

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0627.15
(557 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
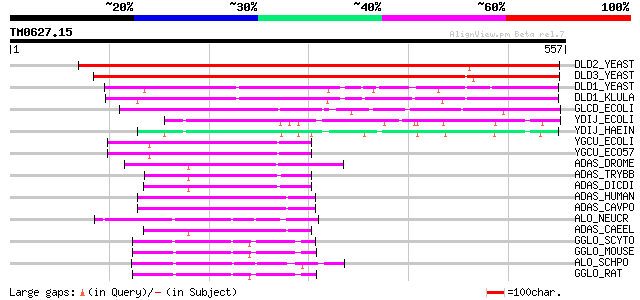
Score E
Sequences producing significant alignments: (bits) Value
DLD2_YEAST (P46681) D-lactate dehydrogenase [cytochrome] 2, mito... 511 e-144
DLD3_YEAST (P39976) D-lactate dehydrogenase [cytochrome] 3 (EC 1... 455 e-127
DLD1_YEAST (P32891) D-lactate dehydrogenase [cytochrome] 1, mito... 162 3e-39
DLD1_KLULA (Q12627) D-lactate dehydrogenase [cytochrome], mitoch... 150 6e-36
GLCD_ECOLI (P52075) Glycolate oxidase subunit glcD 142 3e-33
YDIJ_ECOLI (P77748) Hypothetical protein ydiJ 101 4e-21
YDIJ_HAEIN (Q57252) Protein HI1163 100 1e-20
YGCU_ECOLI (Q46911) Hypothetical flavoprotein ygcU 96 2e-19
YGCU_ECO57 (Q8X7S0) Hypothetical flavoprotein ygcU 96 2e-19
ADAS_DROME (Q9V778) Alkyldihydroxyacetonephosphate synthase (EC ... 82 3e-15
ADAS_TRYBB (O97157) Alkyldihydroxyacetonephosphate synthase (EC ... 74 1e-12
ADAS_DICDI (O96759) Alkyldihydroxyacetonephosphate synthase (EC ... 72 5e-12
ADAS_HUMAN (O00116) Alkyldihydroxyacetonephosphate synthase, per... 70 1e-11
ADAS_CAVPO (P97275) Alkyldihydroxyacetonephosphate synthase, per... 70 2e-11
ALO_NEUCR (Q7SGY1) Putative D-arabinono-1,4-lactone oxidase (EC ... 68 7e-11
ADAS_CAEEL (O45218) Alkyldihydroxyacetonephosphate synthase (EC ... 65 6e-10
GGLO_SCYTO (Q90YK3) L-gulonolactone oxidase (EC 1.1.3.8) (LGO) (... 62 3e-09
GGLO_MOUSE (P58710) L-gulonolactone oxidase (EC 1.1.3.8) (LGO) (... 62 5e-09
ALO_SCHPO (Q9HDX8) D-arabinono-1,4-lactone oxidase (EC 1.1.3.37)... 60 2e-08
GGLO_RAT (P10867) L-gulonolactone oxidase (EC 1.1.3.8) (LGO) (L-... 59 3e-08
>DLD2_YEAST (P46681) D-lactate dehydrogenase [cytochrome] 2,
mitochondrial precursor (EC 1.1.2.4) (D-lactate
ferricytochrome C oxidoreductase) (D-LCR) (Actin
interacting protein 2)
Length = 530
Score = 511 bits (1317), Expect = e-144
Identities = 257/494 (52%), Positives = 352/494 (71%), Gaps = 12/494 (2%)
Query: 70 STGIQHKCYASMPGLVQRNPRFSELNDDDVRYFQEILGQKNVVQ--DEEKLSTVNTDWMH 127
ST IQ + + V R+PRF +L DD+ YF+ IL ++ +++ + E LS N DWM
Sbjct: 37 STKIQTRLTSENYPDVHRDPRFKKLTSDDLNYFKSILSEQEILRASESEDLSFYNEDWMR 96
Query: 128 KYEGSSKLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSSMN 187
KY+G SKL+L+P + ++VS IL YCN +AVVPQGGNTGLVGGSVP+FDE+I+SL+++N
Sbjct: 97 KYKGQSKLVLRPKSVEKVSLILNYCNDEKIAVVPQGGNTGLVGGSVPIFDELILSLANLN 156
Query: 188 KVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLV 247
K+ FD VSGIL C+AG ILEN +++ + ++ PLDLGAKGSC +GG V+TNAGGLRL+
Sbjct: 157 KIRDFDPVSGILKCDAGVILENANNYVMEQNYMFPLDLGAKGSCHVGGVVATNAGGLRLL 216
Query: 248 RYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPP 307
RYGSLHGSVLG+E V+ +G +++ + ++RKDNTGYDLK LFIGSEG++GI+T V+ILT P
Sbjct: 217 RYGSLHGSVLGLEVVMPNGQIVNSMHSMRKDNTGYDLKQLFIGSEGTIGIITGVSILTVP 276
Query: 308 KLSSVNVALLACKDYNCCQKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPLPT 367
K + NV+ L+ + + QK+ A+++L EILSAFEF+D++S L + L+ A PL
Sbjct: 277 KPKAFNVSYLSVESFEDVQKVFVRARQELSEILSAFEFMDAKSQVLAKSQLKDAAFPLED 336
Query: 368 LHDFYVLIETTGSDESSDKQKLEAFLLHSMENELISDGVLAQDINQASSFWRLREGITEA 427
H FY+LIET+GS++ D KLE FL + ME +++DGV+AQD + + W+ RE I EA
Sbjct: 337 EHPFYILIETSGSNKDHDDSKLETFLENVMEEGIVTDGVVAQDETELQNLWKWREMIPEA 396
Query: 428 LMRVGAVYKYDLSIPLENLYNLVEEMRSRLGNA----------ANVVGYGHLGDGNLHLN 477
G VYKYD+S+PL++LY+LVE +RL A +GYGH+GDGNLHLN
Sbjct: 397 SQANGGVYKYDVSLPLKDLYSLVEATNARLSEAELVGDSPKPVVGAIGYGHVGDGNLHLN 456
Query: 478 ISVPQYDDKILSQIEPFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKN 537
++V +Y+ I +EPFVYE+ S +GS+SAEHGLG K N I YSKS E V++M +K
