

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0020.6
(795 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
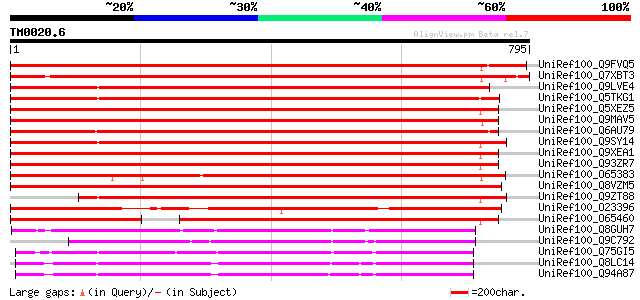
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FVQ5 Hypothetical protein F3C3.11 [Arabidopsis thali... 1189 0.0
UniRef100_Q7XBT3 Hypothetical protein [Oryza sativa] 1108 0.0
UniRef100_Q9LVE4 Gb|AAD36947.1 [Arabidopsis thaliana] 982 0.0
UniRef100_Q5TKG1 Putative early-responsive to dehydration stress... 964 0.0
UniRef100_Q5XEZ5 At4g22120 [Arabidopsis thaliana] 947 0.0
UniRef100_Q9MAV5 F24O1.4 [Arabidopsis thaliana] 943 0.0
UniRef100_Q6AU79 Hypothetical protein OSJNBa0014C03.4 [Oryza sat... 939 0.0
UniRef100_Q9SY14 Hypothetical protein T5J8.22 [Arabidopsis thali... 937 0.0
UniRef100_Q9XEA1 Hypothetical protein T19B17.6 [Arabidopsis thal... 937 0.0
UniRef100_Q93ZR7 Hypothetical protein At4g04340 [Arabidopsis tha... 936 0.0
UniRef100_O65383 F12F1.17 protein [Arabidopsis thaliana] 927 0.0
UniRef100_Q8VZM5 Hypothetical protein At4g15430 [Arabidopsis tha... 920 0.0
UniRef100_Q9ZT88 Hypothetical protein T4I9.22 [Arabidopsis thali... 793 0.0
UniRef100_O23396 Hypothetical protein dl3760w [Arabidopsis thali... 745 0.0
UniRef100_O65460 Hypothetical protein AT4g22120 [Arabidopsis tha... 620 e-176
UniRef100_Q8GUH7 Hypothetical protein At3g01100At3g01110 [Arabid... 435 e-120
UniRef100_Q9C792 Hypothetical protein F10D13_27 [Arabidopsis tha... 381 e-104
UniRef100_Q75GI5 Expressed protein [Oryza sativa] 380 e-104
UniRef100_Q8LC14 Hypothetical protein [Arabidopsis thaliana] 370 e-100
UniRef100_Q94A87 At1g10080/T27I1_10 [Arabidopsis thaliana] 369 e-100
>UniRef100_Q9FVQ5 Hypothetical protein F3C3.11 [Arabidopsis thaliana]
Length = 806
Score = 1189 bits (3077), Expect = 0.0
Identities = 597/807 (73%), Positives = 678/807 (83%), Gaps = 18/807 (2%)
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPR-STGNFVG 59
MATL DIGVSA IN+ AF FL+AFA+LRIQPINDRVYFPKWY+ GER SPR S VG
Sbjct: 1 MATLQDIGVSALINLFGAFLFLIAFAVLRIQPINDRVYFPKWYLTGERNSPRRSDRTLVG 60
Query: 60 KFVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLI 119
KFVNLN+KTY TFLNWMPQA++MSE+EII HAGLDSA+FLRIYTLGLKIF PV V+AL++
Sbjct: 61 KFVNLNYKTYFTFLNWMPQAMKMSESEIIRHAGLDSAIFLRIYTLGLKIFAPVMVLALVV 120
Query: 120 LIPVNVSSGALFFLKRDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEY 179
L+PVNVSSG LFFLK++LVVS+IDKLSISNV KSS+FF HI +EY+FT W CF+LY+EY
Sbjct: 121 LVPVNVSSGTLFFLKKELVVSNIDKLSISNVQPKSSKFFFHIAVEYIFTFWACFMLYREY 180
Query: 180 DNIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVYN 239
+N+A+MRL +LASQRRR EQFTVVVR+VP + GHSV D+V+ FF+TNHP HY+ HQAVYN
Sbjct: 181 NNVAIMRLQYLASQRRRPEQFTVVVRNVPDMPGHSVPDTVDQFFKTNHPEHYLCHQAVYN 240
Query: 240 ANKFAKLVRRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKEL 299
AN +AKLV++R +LQ W DYY LK QR+P K+PT TG LGLWG++VD+IEYY IKE
Sbjct: 241 ANTYAKLVKQRAKLQRWFDYYVLKHQRNPHKQPTCRTGFLGLWGKRVDSIEYYKQQIKEF 300
Query: 300 DKVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYW 359
D M+LERQ+++KD K +LPVAF+SF SRWGA+VCAQTQQSKNPTLWLT APEPRDIYW
Sbjct: 301 DHNMSLERQKVLKDSKLMLPVAFVSFDSRWGAAVCAQTQQSKNPTLWLTSSAPEPRDIYW 360
Query: 360 RNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFI 419
+NLAIPF+SL+IRKLVI +SVFALVFFYMIPIAFVQSLANLEGL+RVAPFLRPV L FI
Sbjct: 361 QNLAIPFISLTIRKLVIGVSVFALVFFYMIPIAFVQSLANLEGLDRVAPFLRPVTRLDFI 420
Query: 420 KSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSI 479
KSFLQGFLPGLALKIFL+ILP VL+IMSKIEG+IA STLER+ AAKYYYFMLVNVFLGSI
Sbjct: 421 KSFLQGFLPGLALKIFLWILPTVLLIMSKIEGYIALSTLERRAAAKYYYFMLVNVFLGSI 480
Query: 480 VTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIY 539
+ GTAFEQLHSFLHQSP+QIPRTIGVSIPMKATFF+TYIMVDGWAGIAGEILRLKPLVI+
Sbjct: 481 IAGTAFEQLHSFLHQSPSQIPRTIGVSIPMKATFFITYIMVDGWAGIAGEILRLKPLVIF 540
Query: 540 HLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALA 599
HLKNMFIVKTE DR +AMDPG VDF ETIPSLQLYFLLGIVY VTPILLPFIL+FFA A
Sbjct: 541 HLKNMFIVKTEEDRVRAMDPGFVDFKETIPSLQLYFLLGIVYTAVTPILLPFILIFFAFA 600
Query: 600 YLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAIL 659
YLVYRHQIINVYNQQYES AFWPHVH RIIASLLISQLL++GLL++KKAA STPLL IL
Sbjct: 601 YLVYRHQIINVYNQQYESCGAFWPHVHGRIIASLLISQLLLMGLLASKKAADSTPLLIIL 660
Query: 660 PILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRSFE 719
PILT +FHKYC+HRFEPAFR+YPLEEAMAKD LEK TEP+LN+KA LADAYLHPIF SFE
Sbjct: 661 PILTLSFHKYCKHRFEPAFRQYPLEEAMAKDKLEKETEPELNMKADLADAYLHPIFHSFE 720
Query: 720 VE---------------EELVEVRVDKQPETQVASPTLSEPSSPSPLHDHVHQPSPPQHV 764
E EE EVRVDK ETQ +SP ++E + S H + SP H
Sbjct: 721 KEVELSSSSSSEKETHQEETPEVRVDKH-ETQSSSP-VTELGTSSHHHHVYNSTSPSSHY 778
Query: 765 NQYSHYPTSPPGYYYHPPPPPHYAYQY 791
+S Y+Y+ + Y+Y
Sbjct: 779 ASAYEQSSSQYEYHYNTHQYEEHEYRY 805
>UniRef100_Q7XBT3 Hypothetical protein [Oryza sativa]
Length = 810
Score = 1108 bits (2866), Expect = 0.0
Identities = 550/816 (67%), Positives = 656/816 (79%), Gaps = 27/816 (3%)
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTS-PRSTGNFVG 59
MATL D+GVSA INIL AF FLL FA LR+QPINDRVYFPK Y+ G+R P G
Sbjct: 1 MATLPDLGVSAFINILGAFVFLLIFAALRLQPINDRVYFPKLYLTGQRRHHPHPHG---- 56
Query: 60 KFVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLI 119
FVNL+ +YL FL W+P ALRMS+ ++I+HAGLDSAV+LRIYTLGLKIF+P+ VALL+
Sbjct: 57 -FVNLDLCSYLRFLAWVPGALRMSQPDLIHHAGLDSAVYLRIYTLGLKIFLPIMTVALLV 115
Query: 120 LIPVNVSSGALFFLKRDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEY 179
LIPVNVS G L L++++V SDIDKLSISNV S+RFF+H+ + Y+FT W CF+LYKEY
Sbjct: 116 LIPVNVSGGTLLNLRKEIVFSDIDKLSISNVNPGSNRFFIHLLMAYVFTFWTCFMLYKEY 175
Query: 180 DNIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVYN 239
N+A MRLHFLASQ+R +QFTV+VR++PH+S HS S++V+ FF+ NHP+HY+G QAVYN
Sbjct: 176 SNVAFMRLHFLASQKRCADQFTVIVRNIPHVSSHSTSETVDEFFRRNHPDHYLGQQAVYN 235
Query: 240 ANKFAKLVRRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKEL 299
AN++AKLV++++RLQNWLDYYQLKF+RHP KRP TG LG GR+VD I+YY I EL
Sbjct: 236 ANRYAKLVKKKERLQNWLDYYQLKFERHPGKRPIGRTGCLGFCGREVDQIDYYRARISEL 295
Query: 300 DKVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYW 359
DK + ERQR++ DPK ++PVAF++F SRWGA+VCAQTQQSKNPT WLTDWAPEPRD+YW
Sbjct: 296 DKKLASERQRVLNDPKAVMPVAFVTFDSRWGAAVCAQTQQSKNPTQWLTDWAPEPRDVYW 355
Query: 360 RNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFI 419
+NLAIPF SLSIRK +IS++VFALVFFYMIPIAFVQSLANLEG+E+VAPFLRPVI+ +
Sbjct: 356 QNLAIPFFSLSIRKFLISIAVFALVFFYMIPIAFVQSLANLEGIEKVAPFLRPVIDTPVV 415
Query: 420 KSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSI 479
KSFLQGFLPGLALKIFLYILP VLMIMSK+EG+++ S+LER+ A+KYYYFMLVNVFLGSI
Sbjct: 416 KSFLQGFLPGLALKIFLYILPTVLMIMSKVEGYVSLSSLERRAASKYYYFMLVNVFLGSI 475
Query: 480 VTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIY 539
+ GTAFEQL++F HQ P+QIPRTIGV+IPMKATFFMTYIMVDGWAGIA EILR+KPLVIY
Sbjct: 476 IAGTAFEQLNAFFHQPPSQIPRTIGVAIPMKATFFMTYIMVDGWAGIANEILRVKPLVIY 535
Query: 540 HLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALA 599
HLKNMFIVKTERDR +AMDPGS+ E +PSLQLYFLLG+VYAVVTPILLPFI++FFA A
Sbjct: 536 HLKNMFIVKTERDRERAMDPGSIGLAENLPSLQLYFLLGLVYAVVTPILLPFIIIFFAFA 595
Query: 600 YLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAIL 659
+LVYRHQIINVYNQ+YESAAAFWP VHSRIIASLLIS + + GL+ST KAA STPLL L
Sbjct: 596 FLVYRHQIINVYNQEYESAAAFWPQVHSRIIASLLISHVTLFGLMSTMKAAYSTPLLIFL 655
Query: 660 PILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRSFE 719
