

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0010.3.5
(80 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
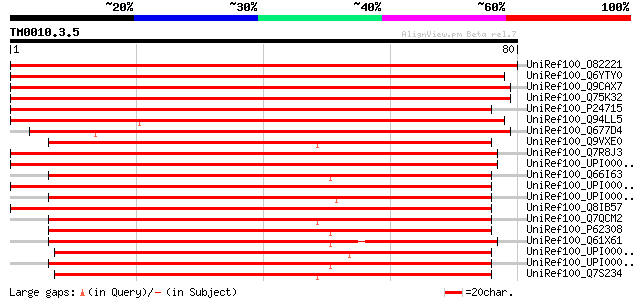
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O82221 Probable small nuclear ribonucleoprotein G [Ara... 145 2e-34
UniRef100_Q6YTY0 Putative small nuclear ribonucleoprotein E [Ory... 143 9e-34
UniRef100_Q9CAX7 Putative small nuclear ribonucleoprotein polype... 143 9e-34
UniRef100_Q75K32 Putative small nuclear ribonucleoprotein polype... 140 6e-33
UniRef100_P24715 Probable small nuclear ribonucleoprotein G [Med... 138 3e-32
UniRef100_Q94LL5 Putative small nuclear ribonucleoprotein G [Ory... 133 1e-30
UniRef100_Q677D4 Small nuclear ribonucleoprotein polypeptide G [... 120 9e-27
UniRef100_Q9VXE0 Probable small nuclear ribonucleoprotein G [Dro... 92 4e-18
UniRef100_Q7R8J3 Putative small nuclear ribonucleoprotein polype... 91 6e-18
UniRef100_UPI000046C1BF UPI000046C1BF UniRef100 entry 91 7e-18
UniRef100_Q66I63 Zgc:103688 [Brachydanio rerio] 90 1e-17
UniRef100_UPI00004679EB UPI00004679EB UniRef100 entry 89 2e-17
UniRef100_UPI000027CF91 UPI000027CF91 UniRef100 entry 89 3e-17
UniRef100_Q8IB57 Small nuclear ribonucleoprotein, putative [Plas... 89 4e-17
UniRef100_Q7QCM2 ENSANGP00000010868 [Anopheles gambiae str. PEST] 88 5e-17
UniRef100_P62308 Small nuclear ribonucleoprotein G [Homo sapiens] 87 8e-17
UniRef100_Q61X61 Hypothetical protein CBG04114 [Caenorhabditis b... 86 2e-16
UniRef100_UPI000023D520 UPI000023D520 UniRef100 entry 84 7e-16
UniRef100_UPI000021F596 UPI000021F596 UniRef100 entry 84 9e-16
UniRef100_Q7S234 Hypothetical protein [Neurospora crassa] 84 1e-15
>UniRef100_O82221 Probable small nuclear ribonucleoprotein G [Arabidopsis thaliana]
Length = 80
Score = 145 bits (366), Expect = 2e-34
Identities = 70/80 (87%), Positives = 76/80 (94%)
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
MSRSGQPPDLKKYMDK LQIKLNANRM+ GTLRG+DQFMN+VVDNTVEVNGN+K DIGMV
Sbjct: 1 MSRSGQPPDLKKYMDKKLQIKLNANRMVTGTLRGFDQFMNLVVDNTVEVNGNDKTDIGMV 60
Query: 61 VIRGNSVVTVEALEPVTRTS 80
VIRGNS+VTVEALEPV R+S
Sbjct: 61 VIRGNSIVTVEALEPVGRSS 80
>UniRef100_Q6YTY0 Putative small nuclear ribonucleoprotein E [Oryza sativa]
Length = 80
Score = 143 bits (361), Expect = 9e-34
Identities = 68/78 (87%), Positives = 75/78 (95%)
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
MSRSGQPPDLKKYMDK LQIKLNANR+++GTLRG+DQFMN+VVDNTVEVNGNEKNDIGMV
Sbjct: 1 MSRSGQPPDLKKYMDKKLQIKLNANRVVIGTLRGFDQFMNLVVDNTVEVNGNEKNDIGMV 60
Query: 61 VIRGNSVVTVEALEPVTR 78
VIRGNSVV +EALEPV +
Sbjct: 61 VIRGNSVVMIEALEPVPK 78
>UniRef100_Q9CAX7 Putative small nuclear ribonucleoprotein polypeptide G;
65009-64161 [Arabidopsis thaliana]
Length = 79
Score = 143 bits (361), Expect = 9e-34
Identities = 69/79 (87%), Positives = 76/79 (95%)
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
MSRSGQPPDLKKYMDK LQIKLNANRM+VGTLRG+DQFMN+VVDNTVEVNG++K DIGMV