Sbjct: 457 VAVREYNKNIEKTLEPFVYEFVSSKHGSVSAEHGLGFQKKNYIGYSKSPEEVKMMKDLKV 516
Query: 538 LLDPNHILNPYKVL 551
DPN ILNPYK +
Sbjct: 517 HYDPNGILNPYKYI 530
>DLD3_YEAST (P39976) D-lactate dehydrogenase [cytochrome] 3 (EC
1.1.2.4) (D-lactate ferricytochrome C oxidoreductase)
(D-LCR)
Length = 496
Score = 455 bits (1171), Expect = e-127
Identities = 228/480 (47%), Positives = 325/480 (67%), Gaps = 14/480 (2%)
Query: 85 VQRNPRFSELNDDDVRYFQEILGQKNVVQDE--EKLSTVNTDWMHKYEGSSKLLLQPWNT 142
V+RNP F L+ +D+ YF+ IL ++ + E+L++ N DWM KY G S L+L P +T
Sbjct: 18 VKRNPNFKVLDSEDLAYFRSILSNDEILNSQAPEELASFNQDWMKKYRGQSNLILLPNST 77
Query: 143 DQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSSMNKVISFDKVSGILVCE 202
D+VS+I+KYCN + LAVVPQGGNT LVG SVPVFDE+++SL +MNKV FD VSG C+
Sbjct: 78 DKVSKIMKYCNDKKLAVVPQGGNTDLVGASVPVFDEIVLSLRNMNKVRDFDPVSGTFKCD 137
Query: 203 AGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEAV 262
AG ++ + FL + I PLDL ++ +CQ+GG VSTNAGGL +RYGSLHG+VLG+E V
Sbjct: 138 AGVVMRDAHQFLHDHDHIFPLDLPSRNNCQVGGVVSTNAGGLNFLRYGSLHGNVLGLEVV 197
Query: 263 LADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPPKLSSVNVALLACKDY 322
L +G ++ + LRKDNTGYDLK LFIG+EG++G+VT V+I+ K ++N +++
Sbjct: 198 LPNGEIISNINALRKDNTGYDLKQLFIGAEGTIGVVTGVSIVAAAKPKALNAVFFGIENF 257
Query: 323 NCCQKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPLPTLHDFYVLIETTGSDE 382
+ QKL +AK +L EILSAFEF+D S+ I +L+ PL H+FYVLIET+GS++
Sbjct: 258 DTVQKLFVKAKSELSEILSAFEFMDRGSIECTIEYLKDLPFPLENQHNFYVLIETSGSNK 317
Query: 383 SSDKQKLEAFLLHSMENELISDGVLAQDINQASSFWRLREGITEALMRVGAVYKYDLSIP 442
D +KL AFL + +++LIS+G++A+D W R+ + A G +YKYD+S+
Sbjct: 318 RHDDEKLTAFLKDTTDSKLISEGMMAKDKADFDRLWTWRKSVPTACNSYGGMYKYDMSLQ 377
Query: 443 LENLYNLVEEMRSRLGNAANVV-----------GYGHLGDGNLHLNISVPQYDDKILSQI 491
L++LY++ + RL NAA ++ GYGH+GDGN+HLNI+V ++ +I +
Sbjct: 378 LKDLYSVSAAVTERL-NAAGLIGDAPKPVVKSCGYGHVGDGNIHLNIAVREFTKQIEDLL 436
Query: 492 EPFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPNHILNPYKVL 551
EPFVYE+ + GSISAEHG+G K ++ Y++S ++ M IKN DPN ILNPYK +
Sbjct: 437 EPFVYEYIASKKGSISAEHGIGFHKKGKLHYTRSDIEIRFMKDIKNHYDPNGILNPYKYI 496
>DLD1_YEAST (P32891) D-lactate dehydrogenase [cytochrome] 1,
mitochondrial precursor (EC 1.1.2.4) (D-lactate
ferricytochrome C oxidoreductase) (D-LCR)
Length = 587
Score = 162 bits (409), Expect = 3e-39
Identities = 135/484 (27%), Positives = 232/484 (47%), Gaps = 46/484 (9%)
Query: 96 DDDVRYFQEILGQK--NVVQDEEKLSTVNTDWMHKYEGSS----KLLLQPWNTDQVSQIL 149
D V +++LG K N + L + + + + S +++L P T++VS+IL
Sbjct: 108 DKVVEDLKQVLGNKPENYSDAKSDLDAHSDTYFNTHHPSPEQRPRIILFPHTTEEVSKIL 167
Query: 150 KYCNSRSLAVVPQGGNTGLVGGSVP--VFDEVIVSLSS-MNKVISFDKVSGILVCEAGCI 206
K C+ ++ VVP G T L G +P + D + V LS MN V+ FDK+ + +AG
Sbjct: 168 KICHDNNMPVVPFSGGTSLEGHFLPTRIGDTITVDLSKFMNNVVKFDKLDLDITVQAGLP 227
Query: 207 LENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEAVLADG 266
E++ +L + G + D G QIGG ++ + G RYG++ +++ + VL DG
Sbjct: 228 WEDLNDYLSDHGLMFGCDPGP--GAQIGGCIANSCSGTNAYRYGTMKENIINMTIVLPDG 285
Query: 267 TVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPPKLSSVNVALLA---CKDYN 323
T++ K RK + GY+L LF+GSEG+LGIVT+ + K + VA+++ KD
Sbjct: 286 TIVKTKKRPRKSSAGYNLNGLFVGSEGTLGIVTEATVKCHVKPKAETVAVVSFDTIKDAA 345
Query: 324 CCQKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLEGARN----PLPTLHDFYVLIETTG 379
C L ++ G L+A E LD M L IN E PT+ F+ + +
Sbjct: 346 ACASNLTQS----GIHLNAMELLDENMMKL-INASESTDRCDWVEKPTM--FFKIGGRSP 398
Query: 380 SDESS--DKQKLEAFLLHSMENELISDGVLAQDINQASSFWRLREGITEALM-------R 430
+ ++ D+ K A L H + A+D ++ W R+ +++ +
Sbjct: 399 NIVNALVDEVKAVAQLNHCNSFQF------AKDDDEKLELWEARKVALWSVLDADKSKDK 452
