

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0002.6
(260 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
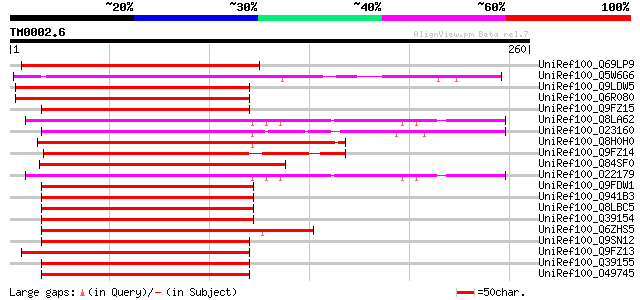
Sequences producing significant alignments: (bits) Value
UniRef100_Q69LP9 Myb-related transcription factor-like [Oryza sa... 195 1e-48
UniRef100_Q5W6G6 Hypothetical protein OSJNBa0077J17.12 [Oryza sa... 180 3e-44
UniRef100_Q9LDW5 MYB transcription factor-like protein [Arabidop... 179 5e-44
UniRef100_Q6R080 MYB transcription factor [Arabidopsis thaliana] 179 5e-44
UniRef100_Q9FZ15 Tuber-specific and sucrose-responsive element b... 179 8e-44
UniRef100_Q8LA62 Putative MYB family transcription factor [Arabi... 178 1e-43
UniRef100_O23160 Myb-related protein [Arabidopsis thaliana] 177 2e-43
UniRef100_Q8H0H0 Myb-like protein [Nicotiana tabacum] 177 2e-43
UniRef100_Q9FZ14 Tuber-specific and sucrose-responsive element b... 176 7e-43
UniRef100_Q84SF0 P0020E09.14 protein [Oryza sativa] 175 1e-42
UniRef100_O22179 MYB family transcription factor [Arabidopsis th... 174 2e-42
UniRef100_Q9FDW1 Myb-related protein, 33.3K [Arabidopsis thaliana] 173 3e-42
UniRef100_Q941B3 AT5g67300/K8K14_2 [Arabidopsis thaliana] 173 3e-42
UniRef100_Q8LBC5 Myb-related protein, 33.3K [Arabidopsis thaliana] 173 3e-42
UniRef100_Q39154 MYB-related protein [Arabidopsis thaliana] 173 3e-42
UniRef100_Q6ZHS5 Putative tuber-specific and sucrose-responsive ... 171 1e-41
UniRef100_Q9SN12 R2R3-MYB transcription factor [Arabidopsis thal... 170 3e-41
UniRef100_Q9FZ13 Tuber-specific and sucrose-responsive element b... 170 3e-41
UniRef100_Q39155 MYB-related protein [Arabidopsis thaliana] 170 3e-41
UniRef100_O49745 R2R3-MYB transcription factor [Arabidopsis thal... 170 3e-41
>UniRef100_Q69LP9 Myb-related transcription factor-like [Oryza sativa]
Length = 318
Score = 195 bits (495), Expect = 1e-48
Identities = 88/119 (73%), Positives = 99/119 (82%)
Query: 7 SSNGGGGGDRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQL 66
S GGGGD DR+KG WSP+EDE L +LVG+HG RNWS+IS IPGRSGKSCRLRWCNQL
Sbjct: 3 SCRRGGGGDVDRIKGPWSPEEDEALQRLVGRHGARNWSLISKSIPGRSGKSCRLRWCNQL 62
Query: 67 SPDVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESL 125
SP V+HRPFTP ED I++AHA GNKWATI+RLL GRTDNAIKNHWNSTL+R+ SL
Sbjct: 63 SPQVEHRPFTPEEDDTILRAHARFGNKWATIARLLAGRTDNAIKNHWNSTLKRKHHSSL 121
>UniRef100_Q5W6G6 Hypothetical protein OSJNBa0077J17.12 [Oryza sativa]
Length = 303
Score = 180 bits (457), Expect = 3e-44
Identities = 117/279 (41%), Positives = 141/279 (49%), Gaps = 55/279 (19%)
Query: 3 GGGDSSNGGGGGDRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRW 62
G G SS+ G + +KGSWSP+EDE L V +HGPRNW+ IS +PGRSGKSCRLRW
Sbjct: 5 GEGSSSSAAAAGGK--IKGSWSPEEDEQLRGAVARHGPRNWTAISEEVPGRSGKSCRLRW 62
