

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0005.1
(346 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
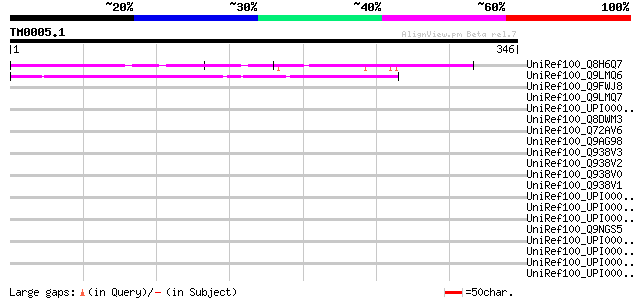
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8H6Q7 CTV.22 [Poncirus trifoliata] 65 4e-09
UniRef100_Q9LMQ6 F7H2.12 protein [Arabidopsis thaliana] 52 2e-05
UniRef100_Q9FWJ8 Hypothetical protein F21N10.2 [Arabidopsis thal... 42 0.019
UniRef100_Q9LMQ7 F7H2.11 [Arabidopsis thaliana] 41 0.055
UniRef100_UPI00003ACAA2 UPI00003ACAA2 UniRef100 entry 40 0.072
UniRef100_Q8DWM3 Putative secreted antigen GbpB/SagA; putative p... 39 0.16
UniRef100_Q72AV6 Methyl-accepting chemotaxis protein [Desulfovib... 39 0.16
UniRef100_Q9AG98 Immunodominant glycoprotein IDG-60 [Streptococc... 39 0.16
UniRef100_Q938V3 Glucan-binding protein B [Streptococcus mutans] 39 0.16
UniRef100_Q938V2 Glucan-binding protein B [Streptococcus mutans] 39 0.16
UniRef100_Q938V0 Glucan-binding protein B [Streptococcus mutans] 39 0.16
UniRef100_Q938V1 Glucan-binding protein B [Streptococcus mutans] 39 0.21
UniRef100_UPI00002F9667 UPI00002F9667 UniRef100 entry 39 0.27
UniRef100_UPI000049A12C UPI000049A12C UniRef100 entry 37 0.61
UniRef100_UPI000026D627 UPI000026D627 UniRef100 entry 37 1.0
UniRef100_Q9NGS5 Prespore protein MF12 [Dictyostelium discoideum] 36 1.8
UniRef100_UPI00003CAC68 UPI00003CAC68 UniRef100 entry 35 2.3
UniRef100_UPI00003CA6DA UPI00003CA6DA UniRef100 entry 35 2.3
UniRef100_UPI00003042AC UPI00003042AC UniRef100 entry 35 2.3
UniRef100_UPI0000258BD1 UPI0000258BD1 UniRef100 entry 35 2.3
>UniRef100_Q8H6Q7 CTV.22 [Poncirus trifoliata]
Length = 1405
Score = 64.7 bits (156), Expect = 4e-09
Identities = 47/180 (26%), Positives = 86/180 (47%), Gaps = 12/180 (6%)
Query: 1 MKDTHFARLNAIYQQICSKLRQHESSPHEPKTFNVEKFRGHKRLYEDIFAMFRLRKSQIT 60
MK+ + LN +YQ+I +KL+QH+S P +PK+ +EK + K + E I + ++ KS I
Sbjct: 626 MKEMYLPELNEMYQKIAAKLQQHDSLPQQPKSDQLEKLKIFKTMLERIISFLQVSKSNIL 685
Query: 61 PEISERVDRVETLINSIIQHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRN 120
P E++ E I + I ++ K + +Q + Q+ P T + +Q SQ +
Sbjct: 686 PSFKEKLGSYEKQIVNFIS----TNRPRKPVSSMQQQGQLPP----THMHSMQQQQSQIS 737
Query: 121 NMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTNDMGFSSSAL 180
+ Q+ Q + + ++++ VQ ++ SGVST+ +S L
Sbjct: 738 QGQPHDNQMNSQIQSMNLAGSMVTMQPNNVTNVQHNSV----PSVSGVSTSQQNMLNSVL 793
Score = 58.9 bits (141), Expect = 2e-07
Identities = 62/205 (30%), Positives = 91/205 (44%), Gaps = 26/205 (12%)
Query: 134 GNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTNDMGFSSSALIK-------GSGN 186
G+SE+ ++ +S G +QT AQA A ++ G S+S L+ GN
Sbjct: 1045 GDSEKPISGISSLSNAGNIGHQQTTSAQAA-APSLAIGTPGISASPLLAEFTGPDGAHGN 1103
