

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0001.6
(381 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
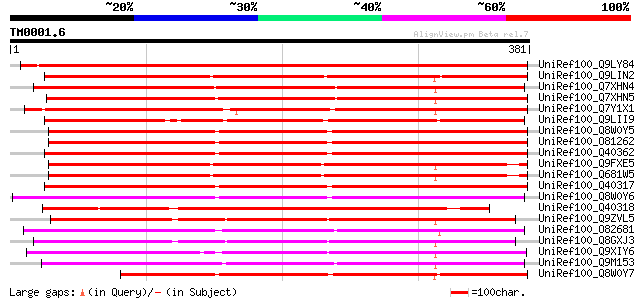
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LY84 Putative early nodule-specific protein [Arabido... 517 e-145
UniRef100_Q9LIN2 Nodulin-like protein protein [Arabidopsis thali... 351 2e-95
UniRef100_Q7XHN4 Putative early nodulin 8 [Oryza sativa] 348 1e-94
UniRef100_Q7XHN5 Putative early nodulin 8 [Oryza sativa] 339 9e-92
UniRef100_Q7Y1X1 Esterase precursor [Hevea brasiliensis] 336 6e-91
UniRef100_Q9LII9 Arabidopsis thaliana genomic DNA, chromosome 3,... 336 7e-91
UniRef100_Q8W0Y5 Enod8.1 [Medicago truncatula] 335 2e-90
UniRef100_O81262 Early nodule-specific protein [Medicago truncat... 335 2e-90
UniRef100_Q40362 Early nodulin [Medicago sativa] 332 1e-89
UniRef100_Q9FXE5 F12A21.4 [Arabidopsis thaliana] 330 3e-89
UniRef100_Q681W5 ENOD8-like protein [Arabidopsis thaliana] 330 3e-89
UniRef100_Q40317 Nodulin [Medicago sativa] 329 7e-89
UniRef100_Q8W0Y6 Enod8.2 [Medicago truncatula] 322 1e-86
UniRef100_Q40318 Coil protein [Medicago sativa] 313 4e-84
UniRef100_Q9ZVL5 T22H22.20 protein [Arabidopsis thaliana] 293 5e-78
UniRef100_O82681 Lanatoside 15'-O-acetylesterase precursor [Digi... 286 7e-76
UniRef100_Q8GXJ3 Hypothetical protein At1g54790/T22H22_20 [Arabi... 278 1e-73
UniRef100_Q9XIY6 Putative lanatoside 15'-O-acetylesterase [Oryza... 278 1e-73
UniRef100_Q9M153 Putative acetyltransferase [Arabidopsis thaliana] 276 9e-73
UniRef100_Q8W0Y7 Enod8.3 [Medicago truncatula] 274 3e-72
>UniRef100_Q9LY84 Putative early nodule-specific protein [Arabidopsis thaliana]
Length = 389
Score = 517 bits (1331), Expect = e-145
Identities = 245/372 (65%), Positives = 288/372 (76%), Gaps = 1/372 (0%)
Query: 9 AFFLSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSA 68
++ L C V + + P C FPAIYNFGDSNSDTGGISAAFEPI PYG+ F +P+
Sbjct: 18 SWLLLCLFAVTT-SVSVQPTCTFPAIYNFGDSNSDTGGISAAFEPIRDPYGQGFFHRPTG 76
Query: 69 RDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPF 128
RD DGRL +DFIAE+L LPYLSAYLNSLG+N+RHGANFATGGSTIRRQNETIFQYGISPF
Sbjct: 77 RDSDGRLTIDFIAERLGLPYLSAYLNSLGSNFRHGANFATGGSTIRRQNETIFQYGISPF 136
Query: 129 SLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRMMN 188
SLDMQ QF QFKAR+ L+ + K+ +R KLP EEF+KALYTFDIGQNDLSVGFR M+
Sbjct: 137 SLDMQIAQFDQFKARSALLFTQIKSRYDREKLPRQEEFAKALYTFDIGQNDLSVGFRTMS 196
Query: 189 FDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYG 248
DQ++ ++PDIVN LASAV+NIY+ GGRTFW+HNT P GCLPVN+FY GYLD G
Sbjct: 197 VDQLKATIPDIVNHLASAVRNIYQQGGRTFWVHNTGPFGCLPVNMFYMGTPAPGYLDKSG 256
Query: 249 CVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKIC 308
CVK QN MA+EFN++LK+ V+ LR EL +AAITYVD+Y AKY ++SN K GF +PLK+C
Sbjct: 257 CVKAQNEMAMEFNRKLKETVINLRKELTQAAITYVDVYTAKYEMMSNPKKLGFANPLKVC 316
Query: 309 CGYHVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFT 368
CGYH HIWCG G N +++G +C P M VSWDGVHY EAAN VA+R LNG T
Sbjct: 317 CGYHEKYDHIWCGNKGKVNNTEIYGGSCPNPVMAVSWDGVHYTEAANKHVADRTLNGLLT 376
Query: 369 DPPTLITQACYR 380
DPP IT+ACYR
Sbjct: 377 DPPVPITRACYR 388
>UniRef100_Q9LIN2 Nodulin-like protein protein [Arabidopsis thaliana]
Length = 380
Score = 351 bits (900), Expect = 2e-95
Identities = 179/361 (49%), Positives = 241/361 (66%), Gaps = 10/361 (2%)
Query: 26 SPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKLN 85
SP C FPAI+NFGDSNSDTGG+SA+F P P G++F PS R DGRLI+DFIAE+L
Sbjct: 24 SPSCNFPAIFNFGDSNSDTGGLSASFGQAPYPNGQTFFHSPSGRFSDGRLIIDFIAEELG 83
Query: 86 LPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKARTK 145
LPYL+A+L+S+G+N+ HGANFAT GST+R N TI Q G+SP SLD+Q VQF F R++
Sbjct: 84 LPYLNAFLDSIGSNFSHGANFATAGSTVRPPNATIAQSGVSPISLDVQLVQFSDFITRSQ 143
Query: 146 QLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRM-MNFDQMRESMPDIVNQLA 204
+ + + + LP E FS+ALYTFDIGQNDL+ G ++ M DQ++ +PD+ +QL+
Sbjct: 144 LI--RNRGGVFKKLLPKKEYFSQALYTFDIGQNDLTAGLKLNMTSDQIKAYIPDVHDQLS 201