P+LT FHKYC+ RFEPAFRKYPLEEAM KD LE+T+EP+LN+K+YL +AYLHPIF FE
Sbjct: 656 PLLTIWFHKYCKSRFEPAFRKYPLEEAMEKDNLERTSEPNLNLKSYLQNAYLHPIFHMFE 715
Query: 720 VE---------EELVEVRVDKQPE---TQVASPTLSEPSSPSPLHDHVHQ-----PSPPQ 762
+ EE VEVR+DK + QV SS + H + H +
Sbjct: 716 QQQQQEQEQQREEKVEVRIDKAQQHHHRQVEEEEEESKSSQATTHYYHHHHEQTTTTTHH 775
Query: 763 HVNQY---SHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
H +Q+ SHY P P PPH+ Y Y +P
Sbjct: 776 HYHQHEHMSHYHMGPSD-TADSPSPPHFVYHYGVDP 810
>UniRef100_Q9LVE4 Gb|AAD36947.1 [Arabidopsis thaliana]
Length = 756
Score = 982 bits (2538), Expect = 0.0
Identities = 464/737 (62%), Positives = 603/737 (80%), Gaps = 4/737 (0%)
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGK 60
MATL DIGV+A INIL+AFAF +AFA+LR+QP+NDRVYFPKWY+ G R+SP TG F K
Sbjct: 1 MATLTDIGVAATINILTAFAFFIAFAILRLQPVNDRVYFPKWYLKGLRSSPIKTGGFASK 60
Query: 61 FVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLIL 120
FVNL+F++Y+ FLNWMPQALRM E E+I+HAGLDS V+LRIY LGLKIF P+ +A ++
Sbjct: 61 FVNLDFRSYIRFLNWMPQALRMPEPELIDHAGLDSVVYLRIYLLGLKIFFPIACIAFTVM 120
Query: 121 IPVNVSSGALFFLKRDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEYD 180
+PVN ++ L LK +L SDIDKLSISN+P+ SSRF+VH+ + Y+ T W CF+L +EY
Sbjct: 121 VPVNWTNSTLDQLK-NLTFSDIDKLSISNIPTGSSRFWVHLCMAYVITFWTCFVLQREYK 179
Query: 181 NIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVYNA 240
+IA MRL FLAS+ RR +QFTV+VR++P SVS+ VE FF+ NHP++Y+ +QAVYNA
Sbjct: 180 HIASMRLQFLASEHRRPDQFTVLVRNIPPDPDESVSELVEHFFKVNHPDYYLTYQAVYNA 239
Query: 241 NKFAKLVRRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKELD 300
NK ++LV++R +LQNWLDYYQ K R+P KRP I G LG WG +VDAI++Y I+ L
Sbjct: 240 NKLSELVQKRMKLQNWLDYYQNKHSRNPSKRPLIKIGFLGCWGEEVDAIDHYIEKIEGLT 299
Query: 301 KVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYWR 360
+ ++ E++ ++ K ++P AF+SF RWGA VC+QTQQS+NPT WLT+WAPEPRDIYW
Sbjct: 300 RKISEEKETVMSSTKSLVPAAFVSFKKRWGAVVCSQTQQSRNPTEWLTEWAPEPRDIYWD 359
Query: 361 NLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFIK 420
NLA+P+V L+IR+LVI+++ F L FF+MIPIAFVQ+LAN+EG+E+ PFL+P+IE+K +K
Sbjct: 360 NLALPYVQLTIRRLVIAVAFFFLTFFFMIPIAFVQTLANIEGIEKAVPFLKPLIEVKTVK 419
Query: 421 SFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSIV 480
SF+QGFLPG+ALKIFL +LP++LM+MSK EG I+KS+LER+ A++YY F +NVFL SI+
Sbjct: 420 SFIQGFLPGIALKIFLIVLPSILMLMSKFEGFISKSSLERRCASRYYMFQFINVFLCSII 479
Query: 481 TGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIYH 540
GTA +QL SFL+QS T+IP+TIGVSIPMKATFF+TYIMVDGWAG+AGEILRLKPL+IYH
Sbjct: 480 AGTALQQLDSFLNQSATEIPKTIGVSIPMKATFFITYIMVDGWAGVAGEILRLKPLIIYH 539
Query: 541 LKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALAY 600
LKN F+VKTE+DR +AMDPG++ F P +QLYF+LG+VYA V+PILLPFILVFFALAY
Sbjct: 540 LKNFFLVKTEKDREEAMDPGTIGFNTGEPQIQLYFILGLVYAAVSPILLPFILVFFALAY 599
Query: 601 LVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAILP 660
+VYRHQIINVYNQ+YESAAAFWP VH R++ +L++SQLL++GLLSTKKAA+STPLL ILP
Sbjct: 600 VVYRHQIINVYNQEYESAAAFWPDVHRRVVIALIVSQLLLMGLLSTKKAARSTPLLFILP 659
Query: 661 ILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRSFE- 719
+LT FHK+CQ R++P F YPL++AM KD LE+ EP+LN+K +L +AY HP+F++ +
Sbjct: 660 VLTIGFHKFCQGRYQPIFVTYPLQDAMVKDTLERMREPNLNLKTFLQNAYAHPVFKAADN 719
Query: 720 VEEELV--EVRVDKQPE 734
+ E+V E DK P+
Sbjct: 720 LANEMVVEEPAPDKTPD 736
>UniRef100_Q5TKG1 Putative early-responsive to dehydration stress protein [Oryza
sativa]
Length = 766
Score = 964 bits (2492), Expect = 0.0
Identities = 460/750 (61%), Positives = 603/750 (80%), Gaps = 4/750 (0%)
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGK 60
MAT++DIG+SAAIN+ A AFLL FA LR+QPINDRVYFPKWY+ G R SP S+G V K
Sbjct: 1 MATVSDIGLSAAINVSMAVAFLLVFAFLRLQPINDRVYFPKWYLRGMRDSPVSSGAAVQK 60
Query: 61 FVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLIL 120
VNLN ++YL FL+WMP AL+M E+E+INHAGLDSAV+LRIY G+KIFVP++++A L+L
Sbjct: 61 VVNLNMRSYLKFLSWMPAALKMPEDELINHAGLDSAVYLRIYLTGIKIFVPISILASLVL 120
Query: 121 IPVNVSSGALFFLKRDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEYD 180
PVN ++ L +K +V S IDKLSISN+P S+RF H+ + Y T W C++L++EY+
Sbjct: 121 FPVNWTNDTLDSMK--VVHSKIDKLSISNIPYGSNRFVTHLVMAYAVTFWTCYVLFREYE 178
Query: 181 NIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVYNA 240
I MRL FLAS++RR +QFTV+VR++P S+S+ VE FF NHP+HY+ HQ VYNA
Sbjct: 179 IITTMRLRFLASEKRRPDQFTVLVRNIPPDPDESISELVEHFFLVNHPDHYLRHQVVYNA 238
Query: 241 NKFAKLVRRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKELD 300
NK A LV ++ +LQNWLDYYQLK++R+P KRPT TG LG +G +VDAIEYY I+++
Sbjct: 239 NKLADLVEKKKKLQNWLDYYQLKYERNPSKRPTTKTGFLGCFGSEVDAIEYYKAEIEKIG 298
Query: 301 KVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYWR 360
K ERQ+I+KDP+ +P AF+SF SRWGA+VCAQTQQ+ NPT+W+T+WAPEPRD+YW
Sbjct: 299 KEEADERQKIMKDPQSAVPAAFVSFRSRWGAAVCAQTQQTSNPTVWITEWAPEPRDVYWN 358
Query: 361 NLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFIK 420
NL+IPFVSL++R+L+++++ F L FFY+IPIAFVQSLA+LEG+E+ PFL+P+I++ IK
Sbjct: 359 NLSIPFVSLTVRRLIVAVAFFFLNFFYVIPIAFVQSLASLEGIEKALPFLKPLIKIDVIK 418
Query: 421 SFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSIV 480
SF+QGFLPG+ALK+FL +LP +LM MSK EG I++S+LER++A+KYY F+ NVFLGSIV
Sbjct: 419 SFIQGFLPGIALKVFLILLPTILMFMSKFEGLISQSSLERRSASKYYIFLFFNVFLGSIV 478
Query: 481 TGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIYH 540
TG+A +QL +++HQS +IPRTIGV+IPM+ATFF+TY+MVDGW G+AGEILRL+ L+I+H
Sbjct: 479 TGSALDQLKAYIHQSANEIPRTIGVAIPMRATFFITYVMVDGWTGVAGEILRLRALIIFH 538
Query: 541 LKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALAY 600
LKN F+VKTE+DR +AMDPGS+ F P +QLYFLLG+VYAVVTP+LLPFILVFF LAY
Sbjct: 539 LKNFFLVKTEKDREEAMDPGSICFDWCEPRIQLYFLLGLVYAVVTPLLLPFILVFFGLAY 598
Query: 601 LVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAILP 660
+VYRHQIINVYNQQYES A FWP VH RII +L++SQLL++GLLSTK ++TP+L +LP
Sbjct: 599 VVYRHQIINVYNQQYESGAQFWPSVHGRIIIALIVSQLLLIGLLSTKGFEETTPVLVVLP 658
Query: 661 ILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRSFEV 720
+LTF F+KYC++RFEPAF + PL+EAM KD LE+ EP ++KAYLA+AYLHP+F+ E
Sbjct: 659 VLTFWFYKYCKNRFEPAFVRNPLQEAMRKDTLERAREPTFDLKAYLANAYLHPVFKGRE- 717
Query: 721 EEELVEVRVDKQPETQVASPTLSEPSSPSP 750
EE+ + + D E +V PT + +P
Sbjct: 718 EEDNMSISEDVGME-EVIVPTKRQSRRNTP 746
>UniRef100_Q5XEZ5 At4g22120 [Arabidopsis thaliana]
Length = 771
Score = 947 bits (2449), Expect = 0.0
Identities = 454/763 (59%), Positives = 593/763 (77%), Gaps = 14/763 (1%)
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGK 60
MATL DIGVSA INILSAF F + FA+LR+QP NDRVYF KWY+ G R+SP G F +
Sbjct: 1 MATLQDIGVSAGINILSAFVFFIIFAVLRLQPFNDRVYFSKWYLKGLRSSPARGGAFAQR 60
Query: 61 FVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLIL 120
FVNL+F++Y+ FLNWMP+AL+M E E+I+HAGLDS V+LRIY LGLKIF P+ V+A +L
Sbjct: 61 FVNLDFRSYMKFLNWMPEALKMPEPELIDHAGLDSVVYLRIYWLGLKIFTPIAVLAWAVL 120
Query: 121 IPVNVSSGALFFLK--RDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKE 178
+PVN ++ L K R++ SDIDKLS+SN+P S RF+ HI + Y FT+W C++L KE
Sbjct: 121 VPVNWTNNTLEMAKQLRNVTSSDIDKLSVSNIPEYSMRFWTHIVMAYAFTIWTCYVLMKE 180
Query: 179 YDNIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVY 238
Y+ IA MRL F+AS+ RR +QFTV+VR+VP + SVS+ VE FF NHP+HY+ HQ V
Sbjct: 181 YETIANMRLQFVASEARRPDQFTVLVRNVPPDADESVSELVEHFFLVNHPDHYLTHQVVC 240
Query: 239 NANKFAKLVRRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKE 298
NANK A LV+++ +LQNWLDYYQLK+ R+ +R + G LGLWG+KVDAIE+Y I +
Sbjct: 241 NANKLADLVKKKKKLQNWLDYYQLKYARNNSQRIMVKLGFLGLWGQKVDAIEHYIAEIDK 300
