

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0009.6
(309 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
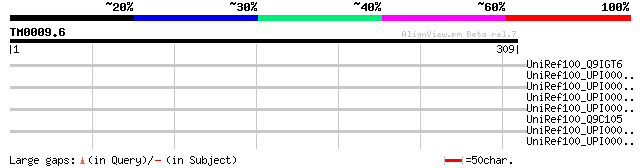
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9IGT6 162R [Porcine adenovirus type 3] 36 1.1
UniRef100_UPI0000364547 UPI0000364547 UniRef100 entry 35 2.5
UniRef100_UPI0000364546 UPI0000364546 UniRef100 entry 35 2.5
UniRef100_UPI0000364545 UPI0000364545 UniRef100 entry 35 2.5
UniRef100_UPI000032EFB4 UPI000032EFB4 UniRef100 entry 35 3.3
UniRef100_Q9C105 SPAPB1E7.04c protein [Schizosaccharomyces pombe] 34 4.3
UniRef100_UPI000041A584 UPI000041A584 UniRef100 entry 34 5.7
UniRef100_UPI000035F026 UPI000035F026 UniRef100 entry 34 5.7
>UniRef100_Q9IGT6 162R [Porcine adenovirus type 3]
Length = 162
Score = 36.2 bits (82), Expect = 1.1
Identities = 30/108 (27%), Positives = 46/108 (41%), Gaps = 3/108 (2%)
Query: 35 GGFSSTLGGSFTQISGFVLVFLLGHHHL---PLTAATLSSTAIPVAAAATLASSSLFRRH 91
GG++ST GS + + V + P AT SST I +++++ AS ++ R
Sbjct: 52 GGWASTPAGSGSPTTVIVRTISSSSSSMVFAPAPGATSSSTTISSSSSSSAASEAVARSC 111
Query: 92 LPPGCQSSRSGTVLLRSRCRVDPYLVPLKHTGFAFRRRSTTSAEPFCC 139
+ +SS S S CR+ P G R +A P CC
Sbjct: 112 ISTTSRSSCSCCRSSASACRIMPTSALRSTDGNTARNTDPVAAVPGCC 159
>UniRef100_UPI0000364547 UPI0000364547 UniRef100 entry
Length = 518
Score = 35.0 bits (79), Expect = 2.5
Identities = 30/117 (25%), Positives = 55/117 (46%), Gaps = 6/117 (5%)
Query: 70 SSTAIPVAAAATLASSSLFRRHL-PPGCQSSRSGTVLLRSRCRVDPYLVPLKHTGFAFRR 128
S + P+AAAA L + + HL PP ++S + +LR C V +P G
Sbjct: 174 SLSGCPIAAAAKLNKTQDKQVHLQPPASEASSNSDRVLRPMCFVKQLEIP--QYGSFRPN 231
Query: 129 RSTTSAEPFCCKEGARVKNVNRTTINHALITRSVFINQTLVPVSLELKDTVVRAYRT 185
TT+ KE ++ ++ + ++A VF +TL+P + +T +A+++
Sbjct: 232 TVTTTPRANLAKE---LEKYSKVSFDYASFDVQVFGKRTLIPKTPSSTETSPKAFKS 285
>UniRef100_UPI0000364546 UPI0000364546 UniRef100 entry
Length = 907
Score = 35.0 bits (79), Expect = 2.5
Identities = 30/117 (25%), Positives = 55/117 (46%), Gaps = 6/117 (5%)
Query: 70 SSTAIPVAAAATLASSSLFRRHL-PPGCQSSRSGTVLLRSRCRVDPYLVPLKHTGFAFRR 128
S + P+AAAA L + + HL PP ++S + +LR C V +P G
Sbjct: 468 SLSGCPIAAAAKLNKTQDKQVHLQPPASEASSNSDRVLRPMCFVKQLEIP--QYGSFRPN 525
Query: 129 RSTTSAEPFCCKEGARVKNVNRTTINHALITRSVFINQTLVPVSLELKDTVVRAYRT 185
TT+ KE ++ ++ + ++A VF +TL+P + +T +A+++
Sbjct: 526 TVTTTPRANLAKE---LEKYSKVSFDYASFDVQVFGKRTLIPKTPSSTETSPKAFKS 579
>UniRef100_UPI0000364545 UPI0000364545 UniRef100 entry
Length = 1022
Score = 35.0 bits (79), Expect = 2.5
Identities = 30/117 (25%), Positives = 55/117 (46%), Gaps = 6/117 (5%)
Query: 70 SSTAIPVAAAATLASSSLFRRHL-PPGCQSSRSGTVLLRSRCRVDPYLVPLKHTGFAFRR 128
S + P+AAAA L + + HL PP ++S + +LR C V +P G
Sbjct: 575 SLSGCPIAAAAKLNKTQDKQVHLQPPASEASSNSDRVLRPMCFVKQLEIP--QYGSFRPN 632
