

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0005.4
(283 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
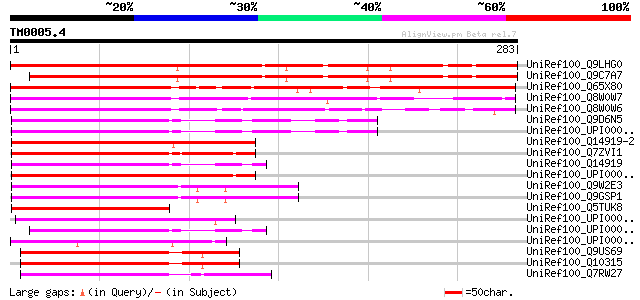
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LHG0 Arabidopsis thaliana genomic DNA, chromosome 3,... 287 2e-76
UniRef100_Q9C7A7 Hypothetical protein T2E22.20 [Arabidopsis thal... 266 3e-70
UniRef100_Q65X80 Hypothetical protein OJ1579_G03.2 [Oryza sativa] 242 7e-63
UniRef100_Q8W0W7 Repressor protein [Oryza sativa] 202 8e-51
UniRef100_Q8W0W6 Repressor protein [Zea mays] 196 7e-49
UniRef100_Q9D6N5 Dr1-associated corepressor [Mus musculus] 119 7e-26
UniRef100_UPI000018197E UPI000018197E UniRef100 entry 116 6e-25
UniRef100_Q14919-2 Splice isoform 2 of Q14919 [Homo sapiens] 115 1e-24
UniRef100_Q7ZVI1 Wu:fb26e10 protein [Brachydanio rerio] 115 1e-24
UniRef100_Q14919 Dr1-associated corepressor [Homo sapiens] 113 6e-24
UniRef100_UPI00003628E8 UPI00003628E8 UniRef100 entry 112 1e-23
UniRef100_Q9W2E3 CG10318-PA, isoform A [Drosophila melanogaster] 105 1e-21
UniRef100_Q9GSP1 NC2alpha [Drosophila melanogaster] 105 1e-21
UniRef100_Q5TUK8 ENSANGP00000026107 [Anopheles gambiae str. PEST] 105 1e-21
UniRef100_UPI000023655D UPI000023655D UniRef100 entry 102 1e-20
UniRef100_UPI000036EEB8 UPI000036EEB8 UniRef100 entry 100 3e-20
UniRef100_UPI00003C1340 UPI00003C1340 UniRef100 entry 100 4e-20
UniRef100_Q9US69 Hypothetical nuclear protein [Schizosaccharomyc... 99 1e-19
UniRef100_Q10315 Hypothetical protein C17G8.03c in chromosome I ... 99 1e-19
UniRef100_Q7RW27 Hypothetical protein [Neurospora crassa] 97 5e-19
>UniRef100_Q9LHG0 Arabidopsis thaliana genomic DNA, chromosome 3, P1 clone: MQC3
[Arabidopsis thaliana]
Length = 293
Score = 287 bits (735), Expect = 2e-76
Identities = 168/300 (56%), Positives = 202/300 (67%), Gaps = 24/300 (8%)
Query: 1 MKKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK 60
M+KKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSK+LELFLQDLCDRTYEITL+RGAK
Sbjct: 1 MRKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKSLELFLQDLCDRTYEITLERGAK 60
Query: 61 TMNSLHLKHCVQSYNVFDFLRDVVSKVPDYSH--GHGHSEPSADDRTIPKRRKAAGDDCN 118
T++SLHLKHCV+ YNVFDFLR+VVSKVPDY H G GH + + DDR+I KRRK D+ N
Sbjct: 61 TVSSLHLKHCVERYNVFDFLREVVSKVPDYGHSQGQGHGDVTMDDRSISKRRKPISDEVN 120
Query: 119 DSDEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRG----RTAVRETPHQAVEPEPCASVQ 174
DSDEE K+ K E+ ++GRG GRGRGRGRGRG + A RE ++ +E E S Q
Sbjct: 121 DSDEEYKKSKTQEIGSAKTSGRG-GRGRGRGRGRGGRAAKAAEREGLNREMEVEAANSGQ 179
Query: 175 QGIQHDTNTDMTIHDSSETKELPK--ENAAVPAESAEFH--------NLDLNANTNE-NE 223
+ N M +SS ++ K + A E + H + DLNA + + NE
Sbjct: 180 P--PPEDNVKMHASESSPQEDEKKGIDGTAASNEDTKQHLQSPKEGIDFDLNAESLDLNE 237