Query: 299 LDKVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIY 358
+ K ++ ER+ ++ DPK I+P AF+SF +RW A+VCAQTQQ++NPT WLT+WAPEPRD++
Sbjct: 301 ISKEISKEREEVVNDPKAIMPAAFVSFKTRWAAAVCAQTQQTRNPTQWLTEWAPEPRDVF 360
Query: 359 WRNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKF 418
W NLAIP+VSL++R+L++ ++ F L FF+++PIAFVQSLA +EG+ + APFL+ +++ KF
Sbjct: 361 WSNLAIPYVSLTVRRLIMHVAFFFLTFFFIVPIAFVQSLATIEGIVKAAPFLKFIVDDKF 420
Query: 419 IKSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGS 478
+KS +QGFLPG+ALK+FL LP++LMIMSK EG + S+LER+ A +YY F LVNVFL S
Sbjct: 421 MKSVIQGFLPGIALKLFLAFLPSILMIMSKFEGFTSISSLERRAAFRYYIFNLVNVFLAS 480
Query: 479 IVTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVI 538
++ G AFEQL+SFL+QS QIP+TIGV+IPMKATFF+TYIMVDGWAG+AGEIL LKPL++
Sbjct: 481 VIAGAAFEQLNSFLNQSANQIPKTIGVAIPMKATFFITYIMVDGWAGVAGEILMLKPLIM 540
Query: 539 YHLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFAL 598
+HLKN F+VKT++DR +AMDPGS+ F P +QLYFLLG+VYA VTP+LLPFILVFFAL
Sbjct: 541 FHLKNAFLVKTDKDREEAMDPGSIGFNTGEPRIQLYFLLGLVYAPVTPMLLPFILVFFAL 600
Query: 599 AYLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAI 658
AY+VYRHQIINVYNQ+YESAAAFWP VH R+IA+L+ISQLL++GLL TK AA + P L
Sbjct: 601 AYIVYRHQIINVYNQEYESAAAFWPDVHGRVIAALVISQLLLMGLLGTKHAALAAPFLIA 660
Query: 659 LPILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRSF 718
LP+LT FH +C+ R+EPAF +YPL+EAM KD LE EP+LN+K YL +AY+HP+F+
Sbjct: 661 LPVLTIGFHHFCKGRYEPAFIRYPLQEAMMKDTLETAREPNLNLKGYLQNAYVHPVFKGD 720
Query: 719 E-----------VEEELVEVRVDKQPETQVASPT-LSEPSSPS 749
E E+E + V +Q +P+ +S SPS
Sbjct: 721 EDDYDIDDKLGKFEDEAIIVPTKRQSRRNTPAPSIISGDDSPS 763
>UniRef100_Q9MAV5 F24O1.4 [Arabidopsis thaliana]
Length = 778
Score = 943 bits (2438), Expect = 0.0
Identities = 454/763 (59%), Positives = 595/763 (77%), Gaps = 14/763 (1%)
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGK 60
MATLADIG++AAINILSA FLL FA+LRIQP NDRVYFPKWY+ G R+SP ++G FV K
Sbjct: 1 MATLADIGLAAAINILSALIFLLLFAILRIQPFNDRVYFPKWYLKGVRSSPVNSGAFVSK 60
Query: 61 FVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLIL 120
+NL+F++Y+ FLNWMP AL+M E E+I+HAGLDSAV+LRIY +GLKIF P+ +++ IL
Sbjct: 61 IMNLDFRSYVRFLNWMPDALKMPEPELIDHAGLDSAVYLRIYLIGLKIFGPIALLSWSIL 120
Query: 121 IPVNVSSGALFFLK-RDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEY 179
+PVN +S L K R++ S+IDKLSISNV S RF+ H+ + Y FT W C++L KEY
Sbjct: 121 VPVNWTSDGLQLAKLRNVTSSNIDKLSISNVERGSDRFWAHLVMAYAFTFWTCYVLMKEY 180
Query: 180 DNIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVYN 239
+ IA MRL FL S++RR +QFTV+VR+VP S S+S++V+ FF NHP+HY+ HQ VYN
Sbjct: 181 EKIAAMRLSFLQSEKRRADQFTVLVRNVPPDSDESISENVQHFFLVNHPDHYLTHQVVYN 240
Query: 240 ANKFAKLVRRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKEL 299
AN+ AKLV + ++QNWLDYYQLK+ R+ ++RP + G LGLWG+KVDA+++Y+ I++L
Sbjct: 241 ANELAKLVEDKKKMQNWLDYYQLKYTRNKEQRPRVKMGFLGLWGKKVDAMDHYTAEIEKL 300
Query: 300 DKVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYW 359
+ + ER+RI KD K ++ AF+SF +RWGA+VCAQTQQ+KNPT WLT+WAPE R++YW
Sbjct: 301 SEQIMEERKRIKKDDKSVMQAAFVSFKTRWGAAVCAQTQQTKNPTEWLTEWAPEAREMYW 360
Query: 360 RNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFI 419
NLA+P+VSL++R+ V+ ++ F L FF++IPIAFVQSLA++EG+E+ APFL P+++ K +
Sbjct: 361 PNLAMPYVSLTVRRFVMHIAFFFLTFFFIIPIAFVQSLASIEGIEKSAPFLSPIVKNKLM 420
Query: 420 KSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSI 479
KS +QGFLPG+ LK+FL LP +LMIMSK EG I+ S+LER+ A +YY F LVNVFLGS+
Sbjct: 421 KSLIQGFLPGIVLKLFLIFLPTILMIMSKFEGFISISSLERRAAFRYYIFNLVNVFLGSV 480
Query: 480 VTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIY 539
+TG+AFEQL SFL QS IPRT+GV+IP+KATFF+TYIMVDGWAG+AGEI RLKPLVI+
Sbjct: 481 ITGSAFEQLDSFLKQSANDIPRTVGVAIPIKATFFITYIMVDGWAGVAGEIFRLKPLVIF 540
Query: 540 HLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALA 599
HLKN F VKTE+DR +AMDPG +DF T P +QLYFLLG+VYA VTP+LLPFI+ FF A
Sbjct: 541 HLKNFFFVKTEKDREEAMDPGQIDFYATEPRIQLYFLLGLVYAPVTPVLLPFIIFFFGFA 600
Query: 600 YLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAIL 659
YLV+RHQIINVYNQ+YESA AFWP VH RII++L+ISQ+L+LGL+STK +STP L +L
Sbjct: 601 YLVFRHQIINVYNQKYESAGAFWPDVHGRIISALIISQILLLGLMSTKGKVQSTPFLLVL 660
Query: 660 PILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRSFE 719
ILTF FH++C+ R+E AF PL+EAM KD LE+ EP+LN+K +L +AY+HP+F+ E
Sbjct: 661 AILTFGFHRFCKGRYESAFVINPLQEAMIKDTLERAREPNLNLKGFLQNAYVHPVFKDEE 720
Query: 720 VEEE-------------LVEVRVDKQPETQVASPTLSEPSSPS 749
+E +V+ + + T VAS S SS S
Sbjct: 721 DSDEEGLIEDSDDEDCVVVQTKRQRSRRTTVASSNASRGSSQS 763
>UniRef100_Q6AU79 Hypothetical protein OSJNBa0014C03.4 [Oryza sativa]
Length = 767
Score = 939 bits (2427), Expect = 0.0
Identities = 449/748 (60%), Positives = 584/748 (78%), Gaps = 3/748 (0%)
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGK 60
M +L DIGV+A INILSA FLLAFA+LRIQPINDRVYFPKWY+ G R+SPRS G K
Sbjct: 1 MGSLTDIGVAAGINILSALGFLLAFAVLRIQPINDRVYFPKWYLKGTRSSPRSMGTVFSK 60
Query: 61 FVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLIL 120
FVN + TY+ FLNWMP AL+M E E+I HAGLDSAV++RIY LGLKIFVP+ V+A ++L
Sbjct: 61 FVNADLSTYIRFLNWMPAALQMPEPELIEHAGLDSAVYVRIYLLGLKIFVPIAVLAFIVL 120
Query: 121 IPVNVSSGALFFLKRDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEYD 180
+P+N +SG L ++ L IDKLSISN+ S RF+ HI + Y+FT W F+LY+EY
Sbjct: 121 VPINWASGTLE-KEKSLSYDQIDKLSISNLGKGSKRFWAHIVMAYVFTFWTFFVLYREYK 179
Query: 181 NIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVYNA 240
+ MRL FLA Q RR +QFTV+VR+VP +VS+ VE FF NH +HY+ HQ VYNA
Sbjct: 180 VVTTMRLRFLAIQNRRADQFTVLVRNVPPDPDETVSEHVEHFFAVNHRDHYLSHQTVYNA 239
Query: 241 NKFAKLVRRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKELD 300
N A LV ++ LQNWL YY+ + ++P K+PT+ TG+ GLWG++VDAIE+Y+ I+EL
Sbjct: 240 NTLAGLVEQKKGLQNWLVYYENQHAKNPAKKPTMKTGLWGLWGKRVDAIEHYTTAIEELC 299
Query: 301 KVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYWR 360
K ER ++I DP I+P AF+SF SRWGA+VCAQTQQ+ NPTLWLT+WAPEPRD++W
Sbjct: 300 KQEDEERHKVITDPNAIMPAAFVSFKSRWGAAVCAQTQQTSNPTLWLTEWAPEPRDVFWP 359
Query: 361 NLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFIK 420
NLAIPFV LS+R+L++++++F L FF+MIPIA VQS+ANL+ +ER+ PFL+P+IE +K
Sbjct: 360 NLAIPFVELSVRRLIMAVALFFLTFFFMIPIAIVQSMANLDDIERMLPFLKPIIERNSLK 419
Query: 421 SFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSIV 480
S +QGFLPG+ALKIFL +LP L++MSKIEGH + S L+R+TA+KYY F+ VNVFLGS++
Sbjct: 420 SIVQGFLPGIALKIFLILLPTFLVMMSKIEGHTSLSGLDRRTASKYYLFLFVNVFLGSVI 479
Query: 481 TGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIYH 540
TGTAF+QL++F+HQS +IP +G SIPMKATFF+TY+MVDGWAG+A E+LRLKPLV++H
Sbjct: 480 TGTAFQQLNNFIHQSANKIPEIVGESIPMKATFFITYVMVDGWAGVAAEVLRLKPLVMFH 539
Query: 541 LKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALAY 600
+KN F+V+TERDR +AMDPGS+DF T P +QLYFLLG+VYAVVTPILLPFI+VFF+LAY
Sbjct: 540 IKNTFLVRTERDREQAMDPGSLDFGTTEPRIQLYFLLGLVYAVVTPILLPFIIVFFSLAY 599
Query: 601 LVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAILP 660
LV+RHQIINVYNQQYES A FWP V R++ +L++SQ+L+LGLLST++A KST L LP
Sbjct: 600 LVFRHQIINVYNQQYESGAQFWPDVQRRLVIALIVSQILLLGLLSTQEAEKSTVALLPLP 659
Query: 661 ILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRSFEV 720
+L+ FH C+ RFEPAF K+PL++AM KD LE+ +P LN++ YL DAY+HP+F+ ++
Sbjct: 660 VLSIWFHYVCKGRFEPAFIKFPLQDAMVKDTLERANDPTLNLREYLKDAYVHPVFQKNDI 719
Query: 721 EEELVEVRVDKQPETQVASPTLSEPSSP 748
E +K P VA+ S ++P