Query: 205 SAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQL 264
+ ++ +Y GGR FWIHNTAP+GCLP + + +PA +D +GC +N +A +N +L
Sbjct: 202 NVIRKVYSKGGRRFWIHNTAPLGCLPY-VLDRFPVPASQIDNHGCAIPRNEIARYYNSEL 260
Query: 265 KDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCG----YHVNDTHIWC 320
K RV++LR EL EAA TYVD+Y+ K LI+ K GF PL CCG Y+ N I C
Sbjct: 261 KRRVIELRKELSEAAFTYVDIYSIKLTLITQAKKLGFRYPLVACCGHGGKYNFNKL-IKC 319
Query: 321 GTIGTANGKD-VFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQACY 379
G GK+ V +C S VSWDG+H+ E N W+ +I +G+F+DPP + AC
Sbjct: 320 GAKVMIKGKEIVLAKSCNDVSFRVSWDGIHFTETTNSWIFQQINDGAFSDPPLPVKSACT 379
Query: 380 R 380
R
Sbjct: 380 R 380
>UniRef100_Q7XHN4 Putative early nodulin 8 [Oryza sativa]
Length = 391
Score = 348 bits (894), Expect = 1e-94
Identities = 183/367 (49%), Positives = 241/367 (64%), Gaps = 8/367 (2%)
Query: 18 VNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIV 77
V+ V + C FPA++NFGDSNSDTGG+SA F PPP G +F P R CDGRL++
Sbjct: 21 VSQVSVAAGADCRFPAVFNFGDSNSDTGGLSATFGAAPPPNGRTFFGMPVGRYCDGRLVI 80
Query: 78 DFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQF 137
DFIAE L LPYLSAYLNS+G+N+ GANFAT GS+IRRQN ++F G SP SLD+Q +F
Sbjct: 81 DFIAESLGLPYLSAYLNSIGSNFTQGANFATAGSSIRRQNTSLFLSGFSPISLDVQSWEF 140
Query: 138 KQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRM-MNFDQMRESM 196
+QF R++ +Y K + R LP E FS+ALYTFDIGQND++ GF + M +Q+ +
Sbjct: 141 EQFINRSQFVYNN-KGGIYRELLPKAEYFSQALYTFDIGQNDITTGFFINMTSEQVIAYI 199
Query: 197 PDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVM 256
PD++ +L + ++N+Y LGGR FWIHNT PIGCLP + ++ +L A D GC N +
Sbjct: 200 PDLMERLTNIIQNVYGLGGRYFWIHNTGPIGCLPYAMVHRPDL-AVVKDGSGCSVAYNEV 258
Query: 257 AVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGY----H 312
A FN++LK+ V LR +AA TYVD+Y+AKY LIS+ K G DP+ CCGY +
Sbjct: 259 AQLFNQRLKETVGHLRKTHADAAFTYVDVYSAKYKLISDAKKLGMDDPMLTCCGYGGGRY 318
Query: 313 VNDTHIWCGTIGTANGK-DVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPP 371
D + CG NG V G +C+ P VSWDGVH+ EAAN +V ++I G +DPP
Sbjct: 319 NFDDRVGCGGKVKVNGTWVVAGKSCDDPLKRVSWDGVHFTEAANKFVFDQIAGGKLSDPP 378
Query: 372 TLITQAC 378
+ QAC
Sbjct: 379 VPLRQAC 385
>UniRef100_Q7XHN5 Putative early nodulin 8 [Oryza sativa]
Length = 405
Score = 339 bits (869), Expect = 9e-92
Identities = 177/359 (49%), Positives = 236/359 (65%), Gaps = 8/359 (2%)
Query: 28 PCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKLNLP 87
PC FPAI+NFGDSNSDTGG+SA +PPP+G ++ P+ R DGRL +DF+A+ L +
Sbjct: 44 PCDFPAIFNFGDSNSDTGGLSALIAVVPPPFGRTYFGMPAGRFSDGRLTIDFMAQSLGIR 103
Query: 88 YLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKARTKQL 147
YLSAYL+S+G+N+ GANFAT ++IR N +IF GISP SLD+Q QF+QF R++ +
Sbjct: 104 YLSAYLDSVGSNFSQGANFATAAASIRPANGSIFVSGISPISLDVQTSQFEQFINRSQFV 163
Query: 148 YQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVG-FRMMNFDQMRESMPDIVNQLASA 206
Y + R LP E FS+ALYTFDIGQNDL++G F M+ +Q+ +PD++ + ++A
Sbjct: 164 YSNI-GGIYREILPKAEYFSRALYTFDIGQNDLTMGYFDNMSTEQVEAYVPDLMERFSAA 222
Query: 207 VKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLKD 266
++ +Y LGGR FW+HNTAP+GCL + L A D GC N A FN +L++
Sbjct: 223 IQKVYSLGGRYFWVHNTAPLGCLTYAVVLLPKL-AAPRDDAGCSVAYNAAARFFNARLRE 281
Query: 267 RVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGY----HVNDTHIWCGT 322
V +LR LP+AA+TYVD+Y+AKY LIS K GF DPL +CCGY + D I CG
Sbjct: 282 TVDRLRAALPDAALTYVDVYSAKYRLISQAKQLGFGDPLLVCCGYGGGEYNFDRDIRCGG 341
Query: 323 IGTANGKDVF-GNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQACYR 380
NG V G +C+ PS VSWDGVH+ EAAN +V I+ G +DPP + QAC R
Sbjct: 342 KVEVNGTSVLAGKSCDDPSRSVSWDGVHFTEAANRFVFELIVGGKLSDPPVPLRQACRR 400
>UniRef100_Q7Y1X1 Esterase precursor [Hevea brasiliensis]
Length = 391
Score = 336 bits (862), Expect = 6e-91
Identities = 181/376 (48%), Positives = 239/376 (63%), Gaps = 15/376 (3%)
Query: 12 LSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDC 71
LS LC+ S+ S C FPAI+NFGDSNSDTGG +AAF P+ PPYGE+F + + R
