

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0005.13
(396 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
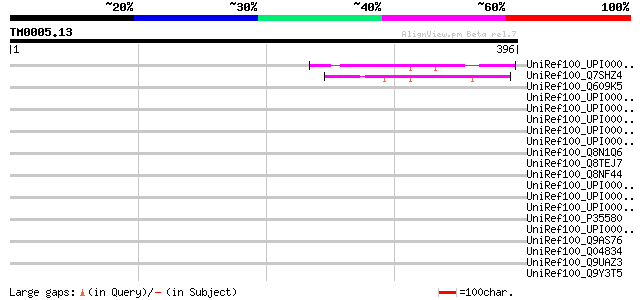
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_UPI00003380AF UPI00003380AF UniRef100 entry 49 2e-04
UniRef100_Q7SHZ4 Hypothetical protein [Neurospora crassa] 49 3e-04
UniRef100_Q609K5 TolA protein, putative [Methylococcus capsulatus] 46 0.002
UniRef100_UPI00002497D4 UPI00002497D4 UniRef100 entry 45 0.003
UniRef100_UPI0000363F8C UPI0000363F8C UniRef100 entry 45 0.003
UniRef100_UPI00003AC153 UPI00003AC153 UniRef100 entry 45 0.003
UniRef100_UPI00002CF509 UPI00002CF509 UniRef100 entry 45 0.003
UniRef100_UPI000029BB48 UPI000029BB48 UniRef100 entry 45 0.003
UniRef100_Q8N1Q6 Hypothetical protein FLJ37970 [Homo sapiens] 45 0.004
UniRef100_Q8TEJ7 FLJ00198 protein [Homo sapiens] 45 0.004
UniRef100_Q8NF44 FLJ00354 protein [Homo sapiens] 45 0.004
UniRef100_UPI000036AB03 UPI000036AB03 UniRef100 entry 45 0.005
UniRef100_UPI000023D00A UPI000023D00A UniRef100 entry 45 0.005
UniRef100_UPI0000235C75 UPI0000235C75 UniRef100 entry 45 0.005
UniRef100_P35580 Myosin heavy chain, nonmuscle type B [Homo sapi... 45 0.005
UniRef100_UPI000042F484 UPI000042F484 UniRef100 entry 44 0.006
UniRef100_Q9AS76 P0028E10.16 protein [Oryza sativa] 44 0.006
UniRef100_Q04834 Nonmuscle myosin heavy chain b [Xenopus laevis] 44 0.008
UniRef100_Q9UAZ3 Hypothetical protein Y38C9A.1 [Caenorhabditis e... 44 0.008
UniRef100_Q9Y3T5 Hypothetical protein DKFZp564G013 [Homo sapiens] 44 0.008
>UniRef100_UPI00003380AF UPI00003380AF UniRef100 entry
Length = 1962
Score = 48.9 bits (115), Expect = 2e-04
Identities = 42/169 (24%), Positives = 78/169 (45%), Gaps = 24/169 (14%)
Query: 235 LSPTPAQVARMDEVVKQQGLDRVSEGAFATAFHAFHHYKRSAQDAEVLRSEVA-RLRALN 293
L + A+++++ E ++Q E + H + K Q+ + L E+ +L
Sbjct: 759 LKDSNAELSKISEKLEQ------CEKDYTDLEHQLNAAKNGCQEKDKLLEELQNQLHQNR 812
Query: 294 AGFVEERKAFLAQLKAKE---TSFAKELAEQSSRAEVSLEF----RDRKISDLEKALGKL 346
+E+ K+F AQL KE TS K+L E+ + E L+ + K+ LE L K
Sbjct: 813 TELLEQEKSFTAQLNTKEEEKTSLKKQLEEEKAAHEKKLQSTVSGMEAKVKALETKLDKF 872
Query: 347 RQEAQEESEAKVDLVASKDQTLSSMETKLEHLRAELAKKDEALSMMRTQ 395
+Q+A++ E+ ++T+ E ++ EL KKD+ + + Q
Sbjct: 873 KQKAKDMHES----------AKKKLQTQEETMKMELEKKDKEIHLKEQQ 911
Score = 34.7 bits (78), Expect = 4.8
Identities = 27/95 (28%), Positives = 49/95 (51%), Gaps = 8/95 (8%)
Query: 298 EERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRDRKISDLEKALGKLRQEAQEESEAK 357
EE+ L +++KE+ K +E + + L ++ I LE+ LRQ+A+E
Sbjct: 1092 EEQIKLLQGVRSKESKDLKTKSESVVQLQAVLNSKEELICTLEE---NLRQQAEENK--- 1145
Query: 358 VDLVASKDQTLSSMETKLEHLRA-ELAKKDEALSM 391
+L S DQ + + ++EH+ A K++ ALS+
Sbjct: 1146 -NLCISLDQLTAQVNAQMEHVTALTQEKENHALSL 1179
>UniRef100_Q7SHZ4 Hypothetical protein [Neurospora crassa]
Length = 4007
Score = 48.5 bits (114), Expect = 3e-04
Identities = 44/155 (28%), Positives = 77/155 (49%), Gaps = 13/155 (8%)
Query: 247 EVVKQQGLDRVSEGAFATAFHAFHHYKRSAQDAEVLRSEVARLRA----LNAGFVEERKA 302
++VK + + +FA A H K D L SEVA+L+ +A + + K
Sbjct: 2694 DIVKLREDVAFKDKSFAKKAEAVDHLKA---DITELNSEVAKLKKEGTNKDAAILGKEKE 2750
Query: 303 FLAQLKAKE--TSFAKELAEQSSRAEVSLEFRDRKISDLEKALGKLRQEAQEESEAKVDL 360
++ KA T+ AK+ A+ S ++ L RD + + EK + +L+QE Q+ + +L
Sbjct: 2751 LVSLRKAVRDLTNQAKQSAQDSKKSAEDLANRDALLKEKEKKIFELQQEIQKVKDTAEEL 2810