Query: 187 LNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVVHLNDAIPAAK----- 241
A ++E+P ++RLI SMS +AL AS+ I VV + D I +
Sbjct: 1104 ALTAISIKASVTEQP---LERLIKAVKSMSPKALSASVSDIGSVVSMIDRIAGSAPGNGS 1160
Query: 242 --FIDEPLEMAGQQNQPSR--IPQG-----RKMARNINAMAFDTSRSCARTHESFNQLTD 292
+ E L + +R I Q RKM R +AM S ++SF QLT
Sbjct: 1161 RAAVGEDLVAMTKCRLQARNFITQDGSSGPRKMRRYTSAMPLSVVSSAGSMNDSFKQLTG 1220
Query: 293 VEEPDLDLVA-SKVKRSRIEVIYLL 316
E DL+ A S +KR R+E + L
Sbjct: 1221 SETSDLESTATSSIKRPRMEANHAL 1245
>UniRef100_Q9LMQ6 F7H2.12 protein [Arabidopsis thaliana]
Length = 1366
Score = 52.4 bits (124), Expect = 2e-05
Identities = 59/268 (22%), Positives = 119/268 (44%), Gaps = 9/268 (3%)
Query: 1 MKDTHFARLNAIYQQICSKLRQHESSPHEPKTFNVEKFRGHKRLYEDIFAMFRLRKSQIT 60
MK+T+ LN IYQ++ +KL+Q +S P + ++ +EK R K + E + + KS I
Sbjct: 635 MKETYLPDLNEIYQRVAAKLQQ-DSMPQQQRSDQLEKLRQFKTMLERMIQFLSVSKSNIM 693
Query: 61 PEISERVDRVE-TLINSIIQHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTS-Q 118
P + ++V E +I + H+ Q Q + PQ Q+ QT+ Q
Sbjct: 694 PALKDKVAYYEKQIIGFLNMHRPRKPVQQGQLPQSQMQPMQQPQSQTVQDQSHDNQTNPQ 753
Query: 119 RNNMCLQ-EQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTNDMGFSS 177
+M +Q A+Q + N LS + PG + +Q + + AS + + +
Sbjct: 754 MQSMSMQGAGPRAQQSSMTNMQSNVLSSR--PGVSAPQQNI-PSSIPASSLESGQGNTLN 810
Query: 178 SALIKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVVHLNDAI 237
+ G++ + + L++ ++A L + ++++ L +S+ + + + D
Sbjct: 811 NGQQVAMGSMQQ--NTSQLVNNSSASAQSGLSTLQSNVNQPQLSSSLLQHQHLKQQQDQQ 868
Query: 238 PAAKFIDEPLEMAGQQNQPSRIPQGRKM 265
K + +M QQ Q + Q +++
Sbjct: 869 MQLKQQFQQRQMQQQQLQARQQQQQQQL 896
Score = 35.0 bits (79), Expect = 3.0
Identities = 57/257 (22%), Positives = 98/257 (37%), Gaps = 38/257 (14%)
Query: 96 SKQQVLPQVLGTQNDHLAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSI---------- 145
S Q+L + HL+ Q Q+N + + Q NS V S
Sbjct: 950 SSPQLLQGASPQMSQHLSPQVDQKNTV--NKMGTPLQPANSPFVVPSPSSTPLAPSPMQV 1007
Query: 146 -KEGPGKA------VQRQTLCAQATEASGVSTNDMGFSSSALIKGSGNLN-EVSHKPALI 197
E PG + + RQ ++ G S+S L++ + + + + +
Sbjct: 1008 DSEKPGSSSLSMGNIARQQATGMQGVVQSLAIGTPGISASPLLQEFTSPDGNILNSSTIT 1067
Query: 198 SEKPSAA---MQRLIGVFTSMSSEALGASIGKIREVVHLNDAIPAAKFIDEPLEMAGQ-- 252
S KPSA ++RLI S+S +AL +++ I VV + D I + + G+
Sbjct: 1068 SGKPSATELPIERLIRAVKSISPQALSSAVSDIGSVVSMVDRIAGSAPGNGSRASVGEDL 1127
Query: 253 --------QNQPSRIPQG----RKMARNINAMAFDTSRSCARTHESFNQLTDVEEPDLDL 300
Q + +G +KM R+ AM + +++ Q E DL+
Sbjct: 1128 VAMTKCRLQARNFMTQEGMMATKKMKRHTTAMPLSVASLGGSVGDNYKQFAGSETSDLES 1187
Query: 301 VA-SKVKRSRIEVIYLL 316
A S K++R E + L
Sbjct: 1188 TATSDGKKARTETEHAL 1204
>UniRef100_Q9FWJ8 Hypothetical protein F21N10.2 [Arabidopsis thaliana]
