

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0019a.4
(310 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
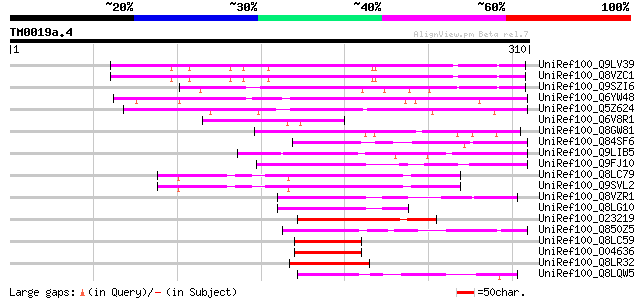
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LV39 Arabidopsis thaliana genomic DNA, chromosome 5,... 138 2e-31
UniRef100_Q8VZC1 Hypothetical protein At5g56860; MPI10.2 [Arabid... 138 2e-31
UniRef100_Q9SZI6 Putative transcription factor [Arabidopsis thal... 130 4e-29
UniRef100_Q6YW48 Zinc finger protein-like [Oryza sativa] 125 1e-27
UniRef100_Q5Z624 Zinc finger protein-like [Oryza sativa] 106 9e-22
UniRef100_Q6V8R1 Putative GATA-type zinc finger protein [Malus d... 87 7e-16
UniRef100_Q8GW81 Hypothetical protein At4g16141 [Arabidopsis tha... 80 9e-14
UniRef100_Q84SF6 P0020E09.8 protein [Oryza sativa] 79 1e-13
UniRef100_Q9LIB5 Arabidopsis thaliana genomic DNA, chromosome 3,... 78 3e-13
UniRef100_Q9FJ10 Gb|AAF63824.1 [Arabidopsis thaliana] 77 8e-13
UniRef100_Q8LC79 Transcription factor-like protein [Arabidopsis ... 75 3e-12
UniRef100_Q9SVL2 Transcription factor-like protein [Arabidopsis ... 74 4e-12
UniRef100_Q8VZR1 Hypothetical protein At3g06740 [Arabidopsis tha... 74 6e-12
UniRef100_Q8LG10 Hypothetical protein [Arabidopsis thaliana] 73 8e-12
UniRef100_O23219 Transcription factor like protein [Arabidopsis ... 72 2e-11
UniRef100_Q850Z5 Putative zinc finger protein [Oryza sativa] 72 2e-11
UniRef100_Q8LC59 Hypothetical protein [Arabidopsis thaliana] 70 7e-11
UniRef100_O04636 F2P16.9 protein [Arabidopsis thaliana] 70 7e-11
UniRef100_Q8LR32 GATA-type zinc finger transcription factor-like... 70 9e-11
UniRef100_Q8LQW5 Hypothetical protein B1045F02.30 [Oryza sativa] 69 2e-10
>UniRef100_Q9LV39 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MPI10
[Arabidopsis thaliana]
Length = 398
Score = 138 bits (348), Expect = 2e-31
Identities = 112/329 (34%), Positives = 151/329 (45%), Gaps = 84/329 (25%)
Query: 61 HDGSSSSSSSNHLYNSTFISPQPVMDNPSSSTCDP---------NLSFSKMEEED----- 106
H +++ ++HL+ S + + + N SS CD L+ K + ED
Sbjct: 72 HVAYNNTYHADHLHLSQPLKAKMFVANGGSSACDHMVPKKETRLKLTIRKKDHEDQPHPL 131
Query: 107 ----IKNVHGSAKW-MSSKMRLMKKMMST--PATDKANN-----STTIPISPRIQ-NQGH 153
K S KW MS KMRL+KK ++ D+ NN S P++ + ++ H
Sbjct: 132 HQNPTKPDSDSDKWLMSPKMRLIKKTITNNKQLIDQTNNNNHKESDHYPLNHKTNFDEDH 191
Query: 154 -------------------ENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPK 194
ENRY+ + +N++ RVCSDCNT+ TPLWRSGP GPK
Sbjct: 192 HEDLNFKNVLTRKTTAATTENRYNTINENGYSNNNGVIRVCSDCNTTKTPLWRSGPRGPK 251