Query: 361 ---VASKDQTLSSMETKLEHLRAELAK-KDEALSM 391
++D TLS +L LR ++ + +DEA S+
Sbjct: 2811 NQTTKTRDSTLSQKNEELRKLREQIKQLEDEANSL 2845
Score = 39.3 bits (90), Expect = 0.20
Identities = 38/129 (29%), Positives = 62/129 (47%), Gaps = 13/129 (10%)
Query: 276 AQDAEV--LRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAE-------QSSRAE 326
A+DAE+ L++E+A A A EE KAF ++ T AK L + Q ++
Sbjct: 2445 ARDAELAKLKAEIASKNAALAKKTEEAKAFEKNVQTL-TDQAKGLNQDVATKTTQLAQDR 2503
Query: 327 VSLEFRDRKISDLEKALGKLRQEAQEESE---AKVDLVASKDQTLSSMETKLEHLRAELA 383
++ ++ I DL+ + KL+QE + K + S+D L+ + +L A LA
Sbjct: 2504 ATISKLNKDIFDLKTDVTKLKQELSTKDANLTQKAGEIGSRDAGLAKLREELRAKEAALA 2563
Query: 384 KKDEALSMM 392
KK E S +
Sbjct: 2564 KKTEEASSL 2572
Score = 36.6 bits (83), Expect = 1.3
Identities = 29/135 (21%), Positives = 62/135 (45%), Gaps = 5/135 (3%)
Query: 250 KQQGLDRVSEGAFATAFHAFHHYKRSAQDAEVLRSEVARLRALNAGFVEERKAFLAQLKA 309
K L +++ G T +++ +D + L+ V +L A EE+ + A
Sbjct: 2925 KAAELSKLNAGQDQTIGEKDASLQKANEDIDNLKGSVQKLENKAATLAEEKAQMGQTIGA 2984
Query: 310 KETSFAKELAEQSSRAEVSLEFRDRKISDLEKALGKLRQEAQEESEAKVDLVASKDQTLS 369
ETS K+ E + +++ + +DL+K + L + ++ A +A K++ +
Sbjct: 2985 HETSLLKK-DEDIKKLTANIQRLTAEANDLKKGIENLTGDIAIQNRA----LAQKEKDIQ 3039
Query: 370 SMETKLEHLRAELAK 384
+ME ++ L E+A+
Sbjct: 3040 NMEKTIQDLNTEVAR 3054
Score = 34.7 bits (78), Expect = 4.8
Identities = 27/132 (20%), Positives = 60/132 (45%), Gaps = 18/132 (13%)
Query: 274 RSAQDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRD 333
+ DA L+++V L + E++ + +L+ K ELAE+ +R + RD
Sbjct: 1790 QKGSDAAKLQADVDSLNKK----ISEKRQKVTELEGKVNKLDSELAEEKAR----VSRRD 1841
Query: 334 RKISDLEKALGKLRQEAQEESEAKVDL----------VASKDQTLSSMETKLEHLRAELA 383
R+I+DL+K + + + DL V+ +D+ ++ ++ + +A
Sbjct: 1842 REITDLKKDVSDEKARTTKRDREITDLKKDVSDEKARVSRRDREVTDLKKDVSDEKARTT 1901
Query: 384 KKDEALSMMRTQ 395
K D + ++++
Sbjct: 1902 KHDNEIGGLQSK 1913
Score = 33.9 bits (76), Expect = 8.2
Identities = 27/99 (27%), Positives = 49/99 (49%), Gaps = 9/99 (9%)
Query: 298 EERKAFLAQLK-----AKETSFAKE-LAEQSSRAEVSLEFRDRKISDLEKALGKLRQEAQ 351
EE + Q+K A KE L + + SLE ++++IS LEK + +L ++A
Sbjct: 2826 EELRKLREQIKQLEDEANSLKMDKETLGRTINTRDSSLEQKEQEISGLEKEIKRLSEQAA 2885
Query: 352 EESEAKVDL---VASKDQTLSSMETKLEHLRAELAKKDE 387
++ KVDL V ++D +L ++ L+ + +E
Sbjct: 2886 NLTQEKVDLGQIVGARDASLLQANKDIDGLKGSIKILEE 2924
>UniRef100_Q609K5 TolA protein, putative [Methylococcus capsulatus]
Length = 467
Score = 46.2 bits (108), Expect = 0.002
Identities = 65/270 (24%), Positives = 105/270 (38%), Gaps = 25/270 (9%)
Query: 135 VLDCARVIEKIPQVSEGLLTVFEKKGNVSPHPVAATSSSDGDRAPVISAAFCGPEETPKP 194
VLD +R+ + +E L T P PV+ + + RA I A EE+ +
Sbjct: 50 VLDDSRIAAE----AERLKTEAAPPAETGPEPVSTPNPEESRRAQQIEAEKAA-EESRRQ 104
Query: 195 NQPGEPSLATPVSSPRAQDDSVDKPPLSAKGFANREPPYSLSPTPAQVARMDEVVKQQGL 254
+ + + R + ++ +K +A+ A ++ A E + +
Sbjct: 105 MEAKKLADQARKDEQRRKAEAEEKARAAAEAAARKKAE----------AEAKEKAEAEAR 154
Query: 255 DRVSEGAFATAFHAFHHYK-----RSAQDAEVLRSEVARLRALNAGFVEERKAFLAQLKA 309
R +E A A A A K R +AE A R A + A A+ K
Sbjct: 155 RRAAEEARAKAAEAEAKRKAAEAARKKAEAEAKEKAEAEARRRAAEEARAKAAAEAEAKR 214
Query: 310 KETSFAKELAEQSSRAEVSLEFRDRKISDLE---KALGKLRQEAQEESEAKVDLVASKDQ 366
K A+E AE +R + + E RK ++ E KA + R+ A EE+ AK A +
Sbjct: 215 KAAEAAREKAEAEAREKAAAEAAARKKAEAEAKEKAEAEARRRAAEEARAKA--AAEAEA 272
