

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0009.7
(1310 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
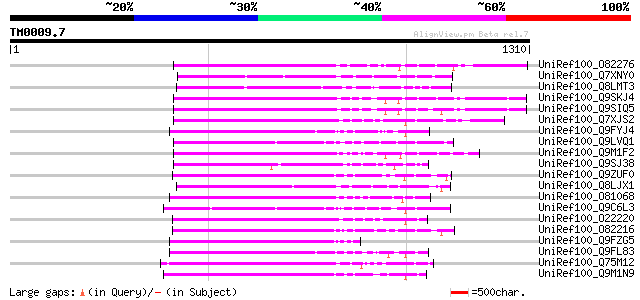
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O82276 Putative non-LTR retroelement reverse transcrip... 333 3e-89
UniRef100_Q7XNY0 OSJNBb0015N08.11 protein [Oryza sativa] 317 2e-84
UniRef100_Q8LMT3 Putative retroelement [Oryza sativa] 315 8e-84
UniRef100_Q9SKJ4 Putative non-LTR retroelement reverse transcrip... 312 4e-83
UniRef100_Q9SIQ5 Putative non-LTR retroelement reverse transcrip... 312 5e-83
UniRef100_Q7XJS2 Putative non-LTR retroelement reverse transcrip... 303 2e-80
UniRef100_Q9FYJ4 F17F8.5 [Arabidopsis thaliana] 301 1e-79
UniRef100_Q9LVQ1 Non-LTR retroelement reverse transcriptase-like... 298 7e-79
UniRef100_Q9M1F2 Hypothetical protein F9K21.130 [Arabidopsis tha... 296 3e-78
UniRef100_Q9SJ38 Putative non-LTR retroelement reverse transcrip... 293 2e-77
UniRef100_Q9ZUF0 Putative non-LTR retroelement reverse transcrip... 290 2e-76
UniRef100_Q8LJX1 Putative reverse transcriptase [Sorghum bicolor] 288 6e-76
UniRef100_O81068 Putative non-LTR retroelement reverse transcrip... 287 1e-75
UniRef100_Q9C6L3 Hypothetical protein F2J7.11 [Arabidopsis thali... 286 3e-75
UniRef100_O22220 Putative non-LTR retroelement reverse transcrip... 285 6e-75
UniRef100_O82216 Putative non-LTR retroelement reverse transcrip... 282 5e-74
UniRef100_Q9FZG5 T2E6.4 [Arabidopsis thaliana] 281 1e-73
UniRef100_Q9FL83 Non-LTR retroelement reverse transcriptase-like... 275 5e-72
UniRef100_Q75M12 Hypothetical protein P0676G05.2 [Oryza sativa] 274 1e-71
UniRef100_Q9M1N9 Hypothetical protein T18B22.50 [Arabidopsis tha... 272 6e-71
>UniRef100_O82276 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1231
Score = 333 bits (853), Expect = 3e-89
Identities = 247/920 (26%), Positives = 415/920 (44%), Gaps = 84/920 (9%)
Query: 413 DFFSSGRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRLQKVMPKL 472
+FF +G + N A + LI K + +RP+SL +K+I K++ TRL+ V+ KL
Sbjct: 337 EFFETGVLPASTNDALLVLIAKVAKPERIQQFRPVSLCNVLFKIITKMMVTRLKNVISKL 396
Query: 473 LSQNQFAFTKERQIADCILLANEVADFLSKKS--EGGLMLKVDLAKAYDQVDWEFLINML 530
+ Q +F R D I+L E + +K +G ++LK+DL KAYD+V W+FL L
Sbjct: 397 IGPAQASFIPGRLSIDNIVLVQEAVHSMRRKKGRKGWMLLKLDLEKAYDRVRWDFLQETL 456
Query: 531 HEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNGLSC 590
+ W S + + ++ S+++L NG TD F +GLRQGDPLSP LF LCL L
Sbjct: 457 EAAGLSEGWTSRIMAGVTDPSMSVLWNGERTDSFVPARGLRQGDPLSPYLFVLCLERLCH 516
Query: 591 LLNQFLGEENYCGVRIS-RNLTINHLLFADDTLLFCENNELQLETLCNVLITFLFASGLK 649
L+ +G+ + + +S ++H+ FADD +LF E + Q+ + VL F ASG K
Sbjct: 517 LIEASVGKREWKPIAVSCGGSKLSHVCFADDLILFAEASVAQIRIIRRVLERFCEASGQK 576
Query: 650 VNNQKSELIGCNVQTVEVERLAHSFGWSVGTLPI-SYLGAPLGGNPRRIVFWNPMLEKLK 708
V+ +KS++ + + E+E+L T + YLG P+ + +LE++
Sbjct: 577 VSLEKSKIFFSHNVSREMEQLISEESGIGCTKELGKYLGMPILQKRMNKETFGEVLERVS 636
Query: 709 CKTRSYNSKFMSLSGRLVITKSVLCSIPLFLMVVFKAPKGVILEVEKLFHAFIWGQEQGR 768
+ + + +SL+GR+ +TK+VL SIP+ +M P + +++ F+WG +
Sbjct: 637 ARLAGWKGRSLSLAGRITLTKAVLSSIPVHVMSAILLPVSTLDTLDRYSRTFLWGSTMEK 696
Query: 769 KKINWIGWDQICLNRKAGGLGLGFIEWRNRGLLLKWAWRFGREINSLWRFFVIERYNLDP 828
KK + + W +IC + GG+GL N+ L+ K WR ++ SLW V ++Y
Sbjct: 697 KKQHLLSWRKICKPKAEGGIGLRSARDMNKALVAKVGWRLLQDKESLWARVVRKKYK--- 753
Query: 829 DSLIWHMASTTPAMGSRAIQDIWRVLNEDHSLSAGFKENLLISI----GDGSRVKFWRDP 884
+ + T+ WR S++ G +E ++ + GDG ++FW D
Sbjct: 754 ---VGGVQDTSWLKPQPRWSSTWR------SVAVGLREVVVKGVGWVPGDGCTIRFWLDR 804
Query: 885 WVSNVALCHLFPRLYAMSENKEARIAEVGTYSRGKWGWSLTFRGQFFAWEEELYQQFLAS 944
W+ L L + E RI Y GW+L G + E ++ L S
Sbjct: 805 WLLQEPLVELGTDMIPEGE----RIKVAADYWLPGSGWNLEILGLYL---PETVKRRLLS 857
Query: 945 INHIIPAAVG-GDQLIWCANQTGLFSVKELCYWLEDHL--------FTHQMWQVPKSIAK 995
+ ++ +G GD++ W Q G F+V+ L+ + F +++W++
Sbjct: 858 V--VVQVFLGNGDEISWKGTQDGAFTVRSAYSLLQGDVGDRPNMGSFFNRIWKL------ 909
Query: 996 LLPPKVALFFWQAQNDKIATRANLLARGIDLNGSTQCLFCNAFDESCNHIFLHCHSIWSL 1055
+ P +V +F W + I T + R + N C CN +E+ H+ C ++ +
Sbjct: 910 ITPERVRVFIWLVSQNVIMTNVERVRRHLSENAI--CSVCNGAEETILHVLRDCPAMEPI 967
Query: 1056 WCVIVNRERFLWVMPASLADLILEWDFLPKISDKSLWSLIPYALSWSIWIERNNICFNHK 1115
W ++ R SL LEW F K +W + W W R F +
Sbjct: 968 WRRLLPLRRHHEFFSQSL----LEWLFTNMDPVKGIWPTLFGMGIWWAWKWRCCDVFGER 1023
Query: 1116 AL----------NISEIWDFHLLRIGWWTHELLRQNSYSIDQLSRNFESIRFASKPSLLR 1165
+ E+ H+ +G R N ++++ R
Sbjct: 1024 KICRDRLKFIKDMAEEVRRVHVGAVG------NRPNGVRVERMIR--------------- 1062
Query: 1166 AASWVTPDAGLIKINVDGAAKGSPGRCGIGGILRNNANVILGFFSKSVGEGYAFEAEVLA 1225
W P G +KI DGA++G+ G GG +RN LG F+ ++G A AE+
Sbjct: 1063 ---WQVPSDGWVKITTDGASRGNHGLAAAGGAIRNGQGEWLGGFALNIGSCAAPLAELWG 1119
Query: 1226 IHEALLFCVQRQLRNIIIESDSSLAVGWVNNRCNRPWKLLNKLNQIDRWLVETNCLKVNH 1285
+ LL + R + ++ D L VG+++ + L + + ++V+H
Sbjct: 1120 AYYGLLIAWDKGFRRVELDLDCKLVVGFLSTGVSNAHPLSFLVRLCQGFFTRDWLVRVSH 1179
Query: 1286 IYREANGEADKLAKRGAEYP 1305
+YREAN AD LA P
Sbjct: 1180 VYREANRLADGLANYAFTLP 1199
>UniRef100_Q7XNY0 OSJNBb0015N08.11 protein [Oryza sativa]
Length = 1026
Score = 317 bits (812), Expect = 2e-84
Identities = 218/706 (30%), Positives = 340/706 (47%), Gaps = 45/706 (6%)
Query: 424 MNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRLQKVMPKLLSQNQFAFTKE 483
+N ITLIPK ++AS + +RPI L+ S+K+I KVL RL M ++S+ Q AF K
Sbjct: 295 LNFGVITLIPKTKDASQIQKFRPICLLNVSFKIITKVLMNRLNGTMAYIISKQQTAFLKN 354
Query: 484 RQIADCILLANEVADFLSKKSEGGLMLKVDLAKAYDQVDWEFLINMLHEMNFGDKWISWM 543
R I + +++ +EV + + +K + G++ KVD KAYD+V+W F+ ML F D+W W+
Sbjct: 355 RFIMEGVVILHEVLNSIHQKKQSGILFKVDFEKAYDKVNWVFIYRMLKAKGFPDQWCDWI 414
Query: 544 KSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNGLSCLLNQFLGEENYC- 602
+ +A+ VN FF KGLRQGDPLSPLLFNL L+ L+ + EEN
Sbjct: 415 MKVVMGGKVAVKVNDQIGSFFKTHKGLRQGDPLSPLLFNLAAEALTLLVQR--AEENSLI 472
Query: 603 -GVRISRNLTINHLLFADDTLLFCENNELQLETLCNVLITFLFASGLKVNNQKSELIGCN 661
G+ + + I L +ADDT+ + + L +L F SGLK+N KSE+
Sbjct: 473 EGLGTNGDNKIAILQYADDTIFLINDKLDHAKNLKYILCLFEQLSGLKINFNKSEVFCFG 532
Query: 662 VQTVEVERLAHSFGWSVGTLPISYLGAPLGGNPRRIV--FWNPMLEKLKCKTRSYNSKFM 719
+ + ++ F VG+LP+ YLG P+ + +RI+ W K++ K + +
Sbjct: 533 EAKEKQDLYSNIFTCKVGSLPLKYLGIPI--DQKRILNKDWKLAENKMEHKLGCWQGRLQ 590
Query: 720 SLSGRLVITKSVLCSIPLFLMVVFKAPKGVILEVEKLFHAFIWGQEQGRKKINWIGWDQI 779
S+ GRL++ S L S+P++++ ++ PKGV ++ F+W ++QG +K + + W +
Sbjct: 591 SIGGRLILLNSTLSSVPMYMISFYRLPKGVQERIDYFRKRFLWQEDQGIRKYHLVNWPLV 650
Query: 780 CLNRKAGGLGLGFIEWRNRGLLLKWAWRFGREINSLWRFFVIERYNLDPDSLIWHMASTT 839
C R GGLG+ +E N+ +L KW WR E W+ + +Y S
Sbjct: 651 CSPRDQGGLGVLDLEAMNKAMLGKWIWRLENE-EGWWQEIIYAKY-----------CSDK 698
Query: 840 PAMGSR---AIQDIWRVLNEDHSLSAGFKENLLISIGDGSRVKFWRDPWVSNVALCHLFP 896
P G R W+ + E F + +G+G + FW D W+ L FP
Sbjct: 699 PLSGLRLKAGSSHFWQGVMEVKDDFFSFCTKI---VGNGEKTLFWEDSWLGGKPLAIQFP 755
Query: 897 RLYAMSENKEARIAEVGTYSRGKWGWSLTFRGQFFAWEEELYQQFLASINHIIPAAVGGD 956
LY + K IA++ K + FR + +++ + S + D
Sbjct: 756 SLYGIVITKRITIADLNR----KGIDCMKFRRDLHGDKLRDWRKIVNSWEGLNLVENCKD 811
Query: 957 QLIWCANQTGLFSVKELCYWLEDHLFTHQMWQVPKSIAKL-LPPKVALFFWQAQNDKIAT 1015
+L W ++ G F+V+ L+ Q K I K +P K+ +F W +KI T
