

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0010.6
(493 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
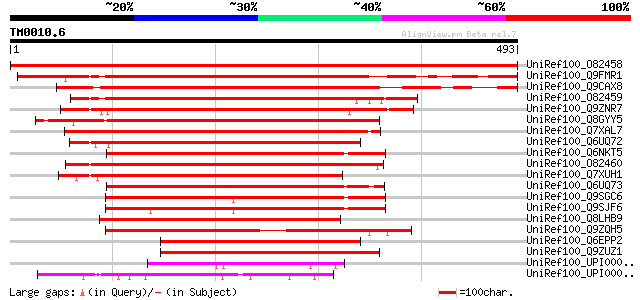
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O82458 Rac GTPase activating protein 1 [Lotus japonicus] 971 0.0
UniRef100_Q9FMR1 Rac GTPase activating protein [Arabidopsis thal... 444 e-123
UniRef100_Q9CAX8 Putative rac GTPase activating protein; 62102-6... 424 e-117
UniRef100_O82459 Rac GTPase activating protein 2 [Lotus japonicus] 384 e-105
UniRef100_Q9ZNR7 Putative rac GTPase activating protein [Arabido... 379 e-103
UniRef100_Q8GYY5 Putative rac GTPase activating protein [Arabido... 372 e-101
UniRef100_Q7XAL7 Rac GTPase activating protein 3-like protein [O... 370 e-101
UniRef100_Q6UQ72 Rho GTPase activating protein 2 [Oryza sativa] 358 1e-97
UniRef100_Q6NKT5 At1g08340 [Arabidopsis thaliana] 357 3e-97
UniRef100_O82460 Rac GTPase activating protein 3 [Lotus japonicus] 356 1e-96
UniRef100_Q7XUH1 OSJNBa0020J04.11 protein [Oryza sativa] 352 1e-95
UniRef100_Q6UQ73 Rho GTPase activating protein 1 [Oryza sativa] 352 1e-95
UniRef100_Q9SGC6 T23G18.20 [Arabidopsis thaliana] 352 2e-95
UniRef100_Q9SJF6 T27G7.4 [Arabidopsis thaliana] 343 7e-93
UniRef100_Q8LHB9 Putative rac GTPase activating protein [Oryza s... 313 9e-84
UniRef100_Q9ZQH5 Putative rac GTPase activating protein [Arabido... 290 5e-77
UniRef100_Q6EPP2 Putative Rho GTPase activating protein 2 [Oryza... 276 7e-73
UniRef100_Q9ZUZ1 Putative rac GTPase activating protein [Arabido... 272 2e-71
UniRef100_UPI00004996C7 UPI00004996C7 UniRef100 entry 99 3e-19
UniRef100_UPI000007CCEF UPI000007CCEF UniRef100 entry 85 5e-15
>UniRef100_O82458 Rac GTPase activating protein 1 [Lotus japonicus]
Length = 493
Score = 971 bits (2511), Expect = 0.0
Identities = 490/493 (99%), Positives = 491/493 (99%)
Query: 1 MTEVLQLPSPSRRCSNDGSHQTALINNSSSVEEGVLQLQTHLDLVEEDEEEEKEKDREGD 60
MTEVLQLPSPSRRCSNDGSHQTALINNSSSVEEGVLQLQTHLDLVEEDEEEEKEKDREGD
Sbjct: 1 MTEVLQLPSPSRRCSNDGSHQTALINNSSSVEEGVLQLQTHLDLVEEDEEEEKEKDREGD 60
Query: 61 QLSLLTLLIATFRKSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRFHGFLGL 120
QLSLLTLLIAT RKSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRFHGFLGL
Sbjct: 61 QLSLLTLLIATLRKSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRFHGFLGL 120
Query: 121 PVEFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGI 180
PVEFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGI
Sbjct: 121 PVEFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGI 180
Query: 181 FRINAENSQEELVREQLNRGVVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQSQS 240
FRINAENSQEELVREQLNRGVVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQSQS
Sbjct: 181 FRINAENSQEELVREQLNRGVVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQSQS 240
Query: 241 EEECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHMADPLTA 300
EEECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHMADPLTA
Sbjct: 241 EEECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHMADPLTA 300
Query: 301 LMYAVQVMSFLKTLVVKTLREREESIVKSNPVPNLNSFDDDGHQSDSQVLPKDGSENGND 360
LMYAVQVM+FLKTLVVKTLR REESIVKSNPVPNLNSFDDDGHQSDSQVLPKDGSENGND
Sbjct: 301 LMYAVQVMNFLKTLVVKTLRVREESIVKSNPVPNLNSFDDDGHQSDSQVLPKDGSENGND 360
Query: 361 CSDEDTVFVSAEPSQPSPTHHTEDGCETESGSETSPTPAENFLSSGSRLLIDSCPCNVVS 420
CSDEDTVFVSAEPSQPSPTHHTEDGCETESGSETSPTPAENFLSSGSRLLIDSCPCNVVS
Sbjct: 361 CSDEDTVFVSAEPSQPSPTHHTEDGCETESGSETSPTPAENFLSSGSRLLIDSCPCNVVS 420
Query: 421 QLCSFAIGLQDSSIATGQAKISRSKSLQMSTSDIDKSFKNVIEFPVVGPAEKNRGTAIIG 480
QLCSFAIGLQDSSIATGQAKISRSKSLQMSTSDIDKSFKNVIEFPVVGPAEKNRGTAIIG