Query: 63 CNQLSPDVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRG 122
CNQLSP V RPFTP ED +I+ AHA +GNKWATI+RLL GRTDN++KNHWNS+LRR R
Sbjct: 63 CNQLSPGVHRRPFTPDEDALIVAAHAKYGNKWATIARLLDGRTDNSVKNHWNSSLRRNRR 122
Query: 123 ESLPLPLASPSPS-------------------PSPSPPLPFSMAKRPHGYELLDRYDGLM 163
+ A+ S S S + P S A G +L
Sbjct: 123 AAAAAAAAAASVSYQSMDLTEEADNDDEGTSDDSVAIPAQSSPAAVVAGVPVLP------ 176
Query: 164 DQGYPFKRHCVEKDDEESRDGVEEAETMTEEEVLASSVPTSLSLFPMGEK---------- 213
P KR CV T E PTSLSL P G
Sbjct: 177 PPPPPAKRLCV------------APPTGVEHRAPPPDPPTSLSLSPPGAAAAAISASTVV 224
Query: 214 --KMVAVAEKE----EKKMDMQEGLMRTMMQRMIAEEVR 246
A AE+E EK Q+ + MM++MI EEV+
Sbjct: 225 GGSSAARAEEEAVAREKARMEQDPWLMAMMRQMITEEVQ 263
>UniRef100_Q9LDW5 MYB transcription factor-like protein [Arabidopsis thaliana]
Length = 399
Score = 179 bits (455), Expect = 5e-44
Identities = 81/117 (69%), Positives = 94/117 (80%)
Query: 4 GGDSSNGGGGGDRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWC 63
GG GGGGG R +VKG WS +ED L KLV + GPRNWS+I+ GIPGRSGKSCRLRWC
Sbjct: 40 GGGGCGGGGGGIRSKVKGPWSTEEDAVLTKLVRKLGPRNWSLIARGIPGRSGKSCRLRWC 99
Query: 64 NQLSPDVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
NQL P ++ +PF+ ED +II AHAVHGNKWA I++LL GRTDNAIKNHWNSTLRR+
Sbjct: 100 NQLDPCLKRKPFSDEEDRMIISAHAVHGNKWAVIAKLLTGRTDNAIKNHWNSTLRRK 156
>UniRef100_Q6R080 MYB transcription factor [Arabidopsis thaliana]
Length = 399
Score = 179 bits (455), Expect = 5e-44
Identities = 81/117 (69%), Positives = 94/117 (80%)
Query: 4 GGDSSNGGGGGDRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWC 63
GG GGGGG R +VKG WS +ED L KLV + GPRNWS+I+ GIPGRSGKSCRLRWC
Sbjct: 40 GGGGCGGGGGGIRSKVKGPWSTEEDAVLTKLVRKLGPRNWSLIARGIPGRSGKSCRLRWC 99
Query: 64 NQLSPDVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
NQL P ++ +PF+ ED +II AHAVHGNKWA I++LL GRTDNAIKNHWNSTLRR+
Sbjct: 100 NQLDPCLKRKPFSDEEDRMIISAHAVHGNKWAVIAKLLTGRTDNAIKNHWNSTLRRK 156
>UniRef100_Q9FZ15 Tuber-specific and sucrose-responsive element binding factor
[Solanum tuberosum]
Length = 364
Score = 179 bits (453), Expect = 8e-44
Identities = 81/104 (77%), Positives = 90/104 (85%)
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
DRVKG WSP+EDE L +LV +HGPRNWS+IS IPGRSGKSCRLRWCNQLSP V+HR FT
Sbjct: 2 DRVKGPWSPEEDELLQQLVQKHGPRNWSLISKSIPGRSGKSCRLRWCNQLSPQVEHRAFT 61
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
P ED II+AHA GNKWATI+RLL GRTDNAIKNHWNSTL+R+
Sbjct: 62 PEEDETIIRAHARFGNKWATIARLLNGRTDNAIKNHWNSTLKRK 105
>UniRef100_Q8LA62 Putative MYB family transcription factor [Arabidopsis thaliana]
Length = 309
Score = 178 bits (451), Expect = 1e-43
Identities = 114/272 (41%), Positives = 150/272 (54%), Gaps = 37/272 (13%)
Query: 9 NGGGGGDRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSP 68