Query: 367 TLSSMETKLEHLRAELAKKDEALSMMRTQA 396
+ E E AE +K A + R +A
Sbjct: 273 KRRAAEAAREKAEAEAREKAAAEAAARKKA 302
Score = 36.2 bits (82), Expect = 1.7
Identities = 32/118 (27%), Positives = 53/118 (44%), Gaps = 12/118 (10%)
Query: 274 RSAQDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRD 333
++A +AE R R E+ A A + K + AKE AE +R + E R
Sbjct: 265 KAAAEAEAKRRAAEAAREKAEAEAREKAAAEAAARKKAEAEAKEKAEAEARRRAAEEARA 324
Query: 334 RKISDLEKALGKLRQEAQEESEAKVDLVASKD-----QTLSSMETKLEHLRAELAKKD 386
R A+ + +E +EE +AK A K + + +E +L + R ELA+++
Sbjct: 325 R-------AMAEATREMEEEVKAKAAAEARKKAVEDARRKAELEEQLNNERRELAERE 375
Score = 35.8 bits (81), Expect = 2.2
Identities = 42/186 (22%), Positives = 75/186 (39%), Gaps = 14/186 (7%)
Query: 203 ATPVSSPRAQDDSVDKPPLSAKGFANREPPYSLSPTPAQVARMDEVVKQQGLDRVSEGAF 262
A + +A+ ++ +K A+ A E + + E +++ E A
Sbjct: 174 AAEAARKKAEAEAKEKAEAEARRRAAEEARAKAAAEAEAKRKAAEAAREKAEAEAREKAA 233
Query: 263 ATAFHAFHHYKRSAQDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQS 322
A A R +AE A R A + A A+ K + A+E AE
Sbjct: 234 AEAA------ARKKAEAEAKEKAEAEARRRAAEEARAKAAAEAEAKRRAAEAAREKAEAE 287
Query: 323 SRAEVSLEFRDRKISDL---EKALGKLRQEAQEESEAKVDLVASKDQTLSSMETKLEHLR 379
+R + + E RK ++ EKA + R+ A EE+ A+ A+++ ME +++
Sbjct: 288 AREKAAAEAAARKKAEAEAKEKAEAEARRRAAEEARARAMAEATRE-----MEEEVKAKA 342
Query: 380 AELAKK 385
A A+K
Sbjct: 343 AAEARK 348
>UniRef100_UPI00002497D4 UPI00002497D4 UniRef100 entry
Length = 877
Score = 45.4 bits (106), Expect = 0.003
Identities = 26/122 (21%), Positives = 63/122 (51%), Gaps = 3/122 (2%)
Query: 278 DAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRD---R 334
D E L+ ++++++ +N E LA++ + +E AE S + ++ D +
Sbjct: 651 DTEGLQKDLSQVQTMNEKLSAENLRLLAEVSELSSQLEEERAEVSDEPDSTVPQADELQQ 710
Query: 335 KISDLEKALGKLRQEAQEESEAKVDLVASKDQTLSSMETKLEHLRAELAKKDEALSMMRT 394
++D + + +LR+E +++ V+ + + L+++ +LE L+ E KDE ++ ++
Sbjct: 711 ALADRDTLISQLREEVGHLQQSQESPVSDQSERLTALSEELEKLKKESKDKDERMNKIKA 770
Query: 395 QA 396
A
Sbjct: 771 VA 772
>UniRef100_UPI0000363F8C UPI0000363F8C UniRef100 entry
Length = 1912
Score = 45.4 bits (106), Expect = 0.003
Identities = 44/178 (24%), Positives = 77/178 (42%), Gaps = 24/178 (13%)
Query: 226 FANREPPYSLSPTPAQVARMDEVVKQQGLDRVSEGAFATAFHAFHHYKRSAQDAEVLRSE 285
F +++ L + A++ ++ E + Q D A H K Q + L E
Sbjct: 733 FQSQQENERLKESNAELRKISENLDQCKKDH------ADLEHQLDASKNDCQQKDALLEE 786
Query: 286 VARLRALNAGFVEER-KAFLAQLKAKE---TSFAKELAEQSSRAEVSLE----FRDRKIS 337
+ N + E+ K+F AQL AKE T K+L E+ + E ++ + K+
Sbjct: 787 LQNQLHQNRNELSEKEKSFTAQLNAKEEEQTCLRKQLEEEKAAHEEKMQNTVSDMEAKVK 846
Query: 338 DLEKALGKLRQEAQEESEAKVDLVASKDQTLSSMETKLEHLRAELAKKDEALSMMRTQ 395
LE L K +Q+A++ E+ + +D+T ++ EL KKD+ L + Q
Sbjct: 847 ALETKLDKFKQKAKDMHESAKKKLQKQDET----------MKVELEKKDKELQLKEQQ 894
Score = 35.0 bits (79), Expect = 3.7
Identities = 37/168 (22%), Positives = 71/168 (42%), Gaps = 20/168 (11%)
Query: 240 AQVARMDEVVKQQGLDRVSEGAFATAFHAFHHYK-------------RSAQDAEVLRSEV 286
AQ++ +E +++Q +R+ E H K + QD ++ +E
Sbjct: 261 AQLSTENETLQEQLQERLQELEKIKELHTTEKTKLITQLGEAKNLIEQLEQDKGMVIAET 320
Query: 287 ARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRDRKISDLEKALGKL 346
R E+ A L + T+ +EL EQ +AE S +LE+ALG
Sbjct: 321 KRQMHETLEMKEDEVAQLRSRLQQVTAHKEELQEQKEKAEKSA------FEELERALGVA 374
Query: 347 RQEAQEESEAKVDLVASKDQTLSSMETKLEHLRAELAK-KDEALSMMR 393
++ + + +V L + + E + + L+ EL + K E +++M+