Sbjct: 812 KLWWTLSKDGKFTVRSFYRALK----LQQTSFPNKKIWKFRVPLKIRIFIWFFTKNKILT 867
Query: 1016 RANLLARGIDLNGSTQCLFCNAFDESCNHIFLHC---HSIWSLWCVIVNRERFLWVMPAS 1072
+ NLL RG G +C FC+ E+ H+F C IW++ +N + L S
Sbjct: 868 KDNLLKRGW-RKGDNKCQFCDKV-ETVQHLFFDCPLARLIWNIIACALNVKPVL-----S 920
Query: 1073 LADLILEWDFLPKISDKSLWSLIPYALSWSIWIERNNICFNHKALN 1118
DL W K+L + A+ WSIW RN CF K N
Sbjct: 921 RQDLFGSWIQSMDKFTKNLVIVGIAAVLWSIWKCRNKACFERKLPN 966
>UniRef100_Q8LMT3 Putative retroelement [Oryza sativa]
Length = 764
Score = 315 bits (806), Expect = 8e-84
Identities = 212/710 (29%), Positives = 341/710 (47%), Gaps = 60/710 (8%)
Query: 422 RGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRLQKVMPKLLSQNQFAFT 481
R +N ITLIPK ++A+++ +RPI L+ +K + K+L RL ++ K++ ++Q AF
Sbjct: 29 RRLNYGVITLIPKVKDANTIKSFRPICLLNVCFKFLTKILTRRLSEIAQKVIGESQTAFI 88
Query: 482 KERQIADCILLANEVADFLSKKSEGGLMLKVDLAKAYDQVDWEFLINMLHEMNFGDKWIS 541
RQ D +++ +EV L K+ + G++ K+D KAYD+V W FL ++LH+ F D+WI
Sbjct: 89 PGRQNLDGVVILHEVLHELKKEKKSGIIFKLDFEKAYDKVQWSFLFDVLHKKGFSDRWIQ 148
Query: 542 WMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNGLSCLLNQFLGEENY 601
W+K +A+ +NG DFF +G+RQGDPLSPLLFNL + LS +LN +
Sbjct: 149 WVKMATIGGKMAVNINGEVKDFFKTYRGVRQGDPLSPLLFNLVADALSEMLN---NAKQA 205
Query: 602 CGVRISRNLTINHLLFADDTLLFCENNELQLETLCNVLITFLFASGLKVNNQKSELIGCN 661
+ LT HL +ADDT+LF N E + T+ +L + SGLK+N QKSE++
Sbjct: 206 VPHLVPGGLT--HLQYADDTILFMTNTEENIVTVKFLLYCYEAMSGLKINYQKSEIMVIG 263
Query: 662 VQTVEVERLAHSFGWSVGTLPISYLGAPLGGNPRRIVFWNPMLEKLKCKTRSYNSKFMSL 721
+E +R+A F G +P +YLG P+ N + K++ + ++ ++S
Sbjct: 264 GDEMETQRVADLFNCQAGKMPFTYLGIPISMNKLTNADLDIPPNKIEKRLATWKCGYLSY 323
Query: 722 SGRLVITKSVLCSIPLFLMVVFKAPKGVILEVEKLFHAFIWGQEQGRKKINWIGWDQICL 781
G+ ++ S L SIPL++M V+ P+GV +++ + F W + ++K + I W+ +C
Sbjct: 324 GGKAILINSCLSSIPLYMMGVYLLPEGVHNKMDSIRARFFWEGLEKKRKYHMIKWEALCR 383
Query: 782 NRKAGGLGLGFIEWR--NRGLLLKWAWRFGREINSLWRFFVIERYNLDPDSLIWHMASTT 839
++ G GLGFI+ R N LL KW +R + +Y D A +
Sbjct: 384 PKEFG--GLGFIDTRKMNIALLCKWIYRLESGKEDPCCVLLRNKYMKDGGGFFQSKAEES 441
Query: 840 PAMGSRAIQDIWRVLNEDH---SLSAGFKENLLISIGDGSRVKFWRDPWVSNVALCHLFP 896
W+ L+E L + +K +G+G FW D W+ L +P
Sbjct: 442 --------SQFWKGLHEVKKWMDLGSSYK------VGNGKATNFWSDVWIGETPLKTQYP 487
Query: 897 RLYAMSENKEARIAEVGTYSRGKWGWSLTFRGQFFAWEEELYQQFLASIN--HIIPAAV- 953
+Y M +KE ++++ G W + L ++ L N H P +
Sbjct: 488 NIYRMCADKEKTVSQM--CLEGDW---------YIELRRSLGERDLNEWNDLHNTPREIH 536
Query: 954 ---GGDQLIWCANQTGLFSVKELCYWLE----DHLFTHQMWQVPKSIAKLLPPKVALFFW 1006
D +IW + G + K L L +W+ +P KV + FW
Sbjct: 537 LKEERDCIIWKLTKNGFYKAKTLYQALSFGGVKDKVMQDLWR------SSIPLKVKILFW 590
Query: 1007 QAQNDKIATRANLLARGIDLNGSTQCLFCNAFDESCNHIFLHCHSIWSLWCVIVNRERFL 1066
+I L + + +GS C C E +H+ CH +WC R+ F
Sbjct: 591 LMLKGRIQAAGQL--KKMKWSGSPNCKLCGQ-TEDVDHLMFRCHISQFMWCCY--RDAFG 645
Query: 1067 W-VMPASLADLILEWDFLPKISDKSLWSLIPYALSWSIWIERNNICFNHK 1115
W +P+S DL LE + K KS+ A +W+IW+ RN+ FN K
Sbjct: 646 WDNIPSSREDL-LEKLNIQKDGQKSVLLACLAAGTWAIWLMRNDWVFNGK 694
>UniRef100_Q9SKJ4 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1750
Score = 312 bits (800), Expect = 4e-83
Identities = 251/923 (27%), Positives = 417/923 (44%), Gaps = 83/923 (8%)
Query: 414 FFSSGRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRLQKVMPKLL 473
FF + + +N I +IPK N ++L+ YRPI+L YK+I+K L RL+ + ++
Sbjct: 856 FFETSFMKPSINHTNICMIPKITNPTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIV 915
Query: 474 SQNQFAFTKERQIADCILLANEVADFLSKK---SEGGLMLKVDLAKAYDQVDWEFLINML 530
S +Q AF R I D +++A+EV L + S+ + +K D++KAYD+V+W+FL +
Sbjct: 916 SDSQAAFIPGRIINDNVMIAHEVMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTM 975
Query: 531 HEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNGLSC 590
F +KWI W+ + + S ++L+NGSP + + +G+RQGDPLSP LF LC + LS
Sbjct: 976 RLFGFCNKWIGWIMAAVKSVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSH 1035
Query: 591 LLNQFLGEENYCGVRISRNL-TINHLLFADDTLLFCENNELQLETLCNVLITFLFASGLK 649
L+N + GVRI I HL FADD+L FC+ N + L +V + + SG K
Sbjct: 1036 LINGRASSGDLRGVRIGNGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQK 1095
Query: 650 VNNQKSEL-IGCNVQTVEVERLAHSFGWSVGTLPISYLGAPLGGNPRRIVFWNPMLEKLK 708
+N QKS + G V RL YLG P ++ + +++++K
Sbjct: 1096 INVQKSMITFGSRVYGSTQSRLKQILEIPNQGGGGKYLGLPEQFGRKKKEMFEYIIDRVK 1155
Query: 709 CKTRSYNSKFMSLSGRLVITKSVLCSIPLFLMVVFKAPKGVILEVEKLFHAFIWGQEQGR 768
+T +++++F+S +G+ ++ KSV ++P++ M FK PKG++ E+E L F W + +
Sbjct: 1156 KRTSTWSARFLSPAGKEIMLKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASNQ 1215
Query: 769 KKINWIGWDQICLNRKAGGLGLGFIEWRNRGLLLKWAWRFGREINSLWRFFVIERYNLDP 828
+ I W+ W ++ ++K GGLG + N LL K AWR + NSL+ + RY D
Sbjct: 1216 RGIPWVAWKRLQYSKKEGGLGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKARYFKD- 1274
Query: 829 DSLIWHMASTTPAMGSRAIQDIWRVLNEDHSLSAGFKENLLISIGDGSRVKFWRDPWVSN 888
S A + W L + +L K+ IGDG ++ D N
Sbjct: 1275 -------VSILDAKVRKQQSYGWASLLDGIAL---LKKGTRHLIGDGQNIRIGLD----N 1320
Query: 889 VALCHLFPRLYAMSENKEARIAEVGTYSRGKWGWSLTFRGQFFAWEEELYQQFLASINH- 947
+ H L KE I + + R +G ++ W++ QF+ +H
Sbjct: 1321 IVDSHPPRPLNTEETYKEMTINNL--FER---------KGSYYFWDDSKISQFVDQSDHG 1369
Query: 948 -----IIPAAVGGDQLIWCANQTGLFSVKELCYWLEDH----------------LFTHQM 986
+ + D++IW N TG ++V+ YWL H ++
Sbjct: 1370 FIHRIYLAKSKKPDKIIWNYNTTGEYTVRS-GYWLLTHDPSTNIPAINPPHGSIDLKTRI 1428
Query: 987 WQVPKSIAKLLPPKVALFFWQAQNDKIATRANLLARGIDLNGSTQCLFCNAFDESCNHIF 1046
W +P + PK+ F W+A + +AT L RG+ ++ S C C+ +ES NH
Sbjct: 1429 WNLP------IMPKLKHFLWRALSQALATTERLTTRGMRIDPS--CPRCHRENESINHAL 1480
Query: 1047 LHCHSIWSLWCV----IVNRERFLWVMPASLADLILEWDFLPKISDKSLWSLIPYALSWS 1102
C W + ++ + ++++++ +F+ + L+P L W
Sbjct: 1481 FTCPFATMAWRLSDSSLIRNQLMSNDFEENISNIL---NFVQDTTMSDFHKLLPVWLIWR 1537
Query: 1103 IWIERNNICFNHKALNISEIWDFHLLRIGWWTHELLRQNSYSIDQLSRNFESIRFASKPS 1162
IW RNN+ FN + S+ +L TH+ L +++ + ++
Sbjct: 1538 IWKARNNVVFNKFRESPSKT----VLSAKAETHDWL--------NATQSHKKTPSPTRQI 1585
Query: 1163 LLRAASWVTPDAGLIKINVDGAAKGSPGRCGIGGILRNNANVILGFFS-KSVGEGYAFEA 1221
W P A +K N D G I+RN+ + + S K EA
Sbjct: 1586 AENKIEWRNPPATYVKCNFDAGFDVQKLEATGGWIIRNHYGTPISWGSMKLAHTSNPLEA 1645
Query: 1222 EVLAIHEALLFCVQRQLRNIIIESDSSLAVGWVNNRCNRPWKLLNKLNQIDRWLVETNCL 1281
E A+ AL R + +E D + +N + L N L I W + +
Sbjct: 1646 ETKALLAALQQTWIRGYTQVFMEGDCQTLINLING-ISFHSSLANHLEDISFWANKFASI 1704
Query: 1282 KVNHIYREANGEADKLAKRGAEY 1304
+ I ++ N A LAK G Y
Sbjct: 1705 QFGFIRKKGNKLAHVLAKYGCTY 1727
>UniRef100_Q9SIQ5 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1524
Score = 312 bits (799), Expect = 5e-83
Identities = 256/930 (27%), Positives = 415/930 (44%), Gaps = 97/930 (10%)
Query: 414 FFSSGRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRLQKVMPKLL 473
FF + + +N I +IPK N ++L+ YRPI+L YK+I+K L RL+ + ++
Sbjct: 630 FFETSFMKPSINHTNICMIPKITNPTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIV 689
Query: 474 SQNQFAFTKERQIADCILLANEVADFLSKK---SEGGLMLKVDLAKAYDQVDWEFLINML 530
S +Q AF R I D +++A+EV L + S+ + +K D++KAYD+V+W+FL +
Sbjct: 690 SDSQAAFIPGRIINDNVMIAHEVMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTM 749
Query: 531 HEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNGLSC 590