Sbjct: 1 MSRSGQPPDLKKYMDKKLQIKLNANRMVVGTLRGFDQFMNLVVDNTVEVNGDDKTDIGMV 60
Query: 61 VIRGNSVVTVEALEPVTRT 79
VIRGNS+VTVEALEPV R+
Sbjct: 61 VIRGNSIVTVEALEPVGRS 79
>UniRef100_Q75K32 Putative small nuclear ribonucleoprotein polypeptide G [Oryza
sativa]
Length = 80
Score = 140 bits (354), Expect = 6e-33
Identities = 67/79 (84%), Positives = 75/79 (94%)
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
MSRSGQPPDLKKYMDK LQIKLNANR+IVGTLRG+DQFMN+VVDNTVEVNGN+K DIGMV
Sbjct: 1 MSRSGQPPDLKKYMDKKLQIKLNANRVIVGTLRGFDQFMNLVVDNTVEVNGNDKTDIGMV 60
Query: 61 VIRGNSVVTVEALEPVTRT 79
V+RGNSVV +EALEPV ++
Sbjct: 61 VVRGNSVVMIEALEPVPKS 79
>UniRef100_P24715 Probable small nuclear ribonucleoprotein G [Medicago sativa]
Length = 81
Score = 138 bits (348), Expect = 3e-32
Identities = 69/76 (90%), Positives = 71/76 (92%)
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
MS SGQPP LKKYMDK LQI L ANRMIVGTLRG+DQFMN+VVDNTVEVNGNEKNDIGMV
Sbjct: 1 MSTSGQPPALKKYMDKQLQINLKANRMIVGTLRGFDQFMNLVVDNTVEVNGNEKNDIGMV 60
Query: 61 VIRGNSVVTVEALEPV 76
VIRGNSVVTVEALEPV
Sbjct: 61 VIRGNSVVTVEALEPV 76
>UniRef100_Q94LL5 Putative small nuclear ribonucleoprotein G [Oryza sativa]
Length = 94
Score = 133 bits (335), Expect = 1e-30
Identities = 67/92 (72%), Positives = 75/92 (80%), Gaps = 14/92 (15%)
Query: 1 MSRSGQPPDLKKYMDKNLQ--------------IKLNANRMIVGTLRGYDQFMNMVVDNT 46
MSRSGQPPDLKKYMDK LQ +KLNANR+++GTLRG+DQFMN+VVDNT
Sbjct: 1 MSRSGQPPDLKKYMDKKLQTLMVYSFTLFSILPVKLNANRVVIGTLRGFDQFMNLVVDNT 60
Query: 47 VEVNGNEKNDIGMVVIRGNSVVTVEALEPVTR 78
VEVNGNEKNDIGMVVIRGNSVV +EALEPV +
Sbjct: 61 VEVNGNEKNDIGMVVIRGNSVVMIEALEPVPK 92
>UniRef100_Q677D4 Small nuclear ribonucleoprotein polypeptide G [Hyacinthus
orientalis]
Length = 96
Score = 120 bits (301), Expect = 9e-27
Identities = 59/77 (76%), Positives = 69/77 (88%), Gaps = 1/77 (1%)
Query: 4 SGQPPDLKK-YMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMVVI 62
SGQPP LK+ + LQIKLNANR++VGTLRG+DQFMN+V+DNT+EVNGN+KNDIGMVVI
Sbjct: 19 SGQPPALKQVHGTSKLQIKLNANRVVVGTLRGFDQFMNLVIDNTMEVNGNDKNDIGMVVI 78
Query: 63 RGNSVVTVEALEPVTRT 79
RGNSVV +EALEPV RT
Sbjct: 79 RGNSVVMIEALEPVART 95
>UniRef100_Q9VXE0 Probable small nuclear ribonucleoprotein G [Drosophila
melanogaster]
Length = 76
Score = 91.7 bits (226), Expect = 4e-18
Identities = 44/71 (61%), Positives = 56/71 (77%), Gaps = 1/71 (1%)
Query: 7 PPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTV-EVNGNEKNDIGMVVIRGN 65
PP++KKYMDK + +KLN R + G LRG+D FMN+V+D+TV E N KN+IGMVVIRGN
Sbjct: 6 PPEVKKYMDKRMMLKLNGGRAVTGILRGFDPFMNVVLDDTVEECKDNTKNNIGMVVIRGN 65
Query: 66 SVVTVEALEPV 76
S+V VEAL+ V
Sbjct: 66 SIVMVEALDRV 76
>UniRef100_Q7R8J3 Putative small nuclear ribonucleoprotein polypeptide G
[Plasmodium yoelii yoelii]
Length = 83
Score = 91.3 bits (225), Expect = 6e-18
Identities = 43/77 (55%), Positives = 58/77 (74%)
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
+ ++G D +K+M+K LQI LN NR+IVG LRGYD FMN+V+DNTVE+ +E +IG+V
Sbjct: 5 VGKAGPASDFRKFMEKRLQIYLNGNRLIVGVLRGYDTFMNLVLDNTVEIKKDENIEIGIV 64