Sbjct: 14 LSFLLCMLSLAYA-SETCDFPAIFNFGDSNSDTGGKAAAFYPLNPPYGETFFHRSTGRYS 72
Query: 72 DGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQY-GISPFSL 130
DGRLI+DFIAE NLPYLS YL+SLG+N++HGA+FAT GSTI+ I + G SPF L
Sbjct: 73 DGRLIIDFIAESFNLPYLSPYLSSLGSNFKHGADFATAGSTIKLPTTIIPAHGGFSPFYL 132
Query: 131 DMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEE--FSKALYTFDIGQNDLSVGFRMMN 188
D+Q+ QF+QF R++ + + E VPEE F KALYTFDIGQNDL+ GF +
Sbjct: 133 DVQYSQFRQFIPRSQFIRETGGIFAEL----VPEEYYFEKALYTFDIGQNDLTEGFLNLT 188
Query: 189 FDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYG 248
+++ ++PD+VN ++ VK IY+LG RTFWIHNT PIGCL L Y P D G
Sbjct: 189 VEEVNATVPDLVNSFSANVKKIYDLGARTFWIHNTGPIGCLSFILTY---FPWAEKDSAG 245
Query: 249 CVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKIC 308
C K N +A FN +LK+ V +LR +LP A +VD+Y+ KY L S + GF PL C
Sbjct: 246 CAKAYNEVAQHFNHKLKEIVAQLRKDLPLATFVHVDIYSVKYSLFSEPEKHGFEFPLITC 305
Query: 309 CGY---HVNDTHIWCG-TIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILN 364
CGY + CG T+ +G + +C PS+ V+WDG HY EAAN + ++I
Sbjct: 306 CGYGGKYNFSVTAPCGDTVTADDGTKIVVGSCACPSVRVNWDGAHYTEAANEYFFDQIST 365
Query: 365 GSFTDPPTLITQACYR 380
G+F+DPP + AC++
Sbjct: 366 GAFSDPPVPLNMACHK 381
>UniRef100_Q9LII9 Arabidopsis thaliana genomic DNA, chromosome 3, TAC clone:K24A2
[Arabidopsis thaliana]
Length = 371
Score = 336 bits (861), Expect = 7e-91
Identities = 174/353 (49%), Positives = 227/353 (64%), Gaps = 11/353 (3%)
Query: 26 SPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKLN 85
S C FPA++NFGDSNSDTG ISAA +PPP G +F + + R DGRLI+DFI E L
Sbjct: 25 SSSCNFPAVFNFGDSNSDTGAISAAIGEVPPPNGVAFFGRSAGRHSDGRLIIDFITENLT 84
Query: 86 LPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKARTK 145
LPYL+ YL+S+G NYRHGANFATGGS IR T+ + SPF L Q QF FK RT
Sbjct: 85 LPYLTPYLDSVGANYRHGANFATGGSCIR---PTLACF--SPFHLGTQVSQFIHFKTRTL 139
Query: 146 QLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRMMNFDQMRESMPDIVNQLAS 205
LY + R L FSKALYT DIGQNDL++GF+ M +Q++ ++P I+
Sbjct: 140 SLYNQTNGKFNR--LSHTNYFSKALYTLDIGQNDLAIGFQNMTEEQLKATIPLIIENFTI 197
Query: 206 AVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLK 265
A+K +Y+ G R F IHNT P GCLP + PA DPYGC+K N +A+EFNKQLK
Sbjct: 198 ALKLLYKEGARFFSIHNTGPTGCLP---YLLKAFPAIPRDPYGCLKPLNNVAIEFNKQLK 254
Query: 266 DRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGYHVNDTHIWCGTIGT 325
+++ +L+ ELP + TYVD+Y+AKY LI+ K GF+DP CC + + CG
Sbjct: 255 NKITQLKKELPSSFFTYVDVYSAKYNLITKAKALGFIDPFDYCCVGAIG-RGMGCGKTIF 313
Query: 326 ANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQAC 378
NG +++ ++C+ ++SWDG+HY E AN VANRIL+GS +DPP +AC
Sbjct: 314 LNGTELYSSSCQNRKNFISWDGIHYTETANMLVANRILDGSISDPPLPTQKAC 366
>UniRef100_Q8W0Y5 Enod8.1 [Medicago truncatula]
Length = 381
Score = 335 bits (858), Expect = 2e-90
Identities = 169/354 (47%), Positives = 224/354 (62%), Gaps = 7/354 (1%)
Query: 29 CAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKLNLPY 88
C FPAI++FG SN DTGG++AAF P PYGE++ + + R DGR+I+DFIA LPY
Sbjct: 32 CDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRSTGRFSDGRIILDFIARSFRLPY 91
Query: 89 LSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKARTKQLY 148
LS YLNSLG+N+ HGANFA+GGSTI + +SPFSL +Q++QFK+F ++TK +
Sbjct: 92 LSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPFSLQIQYIQFKEFISKTKLIR 151
Query: 149 QEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVG-FRMMNFDQMRESMPDIVNQLASAV 207
+ + + +P + FSKALY FDIGQNDL++G F Q+ ++PDIVN +
Sbjct: 152 DQG--GVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTIQQVNATVPDIVNNYIKNI 209
Query: 208 KNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDR 267
KNIY LG R+FWIH T P GC PV L N P+ D YGC K N ++ FN +LK+
Sbjct: 210 KNIYNLGARSFWIHGTGPKGCAPVIL---ANFPSAIKDSYGCAKQYNEVSQYFNFKLKEA 266
Query: 268 VVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGYHVN-DTHIWCGTIGTA 326
+ +LR+ L AAITYVD+Y KY L +N + GF P CCGY + + CG
Sbjct: 267 LAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCGYGGEYNIGVGCGASINI 326
Query: 327 NGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQACYR 380
NG + +C+ PS + WDGVHY EAAN V ++IL G F DPP + +ACYR
Sbjct: 327 NGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQILTGVFNDPPISLDRACYR 380
>UniRef100_O81262 Early nodule-specific protein [Medicago truncatula]
Length = 381
Score = 335 bits (858), Expect = 2e-90
Identities = 169/354 (47%), Positives = 224/354 (62%), Gaps = 7/354 (1%)
Query: 29 CAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKLNLPY 88
C FPAI++FG SN DTGG++AAF P PYGE++ + + R DGR+I+DFIA LPY
Sbjct: 32 CDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRSTGRFSDGRIILDFIARSFRLPY 91
Query: 89 LSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKARTKQLY 148
LS YLNSLG+N+ HGANFA+GGSTI + +SPFSL +Q++QFK+F ++TK +
Sbjct: 92 LSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPFSLQIQYIQFKEFISKTKLIR 151
Query: 149 QEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVG-FRMMNFDQMRESMPDIVNQLASAV 207
+ + + +P + FSKALY FDIGQNDL++G F Q+ ++PDIVN +
Sbjct: 152 DQG--GVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTIQQVNATVPDIVNNYIENI 209
Query: 208 KNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDR 267
KNIY LG R+FWIH T P GC PV L N P+ D YGC K N ++ FN +LK+
Sbjct: 210 KNIYNLGARSFWIHGTGPKGCAPVIL---ANFPSAIKDSYGCAKQYNEVSQYFNFKLKEA 266
Query: 268 VVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGYHVN-DTHIWCGTIGTA 326
+ +LR+ L AAITYVD+Y KY L +N + GF P CCGY + + CG
Sbjct: 267 LAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCGYGGEYNIGVGCGASINI 326
Query: 327 NGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQACYR 380
NG + +C+ PS + WDGVHY EAAN V ++IL G F DPP + +ACYR
Sbjct: 327 NGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQILTGVFNDPPISLDRACYR 380
>UniRef100_Q40362 Early nodulin [Medicago sativa]
Length = 381
Score = 332 bits (851), Expect = 1e-89
Identities = 167/357 (46%), Positives = 226/357 (62%), Gaps = 7/357 (1%)
Query: 26 SPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKLN 85
S C FPAI++FG SN DTGG++AAF P PYG+++ + + R DGR+I+DFIA+
Sbjct: 29 STHCDFPAIFSFGASNVDTGGLAAAFRAPPSPYGQTYFNRSTGRFSDGRIIIDFIAQSFR 88
Query: 86 LPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKARTK 145
LPY S YLNSLG+N+ HGANFAT GSTI + + +SPFSL +Q++QFK F ++TK
Sbjct: 89 LPYPSPYLNSLGSNFTHGANFATAGSTINIPTSILPKGILSPFSLQIQYIQFKDFISKTK 148
Query: 146 QLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVG-FRMMNFDQMRESMPDIVNQLA 204
+ + + + +P + FSKALY FDIGQNDL++G F Q+ ++PD+VN
Sbjct: 149 LIRDQG--GVFATLVPKEDYFSKALYVFDIGQNDLTIGFFGNKTIQQVNATVPDLVNNYI 206
Query: 205 SAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQL 264
+KNIY LG R+FWIH+T P GC PV L N P+ D YGC K N ++ FN +L
Sbjct: 207 ENIKNIYNLGARSFWIHSTGPKGCAPVIL---ANFPSAIKDSYGCAKQYNEVSQYFNLKL 263
Query: 265 KDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGYHVN-DTHIWCGTI 323
K+ + +LR++LP AAITYVD+Y+ KY L +N K GF P CCGY + CG
Sbjct: 264 KEALAQLRSDLPLAAITYVDIYSPKYSLFTNPKKYGFELPYVACCGYGGEYNIGAGCGAT 323
Query: 324 GTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQACYR 380
NG + +C+ PS ++WDG HY EAAN V ++I G+F DPP + ACYR
Sbjct: 324 INVNGTKIVAGSCKNPSTRITWDGTHYTEAANKIVFDQISTGAFNDPPISLDMACYR 380
>UniRef100_Q9FXE5 F12A21.4 [Arabidopsis thaliana]
Length = 372
Score = 330 bits (847), Expect = 3e-89
Identities = 173/357 (48%), Positives = 233/357 (64%), Gaps = 16/357 (4%)
Query: 29 CAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKLNLPY 88
C FPAI+NFGDSNSDTGG+SAAF PP+G SF P+ R CDGRL++DFIAE L LPY
Sbjct: 26 CHFPAIFNFGDSNSDTGGLSAAFGQAGPPHGSSFFGSPAGRYCDGRLVIDFIAESLGLPY 85
Query: 89 LSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKARTKQLY 148
LSA+L+S+G+N+ HGANFAT GS IR N T+ Q G SPFSLD+QFVQF F R++ +
Sbjct: 86 LSAFLDSVGSNFSHGANFATAGSPIRALNSTLRQSGFSPFSLDVQFVQFYNFHNRSQTV- 144
Query: 149 QEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVG-FRMMNFDQMRESMPDIVNQLASAV 207
++ + ++ LP + FSKALYTFDIGQNDL+ G F +Q+ +P+I++Q +A+