Query: 129 RSTTSAEPFCCKEGARVKNVNRTTINHALITRSVFINQTLVPVSLELKDTVVRAYRT 185
TT+ KE ++ ++ + ++A VF +TL+P + +T +A+++
Sbjct: 633 TVTTTPRANLAKE---LEKYSKVSFDYASFDVQVFGKRTLIPKTPSSTETSPKAFKS 686
>UniRef100_UPI000032EFB4 UPI000032EFB4 UniRef100 entry
Length = 949
Score = 34.7 bits (78), Expect = 3.3
Identities = 20/44 (45%), Positives = 26/44 (58%), Gaps = 3/44 (6%)
Query: 7 RIDQRMKKIHNSKDKSNPT---ILGRFLPTSGGFSSTLGGSFTQ 47
R+ +R+ K +N+ + N T L RFLP S G S L GSFTQ
Sbjct: 359 RLQERLSKSNNTSENFNLTGKIDLHRFLPRSLGISIPLNGSFTQ 402
>UniRef100_Q9C105 SPAPB1E7.04c protein [Schizosaccharomyces pombe]
Length = 1236
Score = 34.3 bits (77), Expect = 4.3
Identities = 38/158 (24%), Positives = 70/158 (44%), Gaps = 11/158 (6%)
Query: 17 NSKDKSNPTILGRFLPTSGGFSSTLGGSFTQISGFVLVFLLGHHHLPLTAA--TLSSTAI 74
+S S+ TIL P++ SS + S + ISG + +P++++ T SS+ I
Sbjct: 615 SSSISSSSTILSSPTPST---SSLMISSSSIISGSSSILSSSISTIPISSSLSTYSSSVI 671
Query: 75 PVAAAATLASSSLFRRHLPPGCQSSRSGTVLLRSRCRVDPY---LVPLKHTGFAFRRRST 131
P ++ +SSSL P +S S + + S V Y L + H+ + S+
Sbjct: 672 PSSSTLVSSSSSLIVSSSP---VASSSSSPIPSSSSLVSTYSASLSNITHSSLSLTAMSS 728
Query: 132 TSAEPFCCKEGARVKNVNRTTINHALITRSVFINQTLV 169
+SA P + + T+ ++ + S ++ T V
Sbjct: 729 SSAIPTSVNSSTLITASSSNTLLSSITSSSAIVSSTTV 766
>UniRef100_UPI000041A584 UPI000041A584 UniRef100 entry
Length = 574
Score = 33.9 bits (76), Expect = 5.7
Identities = 18/59 (30%), Positives = 30/59 (50%), Gaps = 1/59 (1%)
Query: 2 VKSTPRIDQRMKKIHNSKDKSNPTILGRFLPTSGGFSSTLGGSFTQISGFVLVFLLGHH 60
+++T R+ M I ++ + GR L GG ST GGSF Q + ++ ++G H
Sbjct: 119 LRNTSRLFAAMGCIEKVQENQKNKVRGR-LSQQGGAQSTAGGSFLQAAALLVTLIIGGH 176
>UniRef100_UPI000035F026 UPI000035F026 UniRef100 entry
Length = 532
Score = 33.9 bits (76), Expect = 5.7
Identities = 24/84 (28%), Positives = 41/84 (48%), Gaps = 12/84 (14%)
Query: 4 STPRIDQRMKKIHNSKDKSNPTILGRFLPTSGGFSSTLGGSFTQISGFVLVFLLGHHHLP 63
S P + K +S P ++ +P+SG FS+T G+ S FV
Sbjct: 192 SPPSATSNSSSVTAGKSESAPAVMSSSVPSSGEFSNTSSGA----SSFV--------DEA 239
Query: 64 LTAATLSSTAIPVAAAATLASSSL 87
+ AT S+A P ++A++++SSS+
Sbjct: 240 SSTATAVSSAAPSSSASSVSSSSV 263
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.323 0.137 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 510,753,280
Number of Sequences: 2790947
Number of extensions: 20541573
Number of successful extensions: 45292
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 45289
Number of HSP's gapped (non-prelim): 11
length of query: 309
length of database: 848,049,833
effective HSP length: 127
effective length of query: 182
effective length of database: 493,599,564
effective search space: 89835120648
effective search space used: 89835120648
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0009.6