Length = 634
Score = 42.4 bits (98), Expect = 0.019
Identities = 32/138 (23%), Positives = 62/138 (44%), Gaps = 6/138 (4%)
Query: 1 MKDTHFARLNAIYQQICSKLRQHESSPHEP-KTFNVEKFRGHKRLYEDIFAMFRLRKSQI 59
+K+ HF L+ ++++I KLRQ ES P +P + ++EK + K E + +++S +
Sbjct: 33 LKEMHFPVLSLMHKRIAEKLRQTESLPPQPMQAQSIEKLKAGKLSMEHLMFFLSVQRSNV 92
Query: 60 TPEISERVDRVETLINSIIQHQNL-----SSHQGKHSADVQSKQQVLPQVLGTQNDHLAK 114
+ + ++ E I + Q + QG+ + Q PQV +Q+ +
Sbjct: 93 SEKHRDKFSLYEHHILKFTKSQTMVQRSTQQQQGQFPPSQTAMQSQSPQVHVSQSLDKEQ 152
Query: 115 QTSQRNNMCLQEQQVAKQ 132
S+ C E + Q
Sbjct: 153 MRSRLMPSCQNEASSSLQ 170
Score = 35.0 bits (79), Expect = 3.0
Identities = 41/170 (24%), Positives = 73/170 (42%), Gaps = 17/170 (10%)
Query: 79 QHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRNNMCLQ--------EQQV- 129
QHQ S+Q DV+ +Q++L +++ + + KQ S+ ++ +Q +QQ+
Sbjct: 234 QHQMQRSNQTNEMNDVRIRQRLLEKLVSSSQLQVPKQVSKVSSPQIQNHSSPQLVDQQIL 293
Query: 130 ---AKQYGNSEQGVNQLSIKEGPGKAV-QRQTLCAQATEASGVSTNDMGFSSSALIKGSG 185
+ G + + P + + + SGV N SSS L
Sbjct: 294 PATVNKTGTPLKSSGSPFVAPAPSPVPGDSEMPISVESPVSGVEINSTLDSSSKLGTQET 353
Query: 186 NLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVVHLND 235
L V P I+E+P + RLI F + S ++L S+ +I V+ + D
Sbjct: 354 PLLSVP-PPEPITERP---IDRLIKAFQAASPKSLAESVSEISSVISMVD 399
>UniRef100_Q9LMQ7 F7H2.11 [Arabidopsis thaliana]
Length = 392
Score = 40.8 bits (94), Expect = 0.055
Identities = 57/307 (18%), Positives = 111/307 (35%), Gaps = 72/307 (23%)
Query: 1 MKDTHFARLNAIYQQICSKLRQHESSPHEPKTFNVEKFRGHKRLYEDIFAMFRLRKSQIT 60
M++ + + +YQ++ KL ES P + ++ + +KF+ K E + + LRK I
Sbjct: 20 MRERYLPHVTDVYQRLADKLNHEESLPQQQRSKHFDKFQSMKTNIEQLIQVLSLRKRNIM 79
Query: 61 P----------------------------EISER-VDRVETLINSIIQHQNLSSHQGKHS 91
P +I +R ++ + + + +Q ++ SHQ + S
Sbjct: 80 PILKDCRWDYYEKRIIYFLITLRTGWEDNQIHQRDLNEIYQRVAAKLQQEDSLSHQKQRS 139
Query: 92 ADVQ------------------SKQQVLP-----------QVLGTQNDHLAKQTSQRNNM 122
+ SK + P ++ N ++T Q+ +
Sbjct: 140 DQFEKLKRGKTVLEGMLRFLSLSKSNIKPDLKDSMDYRKNNIMNFLNMQSLRKTVQKLQL 199
Query: 123 CLQE--------QQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTNDMG 174
E Q + + +Q Q+ G + Q + ++ + T G
Sbjct: 200 TKSEIQPMQQPLSQTVQDQSHDDQTTLQMQSMSMQGAGSRVQQIRQGVLQSLEIGT--PG 257
Query: 175 FSSSALI----KGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREV 230
S+S L+ GN+ S ++RLI S+S +AL +++ IR V
Sbjct: 258 ISASPLLPELTSPDGNIINPLTSTCGKSSATELPIERLIRAMKSISPQALSSAVCDIRSV 317
Query: 231 VHLNDAI 237
V + D I
Sbjct: 318 VSMVDRI 324
>UniRef100_UPI00003ACAA2 UPI00003ACAA2 UniRef100 entry
Length = 732
Score = 40.4 bits (93), Expect = 0.072
Identities = 46/177 (25%), Positives = 74/177 (40%), Gaps = 26/177 (14%)