Sbjct: 421 QLCSFAIGLQDSSIATGQAKISRSKSLQMSTSDIDKSFKNVIEFPVVGPAEKNRGTAIIG 480
Query: 481 RINSRTELTEAWR 493
RINSRTELTEAWR
Sbjct: 481 RINSRTELTEAWR 493
>UniRef100_Q9FMR1 Rac GTPase activating protein [Arabidopsis thaliana]
Length = 466
Score = 444 bits (1143), Expect = e-123
Identities = 259/497 (52%), Positives = 316/497 (63%), Gaps = 65/497 (13%)
Query: 8 PSPSRRCSNDGSHQTALINNSSSVEEGVLQLQTHLDLVEEDEEEE----------KEKDR 57
PSPS S+ T LI++ + V + +D EE++ +E D
Sbjct: 24 PSPSSLSYASRSNATLLISSDHNRRNPVARFDQDVDFHASIEEQDLRRRSSTDGGEEDDG 83
Query: 58 EGDQLSLLTLLIATFRKSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRFHGF 117
DQ+SLL LL+A FR+SLI SC ++ R+ SMEIGWP+NVRHVAHVTFDRF+GF
Sbjct: 84 GEDQISLLALLVAIFRRSLI-SCKSNRREL-----CSMEIGWPTNVRHVAHVTFDRFNGF 137
Query: 118 LGLPVEFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQA 177
LGLPVEFEPEVPRR PSAS +VFGVSTESMQLS+D+RGN VPTILLLMQ LY++GGLQA
Sbjct: 138 LGLPVEFEPEVPRRAPSASATVFGVSTESMQLSYDSRGNCVPTILLLMQNCLYSQGGLQA 197
Query: 178 EGIFRINAENSQEELVREQLNRGVVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQ 237
EGIFR+ AENS+EE VREQLNRG +P +DVHCLAGLIKAWFRELPT +LD LSPE+VMQ
Sbjct: 198 EGIFRLTAENSEEEAVREQLNRGFIPERIDVHCLAGLIKAWFRELPTSVLDSLSPEQVMQ 257
Query: 238 SQSEEECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHMADP 297
Q+EEE +LVRLLPPTEAALLDWAINLMADV Q EH NKMN+RNIAMVFAPNMT M DP
Sbjct: 258 CQTEEENVELVRLLPPTEAALLDWAINLMADVVQYEHLNKMNSRNIAMVFAPNMTQMDDP 317
Query: 298 LTALMYAVQVMSFLKTLVVKTLREREESIVKSNPVPNLNSFDDDGHQSDSQVLPKDGSEN 357
LTALMYAVQVM+FLKTL+ KTLRER++S+V+ L D+ GHQS SQ L
Sbjct: 318 LTALMYAVQVMNFLKTLIEKTLRERQDSVVEQAHAFPLEPSDESGHQSPSQSL------- 370
Query: 358 GNDCSDEDTVFVSAEPSQPSPTHHTEDGCETESGSETSPTPAENFLSSGSRLLIDSCPCN 417
F ++E S+ + + + E+ S SE S L +++ C
Sbjct: 371 ---------AFNTSEQSEETQSDNIENAENQSSSSEIS-----------DELTLENNAC- 409
Query: 418 VVSQLCSFAIGLQDSSIATGQAKISR-SKSLQMSTSDIDKSFKNVIEFPVVGPAEKNRGT 476
+ G+ + R S S Q ++D P P + +G
Sbjct: 410 ------------EQRETDFGKYRTGRLSDSSQQVVLNLDP--------PAQWPVGRTKGL 449
Query: 477 AIIGRINSRTELTEAWR 493
+ R+ SR E TEAWR
Sbjct: 450 TNLSRVGSRVERTEAWR 466
>UniRef100_Q9CAX8 Putative rac GTPase activating protein; 62102-60058 [Arabidopsis
thaliana]
Length = 435
Score = 424 bits (1091), Expect = e-117
Identities = 240/449 (53%), Positives = 293/449 (64%), Gaps = 61/449 (13%)
Query: 46 EEDEEEEKEKDREGDQLS-LLTLLIATFRKSLIGSCSTSPRDSGALSSSSMEIGWPSNVR 104
EE+EEE EK+RE +LS L +L++ R+S+IG C G SMEIG P++VR
Sbjct: 47 EEEEEERSEKERERFELSSALEILVSAIRRSVIGGCV------GEEDLCSMEIGVPTDVR 100
Query: 105 HVAHVTFDRFHGFLGLPVEFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLL 164
HVAHVTFDRFHGFLGLPVEFEPEVPRR PSAS +VFGVSTESMQLS+D RGN VPTILL+
Sbjct: 101 HVAHVTFDRFHGFLGLPVEFEPEVPRRAPSASATVFGVSTESMQLSYDTRGNIVPTILLM 160
Query: 165 MQRHLYARGGLQAEGIFRINAENSQEELVREQLNRGVVPNGVDVHCLAGLIKAWFRELPT 224
MQ HLY+RGGL+ EGIFRIN EN QEE +RE+LN+G++P+ +DVHCLA LIKAWFRELP+
Sbjct: 161 MQSHLYSRGGLRVEGIFRINGENGQEEYIREELNKGIIPDNIDVHCLASLIKAWFRELPS 220
Query: 225 GILDPLSPEEVMQSQSEEECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIA 284
G+LD LSPE+VM+S+SE+EC +LVRLLP TEA+LLDWAINLMADV +ME NKMNARNIA
Sbjct: 221 GVLDSLSPEQVMESESEDECVELVRLLPSTEASLLDWAINLMADVVEMEQLNKMNARNIA 280
Query: 285 MVFAPNMTHMADPLTALMYAVQVMSFLKTLVVKTLREREESIVKSNPVPNLNSFDDDGHQ 344
MVFAPNMT M DPLTALMYAVQVM+FLKTL+VKTL++R+ES K P N + D +G Q
Sbjct: 281 MVFAPNMTQMLDPLTALMYAVQVMNFLKTLIVKTLKDRKESRDKLVPASNPSPRDHNGDQ 340
Query: 345 SDSQVLPKDGSENGNDCSDEDTVFVSAEPSQPSPTHHTEDGCETESGSETSPTPAENFLS 404
S S+ L N + T D E E + E
Sbjct: 341 SSSRQLLHLMKANKEE---------------------TLDNFEAEMKDKEESADEEEEEC 379
Query: 405 SGSRLLIDSCPCNVVSQLCSFAIGLQDSSIATGQAKISRSKSLQMSTSDIDKSFKNVIEF 464