Query: 431 VGAVYKYDLSIPLENLYNLVEEMRSRLGNAANVVG--YGHLGDGNLHLNI--SVPQYDDK 486
++ D+++P+ ++ E + + A+ ++ GH GDGN H I P+ + +
Sbjct: 453 SAKIWTTDVAVPVSQFDKVIHETKKDM-QASKLINAIVGHAGDGNFHAFIVYRTPE-EHE 510
Query: 487 ILSQIEPFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPNHILN 546
SQ+ + + G+ + EHG+G+ K +L V LM IK +DP I+N
Sbjct: 511 TCSQLVDRMVKRALNAEGTCTGEHGVGIGKREYLLEELGEAPVDLMRKIKLAIDPKRIMN 570
Query: 547 PYKV 550
P K+
Sbjct: 571 PDKI 574
>DLD1_KLULA (Q12627) D-lactate dehydrogenase [cytochrome],
mitochondrial precursor (EC 1.1.2.4) (D-lactate
ferricytochrome C oxidoreductase) (D-LCR)
Length = 576
Score = 150 bits (380), Expect = 6e-36
Identities = 125/482 (25%), Positives = 224/482 (45%), Gaps = 39/482 (8%)
Query: 97 DDVRYFQEILGQKNVVQDEEKLSTVNTDWM----------HKYEGSSK--LLLQPWNTDQ 144
D + + ++ K+V++++ + TV D + H E + + ++L P NT+
Sbjct: 96 DKETFAKALVELKDVLENDPENFTVAKDDLDAHSDTYFNSHHAEANQRPEIVLYPRNTED 155
Query: 145 VSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDE--VIVSLSS-MNKVISFDKVSGILVC 201
VS++LK C+ S+ V+P G T L G +P V++ +S +NK+I +K +V
Sbjct: 156 VSKLLKICHKYSIPVIPFSGGTSLEGHFLPTRPGSCVVLDISKYLNKIIQLNKEDLDVVV 215
Query: 202 EAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEA 261
+ G E + +L++ G + D G QI G ++ + G RYG++ +V+ +
Sbjct: 216 QGGVPWEELNEYLNDHGLLFGCDPGP--GAQIAGCIANSCSGTNAYRYGTMKENVVNITM 273
Query: 262 VLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPPKLSSVNVALL---A 318
+ADGT++ + RK + GY+L L IGSEG+LGIVT+ I + + VA++
Sbjct: 274 CMADGTIVKTKRRPRKSSAGYNLNGLIIGSEGTLGIVTEATIKCHVRSTFETVAVVPFPT 333
Query: 319 CKDYNCCQKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPLPTLHDFYVLIETT 378
D C L +A G L+A E LD M ++ + GA + + + +
Sbjct: 334 VSDAASCSSHLIQA----GIQLNAMELLDDNMMKII--NQSGATSKDNWVESPTLFFKIG 387
Query: 379 GSDESSDKQKL-EAFLLHSMENELISDGVLAQDINQASSFWRLREGITEALMRVGA---- 433
G E ++ + E + S N + A D + W R+ + + G
Sbjct: 388 GRSEQIIQEVIKEVEKIASQHNNTKFE--FATDEDSKLELWEARKVALWSTIDTGRKTNP 445
Query: 434 ---VYKYDLSIPLENLYNLVEEMRSRLGNAANVVG--YGHLGDGNLHLNISVPQYDDKIL 488
++ D+++P+ +++ + + NA+ ++ GH GDGN H I K
Sbjct: 446 DANIWTTDVAVPISKFADVINATKEEM-NASGLLTSLVGHAGDGNFHAFIIYNTEQRKTA 504
Query: 489 SQIEPFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPNHILNPY 548
I + + G+ + EHG+G+ K + +L +TV +M +K LDP ILNP
Sbjct: 505 ETIVENMVKRAIDAEGTCTGEHGVGIGKRDYLLEEVGEDTVAVMRKLKLALDPKRILNPD 564
Query: 549 KV 550
K+
Sbjct: 565 KI 566
>GLCD_ECOLI (P52075) Glycolate oxidase subunit glcD
Length = 499
Score = 142 bits (357), Expect = 3e-33
Identities = 124/456 (27%), Positives = 211/456 (46%), Gaps = 30/456 (6%)
Query: 111 VVQDEEKLSTVNTDWMHKYEGSSKLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVG 170
++ +E++ D + Y L++ P +QV+ IL C+ + VV +G TGL G
Sbjct: 34 ILHTDEEIIPYECDGLSAYRTRPLLVVLPKQMEQVTAILAVCHRLRVPVVTRGAGTGLSG 93
Query: 171 GSVPVFDEVIVSLSSMNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGS 230
G++P+ V++ ++ +++ + V + G I + D ++ +
Sbjct: 94 GALPLEKGVLLVMARFKEILDINPVGRRARVQPGVRNLAISQAVAPHNLYYAPDPSSQIA 153
Query: 231 CQIGGNVSTNAGGLRLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIG 290
C IGGNV+ NAGG+ ++YG ++L +E DG L L + D+ G+DL LF G
Sbjct: 154 CSIGGNVAENAGGVHCLKYGLTVHNLLKIEVQTLDGEAL-TLGSDALDSPGFDLLALFTG 212
Query: 291 SEGSLGIVTKVAILTPPKLSSVNVALLACKDYNCCQKLLQEAKRKLGEILS------AFE 344
SEG LG+ T+V + PK V LLA D +++A +G+I++ E
Sbjct: 213 SEGMLGVTTEVTVKLLPKPPVARV-LLASFD------SVEKAGLAVGDIIANGIIPGGLE 265
Query: 345 FLDSQSMNLVINHLEGARNPLPTLHDFYVLIETTG--SDESSDKQKLEAFLLHSMENELI 402
+D+ S+ + + P + +L E G SD D +++ LL +
Sbjct: 266 MMDNLSIRAAEDFIHAG---YPVDAEAILLCELDGVESDVQEDCERVNDILLKAG----A 318