Query: 195 SLCNACGIRQRKARRAMAEAA------------------------------NG-----LA 219
SLCNACGIRQRKARRA AA NG +
Sbjct: 252 SLCNACGIRQRKARRAAMAAAAAAGDQEVAVAPRVQQLPLKKKLQNKKKRSNGGEKYNHS 311
Query: 220 TPINTASTKTRVHHNKEKKPRANHFAQFKNKSKSTSSNAGSSQETKKLECFKDFAISLRN 279
P+ + K ++ +EK+ A A SKST+S+ S K CF D I L
Sbjct: 312 PPMVAKAKKCKIKEEEEKEMEAETVAGDSEISKSTTSSNSSISSNK--FCFDDLTIMLSK 369
Query: 280 NSTFQQVFPRDEVAEAALLLMDLSCGYVH 308
+S +QQVFP+DE EAA+LLM LS G VH
Sbjct: 370 SSAYQQVFPQDE-KEAAVLLMALSYGMVH 397
>UniRef100_Q8VZC1 Hypothetical protein At5g56860; MPI10.2 [Arabidopsis thaliana]
Length = 398
Score = 138 bits (348), Expect = 2e-31
Identities = 112/329 (34%), Positives = 151/329 (45%), Gaps = 84/329 (25%)
Query: 61 HDGSSSSSSSNHLYNSTFISPQPVMDNPSSSTCDP---------NLSFSKMEEED----- 106
H +++ ++HL+ S + + + N SS CD L+ K + ED
Sbjct: 72 HVAYNNTYHADHLHLSQPLKAKMFVANGGSSACDHMVPKKETRLKLTIRKKDHEDQPHPL 131
Query: 107 ----IKNVHGSAKW-MSSKMRLMKKMMST--PATDKANN-----STTIPISPRIQ-NQGH 153
K S KW MS KMRL+KK ++ D+ NN S P++ + ++ H
Sbjct: 132 HQNPTKPDSDSDKWLMSPKMRLIKKTITNNKQLIDQTNNNNHKESDHYPLNHKTNFDEDH 191
Query: 154 -------------------ENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPK 194
ENRY+ + +N++ RVCSDCNT+ TPLWRSGP GPK
Sbjct: 192 HEDLNFKNVLTRKTTAATTENRYNTINENGYSNNNGVIRVCSDCNTTKTPLWRSGPRGPK 251
Query: 195 SLCNACGIRQRKARRAMAEAA------------------------------NG-----LA 219
SLCNACGIRQRKARRA AA NG +
Sbjct: 252 SLCNACGIRQRKARRAAMAAAAAAGDQEVAVAPRVQQLPLKKNLQNKKKRSNGGEKYNHS 311
Query: 220 TPINTASTKTRVHHNKEKKPRANHFAQFKNKSKSTSSNAGSSQETKKLECFKDFAISLRN 279
P+ + K ++ +EK+ A A SKST+S+ S K CF D I L
Sbjct: 312 PPMVAKAKKCKIKEEEEKEMEAETVAGDSEISKSTTSSNSSISSNK--FCFDDLTIMLSK 369
Query: 280 NSTFQQVFPRDEVAEAALLLMDLSCGYVH 308
+S +QQVFP+DE EAA+LLM LS G VH
Sbjct: 370 SSAYQQVFPQDE-KEAAVLLMALSYGMVH 397
>UniRef100_Q9SZI6 Putative transcription factor [Arabidopsis thaliana]
Length = 352
Score = 130 bits (327), Expect = 4e-29
Identities = 88/228 (38%), Positives = 118/228 (51%), Gaps = 31/228 (13%)
Query: 102 MEEEDIKNVHG--SAKWMSSKMRLMKKMMSTPATDKANNSTTIPISPRIQNQGHENRYSQ 159
+ + IK++ G S KW+SSK+RLMKK + T ++ T N + S
Sbjct: 134 LPQSPIKDMTGTNSLKWISSKVRLMKKKKAIITTSDSSKQHT--------NNDQSSNLSN 185
Query: 160 RSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARR-----AMAEA 214
+N N+ R+CSDCNT+ TPLWRSGP GPKSLCNACGIRQRKARR A A A
Sbjct: 186 SERQNGYNNDCVIRICSDCNTTKTPLWRSGPRGPKSLCNACGIRQRKARRAAMATATATA 245
Query: 215 ANGLATPI--NTASTKTRVHHNKEK--KPRANHFAQFKN----------KSKSTSSNAGS 260
+G++ P+ K ++ + K P K + T SN+
Sbjct: 246 VSGVSPPVMKKKMQNKNKISNGVYKILSPLPLKVNTCKRMITLEETALAEDLETQSNSTM 305