+ S L+D ++V+ +SS GQ I + MS
Sbjct: 380 AESVELVDIKKSSLVN----------NSSGGFGQKHIGWEEQRTMS-------------- 415
Query: 465 PVVGPAEKNRGTAIIGRINSRTELTEAWR 493
+ ++I+GR+N R EL EAWR
Sbjct: 416 ---------KASSIVGRVNYRVELFEAWR 435
>UniRef100_O82459 Rac GTPase activating protein 2 [Lotus japonicus]
Length = 424
Score = 384 bits (986), Expect = e-105
Identities = 205/355 (57%), Positives = 256/355 (71%), Gaps = 25/355 (7%)
Query: 60 DQLSLLTLLIATFRKSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRFHGFLG 119
+Q ++L +L+A +KSL+ +CS D SS++I WP+ VRHV+HVTFDRF+GFLG
Sbjct: 21 NQFAILDILVAALKKSLV-TCSVDREDV-----SSLDISWPTEVRHVSHVTFDRFNGFLG 74
Query: 120 LPVEFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEG 179
LP E +PEVP++ P+AS VFGVS +SMQ S+D RGNSVPTILL+MQ LY+ GGL+AEG
Sbjct: 75 LPTELQPEVPQKVPTASAKVFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEG 134
Query: 180 IFRINAENSQEELVREQLNRGVVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQSQ 239
IFRINA+NSQEE VR QLNRG+VP GV+VHCL+GLIKAWFRELPTG+LD L+PE+VM
Sbjct: 135 IFRINADNSQEEFVRCQLNRGLVPRGVEVHCLSGLIKAWFRELPTGVLDSLTPEQVMHCN 194
Query: 240 SEEECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHMADPLT 299
SEE+C LV+LLP TEAALLDWAINLMADV + E FNKMNARNIAMVFAPNMT M DPLT
Sbjct: 195 SEEDCTNLVKLLPSTEAALLDWAINLMADVVEHEQFNKMNARNIAMVFAPNMTQMVDPLT 254
Query: 300 ALMYAVQVMSFLKTLVVKTLREREESIVKSNPVPNL----------NSFDDDGHQSDSQ- 348
AL++AVQVM+FLKTL++KTLRER+ES+ K+ + +L + F D+ +S +Q
Sbjct: 255 ALIHAVQVMNFLKTLILKTLRERDESMAKARQLSSLLNSPSCKGDSHPFKDNREESSAQP 314
Query: 349 -----VLPKDGSENGND--CSDEDTVFVSAEPSQPSPTHHTEDGCETESGSETSP 396
+P D SE C DE V+ S E + G + S E+ P
Sbjct: 315 VDTCATMPPDKSEFSRMEWCVDE-KVWSSEEKGTGGGALESVSGGSSPSRYESGP 368
>UniRef100_Q9ZNR7 Putative rac GTPase activating protein [Arabidopsis thaliana]
Length = 424
Score = 379 bits (973), Expect = e-103
Identities = 202/354 (57%), Positives = 253/354 (71%), Gaps = 13/354 (3%)
Query: 50 EEEKEKDREGDQLSLLTLLIATFRKSLIGSCSTSPRDS-----GALSSS--SMEIGWPSN 102
EEE+E+++ QLSL+ L+ RKS++ SC R G +SS+ MEIGWP+N
Sbjct: 21 EEEEEEEQNQQQLSLVEFLLTALRKSVV-SCRVDNRQDDGGVGGGISSAVHHMEIGWPTN 79
Query: 103 VRHVAHVTFDRFHGFLGLPVEFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTIL 162
VRH+ HVTFDRFHGFLGLP E + E+P R PSAS SVFGVS ESMQ S+D +GNSVPTIL
Sbjct: 80 VRHITHVTFDRFHGFLGLPHELQVEIPCRVPSASVSVFGVSAESMQCSYDEKGNSVPTIL 139
Query: 163 LLMQRHLYARGGLQAEGIFRINAENSQEELVREQLNRGVVPNGVDVHCLAGLIKAWFREL 222
LLMQ LY++ GL+AEGIFRIN ENSQEE VR+QLNRG+VP +DVHCLAGLIKAWFREL
Sbjct: 140 LLMQERLYSQQGLKAEGIFRINPENSQEEHVRDQLNRGIVPENIDVHCLAGLIKAWFREL 199
Query: 223 PTGILDPLSPEEVMQSQSEEECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARN 282
P+G+LD LSPEEV+ +E+E +L++ L PTE+ALL+WA++LMADV + E NKMNARN
Sbjct: 200 PSGVLDGLSPEEVLNCNTEDESVELIKQLKPTESALLNWAVDLMADVVEEEESNKMNARN 259
Query: 283 IAMVFAPNMTHMADPLTALMYAVQVMSFLKTLVVKTLREREESIVKS---NPVPNLNS-F 338
IAMVFAPNMT M DPLTALM+AVQVM+ LKTL+ KTL EREE+ S +P + NS
Sbjct: 260 IAMVFAPNMTQMTDPLTALMHAVQVMNLLKTLITKTLAEREENATGSEGYSPSHSSNSQT 319
Query: 339 DDDGHQSDSQVLPKDGSENGNDCSDEDTVFVSAEPSQPSPTHHTEDGCETESGS 392
D D + + + ++C +E+ V E Q + H+ ET+ GS
Sbjct: 320 DSDSDNAQDMEVSCESQATDSECGEEEEV-EEVEQHQEHLSRHSTHEDETDIGS 372
>UniRef100_Q8GYY5 Putative rac GTPase activating protein [Arabidopsis thaliana]
Length = 455
Score = 372 bits (956), Expect = e-101
Identities = 195/340 (57%), Positives = 243/340 (71%), Gaps = 10/340 (2%)
Query: 26 NNSSSVEEGVLQLQTHLDLVEEDEEEEKEKDREGD-----QLSLLTLLIATFRKSLIGSC 80
N+ EEG T D + R G+ QL+++ LL A RKSL+ SC
Sbjct: 33 NDDDEYEEG--HSTTSTDYYDASTPLSSHASRSGNGSGSGQLTVVDLLAAVLRKSLVMSC 90
Query: 81 STSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRFHGFLGLPVEFEPEVPRRPPSASTSVF 140
