

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0014.18
(181 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
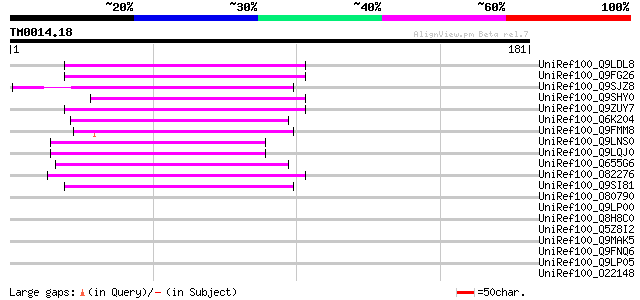
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LDL8 Hypothetical protein AT4g08830 [Arabidopsis tha... 57 2e-07
UniRef100_Q9FG26 Non-LTR retroelement reverse transcriptase-like... 54 2e-06
UniRef100_Q9SJZ8 Putative non-LTR retroelement reverse transcrip... 52 5e-06
UniRef100_Q9SHY0 F1E22.12 [Arabidopsis thaliana] 51 1e-05
UniRef100_Q9ZUY7 Putative reverse transcriptase [Arabidopsis tha... 51 2e-05
UniRef100_Q6K204 Hypothetical protein B1469H02.35 [Oryza sativa] 50 3e-05
UniRef100_Q9FMM8 Non-LTR retroelement reverse transcriptase-like... 50 4e-05
UniRef100_Q9LNS0 F1L3.4 [Arabidopsis thaliana] 48 1e-04
UniRef100_Q9LQJ0 F28G4.15 protein [Arabidopsis thaliana] 48 1e-04
UniRef100_Q655G6 Hypothetical protein P0009H10.11 [Oryza sativa] 48 1e-04
UniRef100_O82276 Putative non-LTR retroelement reverse transcrip... 47 2e-04
UniRef100_Q9SI81 F23N19.5 [Arabidopsis thaliana] 47 3e-04
UniRef100_O80790 Putative non-LTR retroelement reverse transcrip... 44 0.003
UniRef100_Q9LP00 F9C16.18 [Arabidopsis thaliana] 42 0.010
UniRef100_Q8H8C0 Hypothetical protein OJ1134F05.9 [Oryza sativa] 42 0.010
UniRef100_Q5Z8I2 Hypothetical protein P0622F03.7 [Oryza sativa] 38 0.14
UniRef100_Q9MAK5 F27F5.12 [Arabidopsis thaliana] 37 0.24
UniRef100_Q9FNQ6 Gb|AAF19536.1 [Arabidopsis thaliana] 37 0.24
UniRef100_Q9LP05 F9C16.13 [Arabidopsis thaliana] 37 0.31
UniRef100_O22148 Putative non-LTR retroelement reverse transcrip... 36 0.53
>UniRef100_Q9LDL8 Hypothetical protein AT4g08830 [Arabidopsis thaliana]
Length = 947
Score = 57.0 bits (136), Expect = 2e-07
Identities = 29/84 (34%), Positives = 45/84 (53%)
Query: 20 IRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGI 79
+ W P D WVKLN+DGA + +A G++RD + +A +G S AELWG+
Sbjct: 839 VAWKPPDGEWVKLNTDGASRGNLGLATTGGVLRDGIGHWCGGFALDIGVCSAPLAELWGV 898
Query: 80 LCGARLLQERDY*RVLIETNLSIL 103
G + ER + RV +E + ++
Sbjct: 899 YYGLYMAWERRFTRVELEVDSELV 922
>UniRef100_Q9FG26 Non-LTR retroelement reverse transcriptase-like [Arabidopsis
thaliana]
Length = 676
Score = 53.5 bits (127), Expect = 2e-06
Identities = 27/84 (32%), Positives = 45/84 (53%)
Query: 20 IRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGI 79
I W WV +N+DGA + A G+IRD ++++ +A +G S AELWG+
Sbjct: 506 IAWRKPAEGWVTMNTDGASHGNPGQATAGGVIRDEHGSWLVGFALNIGVCSAPLAELWGV 565
Query: 80 LCGARLLQERDY*RVLIETNLSIL 103
G + ER + RV +E + +++
Sbjct: 566 YYGLVVAWERGWRRVRLEVDSALV 589
>UniRef100_Q9SJZ8 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 321
Score = 52.4 bits (124), Expect = 5e-06
Identities = 33/98 (33%), Positives = 50/98 (50%), Gaps = 9/98 (9%)
Query: 2 LRVAQVTRVVSPVRDAPAIRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIA 61
+RV++V R+VS W + WVKLN+DGA + A G++RD A++