Query: 261 SQETKKLECFKDFAISLRNNSTFQQVFPRDEVAEAALLLMDLSCGYVH 308
+ + F D A+ L +S +QQVFP+DE EAA+LLM LS G VH
Sbjct: 306 LSSSDNI-YFDDLALLLSKSSAYQQVFPQDE-KEAAILLMALSHGMVH 351
>UniRef100_Q6YW48 Zinc finger protein-like [Oryza sativa]
Length = 353
Score = 125 bits (315), Expect = 1e-27
Identities = 97/296 (32%), Positives = 135/296 (44%), Gaps = 56/296 (18%)
Query: 63 GSSSSSSSNHLY---NSTFISPQPVMDNPSSSTCDPNLSFS----KMEEEDIKNVHGS-A 114
G S+ ++H+ + F++P P S D S +E + ++ +GS +
Sbjct: 65 GESNQQFNDHMMMGGSDVFLTPSPFRPTIQSIGSDMIQRSSYDPYDIESNNKQHANGSTS 124
Query: 115 KWMSSKMRLMKKMMSTPATDKANNSTTIPISPRIQNQGHENRYSQRSPRNNNNSSNTTRV 174
KWMS+ M+ + ATD + PR + Q H++ Q+ + + RV
Sbjct: 125 KWMSTPPMKMRIIRKGAATDPEGGAVR---KPRRRAQAHQDESQQQLQQ----ALGVVRV 177
Query: 175 CSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPINTASTKTRVHHN 234
CSDCNT+ TPLWRSGP GPKSLCNACGIRQRKARRAMA AANG A S +N
Sbjct: 178 CSDCNTTKTPLWRSGPCGPKSLCNACGIRQRKARRAMAAAANGGAAVAPAKSVAAAPVNN 237
Query: 235 K---EKKPRA---NHFAQFKNKSKSTSSNAGSSQETKKLECFKDFAISLRN--------- 279
K +K+ RA + FK + K A + TK + A + ++
Sbjct: 238 KPAAKKEKRAADVDRSLPFKKRCKMVDHVAAAVAATKPTAAGEVVAAAPKDQDHVIVVGG 297
Query: 280 --------------------------NSTFQQVFPRDEVAEAALLLMDLSCGYVHS 309
+ F PRDE+ +AA+LLM LSCG VHS
Sbjct: 298 ENAAATSMPAQNPISKAAATAAAAAASPAFFHGLPRDEITDAAMLLMTLSCGLVHS 353
>UniRef100_Q5Z624 Zinc finger protein-like [Oryza sativa]
Length = 347
Score = 106 bits (264), Expect = 9e-22
Identities = 86/264 (32%), Positives = 122/264 (45%), Gaps = 33/264 (12%)
Query: 69 SSNHLYNSTFISPQPVMDNP-SSSTCDPNLSFSKMEEEDIKNVHGSAKWMS---SKMRLM 124
SS ++ + F + + + D+ S DP + + V KW + +KM++
Sbjct: 94 SSAGIFATPFPTVKSIRDDMIERSQFDPYDTEKLQASCGLAKVVAGGKWSAVPAAKMKIT 153
Query: 125 KKMMSTPATDKANNSTTI-PISPR------IQNQGHENRYSQRSPRNNNNSSNTTRVCSD 177
+KM + +TT+ P PR ++ GH Q + RVCSD
Sbjct: 154 RKMGEPSSGVTGGAATTVAPKKPRRRPAQAYEDHGHGGAMGQ--------AFGVIRVCSD 205
Query: 178 CNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPINTASTKTRVHHNKEK 237
CNT+ TPLWRSGP GPKSLCNACGIRQRKARRAM A+GL N A K H
Sbjct: 206 CNTTKTPLWRSGPCGPKSLCNACGIRQRKARRAM--MASGLPASPNAAGPKAAAHSGATN 263
Query: 238 KPRANHFAQFKNKS-----KSTSSNAGSSQETKKLECFKDFAISLRNNS-TFQQVFP--- 288
A + + + ++ G+ ++ L K A + + S V P
Sbjct: 264 AAAAAAMEETAESATVAPPPAPTTRGGTLVDSIGLSWSKTHAAATASCSFRPSPVAPGFA 323
Query: 289 ---RDEVAEAALLLMDLSCGYVHS 309
+DE+ +AA+LLM LSCG V S
Sbjct: 324 AAVQDEITDAAMLLMTLSCGLVRS 347
>UniRef100_Q6V8R1 Putative GATA-type zinc finger protein [Malus domestica]
Length = 100