+ + ++S M+IGWP+ V+HV+HVTFDRF+GFLGLP E EPEVP R PSAS SVF
Sbjct: 91 AMERGEDDVVAS--MDIGWPTEVKHVSHVTFDRFNGFLGLPSELEPEVPPRAPSASVSVF 148
Query: 141 GVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVREQLNRG 200
GVS +SMQ S+D RGNSVPTILL MQ+ LY GGL+AEGIFRIN +N +EE VR QLN G
Sbjct: 149 GVSAKSMQCSYDDRGNSVPTILLRMQKRLYTEGGLKAEGIFRINPDNGKEEHVRRQLNCG 208
Query: 201 VVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQSQSEEECDQLVRLLPPTEAALLD 260
VVP G+DVHCLAGLIKAWFRELPTG+LD L+PE+VM+ +EE+C +LV LLPP E+A+LD
Sbjct: 209 VVPRGIDVHCLAGLIKAWFRELPTGVLDVLTPEQVMRCNTEEDCSRLVILLPPVESAILD 268
Query: 261 WAINLMADVAQMEHFNKMNARNIAMVFAPNMTHMADPLTALMYAVQVMSFLKTLVVKTLR 320
WAI LMADV + E FNKMNARN+AMVFAPNMT MADPLTAL++AVQVM+FLKTL++ L+
Sbjct: 269 WAIGLMADVVEHEQFNKMNARNVAMVFAPNMTQMADPLTALIHAVQVMNFLKTLILMNLK 328
Query: 321 EREESIVKSNPVPNLNSFDDDGHQSD-SQVLPKDGSENGN 359
ERE + K+ + S + +S S++L + N N
Sbjct: 329 ERENADAKARWLKKQTSDPSEEWESQHSEILSPEKPNNNN 368
>UniRef100_Q7XAL7 Rac GTPase activating protein 3-like protein [Oryza sativa]
Length = 487
Score = 370 bits (950), Expect = e-101
Identities = 188/307 (61%), Positives = 230/307 (74%), Gaps = 2/307 (0%)
Query: 54 EKDREGDQLSLLTLLIATFRKSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDR 113
E EG Q +L LL+A R+S++ C + D A + MEIGWP++VRHVAHVTFDR
Sbjct: 33 EGKAEGQQGQVLALLLAALRRSVVLPCQMADADDPAAVAWGMEIGWPTDVRHVAHVTFDR 92
Query: 114 FHGFLGLPVEFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARG 173
+GFLGLP EFE E+P PSAS SVFGVS ESMQ FD GNSVP ILLLMQ LYA+
Sbjct: 93 LNGFLGLPAEFELEIPGHVPSASASVFGVSPESMQCCFDDNGNSVPKILLLMQERLYAQD 152
Query: 174 GLQAEGIFRINAENSQEELVREQLNRGVVPNGVDVHCLAGLIKAWFRELPTGILDPLSPE 233
GL+AEGIFRI ENSQEE VREQLNRG+VP+ +DVHCLA LIKAWFRELP G+LD LSPE
Sbjct: 153 GLKAEGIFRITPENSQEENVREQLNRGLVPDDIDVHCLASLIKAWFRELPEGVLDSLSPE 212
Query: 234 EVMQSQSEEECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTH 293
+V+ +EEEC +LVRLLPPT+AALL+W + MADV + E NKMNARN+AMVFAPNMT
Sbjct: 213 QVLHCNTEEECVELVRLLPPTQAALLNWVVEFMADVVEEEESNKMNARNVAMVFAPNMTQ 272
Query: 294 MADPLTALMYAVQVMSFLKTLVVKTLREREESIVKSNPVPNLNSFDDDGHQSDSQVLPKD 353
M+DPLTALM+AVQVM+ LKTL++KTLRERE + + + + +S D+ + V +
Sbjct: 273 MSDPLTALMHAVQVMNLLKTLILKTLREREHDESEYSAISSQSSSSDELDEMHHHV--EQ 330
Query: 354 GSENGND 360
G ++G+D
Sbjct: 331 GGDSGSD 337
>UniRef100_Q6UQ72 Rho GTPase activating protein 2 [Oryza sativa]
Length = 439
Score = 358 bits (920), Expect = 1e-97
Identities = 185/289 (64%), Positives = 228/289 (78%), Gaps = 7/289 (2%)
Query: 59 GDQLSLLTLLIATFRKSLIGSCST--SPRDSGALSSSS----MEIGWPSNVRHVAHVTFD 112
G +S++ + A R+SL+ CS+ + D GA ++++ M+IG P++VRHV+HVTFD
Sbjct: 21 GGGVSVVETVAAALRRSLL-LCSSVRAAEDEGAAAAAAAAAGMQIGRPTDVRHVSHVTFD 79
Query: 113 RFHGFLGLPVEFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYAR 172
RF GFLGLP + EP+VPR PSAS SVFGVS SMQ S+D RGNSVPTILL MQ+ LY
Sbjct: 80 RFVGFLGLPADLEPDVPRPAPSASVSVFGVSPTSMQCSYDNRGNSVPTILLTMQKKLYQL 139
Query: 173 GGLQAEGIFRINAENSQEELVREQLNRGVVPNGVDVHCLAGLIKAWFRELPTGILDPLSP 232
GGLQAEGIFRINA+NSQE VREQLN GVVP+GVD+HCL GLIKAWFRELP+G+LD L+P
Sbjct: 140 GGLQAEGIFRINADNSQELHVREQLNMGVVPDGVDMHCLTGLIKAWFRELPSGVLDSLTP 199
Query: 233 EEVMQSQSEEECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMT 292
E+VM +EEEC L LPP EAALLDWAINLMADV + E++NKMNARNIAMVFAPNMT
Sbjct: 200 EQVMHCNTEEECALLASTLPPVEAALLDWAINLMADVVEHENYNKMNARNIAMVFAPNMT 259
Query: 293 HMADPLTALMYAVQVMSFLKTLVVKTLREREESIVKSNPVPNLNSFDDD 341
MADPLTAL++AVQVM+FLKTL++KT++ REE+ + S+ P+ + D
Sbjct: 260 QMADPLTALIHAVQVMNFLKTLILKTVKGREETAMPSSAFPSSSGSPSD 308
>UniRef100_Q6NKT5 At1g08340 [Arabidopsis thaliana]
Length = 331
Score = 357 bits (917), Expect = 3e-97
