Sbjct: 230 QREPSRRTGCPFSIRISHHKTDNLWHVKVRDPSHNHGPSRP-GSHPSIRR 278
>UniRef100_UPI00002C71B7 UPI00002C71B7 UniRef100 entry
Length = 473
Score = 34.7 bits (78), Expect = 3.4
Identities = 28/108 (25%), Positives = 51/108 (46%), Gaps = 11/108 (10%)
Query: 12 HNQSETQHSILTKVTRSSIYSEQLSEQRSSKKRRKSSSPLLLLKLTGDGKMRENENFAIL 71
HN ++ +TK + I+SE++ + +K + + + +L K++ K +N+ I+
Sbjct: 43 HNGNKFVSQAVTKGVKVVIHSEKIKKNTKTKFIKFADTRDILAKISS--KFFKNKPKNII 100
Query: 72 IFPAPSSVNSKIGFGD------RIPKLKRGFLELSGCRKRKLPSEERL 113
A + N K D ++ LK GF+ GCRK KL + L
Sbjct: 101 ---AVTGTNGKTSISDFFYQIFKLQNLKSGFIGTLGCRKDKLLKKRNL 145
>UniRef100_UPI00003C212D UPI00003C212D UniRef100 entry
Length = 1004
Score = 34.3 bits (77), Expect = 4.4
Identities = 25/64 (39%), Positives = 32/64 (49%)
Query: 15 SETQHSILTKVTRSSIYSEQLSEQRSSKKRRKSSSPLLLLKLTGDGKMRENENFAILIFP 74
S T S + T SS S LS+ S+K RKSS PLL + + D + N N + P
Sbjct: 245 SPTLSSPTSASTPSSSVSSSLSKAASNKPSRKSSFPLLFGRKSLDSQKCANVNNGLHQEP 304
Query: 75 APSS 78
PSS
Sbjct: 305 LPSS 308
>UniRef100_UPI00003607EB UPI00003607EB UniRef100 entry
Length = 790
Score = 33.5 bits (75), Expect = 7.6
Identities = 29/98 (29%), Positives = 44/98 (44%), Gaps = 1/98 (1%)
Query: 20 SILTKVTRSSIYSEQLSEQRSSKKRRKSSSPLLLLKLTGDGKMR-ENENFAILIFPAPSS 78
+I+T VT E+ E+R + R++S LLL L+ + R + EN I
Sbjct: 376 NIITTVTPERKAEEENGERRMPRTMRQTSQTLLLPALSTKKQRRVKGENIFIRHINLMLE 435
Query: 79 VNSKIGFGDRIPKLKRGFLELSGCRKRKLPSEERLETS 116
V + I +LKR FLE + E+RL +S
Sbjct: 436 VAEVVKHQTNISELKRSFLETGDGTQGPTEWEKRLSSS 473
>UniRef100_Q9V3R0 CG7631-PA [Drosophila melanogaster]
Length = 254
Score = 33.5 bits (75), Expect = 7.6
Identities = 27/94 (28%), Positives = 45/94 (47%), Gaps = 13/94 (13%)
Query: 64 ENENFAILIFPAPSSVNSKIGF--GDRIPKLKRGFLELSGCRKRKLPSEERLET------ 115
E+E + + P+ S +SK+G+ G+R KLK G L+ +GC K E L T
Sbjct: 100 EDEENYLFVLPSTSGCSSKVGYQPGERTVKLKPGSLD-TGCFKLGTIQHELLHTLGFHHQ 158
Query: 116 ----SREMGVRVWNLHLKLRIEKVFINQEKPLTG 145
+R+ V++ ++ EK F+ E+ G
Sbjct: 159 QCSPNRDEFVKIVEENISEGHEKNFVKYEEDEVG 192
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 507,833,532
Number of Sequences: 2790947
Number of extensions: 20881957
Number of successful extensions: 46410
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 46397
Number of HSP's gapped (non-prelim): 28
length of query: 313
length of database: 848,049,833
effective HSP length: 127
effective length of query: 186
effective length of database: 493,599,564
effective search space: 91809518904
effective search space used: 91809518904
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0022.15