Score = 86.7 bits (213), Expect = 7e-16
Identities = 47/100 (47%), Positives = 59/100 (59%), Gaps = 15/100 (15%)
Query: 116 WMSSKMRLMKKMMSTPATDKANNSTTI-PISPRIQNQGHENRYSQRSPRN--------NN 166
WMSS MR+M+KM + T ++ S+ PIS ++ + E + Q +N
Sbjct: 1 WMSSNMRMMRKMSNPDQTSSSSTSSDDKPISMKLSSHKFEEQKLQHPSSQLGADMISCSN 60
Query: 167 NSSNTT------RVCSDCNTSSTPLWRSGPNGPKSLCNAC 200
NSSN T RVCSDCNT+ TPLWRSGP GPKSLCNAC
Sbjct: 61 NSSNNTNNIPIIRVCSDCNTTKTPLWRSGPRGPKSLCNAC 100
>UniRef100_Q8GW81 Hypothetical protein At4g16141 [Arabidopsis thaliana]
Length = 197
Score = 79.7 bits (195), Expect = 9e-14
Identities = 60/190 (31%), Positives = 86/190 (44%), Gaps = 34/190 (17%)
Query: 147 RIQNQGHENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRK 206
++++ G + + ++ + +T + C DC TS TPLWR GP GPKSLCNACGI+ RK
Sbjct: 11 KLESAGDSSDVDNGNCSSSGSGGDTKKTCVDCGTSRTPLWRGGPAGPKSLCNACGIKSRK 70
Query: 207 ARRAM-------------------AEAAN---GLATPINTASTKTRVHHNKEKKPRANHF 244
R+A E+ N G P+N K K K +
Sbjct: 71 KRQAALGIRQDDIKIKSKSNNNLGLESRNVKTGKGEPVNVKIAKCEPGIVKIAKGEPGN- 129
Query: 245 AQFKNKSKSTSSNAGSSQETKK----LECFKDFAI---SLRNNSTFQQVFPR--DEVAEA 295
KNK K N+ SS KK + F DF +++ ++ ++ R E A
Sbjct: 130 --VKNKIKRDPENSSSSNNNKKNVKRVGRFLDFGFKVPAMKRSAVEKKRLWRKLGEEERA 187
Query: 296 ALLLMDLSCG 305
A+LLM LSCG
Sbjct: 188 AVLLMALSCG 197
>UniRef100_Q84SF6 P0020E09.8 protein [Oryza sativa]
Length = 140
Score = 79.3 bits (194), Expect = 1e-13
Identities = 51/144 (35%), Positives = 75/144 (51%), Gaps = 24/144 (16%)
Query: 170 NTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPINTASTKT 229
N ++ C+DC+T+ TPLWR GP GPKSLCNACGIR RK RRA A +++++T T
Sbjct: 17 NGSKACADCHTTKTPLWRGGPGGPKSLCNACGIRYRKRRRA--------ALGLDSSATAT 68
Query: 230 RVHHNKEKKPRANHFAQFKNKSKSTSSNAGSSQETKKLECF----KDFAISLRNNSTFQQ 285
A A+ + K+K+ A + T +L KD A+ + ++
Sbjct: 69 -----------ATDGAEQQKKTKAKKEKAQEEEVTMELHTVGFRSKDAAV-FKQRRRMRR 116
Query: 286 VFPRDEVAEAALLLMDLSCGYVHS 309
E AA+LLM LS G +++
Sbjct: 117 RKCLGEEERAAILLMALSSGVIYA 140
>UniRef100_Q9LIB5 Arabidopsis thaliana genomic DNA, chromosome 3, P1 clone:MUH15
[Arabidopsis thaliana]
Length = 190
Score = 78.2 bits (191), Expect = 3e-13
Identities = 62/192 (32%), Positives = 92/192 (47%), Gaps = 27/192 (14%)
Query: 137 NNSTTIPISPRIQNQGHENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSL 196
+ T + + + + +EN S S ++S +T R C DC T TPLWR GP GPKSL
Sbjct: 7 DTKTKLDSAGELSDVDNENCSSSGSG-GGSSSGDTKRTCVDCGTIRTPLWRGGPAGPKSL 65
Query: 197 CNACGIRQRKARRAMAEAANGLATPINTASTKT------RVHHNKEKKPRANHFAQFK-- 248
CNACGI+ RK R +AA G+ + + K+ + H KK + N K