+G + DR+KG WSP+ED+ L LV +HGPRNWS+IS IPGRSGKSCRLRWCNQLSP
Sbjct: 2 SGSTRKEMDRIKGPWSPEEDDLLQSLVQKHGPRNWSLISKSIPGRSGKSCRLRWCNQLSP 61
Query: 69 DVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR-------- 120
+V+HR FT ED II AHA GNKWATI+RLL GRTDNAIKNHWNSTL+R+
Sbjct: 62 EVEHRGFTAEEDDTIILAHARFGNKWATIARLLNGRTDNAIKNHWNSTLKRKCSGGGGGG 121
Query: 121 -RGESLPL--------PLASPSP---SPSPSPPLPFSMAKRPHGYELLDRYDGLMDQGYP 168
G+S L P + S + A P G ++ ++ P
Sbjct: 122 EEGQSCDFGGNGGYDGNLTDEKPLKRTASGGGGVVVVTALSPTGSDVSEQSQS-SGSVLP 180
Query: 169 FKRHC-VEKDDEESRDGVEEAETMTEEE-----VLASSVP------TSLSLFPMGEKKMV 216
C V K + V E+ + EEE L+ S+P T + LFP+ ++
Sbjct: 181 VSSSCHVFKPTARAGGVVIESSSSEEEEKDPMTCLSLSLPWVNESTTPVELFPVKREE-- 238
Query: 217 AVAEKEEKKMDMQEGLMRTMMQRMIAEEVRAY 248
E++E+++ G T++Q MI EVR+Y
Sbjct: 239 --EEEKEREISGLGGDFMTVVQEMIKTEVRSY 268
>UniRef100_O23160 Myb-related protein [Arabidopsis thaliana]
Length = 320
Score = 177 bits (450), Expect = 2e-43
Identities = 112/269 (41%), Positives = 148/269 (54%), Gaps = 43/269 (15%)
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
+R+KG WSP+ED+ L +LV +HGPRNWS+IS IPGRSGKSCRLRWCNQLSP+V+HR F+
Sbjct: 10 ERIKGPWSPEEDDLLQRLVQKHGPRNWSLISKSIPGRSGKSCRLRWCNQLSPEVEHRAFS 69
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR---------------- 120
ED II+AHA GNKWATISRLL GRTDNAIKNHWNSTL+R+
Sbjct: 70 QEEDETIIRAHARFGNKWATISRLLNGRTDNAIKNHWNSTLKRKCSVEGQSCDFGGNGGY 129
Query: 121 ---RGESLPLPLASPSPSPSPSPPLPFSMAKRPHGYELLDRYDGLMDQGYPFKRHCVEKD 177
GE PL + S S L S P G ++ ++ G G + V +
Sbjct: 130 DGNLGEEQPLK-RTASGGGGVSTGLYMSPGS-PSGSDVSEQSSG----GAHVFKPTVRSE 183
Query: 178 DEESRDGVEEAETMT-----EEEVLASSVPTSLS------------LFPMGEKKMVAVAE 220
S G + ++ +E + + P L+ LFP+ +++ V V E
Sbjct: 184 VTASSSGEDPPTYLSLSLPWTDETVRVNEPVQLNQNTVMDGGYTAELFPVRKEEQVEVEE 243
Query: 221 KEEKKMDMQ-EGLMRTMMQRMIAEEVRAY 248
+E K + G T++Q MI EVR+Y
Sbjct: 244 EEAKGISGGFGGEFMTVVQEMIRTEVRSY 272
>UniRef100_Q8H0H0 Myb-like protein [Nicotiana tabacum]
Length = 329
Score = 177 bits (449), Expect = 2e-43
Identities = 90/159 (56%), Positives = 103/159 (64%), Gaps = 6/159 (3%)
Query: 15 DRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRP 74
+ DR+KG WSP+EDE L LV +HGPRNWS+IS +PGRSGKSCRLRWCNQLSP V+HR
Sbjct: 9 ESDRIKGPWSPEEDELLQSLVEKHGPRNWSLISKSVPGRSGKSCRLRWCNQLSPQVEHRA 68
Query: 75 FTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR-----RGESLPLPL 129
FT ED II+AHA GNKWATI+RLL GRTDNAIKNHWNSTL+R+ S P
Sbjct: 69 FTSEEDETIIRAHAKFGNKWATIARLLSGRTDNAIKNHWNSTLKRKCCSMSEDLSFETPQ 128
Query: 130 ASPSPSPSPSPPLPFSMAKRPHGYELLDRYDGLMDQGYP 168
S S P FS P D D + G+P
Sbjct: 129 QPLKRSSSVGPSTNFSSGMNPGSPSGSDLSDSSL-SGFP 166
>UniRef100_Q9FZ14 Tuber-specific and sucrose-responsive element binding factor