Query: 37 KFRGHKRLYEDI---FAMFRLRKSQITPEISERVDRVETLINSIIQHQNLSSHQGKHSAD 93
+F G K L +DI A + K + +I E+ D + N + + QN + + +
Sbjct: 323 EFTGVKEL-DDISQEIAQLQREKYSLEQDIREKEDSIRQKTNEVQELQNDLDRETSNLQE 381
Query: 94 VQSKQQVLPQVLGTQNDHLAKQTSQRNNM---CLQEQQVAKQYGNSEQGVNQLS------ 144
+++++Q L + AK N++ C +E QV K+Y + GVN S
Sbjct: 382 LEAQKQDAQDRLDEMDQQKAKLKDMLNDVRQKCQEETQVVKEYNEALNGVNGGSLTNLAD 441
Query: 145 IKEGPGKAVQRQTLCAQATEASGVSTNDMGFSSSALIKGSGNLNEVSHKPALISEKP 201
I EG G+ TE S D F + AL+ + N E+ H SE P
Sbjct: 442 ISEGLGQ-----------TERSNYGAMDDPFKNKALM-FTNNTQEL-HTDPFQSEDP 485
>UniRef100_Q8DWM3 Putative secreted antigen GbpB/SagA; putative peptidoglycan
hydrolase [Streptococcus mutans]
Length = 431
Score = 39.3 bits (90), Expect = 0.16
Identities = 46/197 (23%), Positives = 82/197 (41%), Gaps = 13/197 (6%)
Query: 112 LAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTN 171
+A Q S+ NN+ Q+Q Q + V+ L ++ +A ++ AT + T
Sbjct: 34 IASQDSKINNLTAQQQAAQAQVNTIQGQVSALQTQQAELQAENQRLEAQSATLGQQIQT- 92
Query: 172 DMGFSSSALIKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVV 231
SS + + NE + A ++K +AA + + S S + IREVV
Sbjct: 93 ---LSSKIVAR-----NESLKQQARSAQKSNAATSYINAIINSKSVSDAINRVSAIREVV 144
Query: 232 HLNDAIPAAKFIDE-PLEMAGQQNQP---SRIPQGRKMARNINAMAFDTSRSCARTHESF 287
N+ + + D+ +E Q+NQ + +A+N NA+ ++ A
Sbjct: 145 SANEKMLQQQEQDKAAVEQKQQENQAAINTVAANQETIAQNTNALNTQQAQLEAAQLNLQ 204
Query: 288 NQLTDVEEPDLDLVASK 304
+LT ++ LVA K
Sbjct: 205 AELTTAQDQKATLVAQK 221
>UniRef100_Q72AV6 Methyl-accepting chemotaxis protein [Desulfovibrio vulgaris]
Length = 800
Score = 39.3 bits (90), Expect = 0.16
Identities = 46/187 (24%), Positives = 84/187 (44%), Gaps = 42/187 (22%)
Query: 125 QEQQVAKQYGNSEQGVNQ-LSIKEGPGKAVQRQTLCAQATEASGVSTNDMGFSSSALIKG 183
+ +++A+Q + G Q +S E G AV++ A+ EA+ ++ +
Sbjct: 535 ETEELARQVEHGADGARQQVSCLENTGAAVRQ---LAELLEAAAQHADEA-------VNS 584
Query: 184 SGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSS---------EALGASIGKIREV---- 230
+G E + + I E+ +AAMQR+ G+ S+SS E++G+ +G I ++
Sbjct: 585 AGEAGEKARQGTEIMERSAAAMQRMHGLSMSLSSSMHQLGEQTESIGSILGVINDIADQT 644
Query: 231 --VHLNDAIPAAKFIDEPLEMAGQQNQPSRIPQGRKMARNINAMAFDTSRSCARTHESFN 288
+ LN AI AA+ AG+ GR A + + R+ A T E N
Sbjct: 645 NLLALNAAIEAAR--------AGE--------SGRGFAVVADEVRKLAERTVAATGEVRN 688
Query: 289 QLTDVEE 295
+ +++E
Sbjct: 689 SIRNIQE 695
>UniRef100_Q9AG98 Immunodominant glycoprotein IDG-60 [Streptococcus mutans]
Length = 431
Score = 39.3 bits (90), Expect = 0.16
Identities = 46/197 (23%), Positives = 82/197 (41%), Gaps = 13/197 (6%)
Query: 112 LAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTN 171
+A Q S+ NN+ Q+Q Q + V+ L ++ +A ++ AT + T