Sbjct: 145 -RSRGGVYKTMLPESDSFSKALYTFDIGQNDLTAGYFANKTVEQVETEVPEIISQFMNAI 203
Query: 208 KNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDR 267
KNIY GGR FWIHNT PIGCL + + A D +GCV N +A +FN LK
Sbjct: 204 KNIYGQGGRYFWIHNTGPIGCL-AYVIERFPNKASDFDSHGCVSPLNHLAQQFNHALKQA 262
Query: 268 VVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGY---HVNDTHIWCGTIG 324
V++LR+ L EAAITYVD+Y+ K+ L + + GF L CCG+ + + I CG
Sbjct: 263 VIELRSSLSEAAITYVDVYSLKHELFVHAQGHGFKGSLVSCCGHGGKYNYNKGIGCGMKK 322
Query: 325 TANGKDVF-GNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQACYR 380
GK+V+ G C++P V WDGVH+ +AAN ++ ++I G +++AC R
Sbjct: 323 IVKGKEVYIGKPCDEPDKAVVWDGVHFTQAANKFIFDKIAPG--------LSKACKR 371
>UniRef100_Q681W5 ENOD8-like protein [Arabidopsis thaliana]
Length = 364
Score = 330 bits (847), Expect = 3e-89
Identities = 173/357 (48%), Positives = 233/357 (64%), Gaps = 16/357 (4%)
Query: 29 CAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKLNLPY 88
C FPAI+NFGDSNSDTGG+SAAF PP+G SF P+ R CDGRL++DFIAE L LPY
Sbjct: 18 CHFPAIFNFGDSNSDTGGLSAAFGQAGPPHGSSFFGSPAGRYCDGRLVIDFIAESLGLPY 77
Query: 89 LSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKARTKQLY 148
LSA+L+S+G+N+ HGANFAT GS IR N T+ Q G SPFSLD+QFVQF F R++ +
Sbjct: 78 LSAFLDSVGSNFSHGANFATAGSPIRALNSTLRQSGFSPFSLDVQFVQFYNFHNRSQTV- 136
Query: 149 QEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVG-FRMMNFDQMRESMPDIVNQLASAV 207
++ + ++ LP + FSKALYTFDIGQNDL+ G F +Q+ +P+I++Q +A+
Sbjct: 137 -RSRGGVYKTMLPESDSFSKALYTFDIGQNDLTAGYFANKTVEQVETEVPEIISQFMNAI 195
Query: 208 KNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDR 267
KNIY GGR FWIHNT PIGCL + + A D +GCV N +A +FN LK
Sbjct: 196 KNIYGQGGRYFWIHNTGPIGCL-AYVIERFPNKASDFDSHGCVSPLNHLAQQFNHALKQA 254
Query: 268 VVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGY---HVNDTHIWCGTIG 324
V++LR+ L EAAITYVD+Y+ K+ L + + GF L CCG+ + + I CG
Sbjct: 255 VIELRSSLSEAAITYVDVYSLKHELFVHAQGHGFKGSLVSCCGHGGKYNYNKGIGCGMKK 314
Query: 325 TANGKDVF-GNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQACYR 380
GK+V+ G C++P V WDGVH+ +AAN ++ ++I G +++AC R
Sbjct: 315 IVKGKEVYIGKPCDEPDKAVVWDGVHFTQAANKFIFDKIAPG--------LSKACKR 363
>UniRef100_Q40317 Nodulin [Medicago sativa]
Length = 381
Score = 329 bits (844), Expect = 7e-89
Identities = 166/357 (46%), Positives = 224/357 (62%), Gaps = 7/357 (1%)
Query: 26 SPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKLN 85
S C FPAI++FG SN DTGG++AAF P PYG+++ + + R DGR+I+DFIA+
Sbjct: 29 STHCDFPAIFSFGASNVDTGGLAAAFRAPPSPYGQTYFNRSTGRFSDGRIIIDFIAQSFR 88
Query: 86 LPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKARTK 145
LPY S YLNSLG+N+ HGANFAT GSTI + + +SPFSL +Q++QFK F ++TK
Sbjct: 89 LPYPSPYLNSLGSNFTHGANFATAGSTINIPTSILPKGILSPFSLQIQYIQFKDFISKTK 148
Query: 146 QLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVG-FRMMNFDQMRESMPDIVNQLA 204
+ + + + +P + FSKALY FDIGQNDL++G F Q+ ++PD+VN
Sbjct: 149 LIRDQG--GVFATLVPKEDYFSKALYVFDIGQNDLTIGFFGNKTIQQVNATVPDLVNNYI 206
Query: 205 SAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQL 264
+KNIY LG R+FWIH+T P GC PV L N P+ D YGC K N ++ FN +L
Sbjct: 207 ENIKNIYNLGARSFWIHSTGPKGCAPVIL---ANFPSAIKDSYGCAKQYNEVSQYFNLKL 263
Query: 265 KDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGYHVN-DTHIWCGTI 323
K+ + +LR++LP AAITYVD+Y+ KY L +N K GF P CCGY + CG
Sbjct: 264 KEALAQLRSDLPLAAITYVDIYSPKYSLFTNPKKYGFELPYVACCGYGGEYNIGAGCGAT 323
Query: 324 GTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQACYR 380
NG + +C+ PS ++WDG HY E AN +V +I G F DPP + ACYR
Sbjct: 324 INVNGTKIVAGSCKNPSTRITWDGTHYTEEANKFVFYQISTGVFNDPPISLDMACYR 380
>UniRef100_Q8W0Y6 Enod8.2 [Medicago truncatula]
Length = 385
Score = 322 bits (824), Expect = 1e-86
Identities = 168/378 (44%), Positives = 228/378 (59%), Gaps = 7/378 (1%)
Query: 3 LRPLFIAFFLSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESF 62
LR + + + LC+ + + + C FPAI++FG SN DTGG++AAF+ P PYGE++
Sbjct: 7 LRHMSLVSLIVLILCIITPPIFATRNCDFPAIFSFGASNVDTGGLAAAFQAPPSPYGETY 66