Query: 403 SDGVLAQDINQASSFWRLREGITEALMRVGA-VYKYDLSIPLENLYNLVEEMRSRLGNA- 460
+D LAQD + FW R+ A+ R+ Y D +IP L ++E + +RL
Sbjct: 319 TDVRLAQDEAERVRFWAGRKNAFPAVGRISPDYYCMDGTIPRRALPGVLEGI-ARLSQQY 377
Query: 461 -ANVVGYGHLGDGNLHLNISVPQYDDKILSQIEPF---VYEWTSKHNGSISAEHGLGLMK 516
V H GDGN+H I + ++ E + E + GSIS EHG+G K
Sbjct: 378 DLRVANVFHAGDGNMHPLILFDANEPGEFARAEELGGKILELCVEVGGSISGEHGIGREK 437
Query: 517 ANEILYSKSLETVQLMASIKNLLDPNHILNPYKVLP 552
N++ + + + ++K DP+ +LNP K +P
Sbjct: 438 INQMCAQFNSDEITTFHAVKAAFDPDGLLNPGKNIP 473
>YDIJ_ECOLI (P77748) Hypothetical protein ydiJ
Length = 1018
Score = 101 bits (252), Expect = 4e-21
Identities = 122/477 (25%), Positives = 200/477 (41%), Gaps = 93/477 (19%)
Query: 156 SLAVVPQGGNTGLVGGSVPVFDEVIVSLSS-MNKVISFDKVSGILVCEAGCILENIISFL 214
SL P+GG TG G ++ +IV +S MN++I + G + EAG I + + +L
Sbjct: 78 SLIFTPRGGGTGTNGQALN--QGIIVDMSRHMNRIIEINPEEGWVRVEAGVIKDQLNQYL 135
Query: 215 DNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEAVLADGTVLD---- 270
G+ +L +GG ++T+A G + YG VLGV AVL G +LD
Sbjct: 136 KPFGYFFAPELSTSNRATLGGMINTDASGQGSLVYGKTSDHVLGVRAVLLGGDILDTQPL 195
Query: 271 ---MLKTLRKDN--------------------------------TGYDLKHLF------- 288
+ +TL K N TGYDL+H+F
Sbjct: 196 PVELAETLGKSNTTIGRIYNTVYQRCRQQRQLIIDNFPKLNRFLTGYDLRHVFNDEMTEF 255
Query: 289 ------IGSEGSLGIVT--KVAILTPPKLSSVNVALLACKDYNCCQKLLQEAKRKLGEIL 340
GSEG+L +T ++ I PK V L Y+ L+ A +
Sbjct: 256 DLTRILTGSEGTLAFITEARLDITRLPK-----VRRLVNVKYDSFDSALRNAPFMVEARA 310
Query: 341 SAFEFLDSQSMNLVINHL--EGARNPLPTLHDFYVL----IETTGSDESSDKQKLEAFLL 394
+ E +DS+ +NL + + + D +L +E G DE+ +++ A L
Sbjct: 311 LSVETVDSKVLNLAREDIVWHSVSELITDVPDQEMLGLNIVEFAGDDEALIDERVNA--L 368
Query: 395 HSMENELISD---GVL----AQDINQASSFWRLREGITEALMRV-GAV----YKYDLSIP 442
+ +ELI+ GV+ +++ + +R+ L GA + D +P
Sbjct: 369 CARLDELIASHQAGVIGWQVCRELAGVERIYAMRKKAVGLLGNAKGAAKPIPFAEDTCVP 428
Query: 443 LENLYNLVEEMRSRLGNAANVVG-YGHLGDGNLHLNISVPQYDDK---ILSQIEPFVYEW 498
E+L + + E R+ L + G +GH+ G LH+ ++ D + ++ QI V
Sbjct: 429 PEHLADYIAEFRALLDSHGLSYGMFGHVDAGVLHVRPALDMCDPQQEILMKQISDDVVAL 488
Query: 499 TSKHNGSISAEHGLGLMKANEILYSKSLETVQLMA---SIKNLLDPNHILNPYKVLP 552
T+K+ G + EHG G YS + +L A +K DP++ LNP K+ P
Sbjct: 489 TAKYGGLLWGEHGKGFRAE----YSPAFFGEELFAELRKVKAAFDPHNRLNPGKICP 541
>YDIJ_HAEIN (Q57252) Protein HI1163
Length = 1027
Score = 100 bits (249), Expect = 1e-20
Identities = 123/511 (24%), Positives = 203/511 (39%), Gaps = 105/511 (20%)
Query: 129 YEGSSKLLLQPWNTDQVSQILKYCNS---RSLAVVPQGGNTGLVGGSVPVFDEVIVSLSS 185
Y+ + +L P + +I K N +S++ P+GG TG G S+ + +IV LS
Sbjct: 48 YQQLPQAILFPKTVADIVRITKLANLPEYQSISFTPRGGGTGTNGQSIN--NNIIVDLSR 105
Query: 186 -MNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGL 244
M ++ + + +AG + + + FL G +L +GG ++T+A G
Sbjct: 106 HMTAILELNVKERWVRVQAGVVKDQLNQFLKPHGLFFAPELSTSNRATLGGMINTDASGQ 165
Query: 245 RLVRYGSLHGSVLGVEAVLADGTVLDM--------------------------------- 271
++YG VL + AVL +G +LD
Sbjct: 166 GSLQYGKTSNHVLALRAVLINGEILDTSAVNSVDVLENIDALELSESSKKLHQTIAQHCK 225
Query: 272 ---------LKTLRKDNTGYDLKHLF-------------IGSEGSLGIVTKV---AILTP 306
L L + TGYDLK++F GSEGSL + + +L P
Sbjct: 226 EKRAAIIKDLPQLNRFLTGYDLKNVFNEDESEFNLTRILTGSEGSLAFICEAKLNLLLIP 285
Query: 307 PKLSSVNVALLACKDYNCCQKLLQEAKRKLGEILSAFEFLDSQSMNLV--------INHL 358
+ +N+ Y L+ A + + E +DS+ +NL +N L
Sbjct: 286 QYRTLINI------KYRSFDAALRNAPFMVKANALSVETVDSKVLNLAKQDIIWHSVNEL 339
Query: 359 --EGARNPLPTLHDFYVLIETTGSD-ESSDKQKLEAFLLHSMENELISDGVL----AQDI 411
E ++P+ L+ ++E G++ E D+Q L + E D ++ D+
Sbjct: 340 LTEDEKDPILGLN----IVEFAGNNKEKIDRQVTALCRLLDEKIEHNQDHIIGYQVCSDL 395