Sbjct: 720 YEFAGIDEEEKNP--MVATKRQSRMNTP 745
>UniRef100_Q9SY14 Hypothetical protein T5J8.22 [Arabidopsis thaliana]
Length = 785
Score = 937 bits (2423), Expect = 0.0
Identities = 458/766 (59%), Positives = 590/766 (76%), Gaps = 7/766 (0%)
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGK 60
MA++ DIG+SAAIN+LSAFAFL AFA+LR+QP+NDRVYFPKWY+ G R SP + + +
Sbjct: 1 MASVQDIGLSAAINLLSAFAFLFAFAMLRLQPVNDRVYFPKWYLKGIRGSPTRSRGIMTR 60
Query: 61 FVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLIL 120
FVNL++ TY+ FLNWMP AL+M E E+I HAGLDSAV++RIY LGLK+FVP+T++A +L
Sbjct: 61 FVNLDWTTYVKFLNWMPAALQMPEPELIEHAGLDSAVYIRIYLLGLKMFVPITLLAFGVL 120
Query: 121 IPVNVSSGALFFLKRDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEYD 180
+PVN + L + DL S++DKLSISNVP S RF+ HI + Y+ T W C++LY EY
Sbjct: 121 VPVNWTGETLENID-DLTFSNVDKLSISNVPPGSPRFWAHITMTYVITFWTCYILYMEYK 179
Query: 181 NIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVYNA 240
+A MRL LA++ RR +Q TV+VR+VP SV++ VE FF NHP+HY+ HQ VYNA
Sbjct: 180 AVANMRLRHLAAESRRPDQLTVLVRNVPPDPDESVNEHVEHFFCVNHPDHYLCHQVVYNA 239
Query: 241 NKFAKLVRRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKELD 300
N AKLV +R +QNWL YY+ KF+R P RPT TG G WG VDAI++Y+ + L
Sbjct: 240 NDLAKLVAQRKAMQNWLTYYENKFERKPSSRPTTKTGYGGFWGTTVDAIDFYTSKMDILA 299
Query: 301 KVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYWR 360
+ +ER++I+ DPK I+P AF+SF SRWG +VCAQTQQ NPT+WLT+WAPEPRD++W
Sbjct: 300 EQEAVEREKIMNDPKAIMPAAFVSFRSRWGTAVCAQTQQCHNPTIWLTEWAPEPRDVFWD 359
Query: 361 NLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFIK 420
NLAIP+V LSIR+L+ ++++F L+F +MIPIAFVQSLANLEG+++V PFL+PVIE+K +K
Sbjct: 360 NLAIPYVELSIRRLLTTVALFFLIFCFMIPIAFVQSLANLEGIQKVLPFLKPVIEMKTVK 419
Query: 421 SFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSIV 480
S +QGFLPG+ALKIFL ILP +LM MS+IEG+ + S L+R++A KY++F++VNVFLGSI+
Sbjct: 420 SVIQGFLPGIALKIFLIILPTILMTMSQIEGYTSLSYLDRRSAEKYFWFIIVNVFLGSII 479
Query: 481 TGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIYH 540
TGTAF+QL SFL Q PT+IP+T+GVSIPMKATFF+TYIMVDGWAGIA EILR+ PLVI+H
Sbjct: 480 TGTAFQQLKSFLEQPPTEIPKTVGVSIPMKATFFITYIMVDGWAGIAAEILRVVPLVIFH 539
Query: 541 LKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALAY 600
LKN F+VKTE+DR +AMDPG +DF + P +Q YFLLG+VYA V PILLPFI+VFFA AY
Sbjct: 540 LKNTFLVKTEQDRQQAMDPGHLDFATSEPRIQFYFLLGLVYAAVAPILLPFIIVFFAFAY 599
Query: 601 LVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAILP 660
+V+RHQ+INVY+Q+YES A +WP VH R+I L+ISQLLM+GLLSTKK AK T LL P
Sbjct: 600 VVFRHQVINVYDQKYESGARYWPDVHRRLIICLIISQLLMMGLLSTKKFAKVTALLLPQP 659
Query: 661 ILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRSFE- 719
ILTF F++YC RFE AF K+PL+EAM KD LEK TEP+LN+K YL DAY+HP+F+ +
Sbjct: 660 ILTFWFYRYCAGRFESAFSKFPLQEAMVKDTLEKATEPNLNLKEYLKDAYVHPVFKGNDF 719
Query: 720 -----VEEELVEVRVDKQPETQVASPTLSEPSSPSPLHDHVHQPSP 760
V+EE V + +Q + SE SS + + SP
Sbjct: 720 DRPRVVDEEESNPLVRTKRTSQGTTRYNSEASSSATTTPVANNDSP 765
>UniRef100_Q9XEA1 Hypothetical protein T19B17.6 [Arabidopsis thaliana]
Length = 772
Score = 937 bits (2421), Expect = 0.0
Identities = 453/763 (59%), Positives = 590/763 (76%), Gaps = 14/763 (1%)
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGK 60
MATL DIGVSA INIL+AF F + FA LR+QP NDRVYF KWY+ G R+SP S G F G+
Sbjct: 1 MATLKDIGVSAGINILTAFIFFIIFAFLRLQPFNDRVYFSKWYLRGLRSSPASGGGFAGR 60
Query: 61 FVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLIL 120
FVNL ++YL FL+WMP+AL+M E E+I+HAGLDS V+LRIY LGLKIF P+ ++A +L
Sbjct: 61 FVNLELRSYLKFLHWMPEALKMPERELIDHAGLDSVVYLRIYWLGLKIFAPIAMLAWAVL 120
Query: 121 IPVNVSSGALFFLK--RDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKE 178
+PVN ++ L K +++ SDIDKL+ISN+P S+RF+ HI + Y FT+W C++L KE
Sbjct: 121 VPVNWTNNELELAKHFKNVTSSDIDKLTISNIPEGSNRFWAHIIMAYAFTIWTCYMLMKE 180
Query: 179 YDNIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVY 238
Y+ +A MRL FLAS+ RR +QFTV+VR+VP +VS+ VE FF NHP++Y+ HQ V
Sbjct: 181 YETVANMRLQFLASEGRRPDQFTVLVRNVPPDPDETVSELVEHFFLVNHPDNYLTHQVVC 240
Query: 239 NANKFAKLVRRRDRLQNWLDYYQLKFQRHPDK-RPTINTGVLGLWGRKVDAIEYYSHTIK 297
NANK A LV ++ +LQNWLDYYQLK+ R+ + RP G LGL G+KVDAIE+Y +
Sbjct: 241 NANKLADLVSKKTKLQNWLDYYQLKYTRNNSQIRPITKLGCLGLCGQKVDAIEHYIAEVD 300
Query: 298 ELDKVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDI 357
+ K + ER+ ++ D K ++P +F+SF +RW A+VCAQT Q++NPT WLT+WA EPRDI
Sbjct: 301 KTSKEIAEERENVVNDQKSVMPASFVSFKTRWAAAVCAQTTQTRNPTEWLTEWAAEPRDI 360
Query: 358 YWRNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELK 417
YW NLAIP+VSL++R+LV++++ F L FF++IPIAFVQSLA +EG+E+VAPFL+ +IE
Sbjct: 361 YWPNLAIPYVSLTVRRLVMNVAFFFLTFFFIIPIAFVQSLATIEGIEKVAPFLKVIIEKD 420
Query: 418 FIKSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLG 477
FIKS +QG L G+ALK+FL LPA+LM MSK EG + S LER++A++YY F LVNVFLG
Sbjct: 421 FIKSLIQGLLAGIALKLFLIFLPAILMTMSKFEGFTSVSFLERRSASRYYIFNLVNVFLG 480
Query: 478 SIVTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLV 537
S++ G AFEQL+SFL+QSP QIP+TIG++IPMKATFF+TYIMVDGWAG+AGEIL LKPL+
Sbjct: 481 SVIAGAAFEQLNSFLNQSPNQIPKTIGMAIPMKATFFITYIMVDGWAGVAGEILMLKPLI 540
Query: 538 IYHLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFA 597
IYHLKN F+VKTE+DR +AM+PGS+ F P +QLYFLLG+VYA VTP+LLPFILVFFA
Sbjct: 541 IYHLKNAFLVKTEKDREEAMNPGSIGFNTGEPQIQLYFLLGLVYAPVTPMLLPFILVFFA 600
Query: 598 LAYLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLA 657
LAY+VYRHQIINVYNQ+YESAAAFWP VH R+I +L+ISQLL++GLL TK AA + P L
Sbjct: 601 LAYVVYRHQIINVYNQEYESAAAFWPDVHGRVITALIISQLLLMGLLGTKHAASAAPFLI 660
Query: 658 ILPILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRS 717
LP++T FH++C+ RFEPAF +YPL+EAM KD LE+ EP+LN+K YL DAY+HP+F+
Sbjct: 661 ALPVITIGFHRFCKGRFEPAFVRYPLQEAMMKDTLERAREPNLNLKGYLQDAYIHPVFKG 720
Query: 718 FE----------VEEELVEVRVDKQPETQVASPT-LSEPSSPS 749
+ +E E++ V +Q +P+ +S SSPS
Sbjct: 721 GDNDDDGDMIGKLENEVIIVPTKRQSRRNTPAPSRISGESSPS 763
>UniRef100_Q93ZR7 Hypothetical protein At4g04340 [Arabidopsis thaliana]
Length = 772
Score = 936 bits (2419), Expect = 0.0
Identities = 453/763 (59%), Positives = 590/763 (76%), Gaps = 14/763 (1%)
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGK 60
MATL DIGVSA INIL+AF F + FA LR+QP NDRVYF KWY+ G R+SP S G F G+
Sbjct: 1 MATLKDIGVSAGINILTAFIFFIIFAFLRLQPFNDRVYFSKWYLRGLRSSPASGGGFAGR 60
Query: 61 FVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLIL 120
FVNL ++YL FL+WMP+AL+M E E+I+HAGLDS V+LRIY LGLKIF P+ ++A +L
Sbjct: 61 FVNLELRSYLKFLHWMPEALKMPERELIDHAGLDSVVYLRIYWLGLKIFAPIAMLAWAVL 120
Query: 121 IPVNVSSGALFFLK--RDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKE 178
+PVN ++ L K +++ SDIDKL+ISN+P S+RF+ HI + Y FT+W C++L KE
Sbjct: 121 VPVNWTNNELELAKHFKNVTSSDIDKLTISNIPEGSNRFWAHIIMAYAFTIWTCYMLMKE 180
Query: 179 YDNIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVY 238
Y+ +A MRL FLAS+ RR +QFTV+VR+VP +VS+ VE FF NHP++Y+ HQ V
Sbjct: 181 YETVANMRLQFLASEGRRPDQFTVLVRNVPPDPDETVSELVEHFFLVNHPDNYLTHQVVC 240
Query: 239 NANKFAKLVRRRDRLQNWLDYYQLKFQRHPDK-RPTINTGVLGLWGRKVDAIEYYSHTIK 297
NANK A LV ++ +LQNWLDYYQLK+ R+ + RP G LGL G+KVDAIE+Y +
Sbjct: 241 NANKLADLVSKKTKLQNWLDYYQLKYTRNNSQIRPITKLGCLGLCGQKVDAIEHYIAEVD 300
Query: 298 ELDKVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDI 357
+ K + ER+ ++ D K ++P +F+SF +RW A+VCAQT Q++NPT WLT+WA EPRDI
Sbjct: 301 KTSKEIAEERENVVNDQKSVMPASFVSFKTRWAAAVCAQTTQTRNPTEWLTEWAAEPRDI 360
Query: 358 YWRNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELK 417