Identities = 179/272 (65%), Positives = 218/272 (79%), Gaps = 4/272 (1%)
Query: 95 MEIGWPSNVRHVAHVTFDRFHGFLGLPVEFEPEVPRRPPSASTSVFGVSTESMQLSFDAR 154
M+IG P+N+RHVAHVTFDRF GFLGLP EFEP+VPR+ PSAS +VFGVSTESMQLS+D+R
Sbjct: 1 MDIGGPTNIRHVAHVTFDRFDGFLGLPSEFEPDVPRKAPSASATVFGVSTESMQLSYDSR 60
Query: 155 GNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVREQLNRGVVPNGVDVHCLAGL 214
GN VP ILLL+Q LY +GGLQAEG+FRI ENS+EE VREQLN+G++P+G+DVHCLAGL
Sbjct: 61 GNCVPVILLLLQSRLYDQGGLQAEGVFRITGENSEEEFVREQLNKGIIPDGIDVHCLAGL 120
Query: 215 IKAWFRELPTGILDPLSPEEVMQSQSEEECDQLVRLLPPTEAALLDWAINLMADVAQMEH 274
IKAWFRELP G+LDPL E+VMQ +S+E+ ++VRLLP TEA+LL+WAINLMADV Q EH
Sbjct: 121 IKAWFRELPRGVLDPLPSEQVMQCESDEDFVKVVRLLPQTEASLLNWAINLMADVIQFEH 180
Query: 275 FNKMNARNIAMVFAPNMTHMADPLTALMYAVQVMSFLKTLVVKTLREREESIVKSNPVPN 334
NKMN+RN+A+VFAPNM+ MADPLTALMYAVQVM LK+L KT+RERE S S+ V
Sbjct: 181 VNKMNSRNLALVFAPNMSQMADPLTALMYAVQVMKLLKSLTEKTVREREAS---SSVVDR 237
Query: 335 LNSFD-DDGHQSDSQVLPKDGSENGNDCSDED 365
S + +DG + ++ E + DED
Sbjct: 238 RCSKEAEDGEKEKDNEEEEEDEEEEEEEEDED 269
>UniRef100_O82460 Rac GTPase activating protein 3 [Lotus japonicus]
Length = 432
Score = 356 bits (913), Expect = 1e-96
Identities = 186/313 (59%), Positives = 233/313 (74%), Gaps = 5/313 (1%)
Query: 55 KDREGDQLSLLTLLIATFRKSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRF 114
++ E +Q S L+A +KS++ +CS D + MEIGWP+NV+HV HVTFDRF
Sbjct: 1 EEAEQNQGSPAAFLLAALKKSMV-ACSVDSPDDVISAVHPMEIGWPTNVKHVTHVTFDRF 59
Query: 115 HGFLGLPVEFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGG 174
+GFLGLP+E E VP PSAS SVFGVS ESM S+D++GNSVPTILLLMQ LY++GG
Sbjct: 60 NGFLGLPLELEVHVPAPVPSASVSVFGVSAESMHCSYDSKGNSVPTILLLMQERLYSQGG 119
Query: 175 LQAEGIFRINAENSQEELVREQLNRGVVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEE 234
L AEGIFRIN EN QEE +R+QLNRGVVP+ +DVHCLAGLIKAWFRELP+G+LD LSPE+
Sbjct: 120 LMAEGIFRINPENGQEEHLRDQLNRGVVPDNIDVHCLAGLIKAWFRELPSGVLDGLSPEQ 179
Query: 235 VMQSQSEEECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHM 294
V++ +EEE QLV+ L PTE ALL+WA++LMADV + E NKM+ARNIAMVFAPNMT M
Sbjct: 180 VLECNTEEEFVQLVKQLKPTELALLNWALDLMADVVEEEEHNKMDARNIAMVFAPNMTQM 239
Query: 295 ADPLTALMYAVQVMSFLKTLVVKTLREREESIVKSNPVPNLNSFDDDGH-QSDSQVLPKD 353
+DPLTALM+AVQVM+ LKTL++KTL EREE+ + +S D + DSQ+
Sbjct: 240 SDPLTALMHAVQVMNLLKTLILKTLSEREEATTAGYSSMSSHSSDRQSEDEYDSQLEMYT 299
Query: 354 GSE---NGNDCSD 363
+E + +DC D
Sbjct: 300 SAELRGSQSDCDD 312
>UniRef100_Q7XUH1 OSJNBa0020J04.11 protein [Oryza sativa]
Length = 479
Score = 352 bits (903), Expect = 1e-95
Identities = 186/304 (61%), Positives = 221/304 (72%), Gaps = 29/304 (9%)
Query: 48 DEEEEKEKDREGDQLS------------------------LLTLLIATFRKSLIGSCSTS 83
DEEEE ++D EG + L+ ++ R+SL+ CS
Sbjct: 19 DEEEEDDEDEEGFEYEEILSDDGTDSPPPLMMQAEKGGGGLVGAVVGALRRSLV-MCSAG 77
Query: 84 P----RDSGALSSSSMEIGWPSNVRHVAHVTFDRFHGFLGLPVEFEPEVPRRPPSASTSV 139
DS MEIG P++VRHV+HVTFDRF GFLGLP + EPEVP PSAS +V
Sbjct: 78 KVGEEEDSEDEEEEGMEIGRPTDVRHVSHVTFDRFGGFLGLPADLEPEVPSPTPSASVNV 137
Query: 140 FGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVREQLNR 199
FGVS S+Q SFD +GNSVPTILL+MQR LY R GL+ EGIFRINAENSQE VR+QLN
Sbjct: 138 FGVSPTSLQCSFDHKGNSVPTILLMMQRKLYEREGLKIEGIFRINAENSQEICVRKQLNS 197
Query: 200 GVVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQSQSEEECDQLVRLLPPTEAALL 259
GVVP+ VD+HCLAGLIKAWFRELPTG+LD L+PE+VM +EE+C L +LPP EAALL
Sbjct: 198 GVVPDEVDLHCLAGLIKAWFRELPTGVLDSLTPEQVMHCNTEEDCALLASMLPPVEAALL 257
Query: 260 DWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHMADPLTALMYAVQVMSFLKTLVVKTL 319
DWAINLMADV + E++NKMNARNIAMVFAPNMT MADPLTAL++AVQVM+FLKTL++KTL