Query: 412 NQASSFWRLREGITEALMRVGAVYK-----YDLSIPLENLYNLVEEMRSRLGNAANVVG- 465
+ +R+ L K D +P ENL + + E R+ L G
Sbjct: 396 PSIERIYAMRKKAVGLLGNAKGAAKPIPFVEDTCVPPENLADYISEFRALLDQHNLQYGM 455
Query: 466 YGHLGDGNLHLNISVPQYDD---KILSQIEPFVYEWTSKHNGSISAEHGLGLMKANEILY 522
+GH+ G LH+ ++ D K+ QI V E T K+ G + EHG G+ Y
Sbjct: 456 FGHVDAGVLHVRPALDLCDKEQVKLFKQISDEVAELTIKYGGLLWGEHGKGVRSH----Y 511
Query: 523 SKSLETVQL---MASIKNLLDPNHILNPYKV 550
+ T +L + IK L DPN+ LNP K+
Sbjct: 512 GEKFFTPELWHELRYIKTLFDPNNRLNPGKI 542
>YGCU_ECOLI (Q46911) Hypothetical flavoprotein ygcU
Length = 484
Score = 96.3 bits (238), Expect = 2e-19
Identities = 65/214 (30%), Positives = 112/214 (51%), Gaps = 10/214 (4%)
Query: 99 VRYFQEILGQKNVVQDEEKLSTVNTDWMHKYEGSSKLLLQP--------WNTDQVSQILK 150
V +EI+G V+ DE L + D K+ + P +T+QVS++L
Sbjct: 9 VDQLKEIVGADRVITDETVLKKNSIDRFRKFPDIHGIYTLPIPAAVVKLGSTEQVSRVLN 68
Query: 151 YCNSRSLAVVPQGGNTGLVGGSVPVFDE-VIVSLSSMNKVISFDKVSGILVCEAGCILEN 209
+ N+ + VP+ G + GG V + V++ S+MN++I+ D + + G LE
Sbjct: 69 FMNAHKINGVPRTGASATEGGLETVVENSVVLDGSAMNQIINIDIENMQATAQCGVPLEV 128
Query: 210 IISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEAVLADGTVL 269
+ + L +G+ +K Q+GG V+T + G YG++ V+G+EAVLADGTV
Sbjct: 129 LENALREKGYTTGHSPQSKPLAQMGGLVATRSIGQFSTLYGAIEDMVVGLEAVLADGTV- 187
Query: 270 DMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAI 303
+K + + G D++H+ IG+EG+L +T+V +
Sbjct: 188 TRIKNVPRRAAGPDIRHIIIGNEGALCYITEVTV 221
>YGCU_ECO57 (Q8X7S0) Hypothetical flavoprotein ygcU
Length = 484
Score = 96.3 bits (238), Expect = 2e-19
Identities = 65/214 (30%), Positives = 112/214 (51%), Gaps = 10/214 (4%)
Query: 99 VRYFQEILGQKNVVQDEEKLSTVNTDWMHKYEGSSKLLLQP--------WNTDQVSQILK 150
V +EI+G V+ DE L + D K+ + P +T+QVS++L
Sbjct: 9 VDQLKEIVGADRVITDETVLKKNSIDRFRKFPDIHGIYTLPIPAAVVKLGSTEQVSRVLN 68
Query: 151 YCNSRSLAVVPQGGNTGLVGGSVPVFDE-VIVSLSSMNKVISFDKVSGILVCEAGCILEN 209
+ N+ + VP+ G + GG V + V++ S+MN++I+ D + + G LE
Sbjct: 69 FMNAHKINGVPRTGASATEGGLETVVENSVVLDGSAMNQIINIDIENMQATAQCGVPLEV 128
Query: 210 IISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEAVLADGTVL 269
+ + L +G+ +K Q+GG V+T + G YG++ V+G+EAVLADGTV
Sbjct: 129 LENALREKGYTTGHSPQSKPLAQMGGLVATRSIGQFSTLYGAIEDMVVGLEAVLADGTV- 187
Query: 270 DMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAI 303
+K + + G D++H+ IG+EG+L +T+V +
Sbjct: 188 TRIKNVPRRAAGPDIRHIIIGNEGALCYITEVTV 221
>ADAS_DROME (Q9V778) Alkyldihydroxyacetonephosphate synthase (EC
2.5.1.26) (Alkyl-DHAP synthase)
(Alkylglycerone-phosphate synthase)
Length = 631
Score = 82.4 bits (202), Expect = 3e-15
Identities = 56/224 (25%), Positives = 112/224 (50%), Gaps = 5/224 (2%)
Query: 116 EKLSTVNTDWMHKYEGSSKLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPV 175
+ L+ + + W HK+ L++ P D+V Q+++ N ++ +VP GG T + G
Sbjct: 142 QTLNDIYSLWHHKFRRIPDLVVWPRCHDEVVQLVRLANKHNVMLVPFGGGTSVSGAITCP 201
Query: 176 FDE----VIVSLSSMNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSC 231
+E + S MN+++ ++ + + E+G + +++ L +EG + + +
Sbjct: 202 QNESRMICALDTSQMNRLLWLNRENLTVCFESGIVGQDLERVLRSEGLTVGHEPDSYEFS 261
Query: 232 QIGGNVSTNAGGLRLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGS 291
+GG V+T A G++ YG++ V+ V V GT L+ + + + G D H+ +GS
Sbjct: 262 TLGGWVATRASGMKKNVYGNIEDLVVRVRMVTPSGT-LERECSAPRVSCGPDFNHVILGS 320
Query: 292 EGSLGIVTKVAILTPPKLSSVNVALLACKDYNCCQKLLQEAKRK 335
EG+LG++T+V + P S LA ++ ++E R+
Sbjct: 321 EGTLGVITEVVLKVRPLPSLRRYGSLAFPNFEQGVLFMREVARR 364
>ADAS_TRYBB (O97157) Alkyldihydroxyacetonephosphate synthase (EC
2.5.1.26) (Alkyl-DHAP synthase)
(Alkylglycerone-phosphate synthase)
Length = 613
Score = 73.9 bits (180), Expect = 1e-12