Sbjct: 142 VRVSRVERLVS---------WVSPEDGWVKLNTDGASRGNPGFATAGGVLRDHNGAWIGG 192
Query: 62 YARRLGSYSTLQAELWGILCGARLLQERDY*RVLIETN 99
+A +G S AELWG+ G + R RV +E +
Sbjct: 193 FAVNIGVCSAPLAELWGVYYGLFIAWGRGARRVELEVD 230
>UniRef100_Q9SHY0 F1E22.12 [Arabidopsis thaliana]
Length = 1055
Score = 51.2 bits (121), Expect = 1e-05
Identities = 25/75 (33%), Positives = 42/75 (55%)
Query: 29 WVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGILCGARLLQE 88
WVK+N+DGA + +A+ G++RD A+ ++ +G S QAELWG+ G E
Sbjct: 568 WVKVNTDGASRGNPGLASAGGVLRDCTGAWCGGFSLNIGRCSAPQAELWGVYYGLYFAWE 627
Query: 89 RDY*RVLIETNLSIL 103
+ RV +E + ++
Sbjct: 628 KKVPRVELEVDSEVI 642
>UniRef100_Q9ZUY7 Putative reverse transcriptase [Arabidopsis thaliana]
Length = 314
Score = 50.8 bits (120), Expect = 2e-05
Identities = 29/84 (34%), Positives = 44/84 (51%)
Query: 20 IRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGI 79
I W + W KLN+DGA + +A+ G++RD A+ +A +G S AELWG+
Sbjct: 144 IAWSKPEEGWWKLNTDGASRGNPGLASAGGVLRDEEGAWRGGFALNIGVCSAPLAELWGV 203
Query: 80 LCGARLLQERDY*RVLIETNLSIL 103
G + ER R+ IE + I+
Sbjct: 204 YYGLYIAWERRVTRLEIEVDSEIV 227
>UniRef100_Q6K204 Hypothetical protein B1469H02.35 [Oryza sativa]
Length = 212
Score = 50.1 bits (118), Expect = 3e-05
Identities = 27/76 (35%), Positives = 41/76 (53%)
Query: 22 WWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGILC 81
W +A W+KLN DG+ + +A+ G+ RD AFV+ YA R+G ++ AEL +
Sbjct: 51 WRKPEAGWIKLNFDGSSKHATKIASIGGVYRDHEGAFVLGYAERIGRATSSVAELAALRR 110
Query: 82 GARLLQERDY*RVLIE 97
G L+ + RV E
Sbjct: 111 GLELVVRNGWRRVWAE 126
>UniRef100_Q9FMM8 Non-LTR retroelement reverse transcriptase-like protein
[Arabidopsis thaliana]
Length = 308
Score = 49.7 bits (117), Expect = 4e-05
Identities = 27/78 (34%), Positives = 42/78 (53%), Gaps = 1/78 (1%)
Query: 23 WPLDAV-WVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGILC 81
W L V WVK+N+DGA + +A+ G++RD A+ ++ +G S AELWG+
Sbjct: 140 WVLPCVGWVKVNTDGASRGNPGLASAGGVLRDCEGAWCGGFSLNIGRCSAQHAELWGVYY 199
Query: 82 GARLLQERDY*RVLIETN 99
G E+ RV +E +
Sbjct: 200 GLYFAWEKKVPRVELEVD 217
>UniRef100_Q9LNS0 F1L3.4 [Arabidopsis thaliana]
Length = 253
Score = 47.8 bits (112), Expect = 1e-04
Identities = 25/75 (33%), Positives = 36/75 (47%)
Query: 15 RDAPAIRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQA 74
R+ I W P W KLN+DGA + +A G++RD + ++ +G S A
Sbjct: 78 REERLIAWSPPRVGWFKLNTDGASRGNPRLATAGGVVRDGDGNWCYGFSLNIGICSAPLA 137
Query: 75 ELWGILCGARLLQER 89
ELWG G + ER
Sbjct: 138 ELWGAYYGLNIAWER 152
>UniRef100_Q9LQJ0 F28G4.15 protein [Arabidopsis thaliana]
Length = 272
Score = 47.8 bits (112), Expect = 1e-04
Identities = 25/75 (33%), Positives = 36/75 (47%)
Query: 15 RDAPAIRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQA 74
R+ I W P W KLN+DGA + +A G++RD + ++ +G S A
Sbjct: 97 REERLIAWSPPRVGWFKLNTDGASRGNPRLATAGGVVRDGDGNWCYGFSLNIGICSAPLA 156
Query: 75 ELWGILCGARLLQER 89
ELWG G + ER