Sbjct: 66 CNACGIKSRKKR----QAALGMRSEEKKKNRKSNCNNDLNLDHRNAKKYKINIVDDGKID 121
Query: 249 ----------NKSKSTSSNAGSSQETKKLECFKDFAISLRNNSTFQQVFPR-DEVAEAAL 297
+S S+SSN G S K L+ + R+ ++++ + E AA+
Sbjct: 122 IDDDPKICNNKRSSSSSSNKGVS---KFLDLGFKVPVMKRSAVEKKRLWRKLGEEERAAV 178
Query: 298 LLMDLSCGYVHS 309
LLM LSC V++
Sbjct: 179 LLMALSCSSVYA 190
>UniRef100_Q9FJ10 Gb|AAF63824.1 [Arabidopsis thaliana]
Length = 139
Score = 76.6 bits (187), Expect = 8e-13
Identities = 52/163 (31%), Positives = 77/163 (46%), Gaps = 35/163 (21%)
Query: 148 IQNQGHENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKA 207
+ ++ + R +NN + ++ + C+DC TS TPLWR GP GPKSLCNACGIR RK
Sbjct: 11 VDSETMKTRAEDMIEQNNTSVNDKKKTCADCGTSKTPLWRGGPVGPKSLCNACGIRNRKK 70
Query: 208 RRAMAEAANGLATPINTASTKTRVHHNKEKKPRANHFAQFKNKSKSTSSNAGSSQETKKL 267
RR E NK+ K K+ S + G S + +
Sbjct: 71 RRGGTE-------------------DNKKLK---------KSSSGGGNRKFGESLKQSLM 102
Query: 268 ECFKDFAISLRNNSTFQQVFPR-DEVAEAALLLMDLSCGYVHS 309
+ + +R ST ++ + E +AA+LLM LS G V++
Sbjct: 103 D------LGIRKRSTVEKQRQKLGEEEQAAVLLMALSYGSVYA 139
>UniRef100_Q8LC79 Transcription factor-like protein [Arabidopsis thaliana]
Length = 294
Score = 74.7 bits (182), Expect = 3e-12
Identities = 57/197 (28%), Positives = 83/197 (41%), Gaps = 31/197 (15%)
Query: 89 SSSTCDPNLSF----SKMEEEDIKNVHGSAKWMSSKMRLMKKMMSTPATDKANNSTTIPI 144
SSS D LS +++ EED K ++ SS + ++ T K NNS T P
Sbjct: 62 SSSLVDCTLSLGTPSTRLCEEDEKRRRSTSSGASSCISNFWDLIHT----KNNNSKTAPY 117
Query: 145 SPRIQNQGHENRYSQRSPRNN------------NNSSNTTRVCSDCNTSSTPLWRSGPNG 192
+ + +S P S R C++C+T+STPLWR+GP G
Sbjct: 118 N-------NVPSFSANKPSRGCSGGGDGGGGGGGGDSLLARRCANCDTTSTPLWRNGPRG 170
Query: 193 PKSLCNACGIRQRKARRAMAEAANGLATPINTASTKTRVHHNKEKKPRANHFAQFKNKSK 252
PKSLCNACGIR +K R A+ T HHN N++ N +
Sbjct: 171 PKSLCNACGIRFKKEERRTTAASGNTVVGAAPVQTDQYGHHNS----GYNNYHAATNNNN 226
Query: 253 STSSNAGSSQETKKLEC 269
+ + T+++ C
Sbjct: 227 NNGTPWAHHHSTQRVPC 243
>UniRef100_Q9SVL2 Transcription factor-like protein [Arabidopsis thaliana]
Length = 294
Score = 74.3 bits (181), Expect = 4e-12
Identities = 57/197 (28%), Positives = 82/197 (40%), Gaps = 31/197 (15%)
Query: 89 SSSTCDPNLSF----SKMEEEDIKNVHGSAKWMSSKMRLMKKMMSTPATDKANNSTTIPI 144
SSS D LS +++ EED K ++ SS + ++ T K NNS T P
Sbjct: 62 SSSLVDCTLSLGTPSTRLCEEDEKRRRSTSSGASSCISNFWDLIHT----KNNNSKTAPY 117
Query: 145 SPRIQNQGHENRYSQRSPRNN------------NNSSNTTRVCSDCNTSSTPLWRSGPNG 192
+ + +S P S R C++C+T+STPLWR+GP G
Sbjct: 118 N-------NVPSFSANKPSRGCSGGGGGGGGGGGGDSLLARRCANCDTTSTPLWRNGPRG 170