Identities = 48/173 (27%), Positives = 79/173 (44%), Gaps = 8/173 (4%)
Query: 136 LLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDE-----VIVSLSSMNKVI 190
++ P N D +I++ ++ VVP GG T + GG P E + + + M +++
Sbjct: 133 VILPNNHDDCVKIMELAQKHNVVVVPFGGGTNVTGGVEPNPFETRRMVISIDMRRMGRML 192
Query: 191 SFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYG 250
D SG V E G + +I L GF+M D + +GG ++ G +YG
Sbjct: 193 HIDTESGTAVFEVGVLGPDIDEQLSRYGFMMGHDPDSYAYSTLGGWIAARGSGAMSNKYG 252
Query: 251 SLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAI 303
+ +L + V G V L G DL +F+GSEG+ G+VT+ +
Sbjct: 253 DIENMILAMRVVTPVGVV---ETPLTSRPCGVDLNAMFVGSEGAFGLVTEAVV 302
Score = 36.2 bits (82), Expect = 0.23
Identities = 20/69 (28%), Positives = 32/69 (45%)
Query: 484 DDKILSQIEPFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPNH 543
D KI Q++ E +H G+++ HG+G + + + K LDP +
Sbjct: 510 DLKIFLQVKKRAMEVMLQHRGNLTHHHGIGYEHVPWMKRYNGEGGLDAIMKFKKALDPKN 569
Query: 544 ILNPYKVLP 552
I NP K+LP
Sbjct: 570 ICNPGKLLP 578
>ADAS_DICDI (O96759) Alkyldihydroxyacetonephosphate synthase (EC
2.5.1.26) (Alkyl-DHAP synthase)
(Alkylglycerone-phosphate synthase)
Length = 611
Score = 71.6 bits (174), Expect = 5e-12
Identities = 42/172 (24%), Positives = 87/172 (50%), Gaps = 5/172 (2%)
Query: 135 LLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDE---VIVSLSSMNKVIS 191
L++ P + ++V ++++ + ++ ++P GG + +VG PV +E V + + MNKV+
Sbjct: 143 LIVLPHSHEEVERLVQLAHKYNVVIIPMGGGSNIVGAIEPVSNERFTVSIDMRRMNKVLW 202
Query: 192 FDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGS 251
D+ + G + + L +G + D + +GG ++T + G + +YG
Sbjct: 203 VDRREMTACIQVGIMGPELEKQLHKQGVSLGHDPDSFEFSTLGGWLATCSSGHQSDKYGD 262
Query: 252 LHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAI 303
+ + V GT+ L+ + G + KH+ +GSEG+LGI+T+ +
Sbjct: 263 IEDMAVSFRTVTPTGTL--ELRNGARSGAGINYKHIILGSEGTLGIITEAVM 312
Score = 34.7 bits (78), Expect = 0.67
Identities = 16/52 (30%), Positives = 27/52 (51%)
Query: 501 KHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPNHILNPYKVLP 552
K+ GS+S HG+G + + + + S+K +DP I NP K++P
Sbjct: 537 KYGGSLSHHHGVGYEHVPWMTRYATRGWINVYRSLKETIDPKDICNPRKLIP 588
>ADAS_HUMAN (O00116) Alkyldihydroxyacetonephosphate synthase,
peroxisomal precursor (EC 2.5.1.26) (Alkyl-DHAP
synthase) (Alkylglycerone-phosphate synthase)
Length = 658
Score = 70.1 bits (170), Expect = 1e-11
Identities = 52/183 (28%), Positives = 90/183 (48%), Gaps = 5/183 (2%)
Query: 129 YEGSSKLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEV--IVSL--S 184
+E ++L P D V +I+ +L ++P GG T + G + DE I+SL S
Sbjct: 202 FERIPDIVLWPTCHDDVVKIVNLACKYNLCIIPIGGGTSVSYGLMCPADETRTIISLDTS 261
Query: 185 SMNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGL 244
MN+++ D+ + EAG + + L G+ + + +GG VST A G+
Sbjct: 262 QMNRILWVDENNLTAHVEAGITGQELERQLKESGYCTGHEPDSLEFSTVGGWVSTRASGM 321
Query: 245 RLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAIL 304
+ YG++ V+ ++ V G + + R +TG D+ H +GSEG+LG++T+ I
Sbjct: 322 KKNIYGNIEDLVVHIKMVTPRGIIEKSCQGPRM-STGPDIHHFIMGSEGTLGVITEATIK 380
Query: 305 TPP 307
P
Sbjct: 381 IRP 383
Score = 32.0 bits (71), Expect = 4.4
Identities = 18/65 (27%), Positives = 31/65 (47%)
Query: 487 ILSQIEPFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPNHILN 546
+ Q E E + GS+S HG+G ++ + S S ++ S+K +DPN+I
Sbjct: 594 VFEQTEAAAREEILANGGSLSHHHGVGKLRKQWLKESISDVGFGMLKSVKEYVDPNNIFG 653
Query: 547 PYKVL 551
+L
Sbjct: 654 NRNLL 658
>ADAS_CAVPO (P97275) Alkyldihydroxyacetonephosphate synthase,
peroxisomal precursor (EC 2.5.1.26) (Alkyl-DHAP
synthase) (Alkylglycerone-phosphate synthase)
Length = 658
Score = 69.7 bits (169), Expect = 2e-11
