

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0593a.3
(562 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
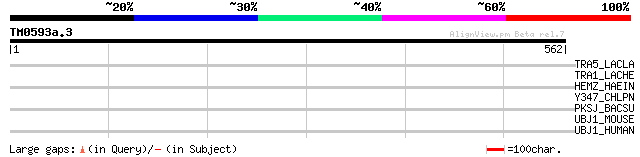
Score E
Sequences producing significant alignments: (bits) Value
TRA5_LACLA (P35881) Transposase for insertion sequence element I... 38 0.080
TRA1_LACHE (P35880) Transposase for insertion sequence element I... 36 0.31
HEMZ_HAEIN (P43868) Ferrochelatase (EC 4.99.1.1) (Protoheme ferr... 33 2.6
Y347_CHLPN (Q9Z8J6) Probable metal transport system membrane pro... 32 5.8
PKSJ_BACSU (P40806) Putative polyketide synthase pksJ (PKS) 31 7.5
UBJ1_MOUSE (Q9JJZ4) Ubiquitin-conjugating enzyme E2 J1 (EC 6.3.2... 31 9.8
UBJ1_HUMAN (Q9Y385) Ubiquitin-conjugating enzyme E2 J1 (EC 6.3.2... 31 9.8
>TRA5_LACLA (P35881) Transposase for insertion sequence element
IS905
Length = 391
Score = 37.7 bits (86), Expect = 0.080
Identities = 39/151 (25%), Positives = 63/151 (40%), Gaps = 19/151 (12%)
Query: 283 KQPKAVVIDGDKSMRAAVKVVFPNATHRLCGWHIQQNCLEKIKIPD---FLNEFKTLIYG 339
+Q VV DG K + + +P A + C HI +N K+K D L +FKT+
Sbjct: 220 QQVSLVVTDGFKGLEQIISQAYPLAKQQRCLIHISRNLASKVKRADRAVILEQFKTI--- 276
Query: 340 NFTPERFETKWLQVIEKYGIGE-----EKWIKQMYETRQMWATAFMRENFFAGIRTTSLC 394
+ E E +Q +E + I E K ++ + T + + I +T+L
Sbjct: 277 -YRAENLEMA-VQALENF-IAEWKPKYRKVMESLENTDNLLTFYQFPYQIWHSIYSTNLI 333
Query: 395 EGINSFIKSYVLCKNSILDFIYNFERAVEEY 425
E +N IK K ++ E A+E Y
Sbjct: 334 ESLNKEIKRQTKKK-----VLFPNEEALERY 359
>TRA1_LACHE (P35880) Transposase for insertion sequence element
IS1201
Length = 369
Score = 35.8 bits (81), Expect = 0.31
Identities = 24/92 (26%), Positives = 45/92 (48%), Gaps = 6/92 (6%)
Query: 248 NHHTETTIFACALVCDETIETYKWVLKALDEAMFGKQPKAVVIDGDKSMRAAVKVVFPNA 307
N H E + A +E IE + +L+++ +Q + + DG M+ A+ +P A
Sbjct: 188 NGHKEVIDYCIAP--NENIEVWTELLQSMKSRGL-EQVELFLSDGVVGMKTALAKTYPQA 244
Query: 308 THRLCGWHIQQNCLEKIKIPD---FLNEFKTL 336
+ C H+ +N K+++ D +NEFK +
Sbjct: 245 HFQRCLVHVMRNICAKVRVEDREAIMNEFKQI 276
>HEMZ_HAEIN (P43868) Ferrochelatase (EC 4.99.1.1) (Protoheme
ferro-lyase) (Heme synthetase)
Length = 323
Score = 32.7 bits (73), Expect = 2.6
Identities = 37/160 (23%), Positives = 61/160 (38%), Gaps = 22/160 (13%)
Query: 314 WHIQQNCL----EKIKIPDFLNEFKTLIY----------GNFTPERFETKWLQVIEKYGI 359
+HI +N + + IK+ +EF Y G++ E + + V+ K G+
Sbjct: 167 YHIDENYINALADSIKVRLKSDEFLLFSYHGIPLRYEKMGDYYREHCKQTTIAVVNKLGL 226
Query: 360 GEEKWIKQMYETRQMWATAFMRENFFAGIRTTSL-CEGINSFIKSYVLCKNSILDFIYNF 418
E +W R + + F RE + L + K V+C +D +
Sbjct: 227 TENQW-------RMTFQSRFGREEWLQPYTDKFLESAAAQNIQKIAVICPGFSVDCLETI 279