Query: 63 PQKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQ 122
+ + R DGR+I+DFIA+ LPYLS YLNSLG+N+ HGANFATGGSTI N I
Sbjct: 67 FHRSTGRFSDGRIILDFIAQSFGLPYLSPYLNSLGSNFTHGANFATGGSTINIPNSIIPN 126
Query: 123 YGISPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSV 182
SPFSL +Q++QFK F ++T + + + + +P + FSKALYTFDIGQNDL
Sbjct: 127 GIFSPFSLQIQYIQFKDFISKTNLIRDQG--GVFATLIPKEDYFSKALYTFDIGQNDLIG 184
Query: 183 G-FRMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPA 241
G F Q+ ++PDIVN +KNIY LG R+FWIH+T P GC P L N P+
Sbjct: 185 GYFGNKTIKQVNATVPDIVNNFIVNIKNIYNLGARSFWIHSTVPSGCTPTIL---ANFPS 241
Query: 242 GYLDPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGF 301
D YGC K N ++ FN +LK + +LR +LP AAITYVD+Y+ Y L N K GF
Sbjct: 242 AIKDSYGCAKQYNEVSQYFNLKLKKALAQLRVDLPLAAITYVDIYSPNYSLFQNPKKYGF 301
Query: 302 VDPLKICCGYHVN-DTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVAN 360
P CCGY + + CG NG + +C+ PS + WDG H+ E V +
Sbjct: 302 ELPHVACCGYGGKYNIRVGCGETLNINGTKIEAGSCKNPSTRIIWDGSHFTERRYKIVFD 361
Query: 361 RILNGSFTDPPTLITQAC 378
+I G+F+DPP + +AC
Sbjct: 362 QISTGAFSDPPISLNRAC 379
>UniRef100_Q40318 Coil protein [Medicago sativa]
Length = 340
Score = 313 bits (803), Expect = 4e-84
Identities = 159/328 (48%), Positives = 211/328 (63%), Gaps = 16/328 (4%)
Query: 25 NSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKL 84
+S C +PAIYNFGDSNSDTG A F+ PP G SF S R DGRLI+D+I E+L
Sbjct: 25 SSHECVYPAIYNFGDSNSDTGTAYAIFKRNQPPNGISFGNI-SGRASDGRLIIDYITEEL 83
Query: 85 NLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKART 144
PYLSAYLNS+G+NYR+GANFA+GG++I + G SPF L +Q QF+QFK++T
Sbjct: 84 KAPYLSAYLNSVGSNYRYGANFASGGASICPGS------GWSPFDLGLQVTQFRQFKSQT 137
Query: 145 KQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRMMNFDQMRESMPDIVNQLA 204
+ L+ +S LP PE+FSKALYT DIG NDL+ GF + +Q++ S P+I+ +
Sbjct: 138 RILFNNETEPSLKSGLPRPEDFSKALYTIDIGLNDLASGFLRFSEEQVQRSFPEILGNFS 197
Query: 205 SAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQL 264
AVK +Y G R FWIHN P+GCLP+N + N G LD CV+ +N + E N +L
Sbjct: 198 QAVKQLYNEGARVFWIHNVGPVGCLPLNYYSNQNKKKGNLDANVCVESENKITQELNNKL 257
Query: 265 KDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGYHVNDTHIWCGTIG 324
KD+V +LR EL +A TYVD+Y AKY LISN K++GFV + CCG + D + CG
Sbjct: 258 KDQVSQLRKELVQAKFTYVDMYKAKYELISNAKSQGFVSLIDFCCGSYTGDYSVNCG--- 314
Query: 325 TANGKDVFGNACEKPSMYVSWDGVHYAE 352
+ N C PS ++SWDG+HY++
Sbjct: 315 ------MNTNLCTNPSQHISWDGIHYSK 336
>UniRef100_Q9ZVL5 T22H22.20 protein [Arabidopsis thaliana]
Length = 383
Score = 293 bits (750), Expect = 5e-78
Identities = 156/347 (44%), Positives = 212/347 (60%), Gaps = 12/347 (3%)
Query: 31 FPAIYNFGDSNSDTGG-ISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKLNLPYL 89
+PAI NFGDSNSDTG ISA E + PPYG+++ PS R CDGRLIVDF+ ++++LP+L
Sbjct: 30 YPAIINFGDSNSDTGNLISAGIENVNPPYGQTYFNLPSGRYCDGRLIVDFLLDEMDLPFL 89
Query: 90 SAYLNSLGT-NYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKARTKQLY 148
+ YL+SLG N++ G NFA GSTI N T +SPFS D+Q QF +FK+R +L
Sbjct: 90 NPYLDSLGLPNFKKGCNFAAAGSTILPANPT----SVSPFSFDLQISQFIRFKSRAIELL 145
Query: 149 QEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRMMNFDQMRESMPDIVNQLASAVK 208
+ E+ LP + +SK LY DIGQND++ F DQ+ S+P I+ + +K
Sbjct: 146 SKTGRKYEKY-LPPIDYYSKGLYMIDIGQNDIAGAFYSKTLDQVLASIPSILETFEAGLK 204
Query: 209 NIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDRV 268
+YE GGR WIHNT P+GCL N+ K + LD +GCV N A FN QL
Sbjct: 205 RLYEEGGRNIWIHNTGPLGCLAQNI-AKFGTDSTKLDEFGCVSSHNQAAKLFNLQLHAMS 263
Query: 269 VKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGY---HVN-DTHIWCGTIG 324
K + + P+A +TYVD+++ K LI+N GF PL CCG +N D+ I CG
Sbjct: 264 NKFQAQYPDANVTYVDIFSIKSNLIANYSRFGFEKPLMACCGVGGAPLNYDSRITCGQTK 323
Query: 325 TANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPP 371
+G V AC S Y++WDG+HY EAAN +V+++IL G ++DPP
Sbjct: 324 VLDGISVTAKACNDSSEYINWDGIHYTEAANEFVSSQILTGKYSDPP 370