Query: 224 DKKASTTAKQEISEPPTESQHEEIPGWSLSDVDKMAIDSVQLANLGTHIEEDEEDYDEEG 283
K A T + T+S EE GW + D+ KM D QLA+LG I+EDEEDYDEEG
Sbjct: 238 TKLAPATGTTTTTTAATDS--EEYSGWPMMDISKM--DPAQLASLGKRIDEDEEDYDEEG 293
>UniRef100_Q9C7A7 Hypothetical protein T2E22.20 [Arabidopsis thaliana]
Length = 297
Score = 266 bits (681), Expect = 3e-70
Identities = 158/289 (54%), Positives = 191/289 (65%), Gaps = 24/289 (8%)
Query: 12 ARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHLKHCV 71
ARIKKIMQADEDVGKIALAVPVLVSK+LELFLQDLCDRTYEITL+RGAKT++SLHLKHCV
Sbjct: 16 ARIKKIMQADEDVGKIALAVPVLVSKSLELFLQDLCDRTYEITLERGAKTVSSLHLKHCV 75
Query: 72 QSYNVFDFLRDVVSKVPDYSH--GHGHSEPSADDRTIPKRRKAAGDDCNDSDEEVKRGKM 129
+ YNVFDFLR+VVSKVPDY H G GH + + DDR+I KRRK D+ NDSDEE K+ K
Sbjct: 76 ERYNVFDFLREVVSKVPDYGHSQGQGHGDVTMDDRSISKRRKPISDEVNDSDEEYKKSKT 135
Query: 130 PELSHTGSTGRGRGRGRGRGRGRG----RTAVRETPHQAVEPEPCASVQQGIQHDTNTDM 185
E+ ++GRG GRGRGRGRGRG + A RE ++ +E E S Q + N M
Sbjct: 136 QEIGSAKTSGRG-GRGRGRGRGRGGRAAKAAEREGLNREMEVEAANSGQP--PPEDNVKM 192
Query: 186 TIHDSSETKELPK--ENAAVPAESAEFH--------NLDLNANTNE-NEDKKASTTAKQE 234
+SS ++ K + A E + H + DLNA + + NE K A T
Sbjct: 193 HASESSPQEDEKKGIDGTAASNEDTKQHLQSPKEGIDFDLNAESLDLNETKLAPATGTTT 252
Query: 235 ISEPPTESQHEEIPGWSLSDVDKMAIDSVQLANLGTHIEEDEEDYDEEG 283
+ T+S EE GW + D+ KM D QLA+LG I+EDEEDYDEEG
Sbjct: 253 TTTAATDS--EEYSGWPMMDISKM--DPAQLASLGKRIDEDEEDYDEEG 297
>UniRef100_Q65X80 Hypothetical protein OJ1579_G03.2 [Oryza sativa]
Length = 290
Score = 242 bits (618), Expect = 7e-63
Identities = 153/305 (50%), Positives = 192/305 (62%), Gaps = 39/305 (12%)
Query: 1 MKKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK 60
M+KKLDTRFPA RIKKIMQADEDVGKIALAVPVLVSKALELFLQDLC+RTY+IT+QRG K
Sbjct: 1 MRKKLDTRFPAPRIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCNRTYDITVQRGVK 60
Query: 61 TMNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDS 120
T++S HLK C+ SYNV+DFLRDVVSKVPD G S+ DD+ + KRRK A D DS
Sbjct: 61 TLSSSHLKQCIHSYNVYDFLRDVVSKVPDM----GTSDAGVDDK-LGKRRKTAED---DS 112
Query: 121 DEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGRTAVRE--TPHQAVE-----PEPCASV 173
+EE KR + S T STGRGRGRGRGRGR GR + RE + ++ E P S
Sbjct: 113 EEESKRTRNEAASQT-STGRGRGRGRGRGRRGGRVSEREIISAYEKFEENHEFPPGQFSK 171
Query: 174 QQGIQHDTNTDMTIHDSSETKE-LPKENAAVPAESAEFHNLDLNANTNENEDKKA----- 227
++ D + D T D+ ETKE P NA A N+DLN + +D+ +
Sbjct: 172 PSQLKVDVSVDGT--DAIETKEATPLSNA-----RASLRNIDLNIELTDYDDEGSAPLEV 224
Query: 228 ----------STTAKQEISEPPTESQHEEIPGWSLSDVDKMAIDSVQLANLGTHIEEDEE 277
+T++ +SE E++ ++ GW L ++ KMA+D VQ A H E++E
Sbjct: 225 QPPAPAAGVVTTSSGPLVSEVNEEAKTKDFLGWQLPELTKMAMDPVQFALSSNHRLEEDE 284
Query: 278 DYDEE 282
DYD E
Sbjct: 285 DYDNE 289