Sbjct: 375 QRAEEARKQLQVQLEEQVKEVERASEEERKSLQQELTRVKQEVVTIMK 422
>UniRef100_UPI00003AC153 UPI00003AC153 UniRef100 entry
Length = 2363
Score = 45.4 bits (106), Expect = 0.003
Identities = 29/121 (23%), Positives = 61/121 (49%)
Query: 273 KRSAQDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFR 332
K + EVL V++L+ N + + A + +L+ E+S A E +S E E +
Sbjct: 1897 KERERKIEVLEKAVSKLQQQNDDTMMQTTAIVQKLQCAESSLAARDQEIASLKEHVQELQ 1956
Query: 333 DRKISDLEKALGKLRQEAQEESEAKVDLVASKDQTLSSMETKLEHLRAELAKKDEALSMM 392
+K S+ ++ + ++ E++A+ + S+D+ + + L++++ L KKDE +
Sbjct: 1957 KQKESEAKQVKKEALHGSEWETKAQGEQKKSEDEEIRGLHEDLQYVQQTLTKKDEEIKYQ 2016
Query: 393 R 393
R
Sbjct: 2017 R 2017
Score = 35.0 bits (79), Expect = 3.7
Identities = 31/127 (24%), Positives = 63/127 (49%), Gaps = 19/127 (14%)
Query: 277 QDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRA-----EVSLEF 331
+D + + E+ L+ L E+ + L+ K E KE ++ + +++L+
Sbjct: 1841 EDDSMQQKEI--LQHLKGALKEQEQETLSLRKQCEAFKEKEEKHKTDQVILQETKITLKE 1898
Query: 332 RDRKISDLEKALGKLRQEAQEESEAKVDLV----------ASKDQTLSSMETKLEHLRAE 381
R+RKI LEKA+ KL+Q+ + +V A++DQ ++S++ ++ L+ +
Sbjct: 1899 RERKIEVLEKAVSKLQQQNDDTMMQTTAIVQKLQCAESSLAARDQEIASLKEHVQELQKQ 1958
Query: 382 LAKKDEA 388
K+ EA
Sbjct: 1959 --KESEA 1963
>UniRef100_UPI00002CF509 UPI00002CF509 UniRef100 entry
Length = 263
Score = 45.4 bits (106), Expect = 0.003
Identities = 42/154 (27%), Positives = 73/154 (47%), Gaps = 21/154 (13%)
Query: 250 KQQGLDRVSEGAFATAFHAFHHYKRSAQDAEVLRSEVARL------RALNAGFVEER-KA 302
++ L+RV A A+ H +R A + E LR RA+N EE K
Sbjct: 1 EESQLERVRAAESALERAAYDHRQRLAAEHEKLREAKEEAHREETERAMNRAREEEDLKR 60
Query: 303 FLAQLKAKETSFAKELAEQSSRAEVSLEFRDRKISDLEKALGKLRQEAQEESEAKVDLVA 362
+++K+KE+S++ AE S+ AE E+AL + R+E + E+E +++++
Sbjct: 61 RESEIKSKESSWSMRYAELSAEAE-------------ERALAR-RRETETEAERRLEMIK 106
Query: 363 SKDQTLSSMETKLEHLRAELAKKDEALSMMRTQA 396
++ L KLE RA+ A + ++ R A
Sbjct: 107 TEMMALEDERAKLELDRAQHAVEIATIAQARRDA 140
>UniRef100_UPI000029BB48 UPI000029BB48 UniRef100 entry
Length = 2252
Score = 45.4 bits (106), Expect = 0.003
Identities = 52/234 (22%), Positives = 90/234 (38%), Gaps = 31/234 (13%)
Query: 189 EETPKPNQPGEPSLATPVSSPRAQDDSVDKPPLSAKGFANREPPYSLSPTPAQVARMDEV 248
E P P Q PS A P P + D +V + A+ F E S +V R+ E
Sbjct: 156 EVEPSPAQAESPSAAIPQPDPSSSDRNVSSGQVFAQAFVE-ELQRKCSDLLLEVERLREA 214
Query: 249 VKQQG----LDRVSEGAFATAFHAFHHYKRS-AQDAEVLRSEVARLRALNAGF------- 296
+ L + A A A A S A++ ++ R ++ +L N
Sbjct: 215 AAESAGKMKLLQEDVEALAAAKEASEVQASSVAEELQLAREQLEKLSRENMSSAEKHGVE 274
Query: 297 --------------VEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRDRKISDLEKA 342
VE ++ + L+ + + K L+ Q E+ +RK+ D+E +
Sbjct: 275 MQLLEEQLDILNATVEAKEEKIQNLQTEREAQVKMLSAQLEDRELVSSQLERKVQDMENS 334
Query: 343 LGKLRQEAQEESEAKVDLVASKDQTLSSMETKLEHLRAELAKKDEALSMMRTQA 396
+ E + SE D ++ KD +S ++ L E++ E++S QA
Sbjct: 335 M----SEYSQTSELNSDALSKKDSEISELQLLLSQKEEEVSTLGESMSAKLLQA 384
>UniRef100_Q8N1Q6 Hypothetical protein FLJ37970 [Homo sapiens]
Length = 621
Score = 45.1 bits (105), Expect = 0.004
Identities = 55/221 (24%), Positives = 88/221 (38%), Gaps = 25/221 (11%)
Query: 187 GPEETPKPNQPGEPSLATPVSSPRAQDDSVDKPPLSAKGFANREPPYSLSPTPAQVARMD 246
GPE P P++P L + D L+ RE L T A
Sbjct: 294 GPEHKPGPSEPSSVQLEEQEGPNQGLD-------LATGQAEAREHDQRLEGTVRDPAWQK 346
Query: 247 EVVKQQGLDRVS--EGAFATAFHAFHHYKRSAQDAEVLRSEVARLRALNAGFVEERKAFL 304