[Solanum tuberosum]
Length = 306
Score = 176 bits (445), Expect = 7e-43
Identities = 88/152 (57%), Positives = 105/152 (68%), Gaps = 12/152 (7%)
Query: 18 RVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
R+KG WSP+EDE L LV +HGPRNW++IS +P RSGKSCRLRWCNQLSP V+HR FTP
Sbjct: 1 RIKGPWSPEEDELLQTLVEKHGPRNWTLISKSVPRRSGKSCRLRWCNQLSPQVEHRAFTP 60
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESLPLPLASPSPS-P 136
ED II+AHA +GNKWATI+RLL GRTDNAIKNHWNSTL+R+ P S S
Sbjct: 61 EEDDTIIRAHAKYGNKWATIARLLSGRTDNAIKNHWNSTLKRK------CPSMSEDLSFE 114
Query: 137 SPSPPLPFSMAKRPHGYELLDRYDGLMDQGYP 168
+P PPL S + P + +M+ G P
Sbjct: 115 TPQPPLKRSSSVGP-----CTNFSSVMNPGSP 141
>UniRef100_Q84SF0 P0020E09.14 protein [Oryza sativa]
Length = 250
Score = 175 bits (443), Expect = 1e-42
Identities = 79/123 (64%), Positives = 88/123 (71%)
Query: 16 RDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPF 75
+ R KGSW +ED L +LV QHGP WS+IS IPGRSGKSCRLRWCNQLSP VQHRPF
Sbjct: 6 KTRKKGSWRAEEDALLTRLVAQHGPHRWSIISGAIPGRSGKSCRLRWCNQLSPAVQHRPF 65
Query: 76 TPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESLPLPLASPSPS 135
TP ED ++ AHA HGNKWATI+RLLPGRTDN++KNHWNS LRR P
Sbjct: 66 TPQEDALLAAAHARHGNKWATIARLLPGRTDNSVKNHWNSNLRRCLRRQAKFKSKDPDLL 125
Query: 136 PSP 138
P P
Sbjct: 126 PDP 128
>UniRef100_O22179 MYB family transcription factor [Arabidopsis thaliana]
Length = 309
Score = 174 bits (442), Expect = 2e-42
Identities = 114/272 (41%), Positives = 147/272 (53%), Gaps = 37/272 (13%)
Query: 9 NGGGGGDRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSP 68
+G + DR+KG WSP+ED+ L LV +HGPRNWS+IS IPGRSGKSCRLRWCNQLSP
Sbjct: 2 SGSTRKEMDRIKGPWSPEEDDLLQSLVQKHGPRNWSLISKSIPGRSGKSCRLRWCNQLSP 61
Query: 69 DVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR-------- 120
+V+HR FT ED II AHA GNKWATI+RLL GRTDNAIKNHWNSTL+R+
Sbjct: 62 EVEHRGFTAEEDDTIILAHARFGNKWATIARLLNGRTDNAIKNHWNSTLKRKCSGGGGGG 121
Query: 121 -RGESLPL--------PLASPSP---SPSPSPPLPFSMAKRPHGYELLDRYDGLMDQGYP 168
G+S L P S + A P G ++ ++ P
Sbjct: 122 EEGQSCDFGGNGGYDGNLTDEKPLKRRASGGGGVVVVTALSPTGSDVSEQSQS-SGSVLP 180
Query: 169 FKRHC-VEKDDEESRDGVEEAETMTEEE-----VLASSVP------TSLSLFPMGEKKMV 216
C V K + V E+ + EEE L S+P T LFP+ ++
Sbjct: 181 VSSSCHVFKPTARAGGVVIESSSPEEEEKDPMTCLRLSLPWVNESTTPPELFPVKREE-- 238
Query: 217 AVAEKEEKKMDMQEGLMRTMMQRMIAEEVRAY 248
E++E+++ G T++Q MI EVR+Y
Sbjct: 239 --EEEKEREISGLGGDFMTVVQEMIKTEVRSY 268
>UniRef100_Q9FDW1 Myb-related protein, 33.3K [Arabidopsis thaliana]
Length = 305
Score = 173 bits (439), Expect = 3e-42
Identities = 77/106 (72%), Positives = 90/106 (84%)
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
DR+KG WSP+EDE L +LV ++GPRNW+VIS IPGRSGKSCRLRWCNQLSP V+HRPF+