F +KWI W+ + + S ++L+NGSP + + +G+RQGDPLSP LF LC + LS
Sbjct: 750 RLFGFCNKWIGWIMAAVKSVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSH 809
Query: 591 LLNQFLGEENYCGVRISRNL-TINHLLFADDTLLFCENNELQLETLCNVLITFLFASGLK 649
L+N + GVRI I HL FADD+L FC+ N + L +V + + SG K
Sbjct: 810 LINGRASSGDLRGVRIGNGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQK 869
Query: 650 VNNQKSEL-IGCNVQTVEVERLAHSFGWSVGTLPISYLGAPLGGNPRRIVFWNPMLEKLK 708
+N QKS + G V +L YLG P ++ + +++++K
Sbjct: 870 INVQKSMITFGSRVYGSTQSKLKQILEIPNQGGGGKYLGLPEQFGRKKKEMFEYIIDRVK 929
Query: 709 CKTRSYNSKFMSLSGRLVITKSVLCSIPLFLMVVFKAPKGVILEVEKLFHAFIWGQEQGR 768
+T +++++F+S +G+ ++ KSV ++P++ M FK PKG++ E+E L F W + +
Sbjct: 930 KRTSTWSARFLSPAGKEIMLKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASNQ 989
Query: 769 KKINWIGWDQICLNRKAGGLGLGFIEWRNRGLLLKWAWRFGREINSLWRFFVIERYNLDP 828
+ I W+ W ++ ++K GGLG + N LL K AWR + NSL+ + RY D
Sbjct: 990 RGIPWVAWKRLQYSKKEGGLGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKARYFKD- 1048
Query: 829 DSLIWHMASTTPAMGSRAIQDIWRVLNEDHSLSAGFKENLLISIGDGSRVKFWRDPWVSN 888
S A + W L + +L K+ IGDG ++ D N
Sbjct: 1049 -------VSILDAKVRKQQSYGWASLLDGIAL---LKKGTRHLIGDGQNIRIGLD----N 1094
Query: 889 VALCHLFPRLYAMSENKEARIAEVGTYSRGKWGWSLTFRGQFFAWEEELYQQFLASINH- 947
+ H L KE I + + R +G ++ W++ QF+ +H
Sbjct: 1095 IVDSHPPRPLNTEETYKEMTINNL--FER---------KGSYYFWDDSKISQFVDQSDHG 1143
Query: 948 -----IIPAAVGGDQLIWCANQTGLFSVKELCYWLEDH----------------LFTHQM 986
+ + D++IW N TG ++V+ YWL H ++
Sbjct: 1144 FIHRIYLAKSKKPDKIIWNYNTTGEYTVRS-GYWLLTHDPSTNIPAINPPHGSIDLKTRI 1202
Query: 987 WQVPKSIAKLLPPKVALFFWQAQNDKIATRANLLARGIDLNGSTQCLFCNAFDESCNHIF 1046
W +P + PK+ F W+A + +AT L RG+ ++ C C+ +ES NH
Sbjct: 1203 WNLP------IMPKLKHFLWRALSQALATTERLTTRGMRID--PICPRCHRENESINHAL 1254
Query: 1047 LHCHSIWSLWCVIVNRERFLWVMPASL-ADLILEWDFLPKISD----------KSLWSLI 1095
C W W+ +SL + ++ DF IS+ L+
Sbjct: 1255 FTCPFATMAW----------WLSDSSLIRNQLMSNDFEENISNILNFVQDTTMSDFHKLL 1304
Query: 1096 PYALSWSIWIERNNICFNHKALNISEIWDFHLLRIGWWTHELLRQNSYSIDQLSRNFESI 1155
P L W IW RNN+ FN + S+ +L TH+ L +++ +
Sbjct: 1305 PVWLIWRIWKARNNVVFNKFRESPSKT----VLSAKAETHDWL--------NATQSHKKT 1352
Query: 1156 RFASKPSLLRAASWVTPDAGLIKINVDGAAKGSPGRCGIGGILRNNANVILGFFS-KSVG 1214
++ W P A +K N D G I+RN+ + + S K
Sbjct: 1353 PSPTRQIAENKIEWRNPPATYVKCNFDAGFDVQKLEATGGWIIRNHYGTPISWGSMKLAH 1412
Query: 1215 EGYAFEAEVLAIHEALLFCVQRQLRNIIIESDSSLAVGWVNNRCNRPWKLLNKLNQIDRW 1274
EAE A+ AL R + +E D + +N + L N L I W
Sbjct: 1413 TSNPLEAETKALLAALQQTWIRGYTQVFMEGDCQTLINLING-ISFHSSLANHLEDISFW 1471
Query: 1275 LVETNCLKVNHIYREANGEADKLAKRGAEY 1304
+ ++ I R+ N A LAK G Y
Sbjct: 1472 ANKFASIQFGFIRRKGNKLAHVLAKYGCTY 1501
>UniRef100_Q7XJS2 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1344
Score = 303 bits (777), Expect = 2e-80
Identities = 240/865 (27%), Positives = 396/865 (45%), Gaps = 86/865 (9%)
Query: 414 FFSSGRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRLQKVMPKLL 473
FF +G + + N I LIPK + ++ RPISL YK+I+K+L RL+K +P ++
Sbjct: 452 FFETGVLPQDWNHTHICLIPKITSPQRMSDLRPISLCSVLYKIISKILTQRLKKHLPAIV 511
Query: 474 SQNQFAFTKERQIADCILLANEVADFL---SKKSEGGLMLKVDLAKAYDQVDWEFLINML 530
S Q AF +R I+D IL+A+E+ L + S+ + K D++KAYD+V+W FL M+
Sbjct: 512 STTQSAFVPQRLISDNILVAHEMIHSLRTNDRISKEHMAFKTDMSKAYDRVEWPFLETMM 571
Query: 531 HEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNGLSC 590
+ F +KWISW+ +C++S S ++L+NG P +G+RQGDPLSP LF LC L
Sbjct: 572 TALGFNNKWISWIMNCVTSVSYSVLINGQPYGHIIPTRGIRQGDPLSPALFVLCTEALIH 631
Query: 591 LLNQFLGEENYCGVRI-SRNLTINHLLFADDTLLFCENNELQLETLCNVLITFLFASGLK 649
+LN+ G++ + +++NHLLFADDTLL C+ + + E L L + SG
Sbjct: 632 ILNKAEQAGKITGIQFQDKKVSVNHLLFADDTLLMCKATKQECEELMQCLSQYGQLSGQM 691
Query: 650 VNNQKSEL-IGCNVQTVEVERLAHSFGWSVGTLPISYLGAP--LGGNPRRIVFWNPMLEK 706
+N KS + G NV + + G S+ YLG P L G+ R + + + EK
Sbjct: 692 INLNKSAITFGKNVDIQIKDWIKSRSGISLEGGTGKYLGLPECLSGSKRDL--FGFIKEK 749
Query: 707 LKCKTRSYNSKFMSLSGRLVITKSVLCSIPLFLMVVFKAPKGVILEVEKLFHAFIWGQEQ 766
L+ + + +K +S G+ V+ KS+ ++P+++M FK PK + ++ + F W Q
Sbjct: 750 LQSRLTGWYAKTLSQGGKEVLLKSIALALPVYVMSCFKLPKNLCQKLTTVMMDFWWNSMQ 809
Query: 767 GRKKINWIGWDQICLNRKAGGLGLGFIEWRNRGLLLKWAWRFGREINSLWRFFVIERYNL 826
++KI+W+ W ++ L + GG G ++ N+ LL K AWR +E SL+ RY
Sbjct: 810 QKRKIHWLSWQRLTLPKDQGGFGFKDLQCFNQALLAKQAWRVLQEKGSLFSRVFQSRYFS 869
Query: 827 DPDSLIWHMASTTPAMGSRAIQDIWRVLNEDHSLSAGFKENLLISIGDGSRVKFWRDPWV 886
+ D L + GSR WR + L + L IG+G + W D W+
Sbjct: 870 NSDFL-------SATRGSRP-SYAWRSILFGREL---LMQGLRTVIGNGQKTFVWTDKWL 918
Query: 887 SNVALCHLFPRLYAMSENKEARIAE-VGTYSRGKWGWSLTFRGQFFAWEEELYQQFLASI 945
+ + + P N + ++++ + SR W+L F W++ + I
Sbjct: 919 HDGS--NRRPLNRRRFINVDLKVSQLIDPTSR---NWNLNMLRDLFPWKD------VEII 967
Query: 946 NHIIPAAVGGDQLIWCANQTGLFSVKELCYWLEDHLFTHQMWQ---VPKSIAKLL----- 997
P D W + GL+SVK +L + H+++Q V S+ L
Sbjct: 968 LKQRPLFFKEDSFCWLHSHNGLYSVKTGYEFLSKQVH-HRLYQEAKVKPSVNSLFDKIWN 1026
Query: 998 ---PPKVALFFWQAQNDKIATRANLLARGIDLNGSTQCLFCNAFDESCNHIFLHC---HS 1051
PK+ +F W+A + I L RGI CL C+ +E+ NHI C
Sbjct: 1027 LHTAPKIRIFLWKALHGAIPVEDRLRTRGI--RSDDGCLMCDTENETINHILFECPLARQ 1084
Query: 1052 IWSLWCVIVNRERF---LWVMPASLADLILEWDFLPKISDKSLWSLIPYALSWSIWIERN 1108
+W++ + F ++ + L DL + D + S W L W +W RN
Sbjct: 1085 VWAITHLSSAGSEFSNSVYTNMSRLIDLTQQNDLPHHLRFVSPWIL------WFLWKNRN 1138
Query: 1109 NICFNHKALNISEIWD-----FHLLRIGWWTHELLRQNSYSIDQLSRNFESIRFASKPSL 1163
+ F K + + D +H W++ + QN
Sbjct: 1139 ALLFEGKGSITTTLVDKAYEAYH----EWFSAQTHMQND------------------EKH 1176
Query: 1164 LRAASWVTPDAGLIKINVDGAAKGSPGRCGIGGILRNNANVILGFFSKSVGEGYA-FEAE 1222
L+ W P G +K N+ A G ++R++ +L +S E ++ + A+
Sbjct: 1177 LKITKWCPPLPGELKCNIGFAWSKQHHFSGASWVVRDSQGKVLLHSRRSFNEVHSPYSAK 1236
Query: 1223 VLAIHEALLFCVQRQLRNIIIESDS 1247
+ + AL +I S +
Sbjct: 1237 IRSWEWALESMTHHHFDRVIFASST 1261
>UniRef100_Q9FYJ4 F17F8.5 [Arabidopsis thaliana]
Length = 872
Score = 301 bits (770), Expect = 1e-79
Identities = 205/672 (30%), Positives = 332/672 (48%), Gaps = 49/672 (7%)
Query: 404 GAWFEWMFEDFFSSGRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLAT 463
G F + FF G + +G+N+ + LIPKK A + YRPIS YK+I+K++A
Sbjct: 136 GQEFTLPVQSFFQKGFLPKGINSIILALIPKKLAAKEMRDYRPISCCNVLYKVISKIIAN 195
Query: 464 RLQKVMPKLLSQNQFAFTKERQIADCILLANE-VADFLSKKSEGGLMLKVDLAKAYDQVD 522
RL+ ++P+ +++NQ AF K+R + + +LLA E V D+ +K+D++KA+D V
Sbjct: 196 RLKLLLPRFIAENQSAFVKDRLLIENLLLATELVKDYHKDSISARCAIKIDISKAFDSVQ 255
Query: 523 WEFLINMLHEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFN 582
W FL N L MNF +I W+ CI++AS ++ VNG +F ++GLRQG LSP LF
Sbjct: 256 WSFLTNTLVAMNFSPTFIHWINLCITTASFSVQVNGDLVGYFQSKRGLRQGCSLSPYLFV 315
Query: 583 LCLNGLSCLLNQFLGEENYCGVRISRNLTINHLLFADDTLLFCENNELQLETLCNVLITF 642
+C++ LS +L++ G + + L + HL FADD ++ + +E + V F
Sbjct: 316 ICMDVLSKMLDKAAGVRKFGFHPKCQRLGLTHLSFADDLMVLSDGKTRSIEGILEVFDEF 375
Query: 643 LFASGLKVNNQKSELIGCNVQTVEVERLAHSFGWSVGTLPISYLGAPLGGNPRRIVFWNP 702
SGL+++ +KS L V + + +A F + VG LP+ YLG PL ++P
Sbjct: 376 CKRSGLRISLEKSTLYMAGVSPIIKQEIAAKFLFDVGQLPVRYLGLPLVTKRLTSADYSP 435
Query: 703 MLEKLKCKTRSYNSKFMSLSGRLVITKSVLCSIPLFLMVVFKAPKGVILEVEKLFHAFIW 762
+LE++K + ++ +F S +GR + KSVL SI F + F+ P+ I E++KL +F+W
