

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0590c.8
(919 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
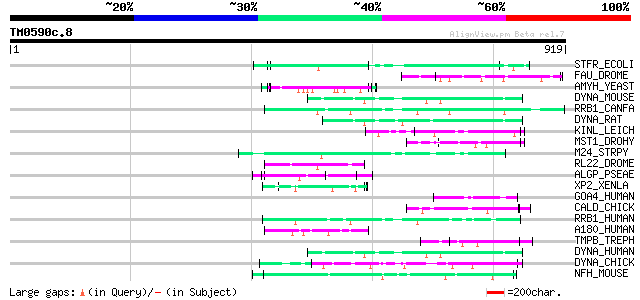
Score E
Sequences producing significant alignments: (bits) Value
STFR_ECOLI (P76072) Side tail fiber protein homolog from lambdoi... 71 1e-11
FAU_DROME (Q9VGX3) Anoxia up-regulated protein 62 9e-09
AMYH_YEAST (P08640) Glucoamylase S1/S2 precursor (EC 3.2.1.3) (G... 58 1e-07
DYNA_MOUSE (O08788) Dynactin 1 (150 kDa dynein-associated polype... 56 4e-07
RRB1_CANFA (Q28298) Ribosome-binding protein 1 (180 kDa ribosome... 55 9e-07
DYNA_RAT (P28023) Dynactin 1 (150 kDa dynein-associated polypept... 55 9e-07
KINL_LEICH (P46865) Kinesin-like protein K39 (Fragment) 54 1e-06
MST1_DROHY (Q08695) Axoneme-associated protein mst101(1) 54 2e-06
M24_STRPY (P12379) M protein, serotype 24 precursor 54 2e-06
RL22_DROME (P50887) 60S ribosomal protein L22 54 2e-06
ALGP_PSEAE (P15276) Transcriptional regulatory protein algP (Alg... 54 2e-06
XP2_XENLA (P17437) Skin secretory protein xP2 precursor (APEG pr... 52 6e-06
GOA4_HUMAN (Q13439) Golgi autoantigen, golgin subfamily A member... 52 6e-06
CALD_CHICK (P12957) Caldesmon (CDM) 52 6e-06
RRB1_HUMAN (Q9P2E9) Ribosome-binding protein 1 (Ribosome recepto... 52 7e-06
A180_HUMAN (O60641) Clathrin coat assembly protein AP180 (Clathr... 52 7e-06
TMPB_TREPH (P29720) Treponemal membrane protein B precursor (Ant... 52 9e-06
DYNA_HUMAN (Q14203) Dynactin 1 (150 kDa dynein-associated polype... 52 9e-06
DYNA_CHICK (P35458) Dynactin 1 (150 kDa dynein-associated polype... 52 9e-06
NFH_MOUSE (P19246) Neurofilament triplet H protein (200 kDa neur... 51 1e-05
>STFR_ECOLI (P76072) Side tail fiber protein homolog from lambdoid
prophage Rac
Length = 1120
Score = 71.2 bits (173), Expect = 1e-11
Identities = 112/442 (25%), Positives = 162/442 (36%), Gaps = 71/442 (16%)
Query: 432 AESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPA 491
A++T A K K++SDA TS A A AA + + ++ AA+ +A ASS A
Sbjct: 121 AQNTAAAK-----KSASDASTSAREAATHAADAADSARAASTSAGQAAS--SAQSASSSA 173
Query: 492 SAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPS 551
+T A A S A +S AT A A+ K +ET S Q A S
Sbjct: 174 GTASTKATEASKSAAAAESSKSAAATSAGAA---------KTSETNASASLQSAATS--- 221
Query: 552 THDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSSYYNMLPNAIEPSEFLLAG 611
A + + S AA + EAA+ + A SS+ S AG