Sbjct: 258 DWAINLMADVVEHENYNKMNARNIAMVFAPNMTQMADPLTALIHAVQVMNFLKTLILKTL 317
Query: 320 RERE 323
+ERE
Sbjct: 318 KERE 321
>UniRef100_Q6UQ73 Rho GTPase activating protein 1 [Oryza sativa]
Length = 364
Score = 352 bits (903), Expect = 1e-95
Identities = 182/270 (67%), Positives = 214/270 (78%), Gaps = 5/270 (1%)
Query: 95 MEIGWPSNVRHVAHVTFDRFHGFLGLPVEFEPEVPRRPPSASTSVFGVSTESMQLSFDAR 154
MEIGWP++VRHVAHVTFDRFHGFLGLPVEFE E+P R PSAS SVFGVS ESMQ ++D +
Sbjct: 1 MEIGWPTDVRHVAHVTFDRFHGFLGLPVEFEVEMPCRVPSASASVFGVSAESMQCTYDGK 60
Query: 155 GNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVREQLNRGVVPNGVDVHCLAGL 214
GNSVPTILL MQ LYA+GGL+AEGIFRIN EN QEE VR+QLN+GVVP +DVHCLA L
Sbjct: 61 GNSVPTILLHMQERLYAQGGLKAEGIFRINPENDQEEHVRDQLNKGVVPEDIDVHCLASL 120
Query: 215 IKAWFRELPTGILDPLSPEEVMQSQSEEECDQLVRLLPPTEAALLDWAINLMADVAQMEH 274
IKAWFRELP G+LD LSPE+V+Q SE E +LV LL PT+AALL+WA+ LMADV + E
Sbjct: 121 IKAWFRELPEGVLDSLSPEQVLQCNSEGEFLELVTLLRPTQAALLNWAVELMADVVEEEE 180
Query: 275 FNKMNARNIAMVFAPNMTHMADPLTALMYAVQVMSFLKTLVVKTLREREESIVKSNPVPN 334
NKMNARNIAMVFAPNMT M+DPLTALM+AVQVM+FLKTL+++TLRER+++ S
Sbjct: 181 LNKMNARNIAMVFAPNMTQMSDPLTALMHAVQVMNFLKTLILRTLRERDDA--ASGDYTP 238
Query: 335 LNSFDDDGHQSDSQVLPKDGSENGNDCSDE 364
+S Q+D++ GSE D S E
Sbjct: 239 YSSPASSSQQNDAEYY---GSERDMDRSCE 265
>UniRef100_Q9SGC6 T23G18.20 [Arabidopsis thaliana]
Length = 409
Score = 352 bits (902), Expect = 2e-95
Identities = 179/278 (64%), Positives = 219/278 (78%), Gaps = 9/278 (3%)
Query: 94 SMEIGWPSNVRHVAHVTFDRFHGFLGLPVEFEPEVPRRPPSASTSVFGVSTESMQLSFDA 153
+M+IG P+N+RHVAHVTFDRF GFLGLP EFEP+VPR+ PSAS +VFGVSTESMQLS+D+
Sbjct: 73 AMDIGGPTNIRHVAHVTFDRFDGFLGLPSEFEPDVPRKAPSASATVFGVSTESMQLSYDS 132
Query: 154 RGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVREQLNRGVVPNGVDVHCLAG 213
RGN VP ILLL+Q LY +GGLQAEG+FRI ENS+EE VREQLN+G++P+G+DVHCLAG
Sbjct: 133 RGNCVPVILLLLQSRLYDQGGLQAEGVFRITGENSEEEFVREQLNKGIIPDGIDVHCLAG 192
Query: 214 LIK-----AWFRELPTGILDPLSPEEVMQSQSEEECDQLVRLLPPTEAALLDWAINLMAD 268
LIK AWFRELP G+LDPL E+VMQ +S+E+ ++VRLLP TEA+LL+WAINLMAD
Sbjct: 193 LIKVLVVIAWFRELPRGVLDPLPSEQVMQCESDEDFVKVVRLLPQTEASLLNWAINLMAD 252
Query: 269 VAQMEHFNKMNARNIAMVFAPNMTHMADPLTALMYAVQVMSFLKTLVVKTLREREESIVK 328
V Q EH NKMN+RN+A+VFAPNM+ MADPLTALMYAVQVM LK+L KT+RERE S
Sbjct: 253 VIQFEHVNKMNSRNLALVFAPNMSQMADPLTALMYAVQVMKLLKSLTEKTVREREAS--- 309
Query: 329 SNPVPNLNSFD-DDGHQSDSQVLPKDGSENGNDCSDED 365
S+ V S + +DG + ++ E + DED
Sbjct: 310 SSVVDRRCSKEAEDGEKEKDNEEEEEDEEEEEEEEDED 347
>UniRef100_Q9SJF6 T27G7.4 [Arabidopsis thaliana]
Length = 420
Score = 343 bits (880), Expect = 7e-93
Identities = 179/289 (61%), Positives = 219/289 (74%), Gaps = 20/289 (6%)
Query: 94 SMEIGWPSNVRHVAHVTFDRFHGFLGLPVEFEPEVPRRPPSA-----------STSVFGV 142
+M+IG P+N+RHVAHVTFDRF GFLGLP EFEP+VPR+ PSA S +VFGV
Sbjct: 73 AMDIGGPTNIRHVAHVTFDRFDGFLGLPSEFEPDVPRKAPSARFHIIILFVFGSATVFGV 132
Query: 143 STESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVREQLNRGVV 202
STESMQLS+D+RGN VP ILLL+Q LY +GGLQAEG+FRI ENS+EE VREQLN+G++
Sbjct: 133 STESMQLSYDSRGNCVPVILLLLQSRLYDQGGLQAEGVFRITGENSEEEFVREQLNKGII 192
Query: 203 PNGVDVHCLAGLIK-----AWFRELPTGILDPLSPEEVMQSQSEEECDQLVRLLPPTEAA 257
P+G+DVHCLAGLIK AWFRELP G+LDPL E+VMQ +S+E+ ++VRLLP TEA+
Sbjct: 193 PDGIDVHCLAGLIKVLVVIAWFRELPRGVLDPLPSEQVMQCESDEDFVKVVRLLPQTEAS 252
Query: 258 LLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHMADPLTALMYAVQVMSFLKTLVVK 317
LL+WAINLMADV Q EH NKMN+RN+A+VFAPNM+ MADPLTALMYAVQVM LK+L K
Sbjct: 253 LLNWAINLMADVIQFEHVNKMNSRNLALVFAPNMSQMADPLTALMYAVQVMKLLKSLTEK 312
Query: 318 TLREREESIVKSNPVPNLNSFD-DDGHQSDSQVLPKDGSENGNDCSDED 365
T+RERE S S+ V S + +DG + ++ E + DED
Sbjct: 313 TVREREAS---SSVVDRRCSKEAEDGEKEKDNEEEEEDEEEEEEEEDED 358