Sbjct: 3 DRIKGPWSPEEDEQLRRLVVKYGPRNWTVISKSIPGRSGKSCRLRWCNQLSPQVEHRPFS 62
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRG 122
ED I +AHA GNKWATI+RLL GRTDNA+KNHWNSTL+R+ G
Sbjct: 63 AEEDETIARAHAQFGNKWATIARLLNGRTDNAVKNHWNSTLKRKCG 108
>UniRef100_Q941B3 AT5g67300/K8K14_2 [Arabidopsis thaliana]
Length = 305
Score = 173 bits (439), Expect = 3e-42
Identities = 77/106 (72%), Positives = 90/106 (84%)
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
DR+KG WSP+EDE L +LV ++GPRNW+VIS IPGRSGKSCRLRWCNQLSP V+HRPF+
Sbjct: 3 DRIKGPWSPEEDEQLRRLVVKYGPRNWTVISKSIPGRSGKSCRLRWCNQLSPQVEHRPFS 62
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRG 122
ED I +AHA GNKWATI+RLL GRTDNA+KNHWNSTL+R+ G
Sbjct: 63 AEEDETIARAHAQFGNKWATIARLLNGRTDNAVKNHWNSTLKRKCG 108
>UniRef100_Q8LBC5 Myb-related protein, 33.3K [Arabidopsis thaliana]
Length = 305
Score = 173 bits (439), Expect = 3e-42
Identities = 77/106 (72%), Positives = 90/106 (84%)
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
DR+KG WSP+EDE L +LV ++GPRNW+VIS IPGRSGKSCRLRWCNQLSP V+HRPF+
Sbjct: 3 DRIKGPWSPEEDEQLRRLVVKYGPRNWTVISKSIPGRSGKSCRLRWCNQLSPQVEHRPFS 62
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRG 122
ED I +AHA GNKWATI+RLL GRTDNA+KNHWNSTL+R+ G
Sbjct: 63 AEEDETIARAHAQFGNKWATIARLLNGRTDNAVKNHWNSTLKRKCG 108
>UniRef100_Q39154 MYB-related protein [Arabidopsis thaliana]
Length = 305
Score = 173 bits (439), Expect = 3e-42
Identities = 77/106 (72%), Positives = 90/106 (84%)
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
DR+KG WSP+EDE L +LV ++GPRNW+VIS IPGRSGKSCRLRWCNQLSP V+HRPF+
Sbjct: 3 DRIKGPWSPEEDEQLRRLVVKYGPRNWTVISKSIPGRSGKSCRLRWCNQLSPQVEHRPFS 62
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRG 122
ED I +AHA GNKWATI+RLL GRTDNA+KNHWNSTL+R+ G
Sbjct: 63 AEEDETIARAHAQFGNKWATIARLLNGRTDNAVKNHWNSTLKRKCG 108
>UniRef100_Q6ZHS5 Putative tuber-specific and sucrose-responsive element binding
factor [Oryza sativa]
Length = 301
Score = 171 bits (434), Expect = 1e-41
Identities = 79/139 (56%), Positives = 97/139 (68%), Gaps = 3/139 (2%)
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
DR+KG WSP+EDE L +LV +HG RNW+ I GIPGRSGKSCRLRWCNQLSP V+ RPFT
Sbjct: 8 DRIKGPWSPEEDEALRRLVERHGARNWTAIGRGIPGRSGKSCRLRWCNQLSPQVERRPFT 67
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESL---PLPLASPS 133
ED I++AHA GN+WA I+RLLPGRTDNA+KNHWNS+L+R+ + +
Sbjct: 68 AEEDAAILRAHARLGNRWAAIARLLPGRTDNAVKNHWNSSLKRKLATATDGGEIDRPCKR 127
Query: 134 PSPSPSPPLPFSMAKRPHG 152
SP P P ++ HG
Sbjct: 128 VSPGPGSPTGSERSELSHG 146
>UniRef100_Q9SN12 R2R3-MYB transcription factor [Arabidopsis thaliana]
Length = 301
Score = 170 bits (431), Expect = 3e-41
Identities = 75/104 (72%), Positives = 88/104 (84%)
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