K +G V EG A + E LR EVA+LR +E +A
Sbjct: 347 PQQKSEGALEVQVWEGPIPGESLA-----SGVAEQEALREEVAQLRRKAEALGDELEAQA 401
Query: 305 AQLKAKETSFA---KELAEQSSRAEVSLE-------FRDRKISDLEKALGKLRQEAQEES 354
+L+A+ T A KELA Q+ RAE + ++ + +A G+ + A +E
Sbjct: 402 RKLEAQNTEAARLSKELA-QARRAEAEAHREAEAQAWEQARLREAVEAAGQELESASQER 460
Query: 355 EAKVDLVASKDQTLSSMETKLEHLRAELAKKDEALSMMRTQ 395
EA V+ +A+ + E + LRA+ +E + ++ ++
Sbjct: 461 EALVEALAAAGRERRQWEREGSRLRAQSEAAEERMQVLESE 501
>UniRef100_Q8TEJ7 FLJ00198 protein [Homo sapiens]
Length = 677
Score = 45.1 bits (105), Expect = 0.004
Identities = 55/221 (24%), Positives = 88/221 (38%), Gaps = 25/221 (11%)
Query: 187 GPEETPKPNQPGEPSLATPVSSPRAQDDSVDKPPLSAKGFANREPPYSLSPTPAQVARMD 246
GPE P P++P L + D L+ RE L T A
Sbjct: 108 GPEHKPGPSEPSSVQLEEQEGPNQGLD-------LATGQAEAREHDQRLEGTVRDPAWQK 160
Query: 247 EVVKQQGLDRVS--EGAFATAFHAFHHYKRSAQDAEVLRSEVARLRALNAGFVEERKAFL 304
K +G V EG A + E LR EVA+LR +E +A
Sbjct: 161 PQQKSEGALEVQVWEGPIPGESLA-----SGVAEQEALREEVAQLRRKAEALGDELEAQA 215
Query: 305 AQLKAKETSFA---KELAEQSSRAEVSLE-------FRDRKISDLEKALGKLRQEAQEES 354
+L+A+ T A KELA Q+ RAE + ++ + +A G+ + A +E
Sbjct: 216 RKLEAQNTEAARLSKELA-QARRAEAEAHREAEAQAWEQARLREAVEAAGQELESASQER 274
Query: 355 EAKVDLVASKDQTLSSMETKLEHLRAELAKKDEALSMMRTQ 395
EA V+ +A+ + E + LRA+ +E + ++ ++
Sbjct: 275 EALVEALAAAGRERRQWEREGSRLRAQSEAAEERMQVLESE 315
>UniRef100_Q8NF44 FLJ00354 protein [Homo sapiens]
Length = 1041
Score = 45.1 bits (105), Expect = 0.004
Identities = 55/221 (24%), Positives = 88/221 (38%), Gaps = 25/221 (11%)
Query: 187 GPEETPKPNQPGEPSLATPVSSPRAQDDSVDKPPLSAKGFANREPPYSLSPTPAQVARMD 246
GPE P P++P L + D L+ RE L T A
Sbjct: 652 GPEHKPGPSEPSSVQLEEQEGPNQGLD-------LATGQAEAREHDQRLEGTVRDPAWQK 704
Query: 247 EVVKQQGLDRVS--EGAFATAFHAFHHYKRSAQDAEVLRSEVARLRALNAGFVEERKAFL 304
K +G V EG A + E LR EVA+LR +E +A
Sbjct: 705 PQQKSEGALEVQVWEGPIPGESLA-----SGVAEQEALREEVAQLRRKAEALGDELEAQA 759
Query: 305 AQLKAKETSFA---KELAEQSSRAEVSLE-------FRDRKISDLEKALGKLRQEAQEES 354
+L+A+ T A KELA Q+ RAE + ++ + +A G+ + A +E
Sbjct: 760 RKLEAQNTEAARLSKELA-QARRAEAEAHREAEAQAWEQARLREAVEAAGQELESASQER 818
Query: 355 EAKVDLVASKDQTLSSMETKLEHLRAELAKKDEALSMMRTQ 395
EA V+ +A+ + E + LRA+ +E + ++ ++
Sbjct: 819 EALVEALAAAGRERRQWEREGSRLRAQSEAAEERMQVLESE 859
>UniRef100_UPI000036AB03 UPI000036AB03 UniRef100 entry
Length = 294
Score = 44.7 bits (104), Expect = 0.005
Identities = 30/114 (26%), Positives = 61/114 (53%), Gaps = 10/114 (8%)
Query: 282 LRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRDRK---ISD 338
+ E+ L N+ F++E+K ++ + + +LAE+ +A+ + R+++ ISD
Sbjct: 180 MEEEILLLEDQNSKFIKEKKL----MEDRIAECSSQLAEEEEKAKNLAKIRNKQEVMISD 235
Query: 339 LEKALGKLRQEAQEESEAKVDL---VASKDQTLSSMETKLEHLRAELAKKDEAL 389
LE+ L K + QE +AK L ++ ++ +++ L+ +LAKK+E L
Sbjct: 236 LEERLKKEEKTRQELEKAKRKLDGETTDLQDQIAELQAQIDELKLQLAKKEEEL 289
>UniRef100_UPI000023D00A UPI000023D00A UniRef100 entry
Length = 774
Score = 44.7 bits (104), Expect = 0.005
Identities = 43/144 (29%), Positives = 68/144 (46%), Gaps = 11/144 (7%)
Query: 251 QQGLDRVS--EGAFATAFHAFHHYKRSAQDAEV----LRSEVARLRALNAGFVEERKAFL 304
Q+ D+V E A A + K A+DAE L +E + + A + +
Sbjct: 553 QEAEDKVKNLESEAAQAKESESELKTKAEDAEARVAALEAEAKKAQDSEAELKTKVEEAE 612
Query: 305 AQLKAKETSFAKELAEQSSRAEVSLEFRDRKISDLEKALGKLRQEAQEESEAKVDLVASK 364
A++K+ E AK AE++ +LE +K D E L K +EAQ +EA+ A K