YW NLAIP+VSL++R+LV++++ F L FF++IPIAFVQSLA +EG+E+VAPFL+ +IE
Sbjct: 361 YWPNLAIPYVSLTVRRLVMNVAFFFLTFFFIIPIAFVQSLATIEGIEKVAPFLKVIIEKD 420
Query: 418 FIKSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLG 477
FIKS +QG L G+ALK+FL LPA+LM MSK EG + S LER++A++YY F LVNVFLG
Sbjct: 421 FIKSLIQGLLAGIALKLFLIFLPAILMTMSKFEGFTSVSFLERRSASRYYIFNLVNVFLG 480
Query: 478 SIVTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLV 537
S++ G AFEQL+SFL+QSP QIP+TIG++IPMKATFF+TYIMVDGWAG+AGEIL LKPL+
Sbjct: 481 SVIAGAAFEQLNSFLNQSPNQIPKTIGMAIPMKATFFITYIMVDGWAGVAGEILMLKPLI 540
Query: 538 IYHLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFA 597
IYHLKN F+VKTE+DR +AM+PGS+ F P +QLYFLLG+VYA VTP+LLPFILVFFA
Sbjct: 541 IYHLKNAFLVKTEKDREEAMNPGSIGFNTGEPQIQLYFLLGLVYAPVTPMLLPFILVFFA 600
Query: 598 LAYLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLA 657
LAY+VYRHQIINVYNQ+YESAAAFWP VH R+I +L+ISQLL++GLL TK AA + P L
Sbjct: 601 LAYVVYRHQIINVYNQEYESAAAFWPDVHGRVITALIISQLLLMGLLGTKHAASAAPFLI 660
Query: 658 ILPILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRS 717
LP++T FH++C+ RFEPAF +YPL+EAM KD LE+ EP+LN+K YL DAY+HP+F+
Sbjct: 661 ALPVITTGFHRFCKGRFEPAFVRYPLQEAMMKDTLERAREPNLNLKGYLQDAYIHPVFKG 720
Query: 718 FE----------VEEELVEVRVDKQPETQVASPT-LSEPSSPS 749
+ +E E++ V +Q +P+ +S SSPS
Sbjct: 721 GDNDDDGDMIGKLENEVIIVPTKRQSRRNTPAPSRISGESSPS 763
>UniRef100_O65383 F12F1.17 protein [Arabidopsis thaliana]
Length = 783
Score = 927 bits (2395), Expect = 0.0
Identities = 450/779 (57%), Positives = 595/779 (75%), Gaps = 23/779 (2%)
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGK 60
MATL DIGV+AAINIL+A FLLAFA+LRIQP NDRVYFPKWY+ G R+SP +G V K
Sbjct: 1 MATLGDIGVAAAINILTAIIFLLAFAILRIQPFNDRVYFPKWYLKGIRSSPLHSGALVSK 60
Query: 61 FVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLIL 120
FVN+N +YL FLNWMP AL+M E E+I+HAGLDSAV+LRIY +GLKIFVP+ ++A IL
Sbjct: 61 FVNVNLGSYLRFLNWMPAALKMPEPELIDHAGLDSAVYLRIYLIGLKIFVPIALLAWSIL 120
Query: 121 IPVNVSSGALFFLK-RDLVVSDIDKLSISNVPSKSS-----RFFVHIGLEYMFTLWICFL 174
+PVN +S L K R++ SDIDKLSISN+ + S RF+ H+ + Y FT W C++
Sbjct: 121 VPVNWTSHGLQLAKLRNVTSSDIDKLSISNIENGSDSLYFGRFWTHLVMAYAFTFWTCYV 180
Query: 175 LYKEYDNIAMMRLHFLASQRRRVEQFT----------VVVRSVPHISGHSVSDSVESFFQ 224
L KEY+ +A MRL FL +++RR +QFT V+VR+VP S+SDSVE FF
Sbjct: 181 LMKEYEKVAAMRLAFLQNEQRRPDQFTNLGLSQLLSQVLVRNVPADPDESISDSVEHFFL 240
Query: 225 TNHPNHYIGHQAVYNANKFAKLVRRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGR 284
NHP+HY+ HQ VYNAN A LV ++ QNWLDYYQLK+ R+ + +P I TG LGLWG+
Sbjct: 241 VNHPDHYLTHQVVYNANDLAALVEQKKSTQNWLDYYQLKYTRNQEHKPRIKTGFLGLWGK 300
Query: 285 KVDAIEYYSHTIKELDKVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPT 344
KVDAI++Y I E++K+ ER+++ KD ++P AF+SF +RWGA+V AQTQQS +PT
Sbjct: 301 KVDAIDHY---IAEIEKLNEQERKKVKKDDTSVMPAAFVSFKTRWGAAVSAQTQQSSDPT 357
Query: 345 LWLTDWAPEPRDIYWRNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLE 404
WLT+WAPE R+++W NLAIP+VSL++R+L++ ++ F L FF+MIPIAFVQSLA++EG+E
Sbjct: 358 EWLTEWAPEAREVFWSNLAIPYVSLTVRRLIMHIAFFFLTFFFMIPIAFVQSLASIEGIE 417
Query: 405 RVAPFLRPVIELKFIKSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAA 464
+ APFL+ +IE KS +QGFLPG+ LK+FL LP++LM+MSK EG ++ S+LER+ A
Sbjct: 418 KNAPFLKSIIENDLFKSVIQGFLPGIVLKLFLIFLPSILMVMSKFEGFVSLSSLERRAAF 477
Query: 465 KYYYFMLVNVFLGSIVTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWA 524
+YY F L+NVFLGS++TG+AFEQL SFL QS +IP+T+GV+IP+KATFF+TYIMVDGWA
Sbjct: 478 RYYIFNLINVFLGSVITGSAFEQLDSFLKQSAKEIPKTVGVAIPIKATFFITYIMVDGWA 537
Query: 525 GIAGEILRLKPLVIYHLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVV 584
GIAGEILRLKPL+ +H+KN +VKTE+DR +AM+PG +++ T P +QLYFLLG+VYA V
Sbjct: 538 GIAGEILRLKPLIFFHIKNSLLVKTEKDREEAMNPGQINYHATEPRIQLYFLLGLVYAPV 597
Query: 585 TPILLPFILVFFALAYLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLL 644
TP+LLPFI++FFALAYLV+RHQIINVYNQ+YESAA FWP VH RII++L+I+Q+L++GLL
Sbjct: 598 TPVLLPFIIIFFALAYLVFRHQIINVYNQEYESAARFWPDVHGRIISALIIAQILLMGLL 657
Query: 645 STKKAAKSTPLLAILPILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKA 704
STK AA+STP L LPI+TF FH+YC+ R+EPAF ++PL+EAM KD LE+ EP+ N+K
Sbjct: 658 STKGAAQSTPFLLFLPIITFFFHRYCKGRYEPAFLRHPLKEAMVKDTLERAREPNFNLKP 717
Query: 705 YLADAYLHPIFRSFEVE----EELVEVRVDKQPETQVASPTLSEPSSPSPLHDHVHQPS 759
YL AY+HP+F+ + E +E+ ++ E V PT + +P H + S
Sbjct: 718 YLQKAYIHPVFKDNDYEDSRFDEISGYCIEDSDEECVTVPTKRQSRINTPAVSHASRGS 776
>UniRef100_Q8VZM5 Hypothetical protein At4g15430 [Arabidopsis thaliana]
Length = 761
Score = 920 bits (2377), Expect = 0.0
Identities = 443/755 (58%), Positives = 584/755 (76%), Gaps = 2/755 (0%)
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGK 60
MAT+ DIGV+AAINI++AFAFLLAFA+ RIQP+NDRVYFPKWY+ G R+S TG F K
Sbjct: 1 MATINDIGVAAAINIVTAFAFLLAFAIFRIQPVNDRVYFPKWYLKGLRSSSIQTGGFGSK 60
Query: 61 FVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLIL 120
F+NL+F++Y+ FLNWMP+AL+M E E+++HAGLDS V+LRIY LGLKIF P+ VA +
Sbjct: 61 FINLDFRSYIRFLNWMPEALKMPEPELVDHAGLDSVVYLRIYLLGLKIFFPIACVAFTTM 120
Query: 121 IPVNVSSGALFFLKR-DLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEY 179
+PVN ++ L L+ ++ SDIDKLS+SN+P+ S RF+VH+ + Y T W CF+L +EY
Sbjct: 121 VPVNWTNKGLDRLRHSNISFSDIDKLSLSNIPNGSPRFWVHLCMAYAITFWTCFILKREY 180
Query: 180 DNIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVYN 239
NIA+MRL FLA+ +RR QFTV+VR++P S+ + VE FF+ NHP+HY+ QAV++
Sbjct: 181 QNIALMRLQFLANDQRRPNQFTVLVRNIPADPHESICELVEHFFKVNHPDHYLTFQAVHD 240
Query: 240 ANKFAKLVRRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKEL 299
A K ++LV R ++QN LDY K R+ RP I G LG G + D I+YY+ ++ L
Sbjct: 241 ATKLSELVLTRKQMQNLLDYNINKHMRNLSNRPVIKMGFLGCCGEEADGIKYYTSVVEGL 300
Query: 300 DKVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYW 359
+ ++ E+QR+ K I+P AF+SF SRWGA+VCAQTQQ++NPT WLT+WA EPRDIY+
Sbjct: 301 TREISEEKQRLRTGTKSIVPAAFVSFKSRWGAAVCAQTQQTRNPTEWLTEWAAEPRDIYY 360
Query: 360 RNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFI 419
NLA+P+V L IR+L++ ++ F L FF+MIPIAFVQSLAN+EG+E+ PFL+P+IE+K +
Sbjct: 361 DNLALPYVDLKIRRLIVGVAYFFLTFFFMIPIAFVQSLANIEGIEKAFPFLKPLIEVKLL 420
Query: 420 KSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSI 479
KS +QGFLPG+ALKIFL LP +LM MSK EG ++ S+LER+ A ++Y F +NVFLGSI
Sbjct: 421 KSIIQGFLPGIALKIFLLFLPRILMQMSKFEGFVSTSSLERRAATRFYMFQFINVFLGSI 480
Query: 480 VTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIY 539
VTGTAF+QL+SFL+QS IP+TIGVSIPMKATFF+TYIMVDGWAG+AGEILRLKPL+IY
Sbjct: 481 VTGTAFQQLNSFLNQSANDIPKTIGVSIPMKATFFITYIMVDGWAGVAGEILRLKPLIIY 540
Query: 540 HLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALA 599
HLKN F+V+TE+DR +A DPG++ F P +QLYFLLG+VYA V+PILLPFILVFF LA
Sbjct: 541 HLKNSFLVRTEKDREEATDPGTIGFNTGEPQIQLYFLLGLVYAAVSPILLPFILVFFGLA 600
Query: 600 YLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAIL 659
++VYRHQ+INVYNQ+YESA FWP VH R++ +L++SQLL++GLLSTK A+KSTPLL +L
Sbjct: 601 FVVYRHQVINVYNQKYESAGKFWPDVHRRVVTALVVSQLLLMGLLSTKHASKSTPLLLVL 660
Query: 660 PILTFAFHKYCQHRFEPAFRKYPL-EEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRSF 718
P+LT FHK+C++R++PAF YPL +EAM KD L++ EP+LN+KA+L DAY HP FR
Sbjct: 661 PLLTIGFHKHCKNRYQPAFVTYPLQQEAMIKDTLDRIREPNLNLKAFLRDAYAHPEFRVG 720
Query: 719 EVEEELVEVRVDKQPETQVASPTLSEPSSPSPLHD 753
E E ++ D P VA+ S ++P P D
Sbjct: 721 EDPEPEEKLESDMSPPDLVATKRWSWRNTPLPSKD 755
>UniRef100_Q9ZT88 Hypothetical protein T4I9.22 [Arabidopsis thaliana]
Length = 680