Query: 419 ERAVEEYRHNELASDFKSRYGEPVLITALSHIEQGASKLL 458
E EE R N L + +S P L +HIE +L
Sbjct: 280 EEIDEENRENFLNNGGQSYQYIPALNVEHAHIEMMGKLIL 319
>Y347_CHLPN (Q9Z8J6) Probable metal transport system membrane
protein CPn0347/CP0413/CPj0347/CpB0354
Length = 448
Score = 31.6 bits (70), Expect = 5.8
Identities = 17/45 (37%), Positives = 24/45 (52%), Gaps = 2/45 (4%)
Query: 314 WHIQQNCLEKIKIPDFLNEFKTLIYGNFTPERFETKWLQVIEKYG 358
WHI N LE I + DF+ +K Y F P+ F +Q++E G
Sbjct: 320 WHISHNRLENISVRDFVCSYKYQEY--FGPKPFPRWRVQILEWRG 362
>PKSJ_BACSU (P40806) Putative polyketide synthase pksJ (PKS)
Length = 5045
Score = 31.2 bits (69), Expect = 7.5
Identities = 28/97 (28%), Positives = 40/97 (40%), Gaps = 10/97 (10%)
Query: 104 PAAHVHFIPSFRGLTDGDKAQVDSLKLYGVRSCNITG------LLMGQKGGHESVGFLKK 157
PAAH+ ++D DS+ + G+ SC G + G ES+ F K
Sbjct: 1743 PAAHIE--DQSTEMSDLPDYYDDSVAIIGI-SCEFPGAKNHDEFWENLRDGKESIAFFNK 1799
Query: 158 DLYNHVDKKKRISLGAGDAVSALSYLQGKGEKDPMFF 194
+ K I+ A D V A + + GK DP FF
Sbjct: 1800 EELQRFGISKEIAENA-DYVPAKASIDGKDRFDPSFF 1835
>UBJ1_MOUSE (Q9JJZ4) Ubiquitin-conjugating enzyme E2 J1 (EC
6.3.2.19) (Non-canonical ubiquitin conjugating enzyme 1)
(NCUBE1)
Length = 318
Score = 30.8 bits (68), Expect = 9.8
Identities = 17/42 (40%), Positives = 23/42 (54%), Gaps = 5/42 (11%)
Query: 481 RSEIGTTVMIKFSICRHPNTQLSSWSFMTRIRTCLIVIVGFL 522
R E+G + + S HP T SWS IRT L+ I+GF+
Sbjct: 83 RFEVGKKICLSIS-GHHPETWQPSWS----IRTALLAIIGFM 119
>UBJ1_HUMAN (Q9Y385) Ubiquitin-conjugating enzyme E2 J1 (EC
6.3.2.19) (Non-canonical ubiquitin conjugating enzyme 1)
(NCUBE1) (Yeast ubiquitin conjugating enzyme UBC6
homolog E) (HSUBC6e) (CGI-76) (HSPC153/HSPC205)
Length = 318
Score = 30.8 bits (68), Expect = 9.8
Identities = 17/42 (40%), Positives = 23/42 (54%), Gaps = 5/42 (11%)
Query: 481 RSEIGTTVMIKFSICRHPNTQLSSWSFMTRIRTCLIVIVGFL 522
R E+G + + S HP T SWS IRT L+ I+GF+
Sbjct: 83 RFEVGKKICLSIS-GHHPETWQPSWS----IRTALLAIIGFM 119
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.331 0.143 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 62,732,104
Number of Sequences: 164201
Number of extensions: 2598000
Number of successful extensions: 6725
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 6724
Number of HSP's gapped (non-prelim): 7
length of query: 562
length of database: 59,974,054
effective HSP length: 115
effective length of query: 447
effective length of database: 41,090,939
effective search space: 18367649733
effective search space used: 18367649733
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0593a.3