Sbjct: 436 LLEQIKKRIATWTFRFFSFAGRFNLIKSVLWSICNFWLAAFRLPRQCIREIDKLCSSFLW 495
Query: 763 -GQEQGRKKINWIGWDQICLNRKAGGLGLGFIEWRNRGLLLKWAWRFGREINSLWRFFVI 821
G E K I WD +C + GGLGL ++ N LK WR NSLW +V
Sbjct: 496 SGSEMSSHKAK-ISWDIVCKPKAEGGLGLRNLKEANDVSCLKLVWRIISNSNSLWTKWVA 554
Query: 822 ERYNLDPDSLIWHMASTTPAMGSRAIQDIWRVLNEDHSLSAGFKENLLISIGDGSRVKFW 881
E Y + S IW + +T +MGS IWR + + ++ F + +G+G FW
Sbjct: 555 E-YLIRKKS-IWSLKQST-SMGSW----IWRKILKIRDVAKSFSR---VEVGNGESASFW 604
Query: 882 RDPWVSNVALCHLF--PRLYAMSENKEARIAEVGT-YSRGKWGWSLTFRGQFFAWEEELY 938
D W ++ L + +EA +A+ T SR + SL + E Y
Sbjct: 605 YDHWSAHGRLIDTVGDKGTIDLGIPREASVADAWTRRSRRRHRTSLLNEIE----EMMAY 660
Query: 939 QQFLASINHIIPAAVGGDQLIWCANQTGLFSVKELCYWLEDHLFTHQMWQVPKSIAKLL- 997
Q+ I+H + D ++W + +F + H T W + K+ + +
Sbjct: 661 QR----IHH----SDAEDTVLW-RGKNDVF---------KPHFSTRDTWHLIKATSSTVS 702
Query: 998 ----------PPKVALFFWQAQNDKIATRANLLARGIDLNGSTQCLFCNAFDESCNHIFL 1047
PK AL W A ++++ T +L + S C+ C ++ H+F
Sbjct: 703 WHKGVWFRHATPKYALCTWLAIHNRLPTGDRMLKWNSSGSVSGNCVLCTNNSKTLEHLFF 762
Query: 1048 HCHSIWSLWCVI 1059
C ++W +
Sbjct: 763 SCSYASTVWAAL 774
>UniRef100_Q9LVQ1 Non-LTR retroelement reverse transcriptase-like [Arabidopsis
thaliana]
Length = 1072
Score = 298 bits (763), Expect = 7e-79
Identities = 228/723 (31%), Positives = 349/723 (47%), Gaps = 51/723 (7%)
Query: 413 DFFSSGRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRLQKVMPKL 472
+FF SG++ + NA + LIPK NA +++ +RPIS + + YK+I+K+L +RLQ ++ +
Sbjct: 352 EFFDSGQLLKQWNATTLVLIPKTSNACTISEFRPISCLNTLYKVISKLLTSRLQGLLSAV 411
Query: 473 LSQNQFAFTKERQIADCILLANEVADFLSKKSEGGL-MLKVDLAKAYDQVDWEFLINMLH 531
+ +Q AF R +A+ +LLA E+ ++ + MLKVDL KA+D V WEF+ L
Sbjct: 412 IGHSQSAFLPGRSLAENVLLATEMVHGYNRLNISPRGMLKVDLKKAFDSVKWEFVTAALR 471
Query: 532 EMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNGLSCL 591
+ +++I+W+ CI++ S I VNG+ FF KGLRQGDPLSP LF L + S L
Sbjct: 472 ALAIPERYINWIHQCITTPSFTISVNGATGGFFRSTKGLRQGDPLSPYLFVLAMEVFSKL 531
Query: 592 LNQFLGEENYCGVRISRNLTINHLLFADDTLLFCENNELQLETLCNVLITFLFASGLKVN 651
L + +L+I+HL+FADD ++F + + +C L F SGLKVN
Sbjct: 532 LYSRYDSGYIHYHPKAGDLSISHLMFADDVMIFFDGGSSSMHGICETLDDFADWSGLKVN 591
Query: 652 NQKSELIGCNVQTVEVERLAHSFGWSVGTLPISYLGAPLGGNPRRIVFWNPMLEKLKCKT 711
KS+L + E A ++G+ GT PI YLG PL RI + P+LEKL +
Sbjct: 592 KDKSQLFQAGLDLSERITSA-AYGFPAGTFPIRYLGLPLMCRKLRIADYGPLLEKLSARL 650
Query: 712 RSYNSKFMSLSGRLVITKSVLCSIPLFLMVVFKAPKGVILEVEKLFHAFIWGQEQGRKKI 771
RS+ SK +S +GR + SV+ + F M F PKG I ++E L F+W +K
Sbjct: 651 RSWVSKALSFAGRTQLISSVIFGLINFWMSTFLLPKGCIKKIESLCSKFLWAGSIDGRKS 710
Query: 772 NWIGWDQICLNRKAGGLGL-GFIEWRNRGLLLKWAWRFGREINSLWRFFVIERYNLDPDS 830
+ + W CL + GGLG F EW N+ LLL+ W SLW + +R++ +
Sbjct: 711 SKVSWVDCCLPKSEGGLGFRSFGEW-NKTLLLRLIWVLFDRDTSLWAQW--QRHHRLGHA 767
Query: 831 LIWHMASTTPAMGSRAIQD---IWRVLNEDHSLSAGFKENLLISIGDGSRVKFWRDPWVS 887
W + A+Q W++L L+ F + +G+G V FW D W S
Sbjct: 768 SFWQV---------NALQTDPWTWKMLLNLRPLAEKF---IKAKVGNGGTVSFWFDCWTS 815
Query: 888 NVALCHLFPRLYAMSE--NKEARI---AEVGTYSRGKWGWSLTFRGQFFAWEEELYQQFL 942
L P + + + ++ RI A+V G GW L A + L
Sbjct: 816 ------LGPLIKYLGDVGSRPLRIPFSAKVADAIDGS-GWRLPLSRSLTA---DSILSHL 865
Query: 943 ASINHIIPAAVGGDQLIWCANQTGLFSVKELCYWLEDHLFTHQMWQVPKSI-AKLLPPKV 1001
AS+ P V D WC + W E + + KS+ K PK
Sbjct: 866 ASLPPPSPLMV-SDSYSWCVDDVDCQGFSAAKTW-EVLRPRRPVKRWAKSVWFKGAVPKH 923
Query: 1002 ALFFWQAQNDKIATRANLLARGIDLNGSTQCLFCNAFDESCNHIFLHCHSIWSLWCVI-- 1059
A FW AQ +++ TR L++ G L S +C C+ E+ +H+ L C +W ++
Sbjct: 924 AFNFWTAQLNRLPTRQRLVSWG--LVSSAECCLCSFDTETRDHLLLLCDFSSQVWRMVFL 981
Query: 1060 --VNRERFLWVMPASLADLILEWDFLPKISDKSLWSLIPYALSWSIWIERNNICFNHKAL 1117
R+R L L+ P + K + L+ Y ++W +RN + H +L
Sbjct: 982 RLCPRQRLLCTWAELLSWTRQSTAAAPSLLRKVVAQLVVY----NLWRQRNLVL--HSSL 1035
Query: 1118 NIS 1120
+S
Sbjct: 1036 RVS 1038
>UniRef100_Q9M1F2 Hypothetical protein F9K21.130 [Arabidopsis thaliana]
Length = 851
Score = 296 bits (758), Expect = 3e-78
Identities = 227/802 (28%), Positives = 373/802 (46%), Gaps = 81/802 (10%)
Query: 414 FFSSGRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRLQKVMPKLL 473
FF S + +N I +IPK +N +L+ YRPI+L YK+I+K + RL+ + ++
Sbjct: 95 FFESSHMKTSVNHTNICMIPKIQNPQTLSDYRPIALCNVLYKVISKCMVNRLKAHLNSIV 154
Query: 474 SQNQFAFTKERQIADCILLANEVADFLSKK---SEGGLMLKVDLAKAYDQVDWEFLINML 530
S +Q AF R I D +++A+E+ L + S+ + +K D++KAYD+V+W+FL +
Sbjct: 155 SDSQAAFIPGRIINDNVMIAHEIMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTM 214
Query: 531 HEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNGLSC 590
F DKWI W+ + + S ++L+NGSP + S +G+RQGDPLSP LF LC + LS
Sbjct: 215 RLFGFCDKWIGWIMAAVKSVHYSVLINGSPHGYISPTRGIRQGDPLSPYLFILCGDILSH 274
Query: 591 LLNQFLGEENYCGVRISRNL-TINHLLFADDTLLFCENNELQLETLCNVLITFLFASGLK 649
L+ + GVRI I HL FADD+L FC+ N + L +V + + SG K
Sbjct: 275 LIKVKASSGDIRGVRIGNGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQK 334
Query: 650 VNNQKSEL-IGCNVQTVEVERLAHSFGWSVGTLPISYLGAPLGGNPRRIVFWNPMLEKLK 708
+N QKS + G V RL YLG P ++ +N +++++K
Sbjct: 335 INVQKSLITFGSRVYGSTQTRLKTLLNIPNQGGGGKYLGLPEQFGRKKKEMFNYIIDRVK 394
Query: 709 CKTRSYNSKFMSLSGRLVITKSVLCSIPLFLMVVFKAPKGVILEVEKLFHAFIWGQEQGR 768
+T S+++KF+S +G+ ++ KSV ++P++ M FK P+G++ E+E L F W + +
Sbjct: 395 ERTASWSAKFLSPAGKEILLKSVALAMPVYAMSCFKLPQGIVSEIESLLMNFWWEKASNK 454
Query: 769 KKINWIGWDQICLNRKAGGLGLGFIEWRNRGLLLKWAWRFGREINSLWRFFVIERYNLDP 828
+ I W+ W ++ ++K GGLG + N LL K AWR + NSL+ + RY D
Sbjct: 455 RGIPWVAWKRLQYSKKEGGLGFRDLAKFNDALLAKQAWRIIQYPNSLFARVMKARYFKD- 513
Query: 829 DSLIWHMASTTPAMGSRAIQDIWRVLNEDHSLSAGFKENLLISIGDGSRVKFWRDPWVSN 888
+S+I + + G W L +L ++ IGDG ++ + N
Sbjct: 514 NSIIDAKTRSQQSYG-------WSSLLSGIAL---LRKGTRYVIGDGKTIRL----GIDN 559
Query: 889 VALCHLFPRLYAMSENKEARIAEVGTYSRGKWGWSLTFRGQFFAWEEELYQQFLASINH- 947
V H P +++ + ++ + RG W+ Q F+ +H
Sbjct: 560 VVDSH--PPRPLLTDEQHNGLSLDNLFQH---------RGHSRCWDNAKLQTFVDQSDHD 608
Query: 948 -----IIPAAVGGDQLIWCANQTGLFSVKELCYWLEDHLFTH----------------QM 986
+ D+LIW N TG ++V+ YWL H ++ ++
Sbjct: 609 YIKRIYLSTRSKTDRLIWSYNSTGDYTVRS-GYWLSTHDPSNTIPTMAKPHGSVDLKTKI 667
Query: 987 WQVPKSIAKLLPPKVALFFWQAQNDKIATRANLLARGIDLNGSTQCLFCNAFDESCNHIF 1046
W +P + PK+ F W+ + + T L RG+ ++ C C +ES NH
Sbjct: 668 WNLP------IMPKLKHFLWRILSKALPTTDRLTTRGMRIDPG--CPRCRRENESINHAL 719
Query: 1047 LHC---HSIWSLWCVIVNRERFLW-VMPASLADLILEWDFLPKISDKSLWSLIPYALSWS 1102
C W L + R L + ++++++L L + LIP+ L W
Sbjct: 720 FTCPFATMAWRLSDTPLYRSSILSNNIEDNISNILL---LLQNTTITDSQKLIPFWLLWR 776
Query: 1103 IWIERNNICFNHKALNISEIWDFHLLRIGWWTHELLRQNSYSIDQLSRNFESIRFASKPS 1162
IW RNN+ FN N+ E ++R T+E L + R + +
Sbjct: 777 IWKARNNVVFN----NLRESPSITVVRAKAETNEWLNATQTQGPR--------RLPKRTT 824
Query: 1163 LLRAASWVTPDAGLIKINVDGA 1184
+WV P IK N D +
Sbjct: 825 AAGNTTWVKPQMPYIKCNFDAS 846
>UniRef100_Q9SJ38 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1229
Score = 293 bits (750), Expect = 2e-77
Identities = 203/663 (30%), Positives = 317/663 (47%), Gaps = 56/663 (8%)
Query: 414 FFSSGRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRLQKVMPKLL 473
FF+S R MN I LIPK + YRPI+L YK++AK++ R+Q ++PKL+
Sbjct: 369 FFTSKNFPRRMNETHIRLIPKDLGPRKVADYRPIALCNIFYKIVAKIMTKRMQLILPKLI 428