>UniRef100_O82681 Lanatoside 15'-O-acetylesterase precursor [Digitalis lanata]
Length = 386
Score = 286 bits (732), Expect = 7e-76
Identities = 151/375 (40%), Positives = 224/375 (59%), Gaps = 13/375 (3%)
Query: 11 FLSCTLCVNSVELKNSPP-CAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSAR 69
F+ + + V +++S C F AI+NFGDSNSDTGG AAF PP G ++ ++P+ R
Sbjct: 12 FMVYVVVLMEVSVRSSESKCDFKAIFNFGDSNSDTGGFWAAFPAENPPNGMTYFKRPAGR 71
Query: 70 DCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFS 129
DGRLI+DF+A+ + +P+LS YL S+G+++RHGANFAT ST+ ++F G+SPFS
Sbjct: 72 VTDGRLIIDFLAQGIGIPFLSPYLLSIGSDFRHGANFATAASTVLLPRTSLFVTGVSPFS 131
Query: 130 LDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRMMNF 189
L +Q Q KQFK + +L+ + LP P+ F K+LYT IGQND + +
Sbjct: 132 LGIQLNQMKQFKLQVDRLHHSP----GKLNLPAPDIFRKSLYTLYIGQNDFTGNLGSLGI 187
Query: 190 DQMRES-MPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLF-YKHNLPAGYLDPY 247
+++ +P +V+Q++S +K +YELGGRTF + N APIGC P+ L HN + +D +
Sbjct: 188 SGVKKKIIPQVVSQISSTIKKLYELGGRTFLVLNLAPIGCYPLFLVDLPHN--SSDIDSF 245
Query: 248 GCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKI 307
GC+ N VE+N LK+ + + R ++ +A + Y D+++ L + + G K
Sbjct: 246 GCMISYNKAVVEYNYMLKEALAQTRKDIQDADVIYTDIHSVMLQLFQHPTSNGLKYGTKA 305
Query: 308 CCGYHVN----DTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRIL 363
CCGY + ++C NGK+V NAC+ P YVSWDG+H EAAN VA+ IL
Sbjct: 306 CCGYGGGAFNFNQQVFCSYSKLINGKNVTANACKDPQNYVSWDGIHATEAANKHVAHAIL 365
Query: 364 NGSFTDPPTLITQAC 378
GS DPP + + C
Sbjct: 366 EGSHFDPPFSLHKLC 380
>UniRef100_Q8GXJ3 Hypothetical protein At1g54790/T22H22_20 [Arabidopsis thaliana]
Length = 382
Score = 278 bits (712), Expect = 1e-73
Identities = 149/360 (41%), Positives = 215/360 (59%), Gaps = 12/360 (3%)
Query: 18 VNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFE-PIPPPYGESFPQKPSARDCDGRLI 76
++S+++ NS +P+ +NFGDSNSDTG + A + P G++ + S R CDGRL+
Sbjct: 16 ISSLQISNSIDFNYPSAFNFGDSNSDTGDLVAGLGIRLDLPNGQNSFKTSSQRFCDGRLV 75
Query: 77 VDFIAEKLNLPYLSAYLNSLGT-NYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFV 135
+DF+ ++++LP+L+ YL+SLG N++ G NFA GSTI N T +SPFS D+Q
Sbjct: 76 IDFLMDEMDLPFLNPYLDSLGLPNFKKGCNFAAAGSTILPANPT----SVSPFSFDLQIS 131
Query: 136 QFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRMMNFDQMRES 195
QF +FK+R +L + E+ LP + +SK LY DIGQND++ F DQ+ S
Sbjct: 132 QFIRFKSRAIELLSKTGRKYEKY-LPPIDYYSKGLYMIDIGQNDIAGAFYSKTLDQVLAS 190
Query: 196 MPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNV 255
+P I+ + +K +YE GGR WIHNT P+GCL N+ K + LD +GCV N
Sbjct: 191 IPSILETFEAGLKRLYEEGGRNIWIHNTGPLGCLAQNI-AKFGTDSTKLDEFGCVSSHNQ 249
Query: 256 MAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGY---H 312
A FN QL K + + P+A +TYVD+++ K LI+N GF PL CCG
Sbjct: 250 AAKLFNLQLHAMSNKFQAQYPDANVTYVDIFSIKSNLIANYSRFGFEKPLMACCGVGGAP 309
Query: 313 VN-DTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPP 371
+N D+ I CG +G V AC S Y++WDG+HY EAAN +V+++IL G ++DPP
Sbjct: 310 LNYDSRITCGQTKVLDGISVTAKACNDSSEYINWDGIHYTEAANEFVSSQILTGKYSDPP 369
>UniRef100_Q9XIY6 Putative lanatoside 15'-O-acetylesterase [Oryza sativa]
Length = 379
Score = 278 bits (712), Expect = 1e-73
Identities = 144/371 (38%), Positives = 216/371 (57%), Gaps = 11/371 (2%)
Query: 13 SCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCD 72
+ LC + C FPA++NFGDSNSDTGG AAF P+G ++ ++P+ R D
Sbjct: 11 AAALCCLVAAASATAQCRFPAVFNFGDSNSDTGGFWAAFPAQQAPFGMTYFRRPAGRASD 70
Query: 73 GRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDM 132
GRL+VDF+ + + LP LS YL S+G+ YRHGANFAT ST + N ++F GISPF L +
Sbjct: 71 GRLVVDFLVQAMGLPLLSPYLQSVGSGYRHGANFATLASTALQPNTSLFVTGISPFFLAV 130
Query: 133 QFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRMMNFDQM 192
Q Q K+ RTK L +LP P+ ALYT DIGQNDL+ + + +
Sbjct: 131 QLNQMKEL--RTKVLTSNG----NNDQLPAPDVLHNALYTIDIGQNDLTSNLGSQSIETV 184
Query: 193 RESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKD 252
++S+P +V++++S V+ +Y +G R + N APIGC P L K + +D YGC+K
Sbjct: 185 KQSLPSVVSKISSTVQELYNIGARNIMVFNMAPIGCYPAFL-TKLPHTSNDMDGYGCMKT 243