Score = 793 bits (2049), Expect = 0.0
Identities = 389/661 (58%), Positives = 503/661 (75%), Gaps = 7/661 (1%)
Query: 106 LKIFVPVTVVALLILIPVNVSSGALFFLKRDLVVSDIDKLSISNVPSKSSRFFVHIGLEY 165
LK+FVP+T++A +L+PVN + L + DL S++DKLSISNVP S RF+ HI + Y
Sbjct: 1 LKMFVPITLLAFGVLVPVNWTGETLENID-DLTFSNVDKLSISNVPPGSPRFWAHITMTY 59
Query: 166 MFTLWICFLLYKEYDNIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQT 225
+ T W C++LY EY +A MRL LA++ RR +Q TV+VR+VP SV++ VE FF
Sbjct: 60 VITFWTCYILYMEYKAVANMRLRHLAAESRRPDQLTVLVRNVPPDPDESVNEHVEHFFCV 119
Query: 226 NHPNHYIGHQAVYNANKFAKLVRRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRK 285
NHP+HY+ HQ VYNAN AKLV +R +QNWL YY+ KF+R P RPT TG G WG
Sbjct: 120 NHPDHYLCHQVVYNANDLAKLVAQRKAMQNWLTYYENKFERKPSSRPTTKTGYGGFWGTT 179
Query: 286 VDAIEYYSHTIKELDKVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTL 345
VDAI++Y+ + L + +ER++I+ DPK I+P AF+SF SRWG +VCAQTQQ NPT+
Sbjct: 180 VDAIDFYTSKMDILAEQEAVEREKIMNDPKAIMPAAFVSFRSRWGTAVCAQTQQCHNPTI 239
Query: 346 WLTDWAPEPRDIYWRNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLER 405
WLT+WAPEPRD++W NLAIP+V LSIR+L+ ++++F L+F +MIPIAFVQSLANLEG+++
Sbjct: 240 WLTEWAPEPRDVFWDNLAIPYVELSIRRLLTTVALFFLIFCFMIPIAFVQSLANLEGIQK 299
Query: 406 VAPFLRPVIELKFIKSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAK 465
V PFL+PVIE+K +KS +QGFLPG+ALKIFL ILP +LM MS+IEG+ + S L+R++A K
Sbjct: 300 VLPFLKPVIEMKTVKSVIQGFLPGIALKIFLIILPTILMTMSQIEGYTSLSYLDRRSAEK 359
Query: 466 YYYFMLVNVFLGSIVTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAG 525
Y++F++VNVFLGSI+TGTAF+QL SFL Q PT+IP+T+GVSIPMKATFF+TYIMVDGWAG
Sbjct: 360 YFWFIIVNVFLGSIITGTAFQQLKSFLEQPPTEIPKTVGVSIPMKATFFITYIMVDGWAG 419
Query: 526 IAGEILRLKPLVIYHLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVT 585
IA EILR+ PLVI+HLKN F+VKTE+DR +AMDPG +DF + P +Q YFLLG+VYA V
Sbjct: 420 IAAEILRVVPLVIFHLKNTFLVKTEQDRQQAMDPGHLDFATSEPRIQFYFLLGLVYAAVA 479
Query: 586 PILLPFILVFFALAYLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLS 645
PILLPFI+VFFA AY+V+RHQ+INVY+Q+YES A +WP VH R+I L+ISQLLM+GLLS
Sbjct: 480 PILLPFIIVFFAFAYVVFRHQVINVYDQKYESGARYWPDVHRRLIICLIISQLLMMGLLS 539
Query: 646 TKKAAKSTPLLAILPILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAY 705
TKK AK T LL PILTF F++YC RFE AF K+PL+EAM KD LEK TEP+LN+K Y
Sbjct: 540 TKKFAKVTALLLPQPILTFWFYRYCAGRFESAFSKFPLQEAMVKDTLEKATEPNLNLKEY 599
Query: 706 LADAYLHPIFRSFE------VEEELVEVRVDKQPETQVASPTLSEPSSPSPLHDHVHQPS 759
L DAY+HP+F+ + V+EE V + +Q + SE SS + + S
Sbjct: 600 LKDAYVHPVFKGNDFDRPRVVDEEESNPLVRTKRTSQGTTRYNSEASSSATTTPVANNDS 659
Query: 760 P 760
P
Sbjct: 660 P 660
>UniRef100_O23396 Hypothetical protein dl3760w [Arabidopsis thaliana]
Length = 680
Score = 745 bits (1924), Expect = 0.0
Identities = 389/762 (51%), Positives = 512/762 (67%), Gaps = 97/762 (12%)
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGK 60
MAT+ DIGV+AAINI++AFAFLLAFA+ RIQP+NDRVYFPKWY+ G R+S TG F K
Sbjct: 1 MATINDIGVAAAINIVTAFAFLLAFAIFRIQPVNDRVYFPKWYLKGLRSSSIQTGGFGSK 60
Query: 61 FVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLIL 120
F+NL+F++Y+ FLNWMP+AL+M E E+++HAGLDS V+LRIY LGLKIF P+ VA +
Sbjct: 61 FINLDFRSYIRFLNWMPEALKMPEPELVDHAGLDSVVYLRIYLLGLKIFFPIACVAFTTM 120
Query: 121 IPVNVSSGALFFLKR-DLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEY 179
+PVN ++ L L+ ++ SDIDKLS+SN+P+ S R H +F I +
Sbjct: 121 VPVNWTNKGLDRLRHSNISFSDIDKLSLSNIPNGSPRSLSHFPGIIIFGFSIVY------ 174
Query: 180 DNIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVYN 239
+SD + + + QAV++
Sbjct: 175 -----------------------------------ISDLISELMMVD----LLLTQAVHD 195
Query: 240 ANKFAKLVRRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKEL 299
A K ++LV R ++QN LDY K R+ RP I
Sbjct: 196 ATKLSELVLTRKQMQNLLDYNINKHMRNLSNRPVIK------------------------ 231
Query: 300 DKVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYW 359
++ E+QR+ K I+P AF+SF SRWGA+VCAQTQQ++NPT WLT+WA EPRDIY+
Sbjct: 232 ---ISEEKQRLRTGTKSIVPAAFVSFKSRWGAAVCAQTQQTRNPTEWLTEWAAEPRDIYY 288
Query: 360 RNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIE---- 415
NLA+P+V L IR+L++ ++ F L FF+MIPIAFVQSLAN+EG+E+ PFL+P+IE
Sbjct: 289 DNLALPYVDLKIRRLIVGVAYFFLTFFFMIPIAFVQSLANIEGIEKAFPFLKPLIEVYVS 348
Query: 416 ----LKFIKSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFML 471
+K +KS +QGFLPG+ALKIFL LP +LM MSK EG ++ S+LER+ A ++Y F
Sbjct: 349 VYLNMKLLKSIIQGFLPGIALKIFLLFLPRILMQMSKFEGFVSTSSLERRAATRFYMFQF 408
Query: 472 VNVFLGSIVTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEIL 531
+NVFLGSIVTGTAF+QL+SFL+QS IP+TIGVSIPMKATFF+TYIMVDGWAG+AGEIL
Sbjct: 409 INVFLGSIVTGTAFQQLNSFLNQSANDIPKTIGVSIPMKATFFITYIMVDGWAGVAGEIL 468
Query: 532 RLKPLVIYHLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPF 591
RLKPL+IYHLKN F+V+TE+DR +A DPG++ F V+PILLPF
Sbjct: 469 RLKPLIIYHLKNSFLVRTEKDREEATDPGTIGF----------------NTAVSPILLPF 512
Query: 592 ILVFFALAYLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAK 651
ILVFF LA++VYRHQ+INVYNQ+YESA FWP VH R++ +L++SQLL++GLLSTK A+K
Sbjct: 513 ILVFFGLAFVVYRHQVINVYNQKYESAGKFWPDVHRRVVTALVVSQLLLMGLLSTKHASK 572
Query: 652 STPLLAILPILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYL 711
STPLL +LP+LT FHK+C++R++PAF YPL+EAM KD L++ EP+LN+KA+L DAY
Sbjct: 573 STPLLLVLPLLTIGFHKHCKNRYQPAFVTYPLQEAMIKDTLDRIREPNLNLKAFLRDAYA 632
Query: 712 HPIFRSFEVEEELVEVRVDKQPETQVASPTLSEPSSPSPLHD 753
HP FR E E ++ D P VA+ S ++P P D
Sbjct: 633 HPEFRVGEDPEPEEKLESDMSPPDLVATKRWSWRNTPLPSKD 674
>UniRef100_O65460 Hypothetical protein AT4g22120 [Arabidopsis thaliana]
Length = 697
Score = 620 bits (1598), Expect = e-176
Identities = 298/502 (59%), Positives = 392/502 (77%), Gaps = 13/502 (2%)
Query: 261 QLKFQRHPDKRPTINT-GVLGLWGRKVDAIEYYSHTIKELDKVMTLERQRIIKDPKCILP 319
+L+F +RP T G LGLWG+KVDAIE+Y I ++ K ++ ER+ ++ DPK I+P
Sbjct: 188 RLQFVASEARRPDQFTLGFLGLWGQKVDAIEHYIAEIDKISKEISKEREEVVNDPKAIMP 247
Query: 320 VAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYWRNLAIPFVSLSIRKLVISLS 379
AF+SF +RW A+VCAQTQQ++NPT WLT+WAPEPRD++W NLAIP+VSL++R+L++ ++
Sbjct: 248 AAFVSFKTRWAAAVCAQTQQTRNPTQWLTEWAPEPRDVFWSNLAIPYVSLTVRRLIMHVA 307
Query: 380 VFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFIKSFLQGFLPGLALKIFLYIL 439
F L FF+++PIAFVQSLA +EG+ + APFL+ +++ KF+KS +QGFLPG+ALK+FL L
Sbjct: 308 FFFLTFFFIVPIAFVQSLATIEGIVKAAPFLKFIVDDKFMKSVIQGFLPGIALKLFLAFL 367
Query: 440 PAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSIVTGTAFEQLHSFLHQSPTQI 499
P++LMIMSK EG + S+LER+ A +YY F LVNVFL S++ G AFEQL+SFL+QS QI
Sbjct: 368 PSILMIMSKFEGFTSISSLERRAAFRYYIFNLVNVFLASVIAGAAFEQLNSFLNQSANQI 427
Query: 500 PRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIYHLKNMFIVKTERDRGKAMDP 559
P+TIGV+IPMKATFF+TYIMVDGWAG+AGEIL LKPL+++HLKN F+VKT++DR +AMDP
Sbjct: 428 PKTIGVAIPMKATFFITYIMVDGWAGVAGEILMLKPLIMFHLKNAFLVKTDKDREEAMDP 487
Query: 560 GSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALAYLVYRHQIINVYNQQYESAA 619
GS+ F P +QLYFLLG+VYA VTP+LLPFILVFFALAY+VYRHQIINVYNQ+YESAA
Sbjct: 488 GSIGFNTGEPRIQLYFLLGLVYAPVTPMLLPFILVFFALAYIVYRHQIINVYNQEYESAA 547
Query: 620 AFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAILPILTFAFHKYCQHRFEPAFR 679
AFWP VH R+IA+L+ISQLL++GLL TK AA + P L LP+LT FH +C+ R+EPAF
Sbjct: 548 AFWPDVHGRVIAALVISQLLLMGLLGTKHAALAAPFLIALPVLTIGFHHFCKGRYEPAFI 607
Query: 680 KYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRSFE-----------VEEELVEVR 728
+YPL+EAM KD LE EP+LN+K YL +AY+HP+F+ E E+E + V