Sbjct: 34 IASQDSKINNLTAQQQAAQAQVNTIQGQVSALQTQQAELQAENQRLEAQSATLGQQIQT- 92
Query: 172 DMGFSSSALIKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVV 231
SS + + NE + A ++K +AA + + S S + IREVV
Sbjct: 93 ---LSSKIVAR-----NESLKQQARSAQKSNAATSYINAIINSKSVSDAINRVSAIREVV 144
Query: 232 HLNDAIPAAKFIDE-PLEMAGQQNQP---SRIPQGRKMARNINAMAFDTSRSCARTHESF 287
N+ + + D+ +E Q+NQ + +A+N NA+ ++ A
Sbjct: 145 SANEKMLQQQEQDKAAVEQKQQENQAAINTVAANQETIAQNTNALNTQQAQLEAAQLNLQ 204
Query: 288 NQLTDVEEPDLDLVASK 304
+LT ++ LVA K
Sbjct: 205 AELTTAQDQKATLVAQK 221
>UniRef100_Q938V3 Glucan-binding protein B [Streptococcus mutans]
Length = 431
Score = 39.3 bits (90), Expect = 0.16
Identities = 46/197 (23%), Positives = 82/197 (41%), Gaps = 13/197 (6%)
Query: 112 LAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTN 171
+A Q S+ NN+ Q+Q Q + V+ L ++ +A ++ AT + T
Sbjct: 34 IASQDSKINNLTAQQQAAQAQVNTIQGQVSALQTQQAELQAENQRLEAQSATLGQQIQT- 92
Query: 172 DMGFSSSALIKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVV 231
SS + + NE + A ++K +AA + + S S + IREVV
Sbjct: 93 ---LSSKIVAR-----NESLKQQARSAQKSNAATSYINAIINSKSVSDAINRVSAIREVV 144
Query: 232 HLNDAIPAAKFIDE-PLEMAGQQNQP---SRIPQGRKMARNINAMAFDTSRSCARTHESF 287
N+ + + D+ +E Q+NQ + +A+N NA+ ++ A
Sbjct: 145 SANEKMLQQQEQDKAAVEQKQQENQAAINTVAANQETIAQNTNALNTQQAQLEAAQLNLQ 204
Query: 288 NQLTDVEEPDLDLVASK 304
+LT ++ LVA K
Sbjct: 205 AELTTAQDQKATLVAQK 221
>UniRef100_Q938V2 Glucan-binding protein B [Streptococcus mutans]
Length = 432
Score = 39.3 bits (90), Expect = 0.16
Identities = 46/197 (23%), Positives = 82/197 (41%), Gaps = 13/197 (6%)
Query: 112 LAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTN 171
+A Q S+ NN+ Q+Q Q + V+ L ++ +A ++ AT + T
Sbjct: 34 IASQDSKINNLTAQQQAAQAQVNTIQGQVSALQTQQAELQAENQRLEAQSATLGQQIQT- 92
Query: 172 DMGFSSSALIKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVV 231
SS + + NE + A ++K +AA + + S S + IREVV
Sbjct: 93 ---LSSKIVAR-----NESLKQQARSAQKSNAATSYINAIINSKSVSDAINRVSAIREVV 144
Query: 232 HLNDAIPAAKFIDE-PLEMAGQQNQP---SRIPQGRKMARNINAMAFDTSRSCARTHESF 287
N+ + + D+ +E Q+NQ + +A+N NA+ ++ A
Sbjct: 145 SANEKMLQQQEQDKAAVEQKQQENQAAINTVAANQETIAQNTNALNTQQAQLEAAQLNLQ 204
Query: 288 NQLTDVEEPDLDLVASK 304
+LT ++ LVA K
Sbjct: 205 AELTTAQDQKATLVAQK 221
>UniRef100_Q938V0 Glucan-binding protein B [Streptococcus mutans]
Length = 431
Score = 39.3 bits (90), Expect = 0.16
Identities = 46/197 (23%), Positives = 82/197 (41%), Gaps = 13/197 (6%)
Query: 112 LAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTN 171
+A Q S+ NN+ Q+Q Q + V+ L ++ +A ++ AT + T
Sbjct: 34 IASQDSKINNLTAQQQAAQAQVNTIQGQVSALQTQQAELQAENQRLEAQSATLGQQIQT- 92
Query: 172 DMGFSSSALIKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVV 231