Query: 474 SQNQFAFTKERQIADCILLANEVADFL---SKKSEGGLMLKVDLAKAYDQVDWEFLINML 530
S+NQ AF R I+D +L+ +EV FL S K + +K D++KAYD+V+W+FL +L
Sbjct: 429 SENQSAFVPGRVISDNVLITHEVLHFLRTSSAKKHCSMAVKTDMSKAYDRVEWDFLKKVL 488
Query: 531 HEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNGLSC 590
F WI W+ C++S S + L+NG+P +GLRQGDPLSP LF LC LS
Sbjct: 489 QRFGFHSIWIDWVLECVTSVSYSFLINGTPQGKVVPTRGLRQGDPLSPCLFILCTEVLSG 548
Query: 591 LLNQFLGEENYCGVRISRN-LTINHLLFADDTLLFCENNELQLETLCNVLITFLFASGLK 649
L + GVR+S N +NHLLFADDT+ F +++ L +L + ASG
Sbjct: 549 LCTRAQRLRQLPGVRVSINGPRVNHLLFADDTMFFSKSDPESCNKLSEILSRYGKASGQS 608
Query: 650 VNNQKSELI-------GCNVQTVEVERLAHSFGWSVGTLPISYLGAPLGGNPRRIVFWNP 702
+N KS + Q + ++ G YLG P R+ +
Sbjct: 609 INFHKSSVTFSSKTPRSVKGQVKRILKIRKEGGTG------KYLGLPEHFGRRKRDIFGA 662
Query: 703 MLEKLKCKTRSYNSKFMSLSGRLVITKSVLCSIPLFLMVVFKAPKGVILEVEKLFHAFIW 762
+++K++ K+ S+ S+F+S +G+ V+ K+VL S+PL+ M FK P + +++ L F W
Sbjct: 663 IIDKIRQKSHSWASRFLSQAGKQVMLKAVLASMPLYSMSCFKLPSALCRKIQSLLTRFWW 722
Query: 763 GQEQGRKKINWIGWDQICLNRKAGGLGLGFIEWRNRGLLLKWAWRFGREINSLWRFFVIE 822
+ +K +W+ W ++ + AGGLG IE N LL K WR SL ++
Sbjct: 723 DTKPDVRKTSWVAWSKLTNPKNAGGLGFRDIERCNDSLLAKLGWRLLNSPESLLSRILLG 782
Query: 823 RYNLDPDSLIWHMASTTPAMGSRAIQDIWRVLNEDHSLSAGFKENLLISIGDGSRVKFWR 882
+Y S + + P+ G WR + + KE L I +G +V W
Sbjct: 783 KY-CHSSSFMECKLPSQPSHG-------WRSIIAGREI---LKEGLGWLITNGEKVSIWN 831
Query: 883 DPWVSNVALCHLFPRLYAMSENKEARIAEVGTYSRGKWGWSLTFRGQFFAWEEELYQQFL 942
DPW+S L P A+ E+++ R++ + + +W W+ E L +Q
Sbjct: 832 DPWLS--ISKPLVPIGPALREHQDLRVSALINQNTLQWDWNKI--AVILPNYENLIKQLP 887
Query: 943 ASINHIIPAAVGGDQLIWCANQTGLF---------SVKELCYWLEDHLFTHQMWQVPKSI 993
A P++ G D+L W ++G + SV + + +W++
Sbjct: 888 A------PSSRGVDKLAWLPVKSGQYTSRSGYGIASVASIPIPQTQFNWQSNLWKLQTL- 940
Query: 994 AKLLPPKVALFFWQAQNDKIATRANLLARGIDLNGSTQCLFCNAFDESCNHIFLHCHSIW 1053
PK+ W+A + + L+ R I + S C C A ES H+F HC
Sbjct: 941 -----PKIKHLMWKAAMEALPVGIQLVRRHI--SPSAACHRCGA-PESTTHLFFHCEFAA 992
Query: 1054 SLW 1056
+W
Sbjct: 993 QVW 995
>UniRef100_Q9ZUF0 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1352
Score = 290 bits (743), Expect = 2e-76
Identities = 212/729 (29%), Positives = 347/729 (47%), Gaps = 73/729 (10%)
Query: 412 EDFFSSGRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRLQKVMPK 471
+ FF G + +G+N + LI KK S + YRPIS YK+++K++A RL++++P
Sbjct: 644 QSFFLKGFLPKGINTTILALISKKHEVSGMKDYRPISCCNVLYKIVSKLMANRLKEILPA 703
Query: 472 LLSQNQFAFTKERQIADCILLANE-VADFLSKKSEGGLMLKVDLAKAYDQVDWEFLINML 530
++ NQ AF K+R + + +LLA+E V D+ + LK+D++KA+D V W FLIN+L
Sbjct: 704 SIAPNQSAFIKDRLMMENLLLASELVKDYHKESISSRSALKIDISKAFDFVQWPFLINVL 763
Query: 531 HEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNGLSC 590
++ + +I W++ CI +AS ++ VNG + FF E+GLRQG LSP L+ +C+N LSC
Sbjct: 764 KAIHLPEMFIHWIELCIGTASFSVQVNGELSGFFRSERGLRQGCSLSPYLYVICMNVLSC 823
Query: 591 LLNQFLGEENYCGVRISRNLTINHLLFADDTLLFCENNELQLETLCNVLITFLFASGLKV 650
+L++ E+ RN+ + HL FADD ++F + ++ + F S LK+
Sbjct: 824 MLDKAAVEKKISYHPRCRNMNLTHLCFADDIMVFSDGTSKSIQGTLAIFEKFAAMSWLKI 883
Query: 651 NNQKSELIGCNVQTVEVERLAHSFGWSVGTLPISYLGAPLGGNPRRIVFWNPMLEKLKCK 710
+ +KS + + + F + +GTLP+ YLG PL + P++EK++ +
Sbjct: 884 SLEKSTIFMAGISPNAKTSILQQFPFELGTLPVKYLGLPLLTKRMTQSDYLPLVEKIRAR 943
Query: 711 TRSYNSKFMSLSGRLVITKSVLCSIPLFLMVVFKAPKGVILEVEKLFHAFIWGQEQGRKK 770
S+ ++F+S +GRL + KSVL SI F + VF+ PK + E+EK+F AF+W K
Sbjct: 944 ITSWTNRFLSFAGRLQLIKSVLSSITNFWLSVFRLPKACLQEIEKMFSAFLWSGPDLNTK 1003
Query: 771 INWIGWDQICLNRKAGGLGLGFIEWRNRGLLLKWAWRFGREINSLWRFFVIERYNLDPDS 830
I W ++C ++ GGLGL ++ N LLK WR +SLW +V +L
Sbjct: 1004 KAKIAWSEVCKLKEEGGLGLKPLKEANEVSLLKLIWRILSARDSLWVKWV--NKHLIRKE 1061
Query: 831 LIWHMASTTPAMGSRAIQDIWRVLNEDHSLSAGFKENLLISIGDGSRVKFWRDPWVSNVA 890
W + T +GS +WR + + + F + + G+ FW D W
Sbjct: 1062 TFWSVKENT-GLGSW----LWRKILKQRDKARLFHR---MEVRSGTFTSFWHDHWCPLGR 1113
Query: 891 L-CHLFPR-LYAMSENKEARIAEVGTYSRGKWGWSLTFRGQFFAWEEELYQQFLASINHI 948
L H+ R + A +AEV R K F Q + + EL +Q
Sbjct: 1114 LHQHMGSRGTIDLGIPNNATVAEVMNTHRRK-RHRADFLNQIKS-QIELARQ-------- 1163
Query: 949 IPAAVGGDQLIWCANQTGLFSVKELCYWLEDHLFTHQMWQVPKSIA-----------KLL 997
+ GD+ +W K+ + + + WQ +SI+
Sbjct: 1164 -DRSTDGDRSLW----------KQKEDTFKSSFSSSKTWQQIRSISLRCDWYRGVWFSAS 1212
Query: 998 PPKVALFFWQAQNDKIATRANLLARGIDLNGSTQCLFCNAFDESCNHIFLHC----HSIW 1053
PK + W A ++++ T + + C+FC E+ +H+F C H +
Sbjct: 1213 TPKYSFVTWLAFHNRLTTSDKICK--WNSGARYDCVFCGEELETRDHLFFSCPYSSHVWF 1270
Query: 1054 SLWCVIVNRERFLWVMPASLADLILEWDFL-PKISDKSLWSLIPYALSW-------SIWI 1105
SL ++N IL W+ + P + D S L + L + S+W
Sbjct: 1271 SLTKGLLNGRN------------ILNWNLITPHLLDSSRPYLHVFTLRYAFQASIHSLWR 1318
Query: 1106 ERNNICFNH 1114
ERN C H
Sbjct: 1319 ERN--CRRH 1325
>UniRef100_Q8LJX1 Putative reverse transcriptase [Sorghum bicolor]
Length = 1998
Score = 288 bits (738), Expect = 6e-76
Identities = 212/705 (30%), Positives = 345/705 (48%), Gaps = 57/705 (8%)
Query: 422 RGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRLQKVMPKLLSQNQFAFT 481
R +N +FITLIPK N S+N YRPISL+ +S KLI K+LA RLQ + L+ +NQ+ F
Sbjct: 1296 RSINTSFITLIPKSHNPVSVNDYRPISLLNTSMKLITKLLANRLQMKITDLVHKNQYGFI 1355
Query: 482 KERQIADCILLANEVADFLSKKSEGGLMLKVDLAKAYDQVDWEFLINMLHEMNFGDKWIS 541
++R I DC+ + E + +++K+D KA+D+++ + ++ ++ FG KW++
Sbjct: 1356 RQRTIQDCLAWSFEYLHLCHHSRKAIVIIKLDFEKAFDKMNHKAMLTIMKAKGFGQKWLN 1415
Query: 542 WMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNGLSCLLNQFLGEENY 601
WMKS +S + +L+NG+P +G+RQGDPLSPLLF L + L L+N+ +
Sbjct: 1416 WMKSIFTSGTSNVLLNGTPGKTIHCLRGVRQGDPLSPLLFVLAADFLQSLINKAKCMDLL 1475
Query: 602 -CGVRISRNLTINHLLFADDTLLFCENNELQLETLCNVLITFLFASGLKVNNQKSELIGC 660
+ + N + +ADDTL+ E + QL L +++ TF A+GLKVN QKS ++
Sbjct: 1476 KLPIPMQTNQDFPIIQYADDTLIIAEGDTRQLLILKSIINTFSEATGLKVNLQKSMMLPI 1535
Query: 661 NVQTVEVERLAHSFGWSVGTLPISYLGAPLGGNPRRIVFWNPMLEKLKCKTRSYNSKFMS 720
N+ ++ LA +FG S G+ P +YLG PLG I + P++ K + + S + F+S
Sbjct: 1536 NMDEERLDTLARTFGCSKGSFPFTYLGLPLGITKPTIQDYQPLINKCEARLGSV-ATFLS 1594
Query: 721 LSGRLVITKSVLCSIPLFLMVVFKAPKGVILEVEKLFHAFIW-GQEQGRKKINWIGWDQI 779
+GRL +T +V ++P F M PK VI +++K +W G +K W +
Sbjct: 1595 EAGRLELTNAVFTALPTFYMCTLAIPKSVIHKIDKFRKHCLWRGNGINARKPPKAAWKLV 1654
Query: 780 CLNRKAGGLGLGFIEWRNRGLLLKWAWRFGREINSLWRFFVIERYNLDPDSLIWHMASTT 839
C + GGLG+ IE +N LL+K +F + ++ W + +++ +
Sbjct: 1655 CKPKNEGGLGVIDIEKQNEALLMKNLDKFFNKKDTPWVQMIWDKH---------YAKGKL 1705
Query: 840 PAM---GSRAIQDIWRVLNEDHSLSAGFKENLLISIGDGSRVKFWRDPWVSNVALCHLFP 896
P GS +DI ++L+ +KE LI + G + FW+D W + P
Sbjct: 1706 PGQIRKGSFWWRDILKLLDP-------YKEMSLIQMKGGESIVFWQDKWGMQKLRLEM-P 1757
Query: 897 RLYAMSENKEARIAEVGTYSRGKWGWSLTFRGQFFAWEEELYQQFLASINHIIPAAVGGD 956
L++ ++K AEV G+ ++ + QQ I+++ +
Sbjct: 1758 ELFSFVQHKFLSAAEV----LGQEDFTRLLQLPISETAFHQLQQLTTRIDNLERTQ---E 1810
Query: 957 QLIWCANQTGLFSVKELCYWLEDHLFTHQMWQVPKSIAK-LLPPKVALFFWQAQNDKIAT 1015
Q W FS + L H Q+ +V K I K PK +FFW D+++T
Sbjct: 1811 QDQWKYKWGNNFSATKAYSELMGH---QQIHEVHKWIWKSFCQPKHKVFFWLLIKDRLST 1867
Query: 1016 RANLLARGIDLNGSTQCLFC-NAFDESCNHIFLHCHSIWSLWCVIVNRERFLWVMPASLA 1074
N+L R S C+ C +E+ +H+FLHC W ++
Sbjct: 1868 -CNILKRKNMQLQSYNCVLCLQNSEETTSHLFLHCAYARQCWQLLD-------------M 1913
