

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0593b.1
(208 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
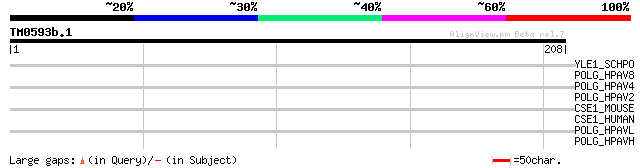
Score E
Sequences producing significant alignments: (bits) Value
YLE1_SCHPO (Q10451) Hypothetical protein C1093.01 in chromosome I 40 0.003
POLG_HPAV8 (P26582) Genome polyprotein [Contains: Coat protein V... 30 4.0
POLG_HPAV4 (P26581) Genome polyprotein [Contains: Coat protein V... 30 4.0
POLG_HPAV2 (P26580) Genome polyprotein [Contains: Coat protein V... 30 4.0
CSE1_MOUSE (Q9ERK4) Importin-alpha re-exporter (Chromosome segre... 30 5.2
CSE1_HUMAN (P55060) Importin-alpha re-exporter (Chromosome segre... 30 5.2
POLG_HPAVL (P06441) Genome polyprotein [Contains: Coat protein V... 29 6.8
POLG_HPAVH (P08617) Genome polyprotein [Contains: Coat protein V... 29 6.8
>YLE1_SCHPO (Q10451) Hypothetical protein C1093.01 in chromosome I
Length = 1261
Score = 40.4 bits (93), Expect = 0.003
Identities = 27/94 (28%), Positives = 45/94 (47%), Gaps = 4/94 (4%)
Query: 97 KPNLLICNRILDYLFKHRRLIKVLDLFSDMRNHMIFPDVCTFNIILNFFCLKSKVDMARE 156
KP++ + N +L L + RR + LF +M+ + P T+ ++N C +A +
Sbjct: 923 KPSVFLYNAVLSKLGRARRTTECWKLFQEMKESGLLPTSVTYGTVINAACRIGDESLAEK 982
Query: 157 LFDGM-IECGFTPDVWCYHIMINLSV---FSRMK 186
LF M + + P V Y+ MI V F+R K
Sbjct: 983 LFAEMENQPNYQPRVAPYNTMIQFEVQTMFNREK 1016
Score = 31.2 bits (69), Expect = 1.8
Identities = 16/52 (30%), Positives = 28/52 (53%), Gaps = 1/52 (1%)
Query: 100 LLICNRILDY-LFKHRRLIKVLDLFSDMRNHMIFPDVCTFNIILNFFCLKSK 150
+L C DY L + + K++ ++ DM N I P+V TF ++ C ++K
Sbjct: 221 VLQCMSTDDYFLSSSKNIQKIIKVYVDMLNSFISPNVTTFETVIFALCRRAK 272
Score = 30.4 bits (67), Expect = 3.1
Identities = 16/59 (27%), Positives = 27/59 (45%)
Query: 120 LDLFSDMRNHMIFPDVCTFNIILNFFCLKSKVDMARELFDGMIECGFTPDVWCYHIMIN 178
L++F + + H + P V +N +L+ + +LF M E G P Y +IN
Sbjct: 911 LNIFEETKRHNVKPSVFLYNAVLSKLGRARRTTECWKLFQEMKESGLLPTSVTYGTVIN 969
>POLG_HPAV8 (P26582) Genome polyprotein [Contains: Coat protein VP4
(P1A); Coat protein VP2 (P1B); Coat protein VP3 (P1C);
Coat protein VP1 (P1D); Picornain 2A (EC 3.4.22.29) (Core
protein P2A); Core protein P2B; Core protein P2C;
Probable protein P3A; Pr
Length = 2226
Score = 30.0 bits (66), Expect = 4.0
Identities = 22/65 (33%), Positives = 35/65 (53%), Gaps = 3/65 (4%)
Query: 64 GFHCDQVMYETLTTDLC--KSGETLLVIQMLHEIDKPNLLICNRILDYLFKHRRLIKVLD 121
GFH V E + T LC KSG L VIQ L++ + +++ R+++Y+ +I
Sbjct: 1005 GFH-HSVTVEIINTVLCFVKSGILLYVIQQLNQDEHSHIIGLLRVMNYVDIGCSVISCGK 1063
Query: 122 LFSDM 126
+FS M
Sbjct: 1064 VFSKM 1068
>POLG_HPAV4 (P26581) Genome polyprotein [Contains: Coat protein VP4
(P1A); Coat protein VP2 (P1B); Coat protein VP3 (P1C);
Coat protein VP1 (P1D); Picornain 2A (EC 3.4.22.29) (Core
protein P2A); Core protein P2B; Core protein P2C;
Probable protein P3A; Pr
Length = 2226
Score = 30.0 bits (66), Expect = 4.0
Identities = 22/65 (33%), Positives = 35/65 (53%), Gaps = 3/65 (4%)
Query: 64 GFHCDQVMYETLTTDLC--KSGETLLVIQMLHEIDKPNLLICNRILDYLFKHRRLIKVLD 121
GFH V E + T LC KSG L VIQ L++ + +++ R+++Y+ +I
Sbjct: 1005 GFH-HSVTVEIINTVLCFVKSGILLYVIQQLNQDEHSHIIGLLRVMNYVDIGCSVISCGK 1063
Query: 122 LFSDM 126
+FS M
Sbjct: 1064 VFSKM 1068
>POLG_HPAV2 (P26580) Genome polyprotein [Contains: Coat protein VP4
(P1A); Coat protein VP2 (P1B); Coat protein VP3 (P1C);
Coat protein VP1 (P1D); Picornain 2A (EC 3.4.22.29) (Core