Query: 193 PKSLCNACGIRQRKARRAMAEAANGLATPINTASTKTRVHHNKEKKPRANHFAQFKNKSK 252
PKSLCNACGIR +K R A T HHN N++ N +
Sbjct: 171 PKSLCNACGIRFKKEERRTTAATGNTVVGAAPVQTDQYGHHNS----GYNNYHAATNNNN 226
Query: 253 STSSNAGSSQETKKLEC 269
+ + T+++ C
Sbjct: 227 NNGTPWAHHHSTQRVPC 243
>UniRef100_Q8VZR1 Hypothetical protein At3g06740 [Arabidopsis thaliana]
Length = 149
Score = 73.6 bits (179), Expect = 6e-12
Identities = 49/143 (34%), Positives = 67/143 (46%), Gaps = 29/143 (20%)
Query: 161 SPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLAT 220
S +N SN + C+ C TS TPLWR GP GPKSLCNACGIR RK RR +
Sbjct: 29 SSSSNEAISNEKKSCAICGTSKTPLWRGGPAGPKSLCNACGIRNRKKRRTLIS------- 81
Query: 221 PINTASTKTRVHHNKEKKPRANHFAQFKNKSKSTSSNAGSSQETKKLECFKDFAISLRNN 280
N + K + HN+ K G S + + +E ++ + R+
Sbjct: 82 --NRSEDKKKKSHNRNPK-------------------FGDSLKQRLMELGREVMMQ-RST 119
Query: 281 STFQQVFPRDEVAEAALLLMDLS 303
+ Q+ E +AA+LLM LS
Sbjct: 120 AENQRRNKLGEEEQAAVLLMALS 142
>UniRef100_Q8LG10 Hypothetical protein [Arabidopsis thaliana]
Length = 136
Score = 73.2 bits (178), Expect = 8e-12
Identities = 36/78 (46%), Positives = 43/78 (54%), Gaps = 9/78 (11%)
Query: 161 SPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLAT 220
S +N SN + C+ C TS TPLWR GP GPKSLCNACGIR RK RR +
Sbjct: 16 SSSSNEAISNEKKSCAICGTSKTPLWRGGPAGPKSLCNACGIRNRKKRRTLIS------- 68
Query: 221 PINTASTKTRVHHNKEKK 238
N + K + HN+ K
Sbjct: 69 --NRSEDKKKKSHNRNPK 84
>UniRef100_O23219 Transcription factor like protein [Arabidopsis thaliana]
Length = 211
Score = 72.0 bits (175), Expect = 2e-11
Identities = 36/83 (43%), Positives = 52/83 (62%), Gaps = 4/83 (4%)
Query: 173 RVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPINTASTKTRVH 232
R C++C+T+STPLWR+GP GPKSLCNACGIR +K R + A N + +TA+ +
Sbjct: 75 RRCANCDTTSTPLWRNGPRGPKSLCNACGIRFKKEERRASTARNSTSGGGSTAAGVPTLD 134
Query: 233 HNKEKKPRANHFAQFKNKSKSTS 255
H + AN++ N+ S+S
Sbjct: 135 H----QASANYYYNNNNQYASSS 153
>UniRef100_Q850Z5 Putative zinc finger protein [Oryza sativa]
Length = 136
Score = 71.6 bits (174), Expect = 2e-11
Identities = 48/146 (32%), Positives = 69/146 (46%), Gaps = 25/146 (17%)
Query: 164 NNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPIN 223
+ +S + C+DC+T+ TPLWR GP+GPKSLCNACGIR RK RR A GL
Sbjct: 16 DERTASGEPKACTDCHTTKTPLWRGGPSGPKSLCNACGIRYRKKRR----EALGLDAGEG 71
Query: 224 TASTKTRVHHNKEKKPRANHFAQFKNKSKSTSSNAGSSQETKKLECFKDFAISLRNNSTF 283
A R K K+ R + E + + K+ + R
Sbjct: 72 GAE---RQEKKKSKRERGEEV----------------TMELRMVGFGKEVVLKQRRRMRR 112
Query: 284 QQVFPRDEVAEAALLLMDLSCGYVHS 309
++ +E +AA+LLM LS G +++