Sbjct: 608 RYPLQEAMMKDTLETAREPNLNLKGYLQNAYVHPVFKGDEDDYDIDDKLGKFEDEAIIVP 667
Query: 729 VDKQPETQVASPT-LSEPSSPS 749
+Q +P+ +S SPS
Sbjct: 668 TKRQSRRNTPAPSIISGDDSPS 689
Score = 251 bits (641), Expect = 6e-65
Identities = 121/204 (59%), Positives = 156/204 (76%), Gaps = 2/204 (0%)
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGK 60
MATL DIGVSA INILSAF F + FA+LR+QP NDRVYF KWY+ G R+SP G F +
Sbjct: 1 MATLQDIGVSAGINILSAFVFFIIFAVLRLQPFNDRVYFSKWYLKGLRSSPARGGAFAQR 60
Query: 61 FVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLIL 120
FVNL+F++Y+ FLNWMP+AL+M E E+I+HAGLDS V+LRIY LGLKIF P+ V+A +L
Sbjct: 61 FVNLDFRSYMKFLNWMPEALKMPEPELIDHAGLDSVVYLRIYWLGLKIFTPIAVLAWAVL 120
Query: 121 IPVNVSSGALFFLK--RDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKE 178
+PVN ++ L K R++ SDIDKLS+SN+P S RF+ HI + Y FT+W C++L KE
Sbjct: 121 VPVNWTNNTLEMAKQLRNVTSSDIDKLSVSNIPEYSMRFWTHIVMAYAFTIWTCYVLMKE 180
Query: 179 YDNIAMMRLHFLASQRRRVEQFTV 202
Y+ IA MRL F+AS+ RR +QFT+
Sbjct: 181 YETIANMRLQFVASEARRPDQFTL 204
>UniRef100_Q8GUH7 Hypothetical protein At3g01100At3g01110 [Arabidopsis thaliana]
Length = 703
Score = 435 bits (1118), Expect = e-120
Identities = 234/710 (32%), Positives = 402/710 (55%), Gaps = 22/710 (3%)
Query: 4 LADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGKFVN 63
L+ + S IN+ F F +++LR QP N VY P+ + + S +S
Sbjct: 3 LSALLTSVGINLGLCFLFFTLYSILRKQPSNVTVYGPR-LVKKDGKSQQSN--------E 53
Query: 64 LNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLILIPV 123
N + L W+ +AL + +EI+++ GLD+ VF+R++ +++F +VV + IL+PV
Sbjct: 54 FNLERLLPTAGWVKRALEPTNDEILSNLGLDALVFIRVFVFSIRVFSFASVVGIFILLPV 113
Query: 124 NVSSGALFFLKRDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEYDNIA 183
N G F DL +D SISNV S++ ++H Y+FT +C LLY E+ I
Sbjct: 114 NYM-GTEFEEFFDLPKKSMDNFSISNVNDGSNKLWIHFCAIYIFTAVVCSLLYYEHKYIL 172
Query: 184 MMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVYNANKF 243
R+ L S + + ++FTV+V VP +SG+S+S++VE+FF+ H + Y+ H V+ +K
Sbjct: 173 TKRIAHLYSSKPQPQEFTVLVSGVPLVSGNSISETVENFFREYHSSSYLSHIVVHRTDKL 232
Query: 244 AKLVRRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKELDKVM 303
L+ ++L L + ++ + G LG++G VD +++Y + +L+ M
Sbjct: 233 KVLMNDAEKLYKKLTRVK---SGSISRQKSRWGGFLGMFGNNVDVVDHYQKKLDKLEDDM 289
Query: 304 TLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYWRNLA 363
L++ + + +P AF+SF +R GA++ QQ +PT WLT+ APEP D++W
Sbjct: 290 RLKQSLLAGEE---VPAAFVSFRTRHGAAIATNIQQGIDPTQWLTEAAPEPEDVHWPFFT 346
Query: 364 IPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFIKSFL 423
FV I +V+ ++ AL+ Y++P+ VQ LANL LE PFL+ ++ +K + +
Sbjct: 347 ASFVRRWISNVVVLVAFVALLILYIVPVVLVQGLANLHQLETWFPFLKGILNMKIVSQVI 406
Query: 424 QGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSIVTGT 483
G+LP L ++FL I+P +++++S ++G I+ S +E+ K F + N F ++++G+
Sbjct: 407 TGYLPSLIFQLFLLIVPPIMLLLSSMQGFISHSQIEKSACIKLLIFTVWNSFFANVLSGS 466
Query: 484 AFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIYHLKN 543
A +++ FL P IPR + ++P +A+FF++Y++ GW G++ EILRL PL+ +
Sbjct: 467 ALYRVNVFL--EPKTIPRVLAAAVPAQASFFVSYVVTSGWTGLSSEILRLVPLLWSFITK 524
Query: 544 MFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALAYLVY 603
+F ++ K + S F + IP + + LLGI Y ++P++LPF+LV++ L Y++Y
Sbjct: 525 LF----GKEDDKEFEVPSTPFCQEIPRILFFGLLGITYFFLSPLILPFLLVYYCLGYIIY 580
Query: 604 RHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAILPILT 663
R+Q++NVY +YE+ FWP VHS I SL++ ++ +GL K+ ++ L LP+LT
Sbjct: 581 RNQLLNVYAAKYETGGKFWPIVHSYTIFSLVLMHIIAVGLFGLKELPVASSLTIPLPVLT 640
Query: 664 FAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHP 713
F YCQ RF P F+ YP + + KD ++ + + L AY P
Sbjct: 641 VLFSIYCQRRFLPNFKSYPTQCLVNKDKADEREQNMSEFYSELVVAYRDP 690
>UniRef100_Q9C792 Hypothetical protein F10D13_27 [Arabidopsis thaliana]
Length = 646
Score = 381 bits (978), Expect = e-104
Identities = 199/624 (31%), Positives = 344/624 (54%), Gaps = 9/624 (1%)
Query: 91 AGLDSAVFLRIYTLGLKIFVPVTVVALLILIPVNVSSGALFFLK-RDLVVSDIDKLSISN 149
+GLD VF+R+ T LK+F+ ++ + +L+PVN L + D + +D S++N
Sbjct: 4 SGLDGVVFMRMITFSLKVFLFAGIIGVFVLLPVNCFGDQLTVIDYADWSANSLDLFSVAN 63
Query: 150 VPSKSSRFFVHIGLEYMFTLWICFLLYKEYDNIAMMRLHFLASQRRRVEQFTVVVRSVPH 209
+ +S +VH G Y+ T+++C LLY E+ IA+ R+ S + + EQFT++VR++P
Sbjct: 64 LKVRSQWLWVHFGAIYLVTVFVCCLLYFEFRYIALKRIEHFYSSKPKPEQFTILVRNIPS 123
Query: 210 ISGHSVSDSVESFFQTNHPNHYIGHQAVYNANKFAKLVRRRDRLQNWLDYYQLKFQRHPD 269
G SVSD+V+ FF NH + Y H ++ +K +V ++ D + ++
Sbjct: 124 SDGSSVSDTVDRFFGENHSSTYFSHVVIHRTSKLRSVVVSLEKAYLPSDKAKKLYKEVKH 183
Query: 270 KRPTINTGVLGLWGRKVDAIEYYSHTIKELDKVMTLERQRIIKDPKCILPVAFLSFSSRW 329
K+P T + + RK + +Y ++E+++ + L Q + P + AF+SF SR+
Sbjct: 184 KKPVKKTP-MRFFSRKDNTEGHYESVLQEMEQNIRLG-QAEVSAPGKEVRAAFVSFKSRY 241
Query: 330 GASVCAQTQQSKNPTLWLTDWAPEPRDIYWRNLAIPFVSLSIRKLVISLSVFALVFFYMI 389
GA+ QS NPT WLT+ APEP D++W + F+ + K+++ + L +++
Sbjct: 242 GAATALHMPQSINPTYWLTEPAPEPHDVHWPFFSASFMQKWLAKILVVFACLLLTILFLV 301
Query: 390 PIAFVQSLANLEGLERVAPFLRPVIELKFIKSFLQGFLPGLALKIFLYILPAVLMIMSKI 449
P+ VQ L NL LE + PFL ++ +K + + G+LP L L+ L ++P + +S I
Sbjct: 302 PVVLVQGLTNLPALEFMFPFLSLILSMKVVSQIITGYLPSLILQTSLKVVPPTMEFLSSI 361
Query: 450 EGHIAKSTLERKTAAKYYYFMLVNVFLGSIVTGTAFEQLHSFLHQSPTQIPRTIGVSIPM 509
+GHI S +++ K +F + NVF ++ +G+AF +L L P QIP + V++P
Sbjct: 362 QGHICHSDIQKSACNKVIWFTIWNVFFATVFSGSAFYKLSVIL--DPKQIPLKLAVAVPA 419
Query: 510 KATFFMTYIMVDGWAGIAGEILRLKPLVIYHLKNMFIVKTERDRGKAMDPGSVDFPETIP 569
+A+FF+ Y++ GW E+ R+ P ++ ++K F D + + P + + P
Sbjct: 420 QASFFIAYVVTTGWTDTLTELFRVVPFMVSYIKRSF---EPSDENEFVVP-PMRYHRDTP 475
Query: 570 SLQLYFLLGIVYAVVTPILLPFILVFFALAYLVYRHQIINVYNQQYESAAAFWPHVHSRI 629
+ + LLGI Y + P++LPFIL++F LAY++YR+Q +NVY ++++ FWP +H +
Sbjct: 476 RVLFFGLLGITYFFLAPLILPFILLYFILAYIIYRNQFMNVYAPKFDTGGMFWPMIHYTM 535
Query: 630 IASLLISQLLMLGLLSTKKAAKSTPLLAILPILTFAFHKYCQHRFEPAFRKYPLEEAMAK 689
I SL++ Q + +GL + KK +T LL LP+ T F+++C+ RF P F YP E +
Sbjct: 536 IFSLVLMQAIAIGLFALKKMELATYLLVPLPVFTLLFNEFCRKRFMPIFTDYPAEVLTKR 595
Query: 690 DLLEKTTEPDLNIKAYLADAYLHP 713
D ++ L AY P
Sbjct: 596 DKEDRNDPTMPEFYNNLVSAYKDP 619
>UniRef100_Q75GI5 Expressed protein [Oryza sativa]
Length = 743
Score = 380 bits (977), Expect = e-104
Identities = 220/702 (31%), Positives = 372/702 (52%), Gaps = 24/702 (3%)
Query: 10 SAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGKFVNLNFKTY 69
SA INI FL +++LR QP N VYF G R + V F + +
Sbjct: 9 SAGINIGLCALFLSLYSVLRKQPHNYGVYF------GRRLAEEKFRQQVDYF---SLERL 59
Query: 70 LTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLILIPVNVSSGA 129
L W+ +A +E EI AGLDS VFLR++ ++IF ++V + ++PVN
Sbjct: 60 LPTAGWIVKAYWCTEEEIRRVAGLDSVVFLRLFIFSIRIFSITSLVCIFGVLPVNYHGKE 119
Query: 130 LFFLKRDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEYDNIAMMRLHF 189
+ + ++ +I+N+ S +VH Y+ T+ C LLY EY I+ RL
Sbjct: 120 TNHGR--IPAESLNVFTIANLKEGSRMLWVHCVALYVITISACILLYYEYKYISRKRLAH 177
Query: 190 LASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVYNANKFAKLVRR 249
+ F+V+VRS+P + D++ +FF H + Y+ HQ +Y K V
Sbjct: 178 ITGSPPGPGHFSVIVRSIPKSDNELLDDTIRNFFVNYHGSSYLSHQMIYRKGSMQKFVDN 237
Query: 250 RDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKELDKVMTLERQR 309
+R+ + + ++K R + + GL G + + + Y + K + +
Sbjct: 238 AERV--YRKFVRVKMSSFGQSRRS-DLSRCGLCGVRASSFQQYRNKFINSKKPDLSDPEV 294
Query: 310 IIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYWRNLAIPFVSL 369
I C P A + F +R+ A V ++ QS NP LW+TD+APEPRD+YW NL IP+ +