>UniRef100_Q8W0W7 Repressor protein [Oryza sativa]
Length = 258
Score = 202 bits (514), Expect = 8e-51
Identities = 127/285 (44%), Positives = 161/285 (55%), Gaps = 31/285 (10%)
Query: 1 MKKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK 60
M+KKL TRFPAARIKKIMQADEDVGKIALAVPVLVS+ALELFLQDL DRTYEITLQ GAK
Sbjct: 1 MRKKLGTRFPAARIKKIMQADEDVGKIALAVPVLVSRALELFLQDLIDRTYEITLQSGAK 60
Query: 61 TMNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDS 120
T+NS HLK CV+ Y+ FDFL +VV+KVPD G ++ DDR +P+RRKA + +
Sbjct: 61 TLNSFHLKQCVRRYSSFDFLTEVVNKVPDL----GGADSCGDDRALPRRRKALPNGSDPE 116
Query: 121 DEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGRTAVRETPHQAVEPEPCASVQQG--IQ 178
+EE + KM S + RGRGRGRGRGRGR T +E + E E QG +
Sbjct: 117 NEESRSSKMAVRS-ANISPRGRGRGRGRGRGRPPTKRKEVGYVQFEDESSMFADQGEALP 175
Query: 179 HDTNTDMTIHDSSETKELPKENAAVPAESAEFHNLDLNANTNENEDKKASTTAKQEISEP 238
+ TIH + A P+ +AE + +N+D +
Sbjct: 176 GEETVPETIHGTESVPPSTHPPAEAPS-AAEIPAPNPKVEEAKNDDHQ------------ 222
Query: 239 PTESQHEEIPGWSLSD-VDKMAIDSVQLANLGTHIEEDEEDYDEE 282
P W + D + + + +L ++ED EDYD E
Sbjct: 223 ---------PDWPMPDAIGNIGVGPSGFGHLTVQVDED-EDYDNE 257
>UniRef100_Q8W0W6 Repressor protein [Zea mays]
Length = 254
Score = 196 bits (497), Expect = 7e-49
Identities = 125/285 (43%), Positives = 169/285 (58%), Gaps = 35/285 (12%)
Query: 1 MKKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK 60
M+KKL TRFPAARIKKIMQADEDVGKIALAVPVLVS++LELFLQDL DRTYEITLQ GAK
Sbjct: 1 MRKKLGTRFPAARIKKIMQADEDVGKIALAVPVLVSRSLELFLQDLIDRTYEITLQSGAK 60
Query: 61 TMNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDS 120
T+NS HLK CV+ Y+ FDFL +VVSKVPD G ++ D+R +P+RRK+ G D
Sbjct: 61 TLNSFHLKQCVKRYSSFDFLTEVVSKVPDL----GGADSCGDERGLPRRRKSNGSD--PE 114
Query: 121 DEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGRTAVRETPHQAVEPEPCASVQQGIQHD 180
++E + KM + + + RGRGRGRGRGRGR T +E + E E +QG
Sbjct: 115 NDESRSSKM-AIRNANISPRGRGRGRGRGRGRPPTKRKEVGYVQFEDESSMFAEQG--EP 171
Query: 181 TNTDMTIHDSSETKELPKENAAVPAESAEFHNLDLNANTNENEDKKASTTAKQEISEPPT 240
+ T+ + + + +P ++ P E+ +A+T++K E
Sbjct: 172 LPGEETVQEINGNETMP-QSTQPPVEAP------------PTALAQATTSSKAE------ 212
Query: 241 ESQHEEIPGWSLSDVDKMAIDSVQLANLG---THIEEDEEDYDEE 282
E+ + W + D AI S+ + G ++ ++EDYD E
Sbjct: 213 EANSDHQSDWPMPD----AIGSIGVVPSGFGHLTVQVEDEDYDNE 253
>UniRef100_Q9D6N5 Dr1-associated corepressor [Mus musculus]
Length = 204
Score = 119 bits (299), Expect = 7e-26
Identities = 76/204 (37%), Positives = 107/204 (52%), Gaps = 41/204 (20%)
Query: 2 KKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKT 61
KKK + RFP ARIKKIMQ DE++GK+A AVPV++S+ALELFL+ L + ++T R AKT
Sbjct: 4 KKKYNARFPPARIKKIMQTDEEIGKVAAAVPVIISRALELFLESLLKKACQVTQSRNAKT 63