Query: 61 VIRGNSVVTVEALEPVT 77
VIRGNS+ E L+ VT
Sbjct: 65 VIRGNSISYWECLDKVT 81
>UniRef100_UPI000046C1BF UPI000046C1BF UniRef100 entry
Length = 83
Score = 90.9 bits (224), Expect = 7e-18
Identities = 42/77 (54%), Positives = 58/77 (74%)
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
+ ++G D +K+M+K LQI LN NR+IVG LRGYD FMN+V+DNT+E+ +E +IG+V
Sbjct: 5 VGKAGPASDFRKFMEKRLQIYLNGNRLIVGVLRGYDTFMNLVLDNTIEIKKDENIEIGIV 64
Query: 61 VIRGNSVVTVEALEPVT 77
VIRGNS+ E L+ VT
Sbjct: 65 VIRGNSISYWECLDKVT 81
>UniRef100_Q66I63 Zgc:103688 [Brachydanio rerio]
Length = 76
Score = 90.1 bits (222), Expect = 1e-17
Identities = 43/71 (60%), Positives = 56/71 (78%), Gaps = 1/71 (1%)
Query: 7 PPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEV-NGNEKNDIGMVVIRGN 65
PP+LKK+MDK L +KLN R + G LRG+D FMN+VVD+T+E+ G + N IGMVVIRGN
Sbjct: 6 PPELKKFMDKKLSLKLNGGRHVQGILRGFDPFMNLVVDDTIEMAPGGQMNTIGMVVIRGN 65
Query: 66 SVVTVEALEPV 76
S++ +EALE V
Sbjct: 66 SIIMLEALERV 76
>UniRef100_UPI00004679EB UPI00004679EB UniRef100 entry
Length = 83
Score = 89.4 bits (220), Expect = 2e-17
Identities = 42/76 (55%), Positives = 57/76 (74%)
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
+ ++G D +K+M+K LQI LN NR+IVG LRGYD FMN+V+DNTVE+ +E +IG+V
Sbjct: 5 VGKAGPASDFRKFMEKRLQIYLNGNRLIVGVLRGYDTFMNLVLDNTVEIKKDENIEIGIV 64
Query: 61 VIRGNSVVTVEALEPV 76
VIRGNS+ E L+ V
Sbjct: 65 VIRGNSISYWECLDKV 80
>UniRef100_UPI000027CF91 UPI000027CF91 UniRef100 entry
Length = 76
Score = 89.0 bits (219), Expect = 3e-17
Identities = 42/71 (59%), Positives = 56/71 (78%), Gaps = 1/71 (1%)
Query: 7 PPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVN-GNEKNDIGMVVIRGN 65
PP+LKK+MDK L +KLN R + G LRG+D FMN+VVD T+E+ G +++ IGMVVIRGN
Sbjct: 6 PPELKKFMDKKLSLKLNGGRHVQGILRGFDPFMNLVVDETLEMGPGGQQSSIGMVVIRGN 65
Query: 66 SVVTVEALEPV 76
S++ +EALE V
Sbjct: 66 SIIMLEALERV 76
>UniRef100_Q8IB57 Small nuclear ribonucleoprotein, putative [Plasmodium falciparum]
Length = 83
Score = 88.6 bits (218), Expect = 4e-17
Identities = 41/76 (53%), Positives = 57/76 (74%)
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
+ ++G D +K+M+K LQI LN NR +VG LRGYD FMN+V+DNT+E+ +E+ DIG+V
Sbjct: 5 VGKAGPASDFRKFMEKRLQIYLNGNRQVVGILRGYDTFMNLVLDNTMEIKKDEQIDIGVV 64
Query: 61 VIRGNSVVTVEALEPV 76
VIRGNS+ E L+ V
Sbjct: 65 VIRGNSISYWECLDKV 80
>UniRef100_Q7QCM2 ENSANGP00000010868 [Anopheles gambiae str. PEST]
Length = 76
Score = 88.2 bits (217), Expect = 5e-17
Identities = 41/71 (57%), Positives = 55/71 (76%), Gaps = 1/71 (1%)
Query: 7 PPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTV-EVNGNEKNDIGMVVIRGN 65
PP+LKKYMDK L +KLN R++ G LRG+D FMN+VVD ++ E +N+IGMVVIRGN
Sbjct: 6 PPELKKYMDKRLSLKLNGGRVVSGILRGFDPFMNVVVDESIEECKDGTRNNIGMVVIRGN 65
Query: 66 SVVTVEALEPV 76
S++ VEAL+ +
Sbjct: 66 SIIMVEALDRI 76
>UniRef100_P62308 Small nuclear ribonucleoprotein G [Homo sapiens]
Length = 76
Score = 87.4 bits (215), Expect = 8e-17
Identities = 41/71 (57%), Positives = 55/71 (76%), Gaps = 1/71 (1%)