Sbjct: 295 IEAQKDC--PGAIVFFKTRYAAIVASRILQSSNPMLWVTDFAPEPRDVYWSNLWIPYRQI 352
Query: 370 SIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFIKSFLQGFLPG 429
+RK+ + A +F +++P+AFVQS+ L+ +E++ P L+ +++ F + G+LP
Sbjct: 353 WLRKIATLAASVAFMFVFIVPVAFVQSMMQLDQIEQLFPSLKNMLKKPFFVKLVTGYLPS 412
Query: 430 LALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSIVTGTAFEQLH 489
+ L + LY +P ++M S IEG I++S ++ K +F + NVF ++++G+ QL+
Sbjct: 413 VVLLLSLYTVPPLMMFFSSIEGSISRSGRKKSACCKILFFTIWNVFFVNVLSGSVLNQLN 472
Query: 490 SFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVI-YHLKNMFIVK 548
F P +P + +P +ATFF+TY++ GWA + EIL++ LV + K +F +
Sbjct: 473 VFTR--PRDMPSMLAELVPKQATFFITYVLTSGWASLCSEILQVYNLVYNFFRKCIFCYR 530
Query: 549 TERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALAYLVYRHQII 608
+ + G S + +P + L+ LLG ++++ P++LPF+LV+F L YLVYR+QI+
Sbjct: 531 DDPEYGY-----SFPYHTEVPKVLLFNLLGFTFSIMAPLILPFLLVYFCLGYLVYRNQIL 585
Query: 609 NVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAILPILTFAFHK 668
NVY +YE WP +HS ++ +L+++Q + LG+ + K A S+ +L I T FH+
Sbjct: 586 NVYYPKYEMGGKLWPIMHSTLVFALVLTQTIALGVFTIKHATISSGFTVLLIIGTVLFHQ 645
Query: 669 YCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAY 710
YC+HRF F + ++ + D ++ + I +L DAY
Sbjct: 646 YCRHRFSSIFNSFSAQDLIEMDRDDEQSGRMEEIHKHLLDAY 687
>UniRef100_Q8LC14 Hypothetical protein [Arabidopsis thaliana]
Length = 762
Score = 370 bits (949), Expect = e-100
Identities = 222/705 (31%), Positives = 368/705 (51%), Gaps = 33/705 (4%)
Query: 10 SAAINILSAFAFLLAFALLRIQPINDRVYFPKWYING--ERTSPRSTGNFVGKFVNLNFK 67
SA INI + +++LR QP N VYF + +G +R PR ++
Sbjct: 9 SAGINIAICVVLVSLYSILRKQPANYCVYFGRLLSDGRVKRHDPRW------------YE 56
Query: 68 TYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLILIPVNVSS 127
+ +W+ +A +E E++ AGLD+ VF+R+ ++IF V VV L ++PVN
Sbjct: 57 RFAPSPSWLVKAWETTEEEMLAAAGLDAVVFIRMVICIIRIFSIVAVVCLAFVLPVNYYG 116
Query: 128 GALFFLKRDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEYDNIAMMRL 187
+ +++ + + +I N+ +S +VH Y+ + C LLY EY NIA RL
Sbjct: 117 QKMEH--KEVHLESLGVFTIENLNPRSRWLWVHCLSLYIISSAACALLYFEYKNIAKKRL 174
Query: 188 HFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVYNANKFAKLV 247
++ + FTV++R++P S S++V +F + Y+ H VY +L+
Sbjct: 175 AHISGSASKPSHFTVLIRAIPQSPDQSYSETVSKYFTNYYAPSYVSHLMVYRDGFIHRLM 234
Query: 248 RRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKELDKVM--TL 305
+R+ + + +P + + G ++ + E S EL ++ T
Sbjct: 235 NETERMCQAIKHVSPDLSCNPSLKSCVLCGPAATNSFQIISNETDSVKGLELGELTLTTT 294
Query: 306 ERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYWRNLAIP 365
E +R PVAF+ F SR+ A V ++ Q+ NP LW+ D APEP D++WRNL IP
Sbjct: 295 EEER---------PVAFVFFKSRYDALVVSEVLQTPNPMLWVADLAPEPHDVHWRNLRIP 345
Query: 366 FVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFIKSFLQG 425
+ L +R++ + A +F ++ P+ FVQ L L L + PFL+ ++ +F++ + G
Sbjct: 346 YRQLWMRRIATLVGAIAFMFVFLFPVTFVQGLTQLPTLSKNFPFLKDLLNRRFMEQVITG 405
Query: 426 FLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSIVTGTAF 485
+LP + L +F Y +P ++M S +EG +++S ++ K YF + NVF +I++G+
Sbjct: 406 YLPSVILVLFFYTVPPLMMYFSTLEGCVSRSQKKKSACLKILYFTIWNVFFVNILSGSVI 465
Query: 486 EQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIYHLKNMF 545
Q F S +P + +P +A FFMTY GWAG+A EI++ L I++L
Sbjct: 466 RQFTVF--NSVRDVPAQLAKLVPAQAGFFMTYCFTSGWAGLACEIMQPVGL-IWNLIAKV 522
Query: 546 IVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALAYLVYRH 605
IVK + + + + + IP L L+ LLG +V+ P++LPF+L++F AYL+Y++
Sbjct: 523 IVKNKEESYETL---RFPYHTEIPRLLLFGLLGFTNSVIAPLILPFLLIYFFFAYLIYKN 579
Query: 606 QIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAILPILTFA 665
QIINVY +YES +WP H+ I SL++SQ++ LG K + ++ L +LT
Sbjct: 580 QIINVYITKYESGGQYWPVFHNTTIFSLILSQVIALGFFGLKLSTVASGFTIPLILLTLL 639
Query: 666 FHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAY 710
F +YC+ RF P F+KYP E +A D ++ T I L AY
Sbjct: 640 FSEYCRQRFAPIFQKYPAEILIAMDRADEMTGKMEEIHNNLKVAY 684
>UniRef100_Q94A87 At1g10080/T27I1_10 [Arabidopsis thaliana]
Length = 762
Score = 369 bits (947), Expect = e-100
Identities = 221/705 (31%), Positives = 368/705 (51%), Gaps = 33/705 (4%)
Query: 10 SAAINILSAFAFLLAFALLRIQPINDRVYFPKWYING--ERTSPRSTGNFVGKFVNLNFK 67
SA INI + +++LR QP N VYF + +G +R PR ++
Sbjct: 9 SAGINIAICVVLVSLYSILRKQPANYCVYFGRLLSDGRVKRHDPRW------------YE 56
Query: 68 TYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLILIPVNVSS 127
+ +W+ +A +E E++ AGLD+ VF+R+ ++IF V VV L ++PVN
Sbjct: 57 RFAPSPSWLVKAWETTEEEMLAAAGLDAVVFIRMVICSIRIFSIVAVVCLAFVLPVNYYG 116
Query: 128 GALFFLKRDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEYDNIAMMRL 187
+ +++ + + +I N+ +S +VH Y+ + C LLY EY NIA RL
Sbjct: 117 QKMEH--KEVHLESLGVFTIENLNPRSRWLWVHCLSLYIISSAACALLYFEYKNIAKKRL 174
Query: 188 HFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVYNANKFAKLV 247
++ + FTV++R++P S S++V +F + Y+ H VY +L+
Sbjct: 175 AHISGSASKPSHFTVLIRAIPQSPDQSYSETVSKYFTNYYAPSYVSHLMVYRDGFIHRLM 234
Query: 248 RRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKELDKVM--TL 305
+R+ + + +P + + G ++ + E S EL ++ T
Sbjct: 235 NETERMCQAIKHVSPDLSCNPSLKSCVLCGPAATNSFQIISNETDSVKGLELGELTLTTT 294
Query: 306 ERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYWRNLAIP 365
E +R PVAF+ F SR+ A V ++ Q+ NP LW+ D APEP D++WRNL IP
Sbjct: 295 EEER---------PVAFVFFKSRYDALVVSEVLQTPNPMLWVADLAPEPHDVHWRNLRIP 345
Query: 366 FVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFIKSFLQG 425
+ L +R++ + A +F ++ P+ FVQ L L L + PFL+ ++ +F++ + G
Sbjct: 346 YRQLWMRRIATLVGAIAFMFVFLFPVTFVQGLTQLPTLSKNFPFLKDLLNRRFMEQVITG 405
Query: 426 FLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSIVTGTAF 485
+LP + L +F Y +P ++M S +EG +++S ++ K YF + NVF +I++G+
Sbjct: 406 YLPSVILVLFFYTVPPLMMYFSTLEGCVSRSQRKKSACLKILYFTIWNVFFVNILSGSVI 465
Query: 486 EQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIYHLKNMF 545
Q + S +P + +P +A FFMTY GWAG+A EI++ L I++L
Sbjct: 466 RQF--TVLNSVRDVPAQLAKLVPAQAGFFMTYCFTSGWAGLACEIMQPVGL-IWNLIAKV 522
Query: 546 IVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALAYLVYRH 605
IVK + + + + + IP L L+ LLG +V+ P++LPF+L++F AYL+Y++
Sbjct: 523 IVKNKEESYETL---RFPYHTEIPRLLLFGLLGFTNSVIAPLILPFLLIYFFFAYLIYKN 579
Query: 606 QIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAILPILTFA 665
QIINVY +YES +WP H+ I SL++SQ++ LG K + ++ L +LT
Sbjct: 580 QIINVYITKYESGGQYWPVFHNTTIFSLILSQVIALGFFGLKLSTVASGFTIPLILLTLL 639
Query: 666 FHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAY 710
F +YC+ RF P F+KYP E +A D ++ T I L AY
Sbjct: 640 FSEYCRQRFAPIFQKYPAEILIAMDRADEMTGKMEEIHNNLKVAY 684
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.326 0.140 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,320,315,278
Number of Sequences: 2790947
Number of extensions: 57236407
Number of successful extensions: 281016
Number of sequences better than 10.0: 598
Number of HSP's better than 10.0 without gapping: 274
Number of HSP's successfully gapped in prelim test: 353
Number of HSP's that attempted gapping in prelim test: 270829
Number of HSP's gapped (non-prelim): 3974
length of query: 795
length of database: 848,049,833
effective HSP length: 136
effective length of query: 659
effective length of database: 468,481,041
effective search space: 308729006019
effective search space used: 308729006019
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 79 (35.0 bits)
Lotus: description of TM0020.6