Query: 62 MNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDSD 121
M + HLK C++ FDFL+D+V+ VPD G G D+ D D
Sbjct: 64 MTTSHLKQCIELEQQFDFLKDLVASVPD-MQGDGE------------------DNHVDGD 104
Query: 122 EEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGRTAVRETPHQAVEPEPCASVQQGIQHDT 181
+ +RG+ P GS+GR G +G+ + + S Q+ DT
Sbjct: 105 KGPRRGRKP-----GSSGRKNGGTGSKGKDKKLSGT-------------DSEQEDESEDT 146
Query: 182 NTDMTIHDSSETKELPKENAAVPA 205
+TD ET +LP + + PA
Sbjct: 147 DTD----GEEETPQLPPQASHPPA 166
>UniRef100_UPI000018197E UPI000018197E UniRef100 entry
Length = 205
Score = 116 bits (291), Expect = 6e-25
Identities = 74/204 (36%), Positives = 105/204 (51%), Gaps = 41/204 (20%)
Query: 2 KKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKT 61
KKK + RFP ARIKKIMQ DE++GK+A AVPV++S+ALELFL+ L + ++T R AKT
Sbjct: 5 KKKYNARFPPARIKKIMQTDEEIGKVAAAVPVIISRALELFLESLLKKACQVTQSRNAKT 64
Query: 62 MNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDSD 121
M + HLK C++ FDFL+D+V+ VPD G G D+ D D
Sbjct: 65 MTTSHLKQCIELEQQFDFLKDLVASVPD-MQGDGE------------------DNHTDGD 105
Query: 122 EEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGRTAVRETPHQAVEPEPCASVQQGIQHDT 181
+ +RG+ P GS+GR G + + + + S Q+ DT
Sbjct: 106 KGPRRGRKP-----GSSGRKNGGTGSKSKDKKLSGT-------------DSEQEDESEDT 147
Query: 182 NTDMTIHDSSETKELPKENAAVPA 205
+TD ET + P + + PA
Sbjct: 148 DTD----GEEETPQAPPQASHPPA 167
>UniRef100_Q14919-2 Splice isoform 2 of Q14919 [Homo sapiens]
Length = 210
Score = 115 bits (289), Expect = 1e-24
Identities = 61/140 (43%), Positives = 85/140 (60%), Gaps = 4/140 (2%)
Query: 2 KKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKT 61
KKK + RFP ARIKKIMQ DE++GK+A AVPV++S+ALELFL+ L + ++T R AKT
Sbjct: 4 KKKYNARFPPARIKKIMQTDEEIGKVAAAVPVIISRALELFLESLLKKACQVTQSRNAKT 63
Query: 62 MNSLHLKHCVQSYNVFDFLRDVVSKVPDY----SHGHGHSEPSADDRTIPKRRKAAGDDC 117
M + HLK C++ FDFL+D+V+ VPD H + A T+P +R+ G
Sbjct: 64 MTTSHLKQCIELEQQFDFLKDLVASVPDMQGDGEDNHMDGDKGARRWTVPSQRRKPGSGG 123
Query: 118 NDSDEEVKRGKMPELSHTGS 137
+ + K +LS T S
Sbjct: 124 RKNGGMGTKSKDKKLSGTDS 143
>UniRef100_Q7ZVI1 Wu:fb26e10 protein [Brachydanio rerio]
Length = 211
Score = 115 bits (288), Expect = 1e-24
Identities = 64/136 (47%), Positives = 86/136 (63%), Gaps = 3/136 (2%)
Query: 2 KKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKT 61
KKK + RFP ARIKKIMQ DE++GK+A AVPV++S+ALELFL+ L + ++T R AKT
Sbjct: 5 KKKYNARFPPARIKKIMQTDEEIGKVAAAVPVIISRALELFLESLLTKACDVTQSRNAKT 64
Query: 62 MNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDSD 121
M + HLK C++ FDFL+D+V+ VPD G G E + IP+R + G +
Sbjct: 65 MTTSHLKQCIELEQQFDFLKDLVATVPD-MQGDG-EENHTEGEKIPRRGRKPGSGRKNGG 122
Query: 122 EEVKRGKMPELSHTGS 137
K GK +LS T S
Sbjct: 123 AGAK-GKDKKLSGTES 137