SS + + NE + A ++K +AA + + S S + IREVV
Sbjct: 93 ---LSSKIVAR-----NESLKQQARSAQKSNAATSYINAIINSKSVSDAINRVSAIREVV 144
Query: 232 HLNDAIPAAKFIDE-PLEMAGQQNQP---SRIPQGRKMARNINAMAFDTSRSCARTHESF 287
N+ + + D+ +E Q+NQ + +A+N NA+ ++ A
Sbjct: 145 SANEKMLQQQEQDKAAVEQKQQENQAAINTVAANQETIAQNTNALNTQQAQLEAAQLNLQ 204
Query: 288 NQLTDVEEPDLDLVASK 304
+LT ++ LVA K
Sbjct: 205 AELTTAQDQKATLVAQK 221
>UniRef100_Q938V1 Glucan-binding protein B [Streptococcus mutans]
Length = 432
Score = 38.9 bits (89), Expect = 0.21
Identities = 46/197 (23%), Positives = 82/197 (41%), Gaps = 13/197 (6%)
Query: 112 LAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTN 171
+A Q S+ NN+ Q+Q Q + V+ L ++ +A ++ AT + T
Sbjct: 34 IASQDSKINNLTAQQQAAQAQVNTIQGQVSALQTQQAELQAENQRLEAQSATLGQQIQT- 92
Query: 172 DMGFSSSALIKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVV 231
SS + + NE + A ++K +AA + + S S + IREVV
Sbjct: 93 ---LSSKIVAR-----NESLKQQARSAQKSNAATSYINAIINSKSVSDAINRVSAIREVV 144
Query: 232 HLNDAIPAAKFIDE-PLEMAGQQNQP---SRIPQGRKMARNINAMAFDTSRSCARTHESF 287
N+ + + D+ +E Q+NQ + +A+N NA+ ++ A
Sbjct: 145 SANEKMLHQQEQDKAAVEQKHQENQAAINTVAANQETIAQNTNALNTQQAQLEAAQLNLQ 204
Query: 288 NQLTDVEEPDLDLVASK 304
+LT ++ LVA K
Sbjct: 205 AELTTAQDQKATLVAQK 221
>UniRef100_UPI00002F9667 UPI00002F9667 UniRef100 entry
Length = 1034
Score = 38.5 bits (88), Expect = 0.27
Identities = 50/251 (19%), Positives = 104/251 (40%), Gaps = 16/251 (6%)
Query: 62 EISERVDRVETLINSIIQHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRNN 121
EI ER+ ++ ++ + N + KH ++++ K++ L Q + T+ D L + +
Sbjct: 611 EIDERLKALQKETAALDRRDNELLAEKKHLSELKGKRRQLEQKISTKQDSLRQMEQNITD 670
Query: 122 M-CLQEQQVAKQYGNSEQGVNQL-----SIKEGPGKAVQRQTLCAQATEASGVST---ND 172
+ ++E+ K + Q V + SIK +++ L + E + T ND
Sbjct: 671 LKKIEEETKGKVSEVNSQKVAMVKVFIDSIKLKAKLTMEKVYLALKMVELTAEKTKLEND 730
Query: 173 MGFSSSALIKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVVH 232
+S L +++ + A ++E+ M+R + + S ++L + +R
Sbjct: 731 FREGASLLRSMDQKCSQLEQRRAQLTEQCKGHMKRAMSICRMQSKDSLPEDLRNVRLCFF 790
Query: 233 LNDAIPAAKFIDEPLEMAGQQNQPSRIPQGRKMARNINAMAFDTSRSCARTHESFNQL-T 291
AK D P E+ SR+ + R A +++ + R + QL
Sbjct: 791 GGGKQAFAKLPDTPDEI------ESRLSEERSRAECFTSLSENVRMRWFRRDQEIKQLEK 844
Query: 292 DVEEPDLDLVA 302
++EE + L A
Sbjct: 845 ELEEKENALEA 855
>UniRef100_UPI000049A12C UPI000049A12C UniRef100 entry
Length = 378
Score = 37.4 bits (85), Expect = 0.61
Identities = 31/120 (25%), Positives = 56/120 (45%), Gaps = 15/120 (12%)
Query: 47 DIFAMFRLRKSQITPEISERVDRVETLINSIIQHQNLSSHQGKHSADVQ----SKQQVLP 102
DI L SQ ++ + V ++T +NS+IQ + + K ++ +K VLP
Sbjct: 3 DITPQQYLELSQKIDQLQKMVQSIQTSVNSLIQITEIQNTLPKTIENIVKATFNKYLVLP 62