Query: 1075 DLILEWDFLPKISD-------KSLWSLIPYALSWSIWIERNNICF 1112
D+ DF P+I+D + + L W+IW RNN+ F
Sbjct: 1914 DIPPNSDF-PEITDHFKHGFNNQFFMVTTILLCWAIWSTRNNLIF 1957
>UniRef100_O81068 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1529
Score = 287 bits (735), Expect = 1e-75
Identities = 200/674 (29%), Positives = 326/674 (47%), Gaps = 46/674 (6%)
Query: 403 SGAWFEWMFEDFFSSGRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLA 462
+G F + FF G + +G+NA + LIPKK+ A + YRPIS YK+I+K+LA
Sbjct: 785 TGPDFIAAIQSFFVKGFLPKGLNATILALIPKKDEAIEMKDYRPISCCNVLYKVISKILA 844
Query: 463 TRLQKVMPKLLSQNQFAFTKERQIADCILLANE-VADFLSKKSEGGLMLKVDLAKAYDQV 521
RL+ ++P + QNQ AF KER + + +LLA E V D+ + +K+D++KA+D V
Sbjct: 845 NRLKLLLPSFILQNQSAFVKERLLMENVLLATELVKDYHKESVTPRCAMKIDISKAFDSV 904
Query: 522 DWEFLINMLHEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLF 581
W+FL+N L +NF + + W+K CIS+A+ ++ VNG FF +GLRQG LSP LF
Sbjct: 905 QWQFLLNTLEALNFPETFRHWIKLCISTATFSVQVNGELAGFFGSSRGLRQGCALSPYLF 964
Query: 582 NLCLNGLSCLLNQFLGEENYCGVRISRNLTINHLLFADDTLLFCENNELQLETLCNVLIT 641
+C+N LS ++++ N + + HL FADD ++F + ++ +E + NV
Sbjct: 965 VICMNVLSHMIDEAAVHRNIGYHPKCEKIGLTHLCFADDLMVFVDGHQWSIEGVINVFKE 1024
Query: 642 FLFASGLKVNNQKSELIGCNVQTVEVERLAHSFGWSVGTLPISYLGAPLGGNPRRIVFWN 701
F SGL+++ +KS + V + + SF ++ G LP+ YLG PL ++
Sbjct: 1025 FAGRSGLQISLEKSTIYLAGVSASDRVQTLSSFPFANGQLPVRYLGLPLLTKQMTTADYS 1084
Query: 702 PMLEKLKCKTRSYNSKFMSLSGRLVITKSVLCSIPLFLMVVFKAPKGVILEVEKLFHAFI 761
P++E +K K S+ ++ +S +GRL + SV+ SI F M ++ P G I E+EKL AF+
Sbjct: 1085 PLIEAVKTKISSWTARSLSYAGRLALLNSVIVSIANFWMSAYRLPAGCIREIEKLCSAFL 1144
Query: 762 WGQEQGRKKINWIGWDQICLNRKAGGLGLGFIEWRNRGLLLKWAWRFGREINSLW----R 817
W K I W IC +K GGLG+ + N+ LK WR SLW
Sbjct: 1145 WSGPVLNPKKAKIAWSSICQPKKEGGLGIKSLAEANKVSCLKLIWRLLSTQPSLWVTWIW 1204
Query: 818 FFVIERYNLDPDSLIWHMASTTPAMGSRAIQDIWRVLNEDHSLSAGFKENLLISIGDGSR 877
F+I + W A+ ++GS +W+ L + L+ + + + +GS
Sbjct: 1205 TFIIRK------GTFW-SANERSSLGSW----MWKKLLKYRELAKSMHK---VEVRNGSS 1250
Query: 878 VKFWRDPWVSNVALCHLFPRLYAMSENKEARIAEVGTYSRGKWGWSLTFRGQFFAWEEEL 937
FW D W HL RL ++ + R+ ++G L Q +
Sbjct: 1251 TSFWYDHW------SHL-GRLLDITGTR--RVIDLGIPLETNLETVLRTH-QHRQHRAAI 1300
Query: 938 YQQFLASINHI--IPAAVGGDQLIWCANQTGLFSVKELCYWLEDHLFTHQMWQ------- 988
Y + A I + G D +W + + F+ + + +++ THQ Q
Sbjct: 1301 YNRINAEIQRLQQQEREAGPDISLWRSLKND-FNKRFITKVTWNNVRTHQPQQNWYKGVW 1359
Query: 989 VPKSIAKLLPPKVALFFWQAQNDKIATRANLLARGIDLNGSTQCLFCNAFDESCNHIFLH 1048
P S PK + W ++++T + + + C CN +E+ +H+F
Sbjct: 1360 FPYS-----TPKYSFLLWLTVQNRLSTGDRI--KAWNSGQLVTCTLCNNAEETRDHLFFS 1412
Query: 1049 CHSIWSLWCVIVNR 1062
C +W + R
Sbjct: 1413 CQYTSYVWEALTQR 1426
>UniRef100_Q9C6L3 Hypothetical protein F2J7.11 [Arabidopsis thaliana]
Length = 1213
Score = 286 bits (732), Expect = 3e-75
Identities = 224/740 (30%), Positives = 352/740 (47%), Gaps = 47/740 (6%)
Query: 389 PQGLTA-LI*NSFRNSGAWFEWMFEDFFSSGRISRGMNAAFITLIPKKENASSLNHYRPI 447
P G TA +S+ GA ++FFSSG + + NA I LIPK N + + +RPI
Sbjct: 467 PDGFTAEFFIDSWSIVGAEVTDAIKEFFSSGCLLKQWNATTIVLIPKIVNPTCTSDFRPI 526
Query: 448 SLIRSSYKLIAKVLATRLQKVMPKLLSQNQFAFTKERQIADCILLANEVADFL--SKKSE 505
S + + YK+IA++L RLQ+++ ++S Q AF R +A+ +LLA ++ S S
Sbjct: 527 SCLNTLYKVIARLLTDRLQRLLSGVISSAQSAFLPGRSLAENVLLATDLVHGYNWSNISP 586
Query: 506 GGLMLKVDLAKAYDQVDWEFLINMLHEMNFGDKWISWMKSCISSASLAILVNGSPTDFFS 565
G MLKVDL KA+D V WEF+I L + +K+I+W+ CIS+ + + +NG FF
Sbjct: 587 RG-MLKVDLKKAFDSVRWEFVIAALRALAIPEKFINWISQCISTPTFTVSINGGNGGFFK 645
Query: 566 IEKGLRQGDPLSPLLFNLCLNGLSCLLNQFLGEENYCGVRISRNLTINHLLFADDTLLFC 625
KGLRQGDPLSP LF L + S LL+ + NL+I+HL+FADD ++F
Sbjct: 646 STKGLRQGDPLSPYLFVLAMEAFSNLLHSRYESGLIHYHPKASNLSISHLMFADDVMIFF 705
Query: 626 ENNELQLETLCNVLITFLFASGLKVNNQKSELIGCNVQTVEVERLAHSFGWSVGTLPISY 685
+ L +C L F SGLKVN KS L + +E A ++G+ +GTLPI Y
Sbjct: 706 DGGSFSLHGICETLDDFASWSGLKVNKDKSHLYLAGLNQLESNANA-AYGFPIGTLPIRY 764
Query: 686 LGAPLGGNPRRIVFWNPMLEKLKCKTRSYNSKFMSLSGRLVITKSVLCSIPLFLMVVFKA 745
LG PL RI + P+LEK+ + RS+ +K +S +GR+ + SV+ F M F
Sbjct: 765 LGLPLMNRKLRIAEYEPLLEKITARFRSWVNKCLSFAGRIQLISSVIFGSINFWMSTFLL 824
Query: 746 PKGVILEVEKLFHAFIWGQEQGRKKINWIGWDQICLNRKAGGLGL-GFIEWRNRGLLLKW 804
PKG I +E L F+W + K + W +CL + GGLGL +EW N+ L ++
Sbjct: 825 PKGCIKRIESLCSRFLWSGNIEQAKGIKVSWAALCLPKSEGGLGLRRLLEW-NKTLSMRL 883
Query: 805 AWRFGREINSLWRFFVIERYNLDPDSLIWHMASTTPAMGSRAIQDIWRVLNEDHSLSAGF 864
WR +SLW + ++L S W + G ++ W+ L L+ F
Sbjct: 884 IWRLFVAKDSLWADWQ-HLHHLSRGS-FWAVE------GGQSDSWTWKRLLSLRPLAHQF 935
Query: 865 KENLLISIGDGSRVKFWRDPWVSNVALCHLFPRLYAMSENKEARIAEVGTYSRGKWGWSL 924
L+ +G+G + +W D W S L + + S + +A+V + + + GW L
Sbjct: 936 ---LVCKVGNGLKADYWYDNWTSLGPLFRIIGDI-GPSSLRVPLLAKVAS-AFSEDGWRL 990
Query: 925 TFRGQFFAWEEELYQQFLASINHIIPAAVGGDQLIWCANQTGLFSVKELCYWLEDHLFTH 984
A + L ++ A D+ W N +L
Sbjct: 991 PVSRSAPA---KGIHDHLCTVPVPSTAQEDVDRYEWSVNG-----------FLCQGFSAA 1036
Query: 985 QMWQV--PKSIAKL---------LPPKVALFFWQAQNDKIATRANLLARGIDLNGSTQCL 1033
+ W+ PK+ K PK A W + +++ TR L + G S C+
Sbjct: 1037 KTWEAIRPKATVKSWASSIWFKGAVPKYAFNMWVSHLNRLLTRQRLASWG--HIQSDACV 1094
Query: 1034 FCNAFDESCNHIFLHCHSIWSLWCVIVNRERFLWVMPASLADLILEWDFLPKISDKSLWS 1093
C+ ES +H+ L C +W ++ R + +S ++L+ + L
Sbjct: 1095 LCSFASESRDHLLLICEFSAQVWRLVFRRICPRQRLFSSWSELLSWVRQSSPEAPPLLRK 1154
Query: 1094 LIPYALSWSIWIERNNICFN 1113
++ + +++W +RNN+ N
Sbjct: 1155 IVSQVVVYNLWRQRNNLLHN 1174
>UniRef100_O22220 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1094
Score = 285 bits (729), Expect = 6e-75
Identities = 196/664 (29%), Positives = 319/664 (47%), Gaps = 45/664 (6%)
Query: 412 EDFFSSGRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRLQKVMPK 471
E FF +G + N + LIPK + + RPISL YK+I+K+L+ RL+K +P
Sbjct: 201 ELFFQTGILPEDWNHTHLCLIPKITKPARMADIRPISLCSVMYKIISKILSARLKKYLPV 260
Query: 472 LLSQNQFAFTKERQIADCILLANEVADFL---SKKSEGGLMLKVDLAKAYDQVDWEFLIN 528
++S Q AF ER ++D I+LA+E+ L K S+ ++ K D++KAYD+V+W FL
Sbjct: 261 IVSPTQSAFVAERLVSDNIILAHEIVHNLRTNEKISKDFMVFKTDMSKAYDRVEWPFLKG 320
Query: 529 MLHEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNGL 588
+L + F WI+WM +C+SS S ++L+NG P + +GLRQGDPLSP LF LC L
Sbjct: 321 ILLALGFNSTWINWMMACVSSVSYSVLINGQPFGHITPHRGLRQGDPLSPFLFVLCTEAL 380
Query: 589 SCLLNQFLGEENYCGVRIS-RNLTINHLLFADDTLLFCENNELQLETLCNVLITFLFASG 647
+LNQ G++ + ++NHLLFADDTLL C+ ++L+ + + L + SG
Sbjct: 381 IHILNQAEKIGKISGIQFNGTGPSVNHLLFADDTLLICKASQLECAEIMHCLSQYGHISG 440
Query: 648 LKVNNQKSEL-IGCNVQTVEVERLAHSFGWSVGTLPISYLGAPLGGNPRRIVFWNPMLEK 706
+N++KS + G V + + + G YLG P + V + + EK
Sbjct: 441 QMINSEKSAITFGAKVNEETKQWIMNRSGIQTEGGTGKYLGLPECFQGSKQVLFGFIKEK 500
Query: 707 LKCKTRSYNSKFMSLSGRLVITKSVLCSIPLFLMVVFKAPKGVILEVEKLFHAFIWGQEQ 766
L+ + + +K +S G+ ++ KS+ + P++ M F+ K + ++ + F W Q
Sbjct: 501 LQSRLSGWYAKTLSQGGKDILLKSIAMAFPVYAMTCFRLSKTLCTKLTSVMMDFWWNSVQ 560
Query: 767 GRKKINWIGWDQICLNRKAGGLGLGFIEWRNRGLLLKWAWRFGREINSLWRFFVIERYNL 826
+KKI+WIG ++ L + GG G ++ N+ LL K A R + +SL + RY +
Sbjct: 561 DKKKIHWIGAQKLMLPKFLGGFGFKDLQCFNQALLAKQASRLHTDSDSLLSQILKSRYYM 620
Query: 827 DPDSLIWHMASTTPAMGSRAIQDIWRVLNEDHSLSAGFKENLLISIGDGSRVKFWRDPWV 886
+ D L + G+R +L L +G K+ IG+G W D W+
Sbjct: 621 NSDFL-------SATKGTRPSYAWQSILYGRELLVSGLKK----IIGNGENTYVWMDNWI 669