>UniRef100_Q14919 Dr1-associated corepressor [Homo sapiens]
Length = 204
Score = 113 bits (282), Expect = 6e-24
Identities = 63/142 (44%), Positives = 84/142 (58%), Gaps = 24/142 (16%)
Query: 2 KKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKT 61
KKK + RFP ARIKKIMQ DE++GK+A AVPV++S+ALELFL+ L + ++T R AKT
Sbjct: 4 KKKYNARFPPARIKKIMQTDEEIGKVAAAVPVIISRALELFLESLLKKACQVTQSRNAKT 63
Query: 62 MNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDSD 121
M + HLK C++ FDFL+D+V+ VPD G G D+ D D
Sbjct: 64 MTTSHLKQCIELEQQFDFLKDLVASVPD-MQGDGE------------------DNHMDGD 104
Query: 122 EEVKRGKMPELSHTGSTGRGRG 143
+ +RG+ P GS GR G
Sbjct: 105 KGARRGRKP-----GSGGRKNG 121
>UniRef100_UPI00003628E8 UPI00003628E8 UniRef100 entry
Length = 215
Score = 112 bits (279), Expect = 1e-23
Identities = 60/136 (44%), Positives = 83/136 (60%), Gaps = 1/136 (0%)
Query: 2 KKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKT 61
KKK + RFP ARIKKIMQ DE++GK+A AVPV++S+ALELFL+ L + ++T R AKT
Sbjct: 5 KKKYNARFPPARIKKIMQTDEEIGKVAAAVPVIISRALELFLESLLTKACQVTHSRNAKT 64
Query: 62 MNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDSD 121
M + HLK C++ FDFL+D+V+ VPD + A + +R + G +
Sbjct: 65 MTTSHLKQCIELEQQFDFLKDLVAAVPDMQGEGEENHNEAVGEKVSRRGRKPGPGRKNGG 124
Query: 122 EEVKRGKMPELSHTGS 137
K GK +LS T S
Sbjct: 125 TGAK-GKDKKLSGTDS 139
>UniRef100_Q9W2E3 CG10318-PA, isoform A [Drosophila melanogaster]
Length = 341
Score = 105 bits (262), Expect = 1e-21
Identities = 62/171 (36%), Positives = 94/171 (54%), Gaps = 12/171 (7%)
Query: 2 KKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKT 61
KKK + RFPA RIKKIMQ+DE++GK+A AVPV++S+ LELF++ L +T IT R AKT
Sbjct: 5 KKKYNARFPAGRIKKIMQSDEEIGKVAQAVPVIISRTLELFVESLLTKTLRITNARNAKT 64
Query: 62 MNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADD--RTIPKRRKAAGDDCND 119
++ H++ C+ S FDFL+++V +PD S + + DD R+ P+ + D D
Sbjct: 65 LSPSHMRQCIVSEKRFDFLKELVRNIPDISVAE-EAAYNEDDVLRSSPEEQYPDSDTPYD 123
Query: 120 ---------SDEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGRTAVRETP 161
S M +S G G G + + + + + +ETP
Sbjct: 124 LSLPSTSMRSQANGTAAYMRSMSLNNGAGSGGGAAATKRQFQSQHSTQETP 174
>UniRef100_Q9GSP1 NC2alpha [Drosophila melanogaster]
Length = 341
Score = 105 bits (262), Expect = 1e-21
Identities = 62/171 (36%), Positives = 94/171 (54%), Gaps = 12/171 (7%)
Query: 2 KKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKT 61
KKK + RFPA RIKKIMQ+DE++GK+A AVPV++S+ LELF++ L +T IT R AKT
Sbjct: 5 KKKYNARFPAGRIKKIMQSDEEIGKVAQAVPVIISRTLELFVESLLTKTLRITNARNAKT 64
Query: 62 MNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADD--RTIPKRRKAAGDDCND 119
++ H++ C+ S FDFL+++V +PD S + + DD R+ P+ + D D
Sbjct: 65 LSPSHMRQCIVSEKRFDFLKELVRNIPDISVAE-EAAYNEDDVLRSSPEEQYPDSDTPYD 123