Sbjct: 613 AKIKSLEADAAK--AEEAEAKVAALESDVKKAQDAEAELKKQLEEAQAATEAEKKESADK 670
Query: 365 DQTLSSMETKLEHLRAELAKKDEA 388
+ S+E +L L+ + AK +EA
Sbjct: 671 TK---SLEDELNELKEKFAKAEEA 691
Score = 35.4 bits (80), Expect = 2.8
Identities = 35/163 (21%), Positives = 66/163 (40%), Gaps = 14/163 (8%)
Query: 246 DEVVKQQGLDRVSEGAFATAFHAFHHYKRSAQDAEVLRSEVARLRALNAGFVEERKAFLA 305
++ K L+ E A + A + S + + L S++A L + A E
Sbjct: 491 EKSTKLADLENQIEEAQSKVAKAEENLNASQTEKKELESKIADLESNAANSKESESGLTT 550
Query: 306 QLKAKETSFAKELAEQSSRAEVSLEFRDRKISDLEKALGKLRQEAQ-------------E 352
+L+ E K L ++++A+ S K D E + L EA+ E
Sbjct: 551 KLQEAEDK-VKNLESEAAQAKESESELKTKAEDAEARVAALEAEAKKAQDSEAELKTKVE 609
Query: 353 ESEAKVDLVASKDQTLSSMETKLEHLRAELAKKDEALSMMRTQ 395
E+EAK+ + + E K+ L +++ K +A + ++ Q
Sbjct: 610 EAEAKIKSLEADAAKAEEAEAKVAALESDVKKAQDAEAELKKQ 652
>UniRef100_UPI0000235C75 UPI0000235C75 UniRef100 entry
Length = 690
Score = 44.7 bits (104), Expect = 0.005
Identities = 63/261 (24%), Positives = 103/261 (39%), Gaps = 33/261 (12%)
Query: 154 TVFEKKGNVSPHPVAATSSSDGDRAPVISAAFCGPEETPKPNQPGEPSLATPVSSPRAQD 213
T + N S P T + R P +S + + + + S P +SP A
Sbjct: 128 TTADTSSNESGKPGTPTPRAGHTRQPSLSIQSKMRSSSFRKSSVSQGS-GVPSTSPSAML 186
Query: 214 DSVDKPPLSAKGFANREPPYSLSPTPAQVARMDEVVKQ-QGLDRVSEGAFATAFHAFHHY 272
S PPLSA G A +E Q R++E+ K+ + L++ E A +
Sbjct: 187 KSPSLPPLSADGEAVQE------VYRKQSGRIEELEKENKRLEKEVEEVTA-------RF 233
Query: 273 KRSAQDAEVLRSEVARLRAL--NAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEV--- 327
K++ E LR L L EE+ A + +LKA+ S ++L +S R
Sbjct: 234 KKTEDQLEDLREANVDLTELKEKLRIAEEKVAGVEELKAEIASLQRQLQTRSHRNNAGIS 293
Query: 328 ------------SLEFRDRKISDLEKALGKLRQEAQEESEAKVDLVASKDQTLSSMETKL 375
LE + + +E + LR + E+S A +++ ++ LS ET L
Sbjct: 294 GPSESPPADLVQQLESKSAAMEAMELEISNLRAQVTEKS-ALESQISALEEKLSRSETAL 352
Query: 376 EHLRAELAKKDEALSMMRTQA 396
E + EL L+ +A
Sbjct: 353 EQTQHELTDAKATLTRASEKA 373
>UniRef100_P35580 Myosin heavy chain, nonmuscle type B [Homo sapiens]
Length = 1976
Score = 44.7 bits (104), Expect = 0.005
Identities = 30/114 (26%), Positives = 61/114 (53%), Gaps = 10/114 (8%)
Query: 282 LRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRDRK---ISD 338
+ E+ L N+ F++E+K ++ + + +LAE+ +A+ + R+++ ISD
Sbjct: 983 MEEEILLLEDQNSKFIKEKKL----MEDRIAECSSQLAEEEEKAKNLAKIRNKQEVMISD 1038
Query: 339 LEKALGKLRQEAQEESEAKVDL---VASKDQTLSSMETKLEHLRAELAKKDEAL 389
LE+ L K + QE +AK L ++ ++ +++ L+ +LAKK+E L
Sbjct: 1039 LEERLKKEEKTRQELEKAKRKLDGETTDLQDQIAELQAQIDELKLQLAKKEEEL 1092
Score = 39.7 bits (91), Expect = 0.15
Identities = 39/140 (27%), Positives = 63/140 (44%), Gaps = 27/140 (19%)
Query: 274 RSAQDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRD 333
R Q+ + L ++ R + + +++K F QL A+E S + AE+ RAE ++
Sbjct: 1424 RLQQELDDLTVDLDHQRQVASNLEKKQKKF-DQLLAEEKSISARYAEERDRAEAEAREKE 1482
Query: 334 RKISDLEKALGKLRQEAQEESEAK--------VDLVASKD-----------------QTL 368
K L +AL + EA+EE E + DL++SKD Q +
Sbjct: 1483 TKALSLARALEE-ALEAKEEFERQNKQLRADMEDLMSSKDDVGKNVHELEKSKRALEQQV 1541
Query: 369 SSMETKLEHLRAELAKKDEA 388
M T+LE L EL ++A
Sbjct: 1542 EEMRTQLEELEDELQATEDA 1561
Score = 38.9 bits (89), Expect = 0.26
Identities = 32/120 (26%), Positives = 58/120 (47%), Gaps = 8/120 (6%)
Query: 274 RSAQDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRD 333
R Q+ E R + ++R L A +ERK + +K+ +L + ++ E + + RD