DRVKG WS +EDE L ++V ++GPRNWS IS IPGRSGKSCRLRWCNQLSP+V+HRPF+
Sbjct: 3 DRVKGPWSQEEDEQLRRMVEKYGPRNWSAISKSIPGRSGKSCRLRWCNQLSPEVEHRPFS 62
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
P ED I+ A A GNKWATI+RLL GRTDNA+KNHWNSTL+R+
Sbjct: 63 PEEDETIVTARAQFGNKWATIARLLNGRTDNAVKNHWNSTLKRK 106
>UniRef100_Q9FZ13 Tuber-specific and sucrose-responsive element binding factor
[Solanum tuberosum]
Length = 255
Score = 170 bits (431), Expect = 3e-41
Identities = 76/114 (66%), Positives = 93/114 (80%)
Query: 7 SSNGGGGGDRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQL 66
S +G +R+KG WS +ED+ L KLV ++G RNWS+IS I GRSGKSCRLRWCNQL
Sbjct: 22 SLSGKNPNKSERIKGPWSAEEDKILTKLVERYGARNWSLISKYIKGRSGKSCRLRWCNQL 81
Query: 67 SPDVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
SP+V+HRPF+P ED I+ AHA +GN+WATI+RLLPGRTDNA+KNHWNSTL+RR
Sbjct: 82 SPNVEHRPFSPAEDEAILAAHAKYGNRWATIARLLPGRTDNAVKNHWNSTLKRR 135
>UniRef100_Q39155 MYB-related protein [Arabidopsis thaliana]
Length = 304
Score = 170 bits (431), Expect = 3e-41
Identities = 75/104 (72%), Positives = 88/104 (84%)
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
DRVKG WS +EDE L ++V ++GPRNWS IS IPGRSGKSCRLRWCNQLSP+V+HRPF+
Sbjct: 3 DRVKGPWSQEEDEQLRRMVEKYGPRNWSAISKSIPGRSGKSCRLRWCNQLSPEVEHRPFS 62
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
P ED I+ A A GNKWATI+RLL GRTDNA+KNHWNSTL+R+
Sbjct: 63 PEEDETIVTARAQFGNKWATIARLLNGRTDNAVKNHWNSTLKRK 106
>UniRef100_O49745 R2R3-MYB transcription factor [Arabidopsis thaliana]
Length = 304
Score = 170 bits (431), Expect = 3e-41
Identities = 75/104 (72%), Positives = 88/104 (84%)
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
DRVKG WS +EDE L ++V ++GPRNWS IS IPGRSGKSCRLRWCNQLSP+V+HRPF+
Sbjct: 3 DRVKGPWSQEEDEQLRRMVEKYGPRNWSAISKSIPGRSGKSCRLRWCNQLSPEVEHRPFS 62
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
P ED I+ A A GNKWATI+RLL GRTDNA+KNHWNSTL+R+
Sbjct: 63 PEEDETIVTARAQFGNKWATIARLLNGRTDNAVKNHWNSTLKRK 106
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.315 0.133 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 491,129,708
Number of Sequences: 2790947
Number of extensions: 23059232
Number of successful extensions: 153892
Number of sequences better than 10.0: 1392
Number of HSP's better than 10.0 without gapping: 1220
Number of HSP's successfully gapped in prelim test: 182
Number of HSP's that attempted gapping in prelim test: 149736
Number of HSP's gapped (non-prelim): 2670
length of query: 260
length of database: 848,049,833
effective HSP length: 125
effective length of query: 135
effective length of database: 499,181,458
effective search space: 67389496830
effective search space used: 67389496830
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0002.6