Query: 120 ---------SDEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGRTAVRETP 161
S M +S G G G + + + + + +ETP
Sbjct: 124 LSLPSTSMRSQANGTAAYMRSMSLNNGAGSGGGAAATKRQFQSQHSTQETP 174
>UniRef100_Q5TUK8 ENSANGP00000026107 [Anopheles gambiae str. PEST]
Length = 93
Score = 105 bits (262), Expect = 1e-21
Identities = 50/88 (56%), Positives = 66/88 (74%)
Query: 2 KKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKT 61
KKK + RFPA RIKKIMQ DE+VGK+A AVPV+V + LELF++ L +T +IT R AKT
Sbjct: 5 KKKYNARFPAGRIKKIMQTDEEVGKVAQAVPVIVPRTLELFVESLLTKTLKITNARNAKT 64
Query: 62 MNSLHLKHCVQSYNVFDFLRDVVSKVPD 89
++ H+K C+ S + FDFLRD+V +PD
Sbjct: 65 LSPSHMKQCIISESRFDFLRDLVKNIPD 92
>UniRef100_UPI000023655D UPI000023655D UniRef100 entry
Length = 1251
Score = 102 bits (254), Expect = 1e-20
Identities = 53/126 (42%), Positives = 74/126 (58%), Gaps = 3/126 (2%)
Query: 4 KLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMN 63
++ T+FP ARIK+IMQADEDVGK+A P+ VSKALELF+ L + + R +K +
Sbjct: 185 EVKTKFPVARIKRIMQADEDVGKVAQVTPIAVSKALELFMISLVTKAAKEAKDRNSKRVT 244
Query: 64 SLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAA---GDDCNDS 120
+ HLK V V DFL D+++KVPD G H + +D PKR++ DD D
Sbjct: 245 ASHLKQAVAKDEVLDFLADIIAKVPDQPTGRKHEDDGSDQNEQPKRKRGGRRPKDDNRDG 304
Query: 121 DEEVKR 126
++R
Sbjct: 305 AHILRR 310
>UniRef100_UPI000036EEB8 UPI000036EEB8 UniRef100 entry
Length = 191
Score = 100 bits (250), Expect = 3e-20
Identities = 57/132 (43%), Positives = 77/132 (58%), Gaps = 24/132 (18%)
Query: 12 ARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHLKHCV 71
ARIKKIMQ DE++GK+A AVPV++S+ALELFL+ L + ++T R AKTM + HLK C+
Sbjct: 1 ARIKKIMQTDEEIGKVAAAVPVIISRALELFLESLLKKACQVTQSRNAKTMTTSHLKQCI 60
Query: 72 QSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDSDEEVKRGKMPE 131
+ FDFL+D+V+ VPD G G D+ D D+ +RG+ P
Sbjct: 61 ELEQQFDFLKDLVASVPD-MQGDGE------------------DNHMDGDKGARRGRKP- 100
Query: 132 LSHTGSTGRGRG 143
GS GR G
Sbjct: 101 ----GSGGRKNG 108
>UniRef100_UPI00003C1340 UPI00003C1340 UniRef100 entry
Length = 133
Score = 100 bits (249), Expect = 4e-20
Identities = 60/133 (45%), Positives = 79/133 (59%), Gaps = 13/133 (9%)
Query: 1 MKKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVS--------KALELFLQDLCDRTYE 52
+K+ +FP ARIKKIMQADEDVGK+A A PVL+S KALELF+ + + T +
Sbjct: 2 VKRSAQCKFPVARIKKIMQADEDVGKVAQATPVLISIWNLGLTAKALELFMASIVEETVK 61
Query: 53 ITLQRGAKTMNSLHLKHCVQSYNVFDFLRDVVSKVPD----YSHGHGHSEPSADDRTIPK 108
T RGAK M H+K V + FDFL+D+V+KVPD S + +A D K
Sbjct: 62 ETRSRGAKKMTPYHVKRTVHTNETFDFLKDIVAKVPDPVETDSSAAATATAAAGDNRGGK 121
Query: 109 RRKAAGDDCNDSD 121
RRK A + ++SD
Sbjct: 122 RRKTA-KNADESD 133
>UniRef100_Q9US69 Hypothetical nuclear protein [Schizosaccharomyces pombe]
Length = 172