Query: 887 SNVALCHLFPRLYAMSENKEARIAE-VGTYSRGKWGWSLTFRGQFFAWEEELYQQFLASI 945
+ P + + + ++++ + +SR W+L F W+E + I
Sbjct: 670 FDDKPRR--PESLQIMVDIQLKVSQLIDPFSRN---WNLNMLRDLFPWKE------IQII 718
Query: 946 NHIIPAAVGGDQLIWCANQTGLFSVKELCYWLEDHLFTHQMWQVPKSIAKLLP------- 998
P A D W GL++VK Y L QM++ + L P
Sbjct: 719 CQQRPMASRQDSFCWFGTNHGLYTVKSE-YDLCSRQVHKQMFKEAEEQPSLNPLFGKIWN 777
Query: 999 ----PKVALFFWQAQNDKIATRANLLARGIDLNGSTQCLFCNAFDESCNHIFLHC---HS 1051
PK+ +F W+ +A L RG+ + C C +E+ NHI C
Sbjct: 778 LNSAPKIKVFLWKVLKGAVAVEDRLRTRGVLIEDG--CSMCPEKNETLNHILFQCPLARQ 835
Query: 1052 IWSL 1055
+W+L
Sbjct: 836 VWAL 839
>UniRef100_O82216 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1216
Score = 282 bits (721), Expect = 5e-74
Identities = 214/732 (29%), Positives = 353/732 (47%), Gaps = 60/732 (8%)
Query: 412 EDFFSSGRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRLQKVMPK 471
+ FF+ G + +G+N+ + LIPKK+ A + YRPIS YK I+K+LA RL++++PK
Sbjct: 218 QSFFAKGFLPKGVNSTILALIPKKKEAREIKDYRPISCCNVLYKAISKILANRLKRILPK 277
Query: 472 LLSQNQFAFTKERQIADCILLANE-VADFLSKKSEGGLMLKVDLAKAYDQVDWEFLINML 530
+ NQ AF K+R + + +LLA E V D+ +K+D++KA+D + W FL ++L
Sbjct: 278 FIVGNQSAFVKDRLLIENVLLATELVKDYHKDSISTRCAMKIDISKAFDSLQWSFLTHVL 337
Query: 531 HEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNGLSC 590
MNF ++I W+ C+S+AS +I VNG +F +GLRQG LSP LF + ++ LS
Sbjct: 338 AAMNFPGEFIHWISLCMSTASFSIQVNGELAGYFRSARGLRQGCSLSPYLFVISMDVLSR 397
Query: 591 LLNQFLGEENYCGVRISRNLTINHLLFADDTLLFCENNELQLETLCNVLITFLFASGLKV 650
+L++ G + + L + HL FADD ++ + ++ + VL F GLK+
Sbjct: 398 MLDKAAGAREFGYHPRCKTLGLTHLCFADDLMILTDGKIRSVDGIVKVLNQFAAKLGLKI 457
Query: 651 NNQKSELIGCNVQTVEVERLAHSFGWSVGTLPISYLGAPLGGNPRRIVFWNPMLEKLKCK 710
+K+ L V + ++ + + VG LP+ YLG PL ++P++++++ +
Sbjct: 458 CMEKTTLYLAGVSDHSRQLMSSRYSFGVGKLPVRYLGLPLVTKRLTTSDYSPLIDQIRRR 517
Query: 711 TRSYNSKFMSLSGRLVITKSVLCSIPLFLMVVFKAPKGVILEVEKLFHAFIWGQEQGRKK 770
+ S+++S +GRL + SVL SI F M F+ P+ I E+ ++ A +W + K
Sbjct: 518 IGMWTSRYLSFAGRLSLINSVLWSITNFWMNAFRLPRECINEINRISSALLWSGPELNPK 577
Query: 771 INWIGWDQICLNRKAGGLGLGFIEWRNRGLLLKWAWRFGREINSLWRFFVIERYNLDPDS 830
+ WD+IC +K GGLGL + N+ LK WR +SLW + R NL
Sbjct: 578 KAKVSWDEICKPKKEGGLGLQSLREANKVSSLKLIWRLLSCQDSLWVKWT--RMNLLKKE 635
Query: 831 LIWHMASTTPAMGSRAIQDIWRVLNEDHSLSAGFKENLLISIGDGSRVKFWRDPWVSNVA 890
W + T +GS IWR L + ++ F + I + +G FW D W
Sbjct: 636 SFWSI-GTHSTLGSW----IWRRLLKHREVAKSFCK---IEVNNGVNTSFWFDNWSEKGP 687
Query: 891 LCHLFPRLYA--MSENKEARIAEVGTYSRGKWGWSLTFRGQFFAWEEELYQQFLASINHI 948
L +L A M ++ +AE + R K R + EE+ Q N
Sbjct: 688 LINLTGARGAIDMGISRHMTLAEAWSRRRRK-----RHRVEILNEFEEILLQKYQHRNIE 742
Query: 949 IPAAVGGDQLIWCANQ---TGLFSVKELCYWLEDHLFT---HQMWQVPKSIAKLLPPKVA 1002
+ D ++W + FS K+ W +H+ T + W A PK +
Sbjct: 743 LE-----DAILWRGKEDVFKARFSTKDT--W--NHIRTSSNQRAWHKGVWFAH-ATPKFS 792
Query: 1003 LFFWQAQNDKIATRANLLARGIDLNGS-TQCLFCNAFDESCNHIFLHC---HSIWSLWCV 1058
W A ++++T ++ NG+ T C+FC++ E+ +H+F C IW+
Sbjct: 793 FCAWLAIRNRLSTGDRMMTWN---NGTPTTCVFCSSPMETRDHLFFQCCYSSEIWTSIAK 849
Query: 1059 IVNRERFLWVMPASLADLILEWDFLPK-ISD------KSLWSLIPYALS-WSIWIERNNI 1110
V ++RF +W + ISD +S S + +S SIW ERN+
Sbjct: 850 NVYKDRF-----------STKWSAVVNYISDSQPDRIQSFLSRYTFQVSIHSIWRERNSR 898
Query: 1111 CFNHKALNISEI 1122
K+ + S +
Sbjct: 899 RHGEKSRSASNL 910
>UniRef100_Q9FZG5 T2E6.4 [Arabidopsis thaliana]
Length = 740
Score = 281 bits (718), Expect = 1e-73
Identities = 163/484 (33%), Positives = 263/484 (53%), Gaps = 11/484 (2%)
Query: 403 SGAWFEWMFEDFFSSGRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLA 462
+G F + FF G + +G+NA + LIPKK+ A+ + YRPIS YK+I+K++A
Sbjct: 61 TGQDFIAAIKSFFIKGFLPKGLNATILALIPKKDEATLMRDYRPISCCNVIYKVISKIIA 120
Query: 463 TRLQKVMPKLLSQNQFAFTKERQIADCILLANE-VADFLSKKSEGGLMLKVDLAKAYDQV 521
RL+ ++P + QNQ AF +ER + + +LLA E V D+ +K+D++KA+D V
Sbjct: 121 NRLKVMLPTFILQNQSAFVRERLLIENVLLATELVKDYHKDSISPRCAMKIDISKAFDSV 180
Query: 522 DWEFLINMLHEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLF 581
W+FL+N L +NF + + W+K CIS+A+ ++ VNG FF ++GLRQG LSP LF
Sbjct: 181 QWQFLLNTLEALNFPENFCHWIKLCISTATFSVQVNGELAGFFGSKRGLRQGCALSPYLF 240
Query: 582 NLCLNGLSCLLNQFLGEENYCGVRISRNLTINHLLFADDTLLFCENNELQLETLCNVLIT 641
+C+N LS +++ N + L++ HL FADD ++F + + +E + N+
Sbjct: 241 VICMNVLSHMIDVAAVHRNIGYHPKCKKLSLTHLCFADDLMVFIDGQQRSVEGVINIFKE 300
Query: 642 FLFASGLKVNNQKSELIGCNVQTVEVERLAHSFGWSVGTLPISYLGAPLGGNPRRIVFWN 701
F SGL ++ +KS L V + + +F ++ G LP+ YLG PL ++
Sbjct: 301 FAGKSGLHISLEKSTLYLAGVSELNRNNILSAFPFASGQLPVRYLGLPLLTKQMTTADYS 360
Query: 702 PMLEKLKCKTRSYNSKFMSLSGRLVITKSVLCSIPLFLMVVFKAPKGVILEVEKLFHAFI 761
P+L+K++ K S+ ++ +S +GRL + SV+ S+ F M ++ P G I E+EKL AF+
Sbjct: 361 PLLDKVRSKISSWTARSLSYAGRLALINSVIVSLSNFWMSAYRLPAGCIKEIEKLCSAFL 420
Query: 762 WGQEQGRKKINWIGWDQICLNRKAGGLGLGFIEWRNRGLLLKWAWRFGREINSLWRFFVI 821
W + K I W +C ++ GGLG+ + N+ LK WR +SLW +V
Sbjct: 421 WSGPELNPKKAKITWTSLCKLKQEGGLGIKSLLEANKVSCLKLIWRLVSRQSSLWVNWV- 479
Query: 822 ERYNLDPDSLIWHMASTTPAMGSRAIQDIWRVLNEDHSLSAGFKENLLISIGDGSRVKFW 881
Y + S W A+ ++GS +W+ L + ++ K + I GS FW
Sbjct: 480 WTYIIRKGS-FW-SANDRSSLGSW----MWKKLLKYRDVA---KSMCKVEIKSGSSTSFW 530
Query: 882 RDPW 885
D W
Sbjct: 531 YDNW 534
>UniRef100_Q9FL83 Non-LTR retroelement reverse transcriptase-like protein [Arabidopsis
thaliana]
Length = 1223
Score = 275 bits (704), Expect = 5e-72
Identities = 192/677 (28%), Positives = 324/677 (47%), Gaps = 67/677 (9%)
Query: 404 GAWFEWMFEDFFSSGRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLAT 463
G F + FF+ G + +G+N+ + LIPKK A + YRPIS YK+I+K++A
Sbjct: 489 GDEFTLAVQSFFTKGFLPKGINSTILALIPKKTEAREMKDYRPISCCNVLYKVISKIIAN 548
Query: 464 RLQKVMPKLLSQNQFAFTKERQIADCILLANE-VADFLSKKSEGGLMLKVDLAKAYDQVD 522
RL+ V+PK ++ NQ AF K+R + + +LLA E V D+ +K+D++KA+D V
Sbjct: 549 RLKLVLPKFIAGNQSAFVKDRLLIENLLLATELVKDYHKDTISTRCAIKIDISKAFDSVQ 608
Query: 523 WEFLINMLHEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFN 582
W FLIN+ + F ++I W+ CI++AS ++ VNG +F +GLRQG LSP LF
Sbjct: 609 WPFLINVFTILGFPREFIHWINICITTASFSVQVNGELAGYFQSSRGLRQGCALSPYLFV 668
Query: 583 LCLNGLSCLLNQFLGEENYCGVRISRNLTINHLLFADDTLLFCENNELQLETLCNVLITF 642
+C++ LS +L++ ++ + + + HL FADD ++ + +E + V F
Sbjct: 669 ICMDVLSKMLDKAAAARHFGYHPKCKTMGLTHLSFADDLMVLSDGKIRSIERIIKVFDEF 728
Query: 643 LFASGLKVNNQKSELIGCNVQTVEVERLAHSFGWSVGTLPISYLGAPLGGNPRRIVFWNP 702
SGL+++ +KS + + +A F +S G LP+ YLG PL P
Sbjct: 729 AKWSGLRISLEKSTVYLAGLSATARNEVADRFPFSSGQLPVRYLGLPLITKRLSTTDCLP 788
Query: 703 MLEKLKCKTRSYNSKFMSLSGRLVITKSVLCSIPLFLMVVFKAPKGVILEVEKLFHAFIW 762
+LE+++ + S+ S+F+S +GRL + SVL SI F + F+ P+ I E+EK+ AF+W
Sbjct: 789 LLEQVRKRIGSWTSRFLSYAGRLNLISSVLWSICNFWLAAFRLPRKCIRELEKMCSAFLW 848
Query: 763 -GQEQGRKKINWIGWDQICLNRKAGGLGLGFIEWRNRGLLLKWAWRFGREINSLWRFFVI 821
G E K I W +C + GGLGL ++ N LK W+ NSLW +V
Sbjct: 849 SGTEMNSNKAK-ISWHMVCKPKDEGGLGLRSLKEANDVCCLKLVWKIVSHSNSLWVKWVD 907
Query: 822 ERYNLDPDSLIWHMASTTPAMGSRAIQDIWRVLNEDHSLSAGFKENLLISIGDGSRVKFW 881
+ +L ++ W + T + GS IW+ L + ++ + + +G+G + FW
Sbjct: 908 Q--HLLRNASFWEVKQTV-SQGSW----IWKKLLKYREVAKTLSK---VEVGNGKQTSFW 957
Query: 882 RDPWVSNVALCHLFPRLYAMSENKEARIAEVGTYSRGKWGWSLTFRGQFFAWEEELYQQF 941
D W L + + + ++G R +T AW ++