Identities = 51/183 (27%), Positives = 90/183 (48%), Gaps = 5/183 (2%)
Query: 129 YEGSSKLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEV--IVSL--S 184
+E ++L P D V +I+ +L ++P GG T + G + DE I+SL S
Sbjct: 202 FERIPDIVLWPTCHDDVVKIVNLACKYNLCIIPIGGGTSVSYGLMCPADETRTIISLDTS 261
Query: 185 SMNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGL 244
MN+++ D+ + EAG + + L G+ + + +GG +ST A G+
Sbjct: 262 QMNRILWVDENNLTAHVEAGITGQELERQLKESGYCTGHEPDSLEFSTVGGWISTRASGM 321
Query: 245 RLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAIL 304
+ YG++ V+ ++ V G + + R +TG D+ H +GSEG+LG++T+ I
Sbjct: 322 KKNIYGNIEDLVVHMKVVTPRGVIEKSCQGPRM-STGPDIHHFIMGSEGTLGVITEATIK 380
Query: 305 TPP 307
P
Sbjct: 381 IRP 383
>ALO_NEUCR (Q7SGY1) Putative D-arabinono-1,4-lactone oxidase (EC
1.1.3.37) (ALO) (L-galactono-gamma-lactone oxidase)
Length = 556
Score = 67.8 bits (164), Expect = 7e-11
Identities = 57/226 (25%), Positives = 101/226 (44%), Gaps = 11/226 (4%)
Query: 86 QRNPRFSELNDDDVRYFQEILGQKNV-VQDEEKLSTVNTDWMHKYEGSSKLLLQPWNTDQ 144
+ NP + L +VR E+ Q++ + K + V+ W + +L +QP + +
Sbjct: 4 ESNPALASL-PSNVREIIELASQRDGGIPFRAKTAHVHRTWAGTFTSLPELYIQPESVSE 62
Query: 145 VSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSSMNKVISFDKVSGILVCEAG 204
+ ++++ V G G + +V+L + NKVIS D ++G++V +AG
Sbjct: 63 IQKVVRLARHARRRVTTTG--CGHSPSDITCTSSWLVNLDNFNKVISVDHLTGLVVVQAG 120
Query: 205 CILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEAVLA 264
L + LD G +P LG+ I G +ST G +++G + S+ ++ LA
Sbjct: 121 IRLYQLSDELDRRGLALP-SLGSINEQSIAGAISTGTHGSG-IKHGLVGESITELKITLA 178
Query: 265 DGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPPKLS 310
+G L DL + S G+LGI+T+V P S
Sbjct: 179 NGETLSC-----SPEDKPDLFRAALISLGALGIITEVTFKAVPAFS 219
>ADAS_CAEEL (O45218) Alkyldihydroxyacetonephosphate synthase (EC
2.5.1.26) (Alkyl-DHAP synthase)
(Alkylglycerone-phosphate synthase)
Length = 597
Score = 64.7 bits (156), Expect = 6e-10
Identities = 40/173 (23%), Positives = 93/173 (53%), Gaps = 5/173 (2%)
Query: 135 LLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGG-SVPVFDE---VIVSLSSMNKVI 190
+++ P + ++ +I++ S + A++P GG T + P ++ + + ++ ++K++
Sbjct: 137 IVVWPKSEHEIVKIIEGAMSHNCAIIPIGGGTSVTNALDTPETEKRAVISMDMALLDKIL 196
Query: 191 SFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYG 250
D+ + +AG + +++ L+ +GF + + +GG VST A G++ +YG
Sbjct: 197 WIDRENLTCRAQAGIVGQSLERQLNKKGFTCGHEPDSIEFSTLGGWVSTRASGMKKNKYG 256
Query: 251 SLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAI 303
++ V+ + V G + + R ++G D+ + +GSEG+LG+V++V I
Sbjct: 257 NIEDLVVHLNFVCPKGIIQKQCQVPRM-SSGPDIHQIILGSEGTLGVVSEVTI 308
Score = 30.8 bits (68), Expect = 9.7
Identities = 15/41 (36%), Positives = 25/41 (60%)
Query: 504 GSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPNHI 544
GSIS HG+G ++ +L + + L+ +IK+ LDP +I
Sbjct: 540 GSISHHHGVGKIRKQWMLTTNGAVGIALLKAIKSELDPANI 580
>GGLO_SCYTO (Q90YK3) L-gulonolactone oxidase (EC 1.1.3.8) (LGO)
(L-gulono-gamma-lactone oxidase) (GLO)
Length = 440
Score = 62.4 bits (150), Expect = 3e-09
Identities = 48/187 (25%), Positives = 84/187 (44%), Gaps = 15/187 (8%)
Query: 124 DWMHKYEGSSKLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSL 183
+W Y +L +P +++ QIL+ N R+ V G G + D +V L
Sbjct: 12 NWATTYSCEPELYFEPTTVEEIRQILELANQRNKRVKVVG--CGHSPSDIACTDNYLVRL 69
Query: 184 SSMNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVST---N 240
+ +N+++ DK + EAG +L ++ LD G + ++GA +GG + T N
Sbjct: 70 NKLNRILQVDKERKWITAEAGILLSDLNEKLDALGLALS-NIGAVSDVALGGVIGTGTHN 128
Query: 241 AGGLRLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTK 300
G +++G L ++ + + A G L+ T+ ++ HL GSLG+V