Query: 7 PPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEV-NGNEKNDIGMVVIRGN 65
PP+LKK+MDK L +KLN R + G LRG+D FMN+V+D VE+ ++N+IGMVVIRGN
Sbjct: 6 PPELKKFMDKKLSLKLNGGRHVQGILRGFDPFMNLVIDECVEMATSGQQNNIGMVVIRGN 65
Query: 66 SVVTVEALEPV 76
S++ +EALE V
Sbjct: 66 SIIMLEALERV 76
>UniRef100_Q61X61 Hypothetical protein CBG04114 [Caenorhabditis briggsae]
Length = 77
Score = 85.9 bits (211), Expect = 2e-16
Identities = 44/73 (60%), Positives = 54/73 (73%), Gaps = 3/73 (4%)
Query: 7 PPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEV--NGNEKNDIGMVVIRG 64
PP+LKKYMDK +++KLN NR + G LRG+D FMNMV+D VE NGN N +GM VIRG
Sbjct: 6 PPELKKYMDKEIELKLNGNRHVSGILRGFDPFMNMVIDEAVEYPKNGNSIN-LGMTVIRG 64
Query: 65 NSVVTVEALEPVT 77
NSVV +E E V+
Sbjct: 65 NSVVIMEPKERVS 77
>UniRef100_UPI000023D520 UPI000023D520 UniRef100 entry
Length = 84
Score = 84.3 bits (207), Expect = 7e-16
Identities = 42/70 (60%), Positives = 54/70 (77%), Gaps = 1/70 (1%)
Query: 8 PDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGN-EKNDIGMVVIRGNS 66
P+LKKY+DK L ++LN +R ++G LRGYD F+N+V+D VE N EK IGMVVIRGNS
Sbjct: 6 PELKKYLDKRLFVQLNGSRKVIGVLRGYDVFLNIVLDEAVEEKDNGEKVRIGMVVIRGNS 65
Query: 67 VVTVEALEPV 76
VV +EALE +
Sbjct: 66 VVMLEALERI 75
>UniRef100_UPI000021F596 UPI000021F596 UniRef100 entry
Length = 76
Score = 84.0 bits (206), Expect = 9e-16
Identities = 39/71 (54%), Positives = 55/71 (76%), Gaps = 1/71 (1%)
Query: 7 PPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEV-NGNEKNDIGMVVIRGN 65
PP+LKK+MDK L +KLN R + G L+G+D FM++V+D VE+ ++N+IGMVVIRGN
Sbjct: 6 PPELKKFMDKKLSLKLNGGRHVQGILQGFDPFMSLVIDECVEMATSGQQNNIGMVVIRGN 65
Query: 66 SVVTVEALEPV 76
S++ +EALE V
Sbjct: 66 SIIMLEALERV 76
>UniRef100_Q7S234 Hypothetical protein [Neurospora crassa]
Length = 85
Score = 83.6 bits (205), Expect = 1e-15
Identities = 40/70 (57%), Positives = 55/70 (78%), Gaps = 1/70 (1%)
Query: 8 PDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTV-EVNGNEKNDIGMVVIRGNS 66
P+LKKY+DK L ++LN +R ++G LRGYD F+N+V+D+ V E +G EK +GMV IRGNS
Sbjct: 6 PELKKYLDKRLFVQLNGSRKVIGVLRGYDVFLNIVLDDAVEEKDGGEKVKLGMVTIRGNS 65
Query: 67 VVTVEALEPV 76
VV +EALE +
Sbjct: 66 VVMMEALERI 75
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.314 0.133 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 125,682,097
Number of Sequences: 2790947
Number of extensions: 4175548
Number of successful extensions: 9193
Number of sequences better than 10.0: 395
Number of HSP's better than 10.0 without gapping: 204
Number of HSP's successfully gapped in prelim test: 191
Number of HSP's that attempted gapping in prelim test: 8862
Number of HSP's gapped (non-prelim): 396
length of query: 80
length of database: 848,049,833
effective HSP length: 56
effective length of query: 24
effective length of database: 691,756,801
effective search space: 16602163224
effective search space used: 16602163224
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0010.3.5