Sbjct: 958 YDNWSDLGQL---------LERTGDRGLIDLGISRR------MTVEE---AWTNRRQRRH 999
Query: 942 LASINHIIPAAV---------GGDQLIWCANQTGLFSVKELCYWLEDHLFTHQMWQVPKS 992
+ ++I A+ D+++W ++ +F T W +S
Sbjct: 1000 RNDVYNVIEDALKKSWDTRTETEDKVLW-RGKSDVFRTT---------FSTRDTWHHTRS 1049
Query: 993 IAKLLP-----------PKVALFFWQAQNDKIATRANLL--ARGIDLNGSTQCLFCNAFD 1039
+ +P PK + W A + ++ T ++ A GI +T C+FC
Sbjct: 1050 TSARVPWHKVIWFSHATPKYSFCSWLAAHGRLPTGDRMINWANGI----ATDCIFCQGTL 1105
Query: 1040 ESCNHIFLHCHSIWSLW 1056
E+ +H+F C +W
Sbjct: 1106 ETRDHLFFTCSFTSVIW 1122
>UniRef100_Q75M12 Hypothetical protein P0676G05.2 [Oryza sativa]
Length = 1936
Score = 274 bits (701), Expect = 1e-71
Identities = 211/715 (29%), Positives = 337/715 (46%), Gaps = 64/715 (8%)
Query: 381 LNPLTAIKPQGLTALI*NSFRNSGAWFEWMF---EDFFSSGRISRGMNAAFITLIPKKEN 437
+ PL + +P G A RN G + +FF SG + +G+N I LIPKK+
Sbjct: 1080 IGPLKSPRPDGFPARFYQ--RNWGTLKSDIILAVRNFFQSGLMPKGVNDTAIVLIPKKDQ 1137
Query: 438 ASSLNHYRPISLIRSSYKLIAKVLATRLQKVMPKLLSQNQFAFTKERQIADCILLANEVA 497
L YRPISL YK+++K L RL+ ++ L+S+ Q AF + R I D LLA E
Sbjct: 1138 PIDLKDYRPISLCNVVYKVVSKCLVNRLRPILDDLVSKEQSAFIQGRMITDNALLAFECF 1197
Query: 498 DFLSKKSEGG---LMLKVDLAKAYDQVDWEFLINMLHEMNFGDKWISWMKSCISSASLAI 554
+ K + K+DL+KAYD+VDW FL L+++ F +W+SW+ C+++ ++
Sbjct: 1198 HSIQKNKKANSAACAYKLDLSKAYDRVDWRFLELALNKLGFAHRWVSWIMLCVTTVRYSV 1257
Query: 555 LVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNGLSCLLNQFLGEENYCGVRISRNLT-IN 613
NG+ F+ +GLRQG+PLSP LF +GLS LL + + + + ++I R I+
Sbjct: 1258 KFNGTLLRSFAPTRGLRQGEPLSPFLFLFVADGLSLLLKEKVAQNSLTPLKICRQAPGIS 1317
Query: 614 HLLFADDTLLFCENNELQLETLCNVLITFLFASGLKVNNQKSELIGCNVQTVEV-ERLAH 672
+LLFADDTLLF + + + E + VL + +G +N K ++ V E + +
Sbjct: 1318 YLLFADDTLLFFKAEKKEAEVVKEVLTNYAQGTGQLINPAKCSILFGEASPSSVSEDIRN 1377
Query: 673 SFGWSVGTLPISYLGAPLGGNPRRIVFWNPMLEKLKCKTRSYNSKFMSLSGRLVITKSVL 732
+ YLG P + + K+ + + F+S G+ ++ K+V+
Sbjct: 1378 TLQVERDNFEDRYLGFPTPEGRMHKGRFQSLQAKIAKRVIQWGENFLSSGGKEILIKAVI 1437
Query: 733 CSIPLFLMVVFKAPKGVILEVEKLFHAFIWGQEQGRKKINWIGWDQICLNRKAGGLGLGF 792
+IP+++M +FK P V E+ K+ F WG + GR++ +W WD + + GGLG
Sbjct: 1438 QAIPVYVMGLFKFPDSVYDELTKMTRNFWWGADNGRRRTHWRAWDSLTKAKINGGLGFRD 1497
Query: 793 IEWRNRGLLLKWAWRFGREINSLWRFFVIERYNLDPDSLIWHMASTTPAMGSRAIQDIWR 852
+ N+ LL + AWR NSL + +Y + S T S W
Sbjct: 1498 YKLFNQALLTRQAWRLIEFPNSLCAQVLKAKY--------FPHGSLTDTTFSANASPTWH 1549
Query: 853 VLNEDHSLSAGFKENLLISIGDGSRVKFWRDPW---------VSNVALCHLFPRLYAMSE 903
+ L K+ ++ IG+G+ V+ WRDPW VS+ A C RL +S+
Sbjct: 1550 GIEYGLDL---LKKGIIWRIGNGNSVRIWRDPWIPRDLSRRPVSSKANC----RLKWVSD 1602
Query: 904 NKEARIAEVGTYSRGKWGWSLTFRGQFFAWEEELYQQFLASINHIIPAAVGGDQLIWCAN 963
IAE GT+ K F + ++ Q+ I A + D + W +
Sbjct: 1603 ----LIAEDGTWDSAK------INQYFLKIDADIIQKI------CISARLEEDFIAWHPD 1646
Query: 964 QTGLFSVK---ELCYWLED--HLFTHQMWQVPKSIAKL----LPPKVALFFWQAQNDKIA 1014
+TG FSV+ +L L D + + ++ KS + +P KV +F W+ ++ +A
Sbjct: 1647 KTGRFSVRSAYKLALQLADMNNCSSSSSSRLNKSWELIWKCNVPQKVRIFAWRVASNSLA 1706
Query: 1015 TRANLLARGIDLNGSTQCLFCNAFDESCNHIFLHCHSIWSLWCVIVNRERFLWVM 1069
T N R +L C C+ E H C SLW VN E+ +V+
Sbjct: 1707 TMENKKKR--NLERFDVCGICDREKEDAGHALCRCVHANSLW---VNLEKGKYVV 1756
Score = 40.4 bits (93), Expect = 0.36
Identities = 40/143 (27%), Positives = 60/143 (40%), Gaps = 2/143 (1%)
Query: 1169 WVTPDAGLIKINVDGAAKGSPGRCGIGGILRN-NANVILGFFSKSVGEGYAFEAEVLAIH 1227
W P G +K+NVDG+ + + GIG ILRN NVI EAE+ A
Sbjct: 1777 WERPRNGWMKLNVDGSFDINSEKGGIGMILRNCLGNVIFSSCRSLDSCSGPLEAELHACV 1836
Query: 1228 EALLFCVQRQLRNIIIESDSSLAVGWVNNRCNRPWKLLNKLNQIDRWLVETNCLKVNHIY 1287
E L + L I +E+D S + +N+ L N + + + ++ +
Sbjct: 1837 EGLHLALHWTLLPIQVETDCSSVIQLLNHPDKDRSVLANIAQEAKSLMAGDRQIAISKVQ 1896
Query: 1288 REANGEADKLA-KRGAEYPNDLW 1309
R N + LA K AE + W
Sbjct: 1897 RSQNVISHFLANKARAESLSSFW 1919
>UniRef100_Q9M1N9 Hypothetical protein T18B22.50 [Arabidopsis thaliana]
Length = 762
Score = 272 bits (695), Expect = 6e-71
Identities = 194/682 (28%), Positives = 321/682 (46%), Gaps = 49/682 (7%)
Query: 389 PQGLTA-LI*NSFRNSGAWFEWMFEDFFSSGRISRGMNAAFITLIPKKENASSLNHYRPI 447
P G T+ S+ G F + FF+ G + +G+N+ + LIPKK + + YRPI
Sbjct: 26 PDGFTSEFFKESWEILGPEFILAIQSFFALGFLPKGVNSTILALIPKKLESKEMKDYRPI 85
Query: 448 SLIRSSYKLIAKVLATRLQKVMPKLLSQNQFAFTKERQIADCILLANE-VADFLSKKSEG 506
S YK+I+K+LA RL+ ++P+ ++ NQ +F K+R + + +LLA + V D+
Sbjct: 86 SCCNVMYKVISKILANRLKLLLPQFIAGNQSSFVKDRLLIENVLLATDLVKDYHKDSISE 145
Query: 507 GLMLKVDLAKAYDQVDWEFLINMLHEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSI 566
+K+D++KA D V W FLIN L M+F + +I W++ CI++ S ++ VNG FF
Sbjct: 146 RCAIKIDISKASDSVQWSFLINTLTAMHFPEMFIHWIRLCITTPSFSVQVNGELAGFFQS 205
Query: 567 EKGLRQGDPLSPLLFNLCLNGLSCLLNQFLGEENYCGVRISRNLTINHLLFADDTLLFCE 626
+GLRQG LSP LF +C++ LS LL++ +G + + + HL FADD ++ +
Sbjct: 206 SRGLRQGCALSPYLFVICMDVLSKLLDKVVGIGRIGYHPHCKRMGLTHLSFADDLMILTD 265
Query: 627 NNELQLETLCNVLITFLFASGLKVNNQKSELIGCNVQTVEVERLAHSFGWSVGTLPISYL 686
+E + V F SGLK++ +KS + + + +L F + VG LPI YL
Sbjct: 266 GQCRSIEGIIEVFDLFSKWSGLKISMEKSTIFSAGLSSTSRAQLHTHFPFEVGELPIRYL 325
Query: 687 GAPLGGNPRRIVFWNPMLEKLKCKTRSYNSKFMSLSGRLVITKSVLCSIPLFLMVVFKAP 746
G PL V + P++E+++ + S++S+F+S +GR + S++ S F + F+ P
Sbjct: 326 GLPLVTKRLSSVDYAPLIEQIRKRIGSWSSRFLSFAGRFNLISSIIWSSCNFWLSAFQLP 385
Query: 747 KGVILEVEKLFHAFIWGQEQGRKKINWIGWDQICLNRKAGGLGLGFIEWRNRGLLLKWAW 806
+ I E+EKL +F+W K I W+Q+C + GGLGL ++ N LK W
Sbjct: 386 RACIQEIEKLCSSFLWSGTNLNSKKAKISWNQVCKPKSEGGLGLRSLKEANDVCCLKLVW 445
Query: 807 RFGREINSLWRFFVIERYNLDPDSLIWHMASTTPAMGSRAIQDIWRVLNEDHSLSAGFKE 866
R +SLW +V +NL + W + +GS IW+ + + ++ F +
Sbjct: 446 RIISHGDSLWVKWV--EHNLLKREIFW-IVKENANLGSW----IWKKILKYRGVAKRFCK 498
Query: 867 NLLISIGDGSRVKFWRDPWVSNVALCHL--FPRLYAMSENKEARIAEVGTYSRGKWGWSL 924
+G+G FW D W L + M ++ +A+ T R
Sbjct: 499 ---AEVGNGESTSFWFDDWSLLGRLIDVAGIRGTIDMGISRTMSVADAWTSRR------- 548
Query: 925 TFRGQFFAWEEELYQQFLASINHIIPAAVGGDQLIWCANQTGLFSVKELCYWLEDHLFTH 984
Q+ L +I ++ Q Q G K +D T
Sbjct: 549 ---------RRHHRQEILNTIEEVLSTQ---HQKRTQQQQQGRVLWKGKNDIYKDKFSTK 596
Query: 985 QMWQVPKSIAKLL-----------PPKVALFFWQAQNDKIATRANLLARGIDLNGSTQCL 1033
W ++ + + PK + W A +D++AT A ++ G C
Sbjct: 597 NTWNYLRTTSNEVAWHKGVWFPHATPKYSFCLWLAAHDRLATGARMIKWNRGETG--DCT 654
Query: 1034 FCNAFDESCNHIF---LHCHSI 1052
FC E+ +H+F LH S+
Sbjct: 655 FCRQGIETRDHLFFMQLHIGSL 676
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.335 0.145 0.488
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,152,357,513
Number of Sequences: 2790947
Number of extensions: 88401014
Number of successful extensions: 283142
Number of sequences better than 10.0: 1373
Number of HSP's better than 10.0 without gapping: 856
Number of HSP's successfully gapped in prelim test: 517
Number of HSP's that attempted gapping in prelim test: 279869
Number of HSP's gapped (non-prelim): 2111
length of query: 1310
length of database: 848,049,833
effective HSP length: 139
effective length of query: 1171
effective length of database: 460,108,200
effective search space: 538786702200
effective search space used: 538786702200
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 81 (35.8 bits)
Lotus: description of TM0009.7