Score = 99.4 bits (246), Expect = 1e-19
Identities = 56/124 (45%), Positives = 76/124 (61%), Gaps = 9/124 (7%)
Query: 7 TRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLH 66
+RFP ARIKKIMQAD+DVGK+A PV++SKALELF+Q + + + T AK + H
Sbjct: 22 SRFPVARIKKIMQADQDVGKVAQVTPVIMSKALELFMQSIIQESCKQTRLHQAKRVTVSH 81
Query: 67 LKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTI--PKRRKAAGDDCNDSDEEV 124
LKH VQS FDFL+D+V KVPD + P +R P+ R+AA + ++
Sbjct: 82 LKHAVQSVEQFDFLQDIVEKVPD-------APPIKAERKTKRPRARRAANEGEHNESVPA 134
Query: 125 KRGK 128
K+ K
Sbjct: 135 KKVK 138
>UniRef100_Q10315 Hypothetical protein C17G8.03c in chromosome I [Schizosaccharomyces
pombe]
Length = 199
Score = 99.4 bits (246), Expect = 1e-19
Identities = 56/124 (45%), Positives = 76/124 (61%), Gaps = 9/124 (7%)
Query: 7 TRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLH 66
+RFP ARIKKIMQAD+DVGK+A PV++SKALELF+Q + + + T AK + H
Sbjct: 22 SRFPVARIKKIMQADQDVGKVAQVTPVIMSKALELFMQSIIQESCKQTRLHQAKRVTVSH 81
Query: 67 LKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTI--PKRRKAAGDDCNDSDEEV 124
LKH VQS FDFL+D+V KVPD + P +R P+ R+AA + ++
Sbjct: 82 LKHAVQSVEQFDFLQDIVEKVPD-------APPIKAERKTKRPRARRAANEGEHNESVPA 134
Query: 125 KRGK 128
K+ K
Sbjct: 135 KKVK 138
>UniRef100_Q7RW27 Hypothetical protein [Neurospora crassa]
Length = 332
Score = 97.1 bits (240), Expect = 5e-19
Identities = 55/140 (39%), Positives = 81/140 (57%), Gaps = 13/140 (9%)
Query: 7 TRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLH 66
T+FP ARIK+IMQADE+VGK+A P+ V KALELF+ L ++ +I +R +K +++
Sbjct: 201 TKFPTARIKRIMQADEEVGKVAQQTPIAVGKALELFMVQLVTKSADIARERNSKRVSAQM 260
Query: 67 LKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDSDEEVKR 126
LK V+S + +DFLR++ SK+ + DD+ P A+G ++ E +
Sbjct: 261 LKQVVESDDQWDFLREITSKIEN------------DDKEKP-AASASGRGRTKAESESEE 307
Query: 127 GKMPELSHTGSTGRGRGRGR 146
PE G GRG GRGR
Sbjct: 308 EAQPEKKGRGGAGRGGGRGR 327
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.310 0.129 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 487,064,217
Number of Sequences: 2790947
Number of extensions: 21808749
Number of successful extensions: 103555
Number of sequences better than 10.0: 936
Number of HSP's better than 10.0 without gapping: 419
Number of HSP's successfully gapped in prelim test: 548
Number of HSP's that attempted gapping in prelim test: 98699
Number of HSP's gapped (non-prelim): 2449
length of query: 283
length of database: 848,049,833
effective HSP length: 126
effective length of query: 157
effective length of database: 496,390,511
effective search space: 77933310227
effective search space used: 77933310227
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0005.4