Sbjct: 1583 RDEQNEEKKRLLIKQVRELEAELEDERKQRALAVASKK-KMEIDLKDLEAQIEAANKARD 1641
Query: 334 RKISDLEKALGKLRQEAQEESEAKVDLVASKDQTLS---SMETKLEHLRAELAKKDEALS 390
I L K +++ +E EA+ AS+D+ + E KL+ L AE+ + E L+
Sbjct: 1642 EVIKQLRKLQAQMKDYQRELEEAR----ASRDEIFAQSKESEKKLKSLEAEILQLQEELA 1697
Score = 34.3 bits (77), Expect = 6.3
Identities = 29/111 (26%), Positives = 55/111 (49%), Gaps = 6/111 (5%)
Query: 285 EVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQ---SSRAEVSLEFRDRKISDLEK 341
EVA L+ + +A + ++ + + +EL+EQ + R + +LE + + K
Sbjct: 1175 EVAELKKALEEETKNHEAQIQDMRQRHATALEELSEQLEQAKRFKANLEKNKQGLETDNK 1234
Query: 342 ALG---KLRQEAQEESEAKVDLVASKDQTLSSMETKLEHLRAELAKKDEAL 389
L K+ Q+ + ESE K + ++ Q L + ++ + LR ELA+K L
Sbjct: 1235 ELACEVKVLQQVKAESEHKRKKLDAQVQELHAKVSEGDRLRVELAEKASKL 1285
>UniRef100_UPI000042F484 UPI000042F484 UniRef100 entry
Length = 1057
Score = 44.3 bits (103), Expect = 0.006
Identities = 51/213 (23%), Positives = 87/213 (39%), Gaps = 26/213 (12%)
Query: 167 VAATSSSDGDRAPVISAAFCGPEETPKPNQPGEPSLATPVSSPRAQDDSVDKPPLSAKGF 226
+AA++S+ + P +S G TP ++ L TP PRA D PP + G
Sbjct: 377 LAASTSTVNGQTPRVSRVSMGMGGTPARSRQSMGGLVTPRVRPRASGIG-DMPPPPSPGN 435
Query: 227 ANREPPYSLSPTPAQVARMDEVVKQQGLDRVSEGAFATAFHAFHHYK-RSAQDAEVLRSE 285
NR T Q+ ++E +++ + G + + E LR E
Sbjct: 436 INRVL------TARQIEALEEEIREL---KRRNGELEEDLKRMPELRDEDVTEMETLRQE 486
Query: 286 VARLRALNAGFVEERKAFLAQLKAKETS------FAKELAEQSSRAEVSLEFRDRKISDL 339
R + +E + QL+ +T+ +EL + + + LE + R+I DL
Sbjct: 487 TERSK-------QELEFLKTQLETSDTNATDAVRILEELQAEHTAQQEELEAKLREIGDL 539
Query: 340 EKALGKLRQEAQEESEAKVDLVASKDQTLSSME 372
+K L + A+EE A + A KD+ +E
Sbjct: 540 KKELKLANERAEEELAAGAE--ARKDEVQKMLE 570
>UniRef100_Q9AS76 P0028E10.16 protein [Oryza sativa]
Length = 593
Score = 44.3 bits (103), Expect = 0.006
Identities = 52/202 (25%), Positives = 89/202 (43%), Gaps = 26/202 (12%)
Query: 215 SVDKPPLSAKGFANREPPYSLSPTPAQVARMDEVVK--QQGLDRVSEGAFATAFHAFHHY 272
S +K L A+ + SL + AQ+ ++ E++K Q+ LD S A +
Sbjct: 294 SQEKLQLKAQVKELEQASRSLDDSSAQIMKLQEIIKDLQRRLDNDSNEKKMLEERAIE-F 352
Query: 273 KRSAQDAEVLRSEVARLRA----LNAGF---VEERKAFLAQLKAKETSFAKELAEQSSR- 324
++ ++ E R+EVA L+A L A +EE+ +++ E + A L E S
Sbjct: 353 EQVRKELEGSRTEVAELQATINNLKADLGRALEEKSQLESRINDLEHTIACNLEEFSQEK 412
Query: 325 ------------AEVSLEFRDRKISDLEKALGKLRQEAQEESEAKVDLVASKDQTLSSME 372
A SLE K++ E L +L E E S + ++ +Q ++ +E
Sbjct: 413 SSLGAEIQKLKEANASLE---GKLTSTESQLQQLHAEKSEASISSEKQISDLNQAIADLE 469
Query: 373 TKLEHLRAELAKKDEALSMMRT 394
TKLE L +E D ++ + T
Sbjct: 470 TKLELLSSEKTTVDNKVASLLT 491
>UniRef100_Q04834 Nonmuscle myosin heavy chain b [Xenopus laevis]
Length = 1992
Score = 43.9 bits (102), Expect = 0.008
Identities = 35/115 (30%), Positives = 66/115 (56%), Gaps = 12/115 (10%)
Query: 282 LRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRDRK---ISD 338
+ ++ L N+ F++E+K L + A+ TS +LAE+ +A+ + ++++ ISD
Sbjct: 998 MEEDILVLEDQNSKFLKEKK-LLEERIAESTS---QLAEEEEKAKNLAKLKNKQEMMISD 1053
Query: 339 LEKALGKLRQEAQEESEAKVDLVAS----KDQTLSSMETKLEHLRAELAKKDEAL 389
LE+ L K + QE +AK L +DQ ++ ++ ++E L+ +LAKK+E L
Sbjct: 1054 LEERLKKEEKTRQELEKAKRKLDGETTDFQDQ-IAELQAQIEELKLQLAKKEEEL 1107
Score = 37.4 bits (85), Expect = 0.74
Identities = 38/140 (27%), Positives = 62/140 (44%), Gaps = 27/140 (19%)
Query: 274 RSAQDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRD 333