Query: 103 --------QVLGTQNDHLAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQ 154
+++ +N +++ ++S +QQ KQ S +NQ S+KE P K +Q
Sbjct: 63 EDYDASIQEMIDKKNKNISTRSSSSYKDTEHKQQEVKQ---SSSSINQNSVKEEPKKVLQ 119
>UniRef100_UPI000026D627 UPI000026D627 UniRef100 entry
Length = 272
Score = 36.6 bits (83), Expect = 1.0
Identities = 26/97 (26%), Positives = 41/97 (41%), Gaps = 1/97 (1%)
Query: 65 ERVDRVETLINSIIQHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRNNMCL 124
E +D + IIQ Q + QG + +QQ + Q Q A Q Q+ M
Sbjct: 76 ELIDNIYNDYQKIIQQQQ-QAMQGGMAQGGMEQQQAMQQQAMQQQQQQAMQQQQQQAMQQ 134
Query: 125 QEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQ 161
Q+QQ +Q +Q + Q + +A+Q+Q Q
Sbjct: 135 QQQQAMQQQQQQQQAMQQQQAMQQQQQAMQQQQQAMQ 171
>UniRef100_Q9NGS5 Prespore protein MF12 [Dictyostelium discoideum]
Length = 1287
Score = 35.8 bits (81), Expect = 1.8
Identities = 30/139 (21%), Positives = 61/139 (43%), Gaps = 8/139 (5%)
Query: 19 KLRQHESSPHEPKTFNVEKFRGHKRLYEDIFAMFRLRKSQITPEISERVDRVETLINSII 78
KL + E E K +E+ R RL ++ K ++T + E + V+ +++ +I
Sbjct: 642 KLLERERERKE-KEKQLEREREQLRLEKE-------EKERLTQQSEELDNLVDEILDELI 693
Query: 79 QHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRNNMCLQEQQVAKQYGNSEQ 138
Q + + +K + Q+L Q +Q Q+ LQ+QQ+ +Q +Q
Sbjct: 694 FDQVSEIATTTYQEYLSNKNKQQQQILQQQQQQQQQQQQQQQQQILQQQQLQQQQQQQQQ 753
Query: 139 GVNQLSIKEGPGKAVQRQT 157
+ Q +++ + Q+ T
Sbjct: 754 QLQQQQLQQQQQQLQQQST 772
>UniRef100_UPI00003CAC68 UPI00003CAC68 UniRef100 entry
Length = 1476
Score = 35.4 bits (80), Expect = 2.3
Identities = 44/192 (22%), Positives = 78/192 (39%), Gaps = 20/192 (10%)
Query: 100 VLPQVLGTQNDHLAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLC 159
+LP + Q D L ++ N M + ++ G++ G++ G + L
Sbjct: 283 ILPALFEKQED-LEAFRARHNQMKAKRAKLEDYEGDAYLGIDA-------GSTTTKLILM 334
Query: 160 AQATEA--SGVSTNDMGFSSSALIKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSS 217
+Q E S S+N+ G ++I + +L E+ + EK A + G S+
Sbjct: 335 SQTNEILYSFYSSNN-GNPLQSVIDATSDLYEI------LPEKVRVAQSGITGYGESLIK 387
Query: 218 EALGASIGKIREVVHLNDAIPAAKFIDEPLEMAGQQNQPSRIPQGRKMARNINAMAFDTS 277
AL +G+I V H A +D L++ GQ + +I +G + +N S
Sbjct: 388 AALKIDVGEIETVAHYRAAREFLPDVDFILDIGGQDMKCMKIKKGALDSLMLNEAC---S 444
Query: 278 RSCARTHESFNQ 289
C E+F Q
Sbjct: 445 AGCGSFLETFAQ 456
>UniRef100_UPI00003CA6DA UPI00003CA6DA UniRef100 entry
Length = 1476
Score = 35.4 bits (80), Expect = 2.3
Identities = 44/192 (22%), Positives = 78/192 (39%), Gaps = 20/192 (10%)
Query: 100 VLPQVLGTQNDHLAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLC 159
+LP + Q D L ++ N M + ++ G++ G++ G + L
Sbjct: 283 ILPALFEKQED-LEDFRARHNQMKAKRAKLEDYEGDAYLGIDA-------GSTTTKLILM 334