Sbjct: 113 RRRLGEEE--KAAILLMALSSGVIYA 136
>UniRef100_Q8LC59 Hypothetical protein [Arabidopsis thaliana]
Length = 120
Score = 70.1 bits (170), Expect = 7e-11
Identities = 28/40 (70%), Positives = 32/40 (80%)
Query: 171 TTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRA 210
T R CS+C T+ TP+WR GP GPKSLCNACGIR RK RR+
Sbjct: 24 TIRCCSECKTTKTPMWRGGPTGPKSLCNACGIRHRKQRRS 63
>UniRef100_O04636 F2P16.9 protein [Arabidopsis thaliana]
Length = 550
Score = 70.1 bits (170), Expect = 7e-11
Identities = 28/40 (70%), Positives = 32/40 (80%)
Query: 171 TTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRA 210
T R CS+C T+ TP+WR GP GPKSLCNACGIR RK RR+
Sbjct: 454 TIRCCSECKTTKTPMWRGGPTGPKSLCNACGIRHRKQRRS 493
>UniRef100_Q8LR32 GATA-type zinc finger transcription factor-like [Oryza sativa]
Length = 242
Score = 69.7 bits (169), Expect = 9e-11
Identities = 29/48 (60%), Positives = 38/48 (78%)
Query: 168 SSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAA 215
+++ R C++C+T+STPLWR+GP GPKSLCNACGIR +K R A AA
Sbjct: 117 AASAPRRCANCDTTSTPLWRNGPRGPKSLCNACGIRYKKEERRAAAAA 164
>UniRef100_Q8LQW5 Hypothetical protein B1045F02.30 [Oryza sativa]
Length = 131
Score = 68.6 bits (166), Expect = 2e-10
Identities = 49/136 (36%), Positives = 64/136 (47%), Gaps = 38/136 (27%)
Query: 173 RVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPINTASTKTRVH 232
R C +C ++TP+WRSGP GP+SLCNACGIR RK RR L +N
Sbjct: 19 RSCVECRATTTPMWRSGPTGPRSLCNACGIRYRKKRR------QDLGLDLN--------- 63
Query: 233 HNKEKKPRANHFAQFK-NKSKSTSSNAGSSQETKKLECFKDFAISLRNNSTFQQVFPRDE 291
+K+ + K + S S + N+GS N+S+ QV P+
Sbjct: 64 -QPQKQEHGEVIPEVKDSNSNSNNCNSGSG-----------------NSSSNLQVVPKRR 105
Query: 292 ----VAEAALLLMDLS 303
V EAALLLM LS
Sbjct: 106 LLMGVEEAALLLMTLS 121
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.311 0.125 0.363
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 541,716,716
Number of Sequences: 2790947
Number of extensions: 22226237
Number of successful extensions: 98697
Number of sequences better than 10.0: 881
Number of HSP's better than 10.0 without gapping: 235
Number of HSP's successfully gapped in prelim test: 669
Number of HSP's that attempted gapping in prelim test: 87725
Number of HSP's gapped (non-prelim): 3987
length of query: 310
length of database: 848,049,833
effective HSP length: 127
effective length of query: 183
effective length of database: 493,599,564
effective search space: 90328720212
effective search space used: 90328720212
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0019a.4