R Q+ + L ++ R + + +++K F QL A+E + + AE+ RAE ++
Sbjct: 1439 RLQQELDDLMVDLDHQRQIVSNLEKKQKKF-DQLLAEEKNISARHAEERDRAEADAREKE 1497
Query: 334 RKISDLEKALGKLRQEAQEESE--------AKVDLVASK-----------------DQTL 368
K L +AL + EAQ+E E DL++SK DQ +
Sbjct: 1498 TKALSLARALDE-ALEAQDEFERLNKQLRAEMEDLMSSKDDVGKNVHELEKSKRALDQQV 1556
Query: 369 SSMETKLEHLRAELAKKDEA 388
M T+LE L EL ++A
Sbjct: 1557 EEMRTQLEELEDELQGTEDA 1576
Score = 35.4 bits (80), Expect = 2.8
Identities = 29/111 (26%), Positives = 54/111 (48%), Gaps = 6/111 (5%)
Query: 285 EVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQ---SSRAEVSLEFRDRKISDLEK 341
EVA LR +A + +++ ++ + +EL+EQ + R +V+LE + + K
Sbjct: 1190 EVAELRKSIEEETRNHEAQIQEMRQRQATALEELSEQLEQAKRFKVNLEKNKQSLESDNK 1249
Query: 342 ALG---KLRQEAQEESEAKVDLVASKDQTLSSMETKLEHLRAELAKKDEAL 389
L K Q+ + ESE K + + Q L + + + LRA++ +K L
Sbjct: 1250 ELATEVKSLQQMKAESEYKRKKLEGQVQELHAKVLEGDRLRADMVEKSSKL 1300
>UniRef100_Q9UAZ3 Hypothetical protein Y38C9A.1 [Caenorhabditis elegans]
Length = 1484
Score = 43.9 bits (102), Expect = 0.008
Identities = 54/213 (25%), Positives = 86/213 (40%), Gaps = 27/213 (12%)
Query: 166 PVAATSSSDGDRAPVISAAFCGPEETPKPNQPGEPSLATPVSSPRAQDDSVDKPP----- 220
P +TS APV+SAA P + P P P+ +P+SS + ++ PP
Sbjct: 1076 PNTSTSIPKAPSAPVLSAA---PPKIVAPAAPAAPTHYSPISSDDDEIQCIEPPPAKKPA 1132
Query: 221 -------LSAKGFANREP--PYSLSPTPA-------QVARMDEVVKQQGLDRVSEGAFAT 264
LSA G P P +P P + M ++ G D VS+ A
Sbjct: 1133 PSASAHNLSALGHQKPGPTAPKVTAPKPTTSTASSQAASTMAALLASSGPDNVSKFILAN 1192
Query: 265 AFHAFHHYKRSAQDAEVLRSEVARLRALNAGFVEERKAFLAQL--KAKETSFAKELAEQS 322
A A +R A + ++ + A A + A L Q AK+ AK +A++
Sbjct: 1193 ANSADSQKQRLALQL-LAHAQASAPAAPTASSTNQIAAQLIQALQAAKDAQTAKLVAQEK 1251
Query: 323 SRAEVSLEFRDRKISDLEKALGKLRQEAQEESE 355
++ E + R + E L +L + A+E S+
Sbjct: 1252 AKKEAQEKARREAEAKKEAQLAQLMKIAEERSK 1284
>UniRef100_Q9Y3T5 Hypothetical protein DKFZp564G013 [Homo sapiens]
Length = 344
Score = 43.9 bits (102), Expect = 0.008
Identities = 37/159 (23%), Positives = 74/159 (46%), Gaps = 15/159 (9%)
Query: 241 QVARMDEVVKQQGLDRVSEGAFATAFHAFHHYKRSAQDAEVLRSEVARLRALNAGFVEER 300
+ A D +V + +++ + K S ++ E + SEV ++R+ + E+
Sbjct: 140 KAAMTDAMVPRSSYEKLQSSLESEVSVLASKLKESVKEKEKVHSEVVQIRSEVSQVKREK 199
Query: 301 KAFLAQLKAKETSF----------AKELAEQSSRAEVSL---EFRDRKISDLEKALGKLR 347
+ LK+KE +ELAE AE S E +D+KI+++ K + KL+
Sbjct: 200 ENIQTLLKSKEQEVNELLQKFQQAQEELAEMKRYAESSSKLEEDKDKKINEMSKEVTKLK 259
Query: 348 QEAQEESEAKVDLVASK--DQTLSSMETKLEHLRAELAK 384
+ S+ +SK Q L +++ +++ L+ +LA+
Sbjct: 260 EALNSLSQLSYSTSSSKRQSQQLEALQQQVKQLQNQLAE 298
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.315 0.130 0.373
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 629,147,160
Number of Sequences: 2790947
Number of extensions: 26880110
Number of successful extensions: 132231
Number of sequences better than 10.0: 1680
Number of HSP's better than 10.0 without gapping: 162
Number of HSP's successfully gapped in prelim test: 1574
Number of HSP's that attempted gapping in prelim test: 128217
Number of HSP's gapped (non-prelim): 5129
length of query: 396
length of database: 848,049,833
effective HSP length: 129
effective length of query: 267
effective length of database: 488,017,670
effective search space: 130300717890
effective search space used: 130300717890
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0005.13