Sbjct: 129 TG----IQHGILATQIVAMTLMTAAGDTLECSNTVNREIFQATRLHL-----GSLGVVLN 179
Query: 301 VAILTPP 307
V I P
Sbjct: 180 VTIQCVP 186
>GGLO_MOUSE (P58710) L-gulonolactone oxidase (EC 1.1.3.8) (LGO)
(L-gulono-gamma-lactone oxidase) (GLO)
Length = 439
Score = 61.6 bits (148), Expect = 5e-09
Identities = 48/188 (25%), Positives = 86/188 (45%), Gaps = 15/188 (7%)
Query: 124 DWMHKYEGSSKLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSL 183
+W Y S ++ QP + +V ++L ++ V GG G + D ++ +
Sbjct: 11 NWAKTYGCSPEMYYQPTSVGEVREVLALARQQNKKVKVVGG--GHSPSDIACTDGFMIHM 68
Query: 184 SSMNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVST---N 240
MN+V+ DK + EAG +L ++ LD G + +LGA +GG + + N
Sbjct: 69 GKMNRVLQVDKEKKQVTVEAGILLTDLHPQLDKHGLALS-NLGAVSDVTVGGVIGSGTHN 127
Query: 241 AGGLRLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTK 300
G +++G L V+ + + ADGTVL+ ++ + D HL G LG++
Sbjct: 128 TG----IKHGILATQVVALTLMKADGTVLECSESSKADVFQAARVHL-----GCLGVILT 178
Query: 301 VAILTPPK 308
V + P+
Sbjct: 179 VTLQCVPQ 186
>ALO_SCHPO (Q9HDX8) D-arabinono-1,4-lactone oxidase (EC 1.1.3.37)
(ALO) (L-galactono-gamma-lactone oxidase)
Length = 461
Score = 59.7 bits (143), Expect = 2e-08
Identities = 54/217 (24%), Positives = 95/217 (42%), Gaps = 20/217 (9%)
Query: 123 TDWMHKYEGSSKLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVS 182
++W + S L P +Q+ +IL NS + G G + ++S
Sbjct: 18 SNWAKTFSAISLGLRCPKTEEQLREILVDANSNGKKIRVVGA--GHSPSDIVCTSGYLLS 75
Query: 183 LSSMNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAG 242
L MNKV+SFD S + +AG + L N G+ +P+ +G+ + G +ST
Sbjct: 76 LDKMNKVVSFDPDSLSITVQAGIRFYQVQEILQNLGYSLPI-VGSISETSVSGIMSTCTH 134
Query: 243 GLRLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSE---GSLGIVT 299
G L ++ L + + +LADG+++ + L+KD +F ++ G+LG++
Sbjct: 135 GSSL-QHQVLPHYIKSMRIMLADGSIVTCSRELQKD--------MFAAAQVSLGALGVIV 185
Query: 300 KVAILTPPKLSSVNVALLACKDYNCCQKLLQEAKRKL 336
+ I P L+A +D L Q+ K L
Sbjct: 186 DITISVVPAFD-----LVATEDVTTVTDLFQDWKNNL 217
>GGLO_RAT (P10867) L-gulonolactone oxidase (EC 1.1.3.8) (LGO)
(L-gulono-gamma-lactone oxidase) (GLO)
Length = 439
Score = 58.9 bits (141), Expect = 3e-08
Identities = 48/188 (25%), Positives = 83/188 (43%), Gaps = 15/188 (7%)
Query: 124 DWMHKYEGSSKLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSL 183
+W Y S ++ QP + ++V ++L + V GG G + D ++ +
Sbjct: 11 NWAKTYGCSPEVYYQPTSVEEVREVLALAREQKKKVKVVGG--GHSPSDIACTDGFMIHM 68
Query: 184 SSMNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVST---N 240
MN+V+ DK + EAG +L ++ LD G M +LGA + G + + N
Sbjct: 69 GKMNRVLQVDKEKKQITVEAGILLADLHPQLDEHGLAMS-NLGAVSDVTVAGVIGSGTHN 127
Query: 241 AGGLRLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTK 300
G +++G L V+ + + ADG VL+ ++ D HL G LGI+
Sbjct: 128 TG----IKHGILATQVVALTLMTADGEVLECSESRNADVFQAARVHL-----GCLGIILT 178
Query: 301 VAILTPPK 308
V + P+
Sbjct: 179 VTLQCVPQ 186
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.136 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 66,169,669
Number of Sequences: 164201
Number of extensions: 2851859
Number of successful extensions: 7416
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 22
Number of HSP's successfully gapped in prelim test: 26
Number of HSP's that attempted gapping in prelim test: 7331
Number of HSP's gapped (non-prelim): 64
length of query: 557
length of database: 59,974,054
effective HSP length: 115
effective length of query: 442
effective length of database: 41,090,939
effective search space: 18162195038
effective search space used: 18162195038
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0627.15