>UniRef100_Q8LHB9 Putative rac GTPase activating protein [Oryza sativa]
Length = 258
Score = 313 bits (801), Expect = 9e-84
Identities = 157/235 (66%), Positives = 185/235 (77%), Gaps = 1/235 (0%)
Query: 88 GALSSSSMEIGWPSNVRHVAHVTFDRFHGFLGLPVEFEPEVPRRPPSASTSVFGVSTESM 147
G MEIGWP++VRHVAHVTFDRFHGF GLPVE +PEV PSAS +VFGVSTESM
Sbjct: 18 GGEQQPPMEIGWPTDVRHVAHVTFDRFHGFQGLPVELQPEVAGNAPSASKTVFGVSTESM 77
Query: 148 QLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVREQLNRGVVP-NGV 206
Q S+DARGNSVP+ILLLMQR LY +GGL+AEGIFRI A+++QE+ VREQLN GV+P GV
Sbjct: 78 QCSYDARGNSVPSILLLMQRRLYEQGGLKAEGIFRIAADDAQEQAVREQLNSGVLPEGGV 137
Query: 207 DVHCLAGLIKAWFRELPTGILDPLSPEEVMQSQSEEECDQLVRLLPPTEAALLDWAINLM 266
DVHCLAGLIKAWFRELP G+LD L EV + QS ++C +L LP +AALLDWA+ LM
Sbjct: 138 DVHCLAGLIKAWFRELPGGMLDSLPAAEVTRCQSADDCARLCARLPAAKAALLDWAVQLM 197
Query: 267 ADVAQMEHFNKMNARNIAMVFAPNMTHMADPLTALMYAVQVMSFLKTLVVKTLRE 321
ADVA+ E NKM +RN+AMVFAPNMTH DP TAL +AV VM+FL L+ + L +
Sbjct: 198 ADVAREERSNKMGSRNVAMVFAPNMTHAMDPFTALKHAVHVMNFLTMLIDRALND 252
>UniRef100_Q9ZQH5 Putative rac GTPase activating protein [Arabidopsis thaliana]
Length = 368
Score = 290 bits (743), Expect = 5e-77
Identities = 155/307 (50%), Positives = 202/307 (65%), Gaps = 34/307 (11%)
Query: 94 SMEIGWPSNVRHVAHVTFDRFHGFLGLPVEFEPEVPRRPPSASTSVFGVSTESMQLSFDA 153
+M+I P+N+ HVAHVT+DRF GFLGLP EFEP+VP++PPSAS +VFGVSTESMQLS+D+
Sbjct: 86 AMDISRPTNISHVAHVTYDRFDGFLGLPSEFEPDVPKKPPSASATVFGVSTESMQLSYDS 145
Query: 154 RGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVREQLNRGVVPNGVDVHCLAG 213
RGN VPTIL L+Q LY +GGLQ EGIFRI +NS+EE +RE+LN+GV+P G+D+HCLAG
Sbjct: 146 RGNCVPTILTLLQSRLYDQGGLQVEGIFRITGDNSEEEFIREELNKGVLPEGIDIHCLAG 205
Query: 214 LIKAWFRELPTGILDPLSPEEVMQSQSEEECDQLVRLLPPTEAALLDWAINLMADVAQME 273
LIKAWFRELP G+LD L ++VMQ +S E+ V E
Sbjct: 206 LIKAWFRELPKGVLDSLPSQQVMQCESGEDF------------------------VKVFE 241
Query: 274 HFNKMNARNIAMVFAPNMTHMADPLTALMYAVQVMSFLKTLVVKTLREREESIVKSNPVP 333
NKM +RN+A+VFAPNM+ MADPLTALMYAVQVM+ L+ L KTLRER+ + +P
Sbjct: 242 VVNKMTSRNLALVFAPNMSQMADPLTALMYAVQVMNLLRNLTDKTLRERKIATSNVDPCD 301
Query: 334 NLNSFDDDGHQSDSQ-----VLPKDGSENGNDCSDEDT-----VFVSAEPSQPSPTHHTE 383
N + +D + +Q VL + E G D D D + + A+ +PS +
Sbjct: 302 NRSEAEDGNVEEYNQEVEIYVLEEVEEEEGEDVDDLDNEESERITLLADEHKPSSAVNAN 361
Query: 384 DGCETES 390
D + E+
Sbjct: 362 DRKKQET 368
>UniRef100_Q6EPP2 Putative Rho GTPase activating protein 2 [Oryza sativa]
Length = 326
Score = 276 bits (707), Expect = 7e-73
Identities = 137/195 (70%), Positives = 162/195 (82%)
Query: 147 MQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVREQLNRGVVPNGV 206
MQ S+D RGNSVPTILL MQ+ LY GGLQAEGIFRINA+NSQE VREQLN GVVP+GV
Sbjct: 1 MQCSYDNRGNSVPTILLTMQKKLYQLGGLQAEGIFRINADNSQELHVREQLNMGVVPDGV 60
Query: 207 DVHCLAGLIKAWFRELPTGILDPLSPEEVMQSQSEEECDQLVRLLPPTEAALLDWAINLM 266
D+HCL GLIKAWFRELP+G+LD L+PE+VM +EEEC L LPP EAALLDWAINLM
Sbjct: 61 DMHCLTGLIKAWFRELPSGVLDSLTPEQVMHCNTEEECALLASTLPPVEAALLDWAINLM 120
Query: 267 ADVAQMEHFNKMNARNIAMVFAPNMTHMADPLTALMYAVQVMSFLKTLVVKTLREREESI 326
ADV + E++NKMNARNIAMVFAPNMT MADPLTAL++AVQVM+FLKTL++KT++ REE+
Sbjct: 121 ADVVEHENYNKMNARNIAMVFAPNMTQMADPLTALIHAVQVMNFLKTLILKTVKGREETA 180
Query: 327 VKSNPVPNLNSFDDD 341
+ S+ P+ + D
Sbjct: 181 MPSSAFPSSSGSPSD 195
>UniRef100_Q9ZUZ1 Putative rac GTPase activating protein [Arabidopsis thaliana]
Length = 301
Score = 272 bits (695), Expect = 2e-71
Identities = 136/214 (63%), Positives = 167/214 (77%), Gaps = 1/214 (0%)
Query: 147 MQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVREQLNRGVVPNGV 206
MQ S+D RGNSVPTILL MQ+ LY GGL+AEGIFRIN +N +EE VR QLN GVVP G+