Query: 253 QNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGY- 311
N +N+ L + + ++R +L +A+I Y+D +A L + K G K CCGY
Sbjct: 244 YNSAVTYYNELLNNSLAEVRKKLQDASIVYLDKHAVTLELFRHPKAHGLKYGTKACCGYG 303
Query: 312 ---HVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFT 368
+ + ++CG+ NG+ V AC P YVSWDG+H EAAN +A+ +++GS++
Sbjct: 304 DGAYNFNPDVYCGSSKLLNGQTVTAKACADPQNYVSWDGIHATEAANKIIASSLMSGSYS 363
Query: 369 DPPTLITQACY 379
PP +++ C+
Sbjct: 364 YPPFDLSKLCH 374
>UniRef100_Q9M153 Putative acetyltransferase [Arabidopsis thaliana]
Length = 382
Score = 276 bits (705), Expect = 9e-73
Identities = 146/360 (40%), Positives = 205/360 (56%), Gaps = 9/360 (2%)
Query: 24 KNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEK 83
K C F AI+NFGDSNSDTGG AAF P+G ++ +KP+ R DGRLI+DF+A+
Sbjct: 25 KGDSKCDFEAIFNFGDSNSDTGGFWAAFPAQSGPWGMTYFKKPAGRASDGRLIIDFLAKS 84
Query: 84 LNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKAR 143
L +P+LS YL S+G+++RHGANFAT ST+ N ++F GISPFSL +Q Q KQFK
Sbjct: 85 LGMPFLSPYLQSIGSDFRHGANFATLASTVLLPNTSLFVSGISPFSLAIQLNQMKQFKVN 144
Query: 144 TKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRMMNFDQMRESMPDIVNQL 203
+ + + L+ LP F K+LYTF IGQND + + ++++ +P ++ Q+
Sbjct: 145 VDESHSLDRPGLK--ILPSKIVFGKSLYTFYIGQNDFTSNLASIGVERVKLYLPQVIGQI 202
Query: 204 ASAVKNIYELGGRTFWIHNTAPIGCLPVNLF-YKHNLPAGYLDPYGCVKDQNVMAVEFNK 262
A +K IY +GGRTF + N AP+GC P L Y H LD YGC+ N +N
Sbjct: 203 AGTIKEIYGIGGRTFLVLNLAPVGCYPAILTGYTHT--DADLDKYGCLIPVNKAVKYYNT 260
Query: 263 QLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGY----HVNDTHI 318
L + + RTEL A + Y+D + L + K+ G +K CCGY + + +
Sbjct: 261 LLNKTLSQTRTELKNATVIYLDTHKILLDLFQHPKSYGMKHGIKACCGYGGRPYNFNQKL 320
Query: 319 WCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQAC 378
+CG AC P YVSWDG+H EAANH ++ IL+GS + PP ++ C
Sbjct: 321 FCGNTKVIGNFSTTAKACHDPHNYVSWDGIHATEAANHHISMAILDGSISYPPFILNNLC 380
>UniRef100_Q8W0Y7 Enod8.3 [Medicago truncatula]
Length = 299
Score = 274 bits (701), Expect = 3e-72
Identities = 144/304 (47%), Positives = 187/304 (61%), Gaps = 11/304 (3%)
Query: 82 EKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFK 141
+ LPYLS YLNSLG+N+ HGANFAT GSTI+ N I SPFSL +Q +QFK F
Sbjct: 1 QSFGLPYLSPYLNSLGSNFTHGANFATAGSTIKIPNSIIPNGMFSPFSLQIQSIQFKDFI 60
Query: 142 ARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGF-RMMNFDQMRESMPDIV 200
+ K + + + + +P + +SKALYTFDIGQNDL+ GF Q+ ++PDIV
Sbjct: 61 PKAKFIRDQG--GVFATLIPKEDYYSKALYTFDIGQNDLTAGFFGNKTIQQVNTTVPDIV 118
Query: 201 NQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEF 260
+KNIY LG R+FWIHNT PIGC+P+ L N P+ D YGC K N ++ F
Sbjct: 119 KSFIDNIKNIYNLGARSFWIHNTGPIGCVPLILA---NFPSAIKDRYGCAKQYNEVSQYF 175
Query: 261 NKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCG----YHVNDT 316
N +LK+ + +LR +LP AAITYVD+Y+ KY L N K GF PL CCG Y+ N
Sbjct: 176 NLKLKEALAQLRKDLPLAAITYVDIYSPKYSLFQNPKKYGFELPLVACCGNGGKYNYN-I 234
Query: 317 HIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQ 376
CG NG + +C+KPS + WDG HY EAAN V ++I NG+FTDPP + +
Sbjct: 235 RAGCGATININGTNTVVGSCKKPSTRIIWDGTHYTEAANKIVFDQISNGAFTDPPIPLNR 294
Query: 377 ACYR 380
ACY+
Sbjct: 295 ACYK 298
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.138 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 688,547,359
Number of Sequences: 2790947
Number of extensions: 30141624
Number of successful extensions: 66759
Number of sequences better than 10.0: 323
Number of HSP's better than 10.0 without gapping: 222
Number of HSP's successfully gapped in prelim test: 101
Number of HSP's that attempted gapping in prelim test: 65524
Number of HSP's gapped (non-prelim): 367
length of query: 381
length of database: 848,049,833
effective HSP length: 129
effective length of query: 252
effective length of database: 488,017,670
effective search space: 122980452840
effective search space used: 122980452840
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0001.6