protein P2A); Core protein P2B; Core protein P2C;
Probable protein P3A; Pr
Length = 2226
Score = 30.0 bits (66), Expect = 4.0
Identities = 22/65 (33%), Positives = 35/65 (53%), Gaps = 3/65 (4%)
Query: 64 GFHCDQVMYETLTTDLC--KSGETLLVIQMLHEIDKPNLLICNRILDYLFKHRRLIKVLD 121
GFH V E + T LC KSG L VIQ L++ + +++ R+++Y+ +I
Sbjct: 1005 GFH-HSVTVEIINTVLCFVKSGILLYVIQQLNQDEHSHIIGLLRVMNYVDIGCSVISCGK 1063
Query: 122 LFSDM 126
+FS M
Sbjct: 1064 VFSKM 1068
>CSE1_MOUSE (Q9ERK4) Importin-alpha re-exporter (Chromosome
segregation 1-like protein) (Cellular apoptosis
susceptibility protein)
Length = 971
Score = 29.6 bits (65), Expect = 5.2
Identities = 17/71 (23%), Positives = 37/71 (51%), Gaps = 6/71 (8%)
Query: 96 DKPNLLICNRILDYL-----FKHRRLIKVLDLFSDMRNHMIFPDVCTFNIILNFFCLKSK 150
D + N I++++ ++R+ I +L LF ++N + +F + +N +C+K
Sbjct: 744 DHQGFYLLNSIIEHMPPESVDQYRKQIFIL-LFQRLQNSKTTKFIKSFLVFINLYCIKYG 802
Query: 151 VDMARELFDGM 161
+E+FDG+
Sbjct: 803 ALALQEIFDGI 813
>CSE1_HUMAN (P55060) Importin-alpha re-exporter (Chromosome
segregation 1-like protein) (Cellular apoptosis
susceptibility protein)
Length = 971
Score = 29.6 bits (65), Expect = 5.2
Identities = 17/71 (23%), Positives = 37/71 (51%), Gaps = 6/71 (8%)
Query: 96 DKPNLLICNRILDYL-----FKHRRLIKVLDLFSDMRNHMIFPDVCTFNIILNFFCLKSK 150
D + N I++++ ++R+ I +L LF ++N + +F + +N +C+K
Sbjct: 744 DHQGFYLLNSIIEHMPPESVDQYRKQIFIL-LFQRLQNSKTTKFIKSFLVFINLYCIKYG 802
Query: 151 VDMARELFDGM 161
+E+FDG+
Sbjct: 803 ALALQEIFDGI 813
>POLG_HPAVL (P06441) Genome polyprotein [Contains: Coat protein VP4
(P1A); Coat protein VP2 (P1B); Coat protein VP3 (P1C);
Coat protein VP1 (P1D); Picornain 2A (EC 3.4.22.29) (Core
protein P2A); Core protein P2B; Core protein P2C;
Probable protein P3A; Pr
Length = 2227
Score = 29.3 bits (64), Expect = 6.8
Identities = 22/65 (33%), Positives = 34/65 (51%), Gaps = 3/65 (4%)
Query: 64 GFHCDQVMYETLTTDLC--KSGETLLVIQMLHEIDKPNLLICNRILDYLFKHRRLIKVLD 121
GFH V E + T LC KSG L VIQ L++ + +++ R+++Y +I
Sbjct: 1005 GFH-HSVTVEIINTVLCFVKSGILLYVIQQLNQDEHSHIIGLLRVMNYADIGCSVISCGK 1063
Query: 122 LFSDM 126
+FS M
Sbjct: 1064 VFSKM 1068
>POLG_HPAVH (P08617) Genome polyprotein [Contains: Coat protein VP4
(P1A); Coat protein VP2 (P1B); Coat protein VP3 (P1C);
Coat protein VP1 (P1D); Picornain 2A (EC 3.4.22.29) (Core
protein P2A); Core protein P2B; Core protein P2C;
Probable protein P3A; Pr
Length = 2227
Score = 29.3 bits (64), Expect = 6.8
Identities = 22/65 (33%), Positives = 34/65 (51%), Gaps = 3/65 (4%)
Query: 64 GFHCDQVMYETLTTDLC--KSGETLLVIQMLHEIDKPNLLICNRILDYLFKHRRLIKVLD 121
GFH V E + T LC KSG L VIQ L++ + +++ R+++Y +I
Sbjct: 1005 GFH-HSVTVEIINTVLCFVKSGILLYVIQQLNQDEHSHIIGLLRVMNYADIGCSVISCGK 1063
Query: 122 LFSDM 126
+FS M
Sbjct: 1064 VFSKM 1068
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.342 0.152 0.492
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,141,340
Number of Sequences: 164201
Number of extensions: 872916
Number of successful extensions: 3443
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 3439
Number of HSP's gapped (non-prelim): 11
length of query: 208
length of database: 59,974,054
effective HSP length: 105
effective length of query: 103
effective length of database: 42,732,949
effective search space: 4401493747
effective search space used: 4401493747
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0593b.1