Sbjct: 1 MQCSYDDRGNSVPTILLRMQKRLYTEGGLKAEGIFRINPDNGKEEHVRRQLNCGVVPRGI 60
Query: 207 DVHCLAGLIKAWFRELPTGILDPLSPEEVMQSQSEEECDQLVRLLPPTEAALLDWAINLM 266
DVHCLAGLIKAWFRELPTG+LD L+PE+VM+ +EE+C +LV LLPP E+A+LDWAI LM
Sbjct: 61 DVHCLAGLIKAWFRELPTGVLDVLTPEQVMRCNTEEDCSRLVILLPPVESAILDWAIGLM 120
Query: 267 ADVAQMEHFNKMNARNIAMVFAPNMTHMADPLTALMYAVQVMSFLKTLVVKTLREREESI 326
ADV + E FNKMNARN+AMVFAPNMT MADPLTAL++AVQVM+FLKTL++ L+ERE +
Sbjct: 121 ADVVEHEQFNKMNARNVAMVFAPNMTQMADPLTALIHAVQVMNFLKTLILMNLKERENAD 180
Query: 327 VKSNPVPNLNSFDDDGHQSD-SQVLPKDGSENGN 359
K+ + S + +S S++L + N N
Sbjct: 181 AKARWLKKQTSDPSEEWESQHSEILSPEKPNNNN 214
>UniRef100_UPI00004996C7 UPI00004996C7 UniRef100 entry
Length = 324
Score = 99.0 bits (245), Expect = 3e-19
Identities = 61/201 (30%), Positives = 108/201 (53%), Gaps = 10/201 (4%)
Query: 135 ASTSVFGVSTESMQ-LSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELV 193
+ + VFGV ES++ R +P I++ + +GG +EG+FR+ E + +
Sbjct: 121 SESGVFGVDPESLEWFKHPTRDLILPQIIITLDIAFREKGGFTSEGVFRLAGEQGMVKSL 180
Query: 194 REQLNR--GVVPNGV---DVHCLAGLIKAWFRELPTGILDPLSPEEVMQSQSEEECDQLV 248
+E+LN+ G++ + V ++ LIK WFRELP IL+ LS +++ S E +
Sbjct: 181 KERLNKSNGIITEDMMDATVDDISNLIKLWFRELPRPILNVLSCDQIFYSTEPAESYKAF 240
Query: 249 RLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNM--THMADPLTALMYAVQ 306
L ALL W +LM +VA+ NKM +N+A+V APN+ DP+ L+ + +
Sbjct: 241 ESLNEKSRALLTWLFDLMIEVAKNRETNKMTIQNLAIVIAPNLYEPESTDPMEGLVMSQK 300
Query: 307 VMSFLKTLV--VKTLREREES 325
+ F++ ++ ++ L+E+ S
Sbjct: 301 AVQFVQNILNYLEALKEQNGS 321
>UniRef100_UPI000007CCEF UPI000007CCEF UniRef100 entry
Length = 1309
Score = 84.7 bits (208), Expect = 5e-15
Identities = 89/316 (28%), Positives = 150/316 (47%), Gaps = 34/316 (10%)
Query: 28 SSSVEEGVLQLQTHLDLVEEDEEEEKEKDR-EGDQLSLLTLLIA----TFRKSLIGSCST 82
S+S+ + L+ + ++ L + E K G +L + L TF+ +L+ +
Sbjct: 988 SASLPQQQLRDELYVQLCRQTTENPKRDSLIRGWELMAICLSYVPPSPTFQPTLLNYVNR 1047
Query: 83 SPRDSGALSSSSMEIG-WPSNVR--HVAHVTFDRFH--GFLGLPVEFEPEVPR----RPP 133
RD + ++ ME+ WP +V+ H A V R G G + +P V R
Sbjct: 1048 H-RDP-SFATCFMEVSKWPIHVQISHYATVCCRRLDRIGSSGRRMAKKPTVDEVEQARQQ 1105
Query: 134 SASTSVFGVS-TESMQLSFDARG-NSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEE 191
S+FG + +E M L + +P I + H+ G Q EGIFR++A+ +
Sbjct: 1106 ILRNSMFGNTLSEVMDLQKEKFPFRKLPWIQTTLSEHVLLLNGKQTEGIFRVSADVDEVG 1165
Query: 192 LVREQLNRGVVPNG----VDVHCLAGLIKAWFRELPTGILDPLSP----EEVMQSQSEEE 243
++ +L+R VP+ VD H A L+K W+REL DPL P E+ + ++ ++
Sbjct: 1166 CMKNRLDRWDVPDYKNTLVDAHTPASLLKLWYREL----YDPLIPDAYYEDCVNTEDPDK 1221
Query: 244 CDQLVRLLPPTEAALLDWAINLMADVA--QMEHFNKMNARNIAMVFAPNMTHMA--DPLT 299
++V LP +L + I+ + A ++ KM++ N+AMVFAPN DP
Sbjct: 1222 AKEIVNKLPEINQLVLTYLIHFLQQFAIPEVVSCTKMDSSNLAMVFAPNCLRCTSEDPKV 1281
Query: 300 ALMYAVQVMSFLKTLV 315
L A + MSF++TL+
Sbjct: 1282 ILENARKEMSFMRTLI 1297
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.313 0.130 0.370
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 800,068,538
Number of Sequences: 2790947
Number of extensions: 33872109
Number of successful extensions: 126350
Number of sequences better than 10.0: 1177
Number of HSP's better than 10.0 without gapping: 189
Number of HSP's successfully gapped in prelim test: 996
Number of HSP's that attempted gapping in prelim test: 124589
Number of HSP's gapped (non-prelim): 1486
length of query: 493
length of database: 848,049,833
effective HSP length: 131
effective length of query: 362
effective length of database: 482,435,776
effective search space: 174641750912
effective search space used: 174641750912
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 77 (34.3 bits)
Lotus: description of TM0010.6