Sbjct: 157 ELWGAYYGLNIAWER 171
>UniRef100_Q655G6 Hypothetical protein P0009H10.11 [Oryza sativa]
Length = 240
Score = 47.8 bits (112), Expect = 1e-04
Identities = 28/81 (34%), Positives = 42/81 (51%)
Query: 17 APAIRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAEL 76
A A W + W+KLN DG+ S +A+ G+ RD AF++ YA R+G+ ++ AEL
Sbjct: 74 ASARIWRKPEEGWMKLNFDGSSKHSTGIASIGGVYRDHDGAFLLGYAERIGTATSSVAEL 133
Query: 77 WGILCGARLLQERDY*RVLIE 97
+ G L + RV E
Sbjct: 134 AALRRGLELAVRNGWRRVWAE 154
>UniRef100_O82276 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1231
Score = 47.0 bits (110), Expect = 2e-04
Identities = 27/90 (30%), Positives = 46/90 (51%)
Query: 14 VRDAPAIRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQ 73
VR IRW WVK+ +DGA + +AA G IR+ ++ +A +GS +
Sbjct: 1055 VRVERMIRWQVPSDGWVKITTDGASRGNHGLAAAGGAIRNGQGEWLGGFALNIGSCAAPL 1114
Query: 74 AELWGILCGARLLQERDY*RVLIETNLSIL 103
AELWG G + ++ + RV ++ + ++
Sbjct: 1115 AELWGAYYGLLIAWDKGFRRVELDLDCKLV 1144
>UniRef100_Q9SI81 F23N19.5 [Arabidopsis thaliana]
Length = 233
Score = 46.6 bits (109), Expect = 3e-04
Identities = 23/80 (28%), Positives = 39/80 (48%)
Query: 20 IRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGI 79
+RW W KLN+DGA + +A G +R+ + + +A +G AELWG+
Sbjct: 63 VRWSKPSLGWCKLNTDGASHGNPGLATAGGALRNEYGEWCFGFALNIGRCLAPLAELWGV 122
Query: 80 LCGARLLQERDY*RVLIETN 99
G + +R R+ +E +
Sbjct: 123 YYGLFMAWDRGITRLELEVD 142
>UniRef100_O80790 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 970
Score = 43.5 bits (101), Expect = 0.003
Identities = 31/97 (31%), Positives = 43/97 (43%), Gaps = 2/97 (2%)
Query: 4 VAQVTRVVSPVRDAPAIRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYA 63
V + V R IRW WVKL +DGA +AA G I + ++ +A
Sbjct: 784 VGTLNNHVKRARVERMIRWKAPSDRWVKLTTDGASRGHQGLAAASGAILNLQGEWLGGFA 843
Query: 64 RRLGSYSTLQAELWGILCGARLLQERDY*RVLIETNL 100
+GS AELWG G + ++ + RV E NL
Sbjct: 844 LNIGSCDAPLAELWGAYYGLLIAWDKGFRRV--ELNL 878
>UniRef100_Q9LP00 F9C16.18 [Arabidopsis thaliana]
Length = 196
Score = 41.6 bits (96), Expect = 0.010
Identities = 17/51 (33%), Positives = 31/51 (60%)
Query: 29 WVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGI 79
W+KLN+DGA + ++ G++RDR ++ ++ +G + AELWG+
Sbjct: 65 WLKLNTDGASRGNPTLSTAGGVLRDREGEWLGGFSLNIGRRTAPVAELWGV 115
>UniRef100_Q8H8C0 Hypothetical protein OJ1134F05.9 [Oryza sativa]
Length = 210
Score = 41.6 bits (96), Expect = 0.010
Identities = 27/78 (34%), Positives = 36/78 (45%), Gaps = 2/78 (2%)
Query: 22 WWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRL--GSYSTLQAELWGI 79
W + W+KLN DG+ + A+ G RD AFV YA R+ G S+ AEL +
Sbjct: 47 WRKPEVGWMKLNFDGSRNDATGAASIGGAFRDHEGAFVAGYAERMIVGGASSFTAELAAL 106
Query: 80 LCGARLLQERDY*RVLIE 97
G L + RV E
Sbjct: 107 RRGLELAARYGWRRVWAE 124
>UniRef100_Q5Z8I2 Hypothetical protein P0622F03.7 [Oryza sativa]
Length = 217
Score = 37.7 bits (86), Expect = 0.14
Identities = 26/88 (29%), Positives = 43/88 (48%), Gaps = 2/88 (2%)