Query: 160 AQATEA--SGVSTNDMGFSSSALIKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSS 217
+Q E S S+N+ G ++I + +L E+ + EK A + G S+
Sbjct: 335 SQTNEILYSFYSSNN-GNPLQSVIDATSDLYEI------LPEKVRVAQSGITGYGESLIK 387
Query: 218 EALGASIGKIREVVHLNDAIPAAKFIDEPLEMAGQQNQPSRIPQGRKMARNINAMAFDTS 277
AL +G+I V H A +D L++ GQ + +I +G + +N S
Sbjct: 388 AALKIDVGEIETVAHYRAAREFLPDVDFILDIGGQDMKCMKIKKGALDSLMLNEAC---S 444
Query: 278 RSCARTHESFNQ 289
C E+F Q
Sbjct: 445 AGCGSFLETFAQ 456
>UniRef100_UPI00003042AC UPI00003042AC UniRef100 entry
Length = 2006
Score = 35.4 bits (80), Expect = 2.3
Identities = 47/182 (25%), Positives = 76/182 (40%), Gaps = 34/182 (18%)
Query: 139 GVNQLSIKEGPGKAVQRQ-------TLCAQATEASGVSTNDMGF-SSSALIKGSGNLNEV 190
G QLSI+ G A T+ T A+G +TND+ + SSS+L+ G + +
Sbjct: 1052 GTPQLSIETGANDATLNYSSGSGSPTITFNYTVAAGHTTNDLEYISSSSLVLNGGAIKDA 1111
Query: 191 SHKPA-----------LISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVVHLNDAIPA 239
+ A +S S A+ ++ V TS++S S + D IP
Sbjct: 1112 AGNNASLSMADPGSSGSLSSNKSFAVDGIVPVMTSVTSSLADGS-------YMVGDTIPV 1164
Query: 240 AKFIDEPLEMAGQQNQPSRIPQGRKMARNINAMAFDTSRSCARTHESFNQLTDV--EEPD 297
A D+ + + G+ Q S + +A N + + D S + FN + +V E D
Sbjct: 1165 AIVFDDTIYVIGRP-QVSLV----TLANNSSTLV-DYSSGSGSSSIIFNYVIEVGHESND 1218
Query: 298 LD 299
LD
Sbjct: 1219 LD 1220
>UniRef100_UPI0000258BD1 UPI0000258BD1 UniRef100 entry
Length = 278
Score = 35.4 bits (80), Expect = 2.3
Identities = 34/148 (22%), Positives = 67/148 (44%), Gaps = 13/148 (8%)
Query: 57 SQITPEI---SERVDRVETLINSIIQHQNLSSHQGK---HSADVQSKQQVLPQVLGTQND 110
++I PE+ + ++R++ LI++++ H +LS + D++ + + L +L Q D
Sbjct: 138 AKIRPEVLSQARSIERLDKLIDTLVGHLSLSLEDRQTLLEEIDLEERSKNLVSLLENQLD 197
Query: 111 HLAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVST 170
+ R+ + Q ++ K+Y SEQ + L + G G+ V + E
Sbjct: 198 LTQMEKKIRDRVKKQMEKSQKEYYLSEQ-MKALKKEMGEGEDVDDE------IEVIEKKI 250
Query: 171 NDMGFSSSALIKGSGNLNEVSHKPALIS 198
D G A+ K + LN+ P +S
Sbjct: 251 KDSGMPEEAVEKVNTELNKFKLMPPNVS 278
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.130 0.362
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 512,357,617
Number of Sequences: 2790947
Number of extensions: 19279999
Number of successful extensions: 62574
Number of sequences better than 10.0: 78
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 79
Number of HSP's that attempted gapping in prelim test: 62223
Number of HSP's gapped (non-prelim): 239
length of query: 346
length of database: 848,049,833
effective HSP length: 128
effective length of query: 218
effective length of database: 490,808,617
effective search space: 106996278506
effective search space used: 106996278506
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0005.1