Query: 12 SPVRDAPAIRWWPLDAVWVKLNSDGA-YISSADMAACRGIIRDRFSAFVIAYARRLGSYS 70
SP +RW P WVKLN DG+ + + A+ G+IRD ++A+A R +
Sbjct: 34 SPPTRITLVRWTPPRHGWVKLNFDGSVHNDGSGRASIGGVIRDDHGRVLLAFAERTPHAT 93
Query: 71 TLQAELWGILCGARLLQERDY-*RVLIE 97
E ++ G +L + + R+L+E
Sbjct: 94 IGVVEARALIRGLQLALDHGWNDRLLVE 121
>UniRef100_Q9MAK5 F27F5.12 [Arabidopsis thaliana]
Length = 205
Score = 37.0 bits (84), Expect = 0.24
Identities = 21/74 (28%), Positives = 36/74 (48%)
Query: 30 VKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGILCGARLLQER 89
VK D ++ + + G+IRD ++ +A +G S AELWG+ G L ER
Sbjct: 58 VKARYDWSFHKNPVLTTAGGLIRDATGSWCGGFALNIGVCSAPLAELWGVYYGLYLTWER 117
Query: 90 DY*RVLIETNLSIL 103
R+ +E + ++
Sbjct: 118 HITRLEVEVDSELV 131
>UniRef100_Q9FNQ6 Gb|AAF19536.1 [Arabidopsis thaliana]
Length = 657
Score = 37.0 bits (84), Expect = 0.24
Identities = 30/103 (29%), Positives = 44/103 (42%), Gaps = 9/103 (8%)
Query: 30 VKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGILCGARLLQER 89
VKLN G +D+ GI+RD+ +V Y R S + A L I G + L +
Sbjct: 504 VKLNIQGTSNPLSDLTRSAGIVRDQSGKWVFGYIRCHKSIPEVVAGLLAIYQGLKYLWDS 563
Query: 90 DY*RVLIETNLSILGVVLVLIPVLLLLTK*SRLFSRVSPRLGA 132
+ R+ +ET ++ LT S LF + LGA
Sbjct: 564 GFRRIHLET---------TSFEIINALTTKSSLFCKSKTLLGA 597
>UniRef100_Q9LP05 F9C16.13 [Arabidopsis thaliana]
Length = 1172
Score = 36.6 bits (83), Expect = 0.31
Identities = 16/53 (30%), Positives = 31/53 (58%)
Query: 30 VKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGILCG 82
+KLN+DGA + ++ G++RDR ++ ++ +G + AELW ++ G
Sbjct: 214 LKLNTDGASRGNPALSTAGGVLRDREGEWLGGFSLNIGRRTAPVAELWALVVG 266
>UniRef100_O22148 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1374
Score = 35.8 bits (81), Expect = 0.53
Identities = 26/96 (27%), Positives = 42/96 (43%), Gaps = 2/96 (2%)
Query: 9 RVVSPVRDAPAIRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGS 68
+V S RD ++W P WVK N+DGA+ ++R+ + R L S
Sbjct: 1203 QVTSSTRDR-CVKWQPPSHGWVKCNTDGAWSKDLGNCGVGWVLRNHTGRLLWLGLRALPS 1261
Query: 69 -YSTLQAELWGILCGARLLQERDY*RVLIETNLSIL 103
S L+ E+ + L +Y RV+ E++ L
Sbjct: 1262 QQSVLETEVEALRWAVLSLSRFNYRRVIFESDSQYL 1297
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.336 0.145 0.454
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 250,212,037
Number of Sequences: 2790947
Number of extensions: 8584737
Number of successful extensions: 44789
Number of sequences better than 10.0: 31
Number of HSP's better than 10.0 without gapping: 20
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 44767
Number of HSP's gapped (non-prelim): 31
length of query: 181
length of database: 848,049,833
effective HSP length: 119
effective length of query: 62
effective length of database: 515,927,140
effective search space: 31987482680
effective search space used: 31987482680
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 71 (32.0 bits)
Lotus: description of TM0014.18