Sbjct: 222 ---ASTATTKASEAATSARDAAASKEAAKSSETNASSSASSA----------ASSATAAG 268
Query: 612 LNRDVIEKEVLSQGLNVTKEETLACLLRAGCIFAHTFEKFNAANVEAERLKAESAKHQEA 671
+ K + N ET A + A K +A A
Sbjct: 269 NSA----KAAKTSETNARSSETAA----------------GQSASAAAGSKTAAASSASA 308
Query: 672 AAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKEEADELDASLQACKKE 731
A+ + AT AGK A TA K GE +Q + A S K
Sbjct: 309 ASTSAGQASASATAAGKSAESAASSASTATTKAGEATEQASAAARSASAAKTSETNAKAS 368
Query: 732 KEQAEKD-LIARGEALIAKESELAILCAELELVKKALAEQEKKSAESLALAKSDMEAVMQ 790
+ AE A A A S + ++ E ++A A + + S ++ A
Sbjct: 369 ETSAESSKTAAASSASSAASSASSASASKDEATRQASAAKSSATTASTKATEAAGSATAA 428
Query: 791 ATSEEIKKATETHAEALATKDAEIASQLAKIKSLEDELATEK------------AKAIEA 838
A S+ ++ T AE A + +IAS +A LED T+K ++ + A
Sbjct: 429 AQSKSTAESAATRAETAAKRAEDIASAVA----LEDASTTKKGIVQLSSATNSTSETLAA 484
Query: 839 REQAADIALDNRERGFYLAKDQ 860
+A A DN E+ L KDQ
Sbjct: 485 TPKAVKSAYDNAEK--RLQKDQ 504
Score = 54.7 bits (130), Expect = 1e-06
Identities = 89/387 (22%), Positives = 140/387 (35%), Gaps = 31/387 (8%)
Query: 428 AENQAESTKAPKRRRLTKASS-DAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVP 486
A + A+S +A ASS + +S G T A A K AE+S +AA
Sbjct: 145 AADAADSARAASTSAGQAASSAQSASSSAGTASTKATEASKSAAAAESSKSAAATSAGAA 204
Query: 487 ASSPASAGATAAFAAESSIGAT--AASAGVNATKAAASSDTPIGDKEKENETPKSPPRQD 544
+S +A A+ AA S+ AT A+ A +A AAAS + K +ET S
Sbjct: 205 KTSETNASASLQSAATSASTATTKASEAATSARDAAASKEA-----AKSSETNASSSASS 259
Query: 545 APPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSSYYNMLPNAIEP 604
A S + ++ +S AA S +A A +S+ A
Sbjct: 260 AASSATAAGNSAKAAKTSETNARSSETAAGQSASAAAGSKTAAASSASAASTSAGQA--S 317
Query: 605 SEFLLAGLNRDVIEKEVLSQGLNVTKEETLACLLRAGCIFAHTFEKFNAANVEAERLKAE 664
+ AG + + S T + A + + + K + N +A AE
Sbjct: 318 ASATAAGKSAE----SAASSASTATTKAGEATEQASAAARSASAAKTSETNAKASETSAE 373
Query: 665 SAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKEEADELDAS 724
S+K A++A + A KD+ A + + +A E S
Sbjct: 374 SSKTAAASSASSAASSASSASASKDE---------ATRQASAAKSSATTASTKATEAAGS 424
Query: 725 LQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKALAEQEKKSAESLALAKSD 784
A + K AE R E + ++A A + A KK L+ A +
Sbjct: 425 ATAAAQSKSTAE-SAATRAETAAKRAEDIASAVALED------ASTTKKGIVQLSSATNS 477
Query: 785 MEAVMQATSEEIKKATETHAEALATKD 811
+ AT + +K A + +AE KD
Sbjct: 478 TSETLAATPKAVKSAYD-NAEKRLQKD 503
Score = 49.7 bits (117), Expect = 4e-05
Identities = 49/199 (24%), Positives = 75/199 (37%), Gaps = 9/199 (4%)
Query: 404 ARERRMARFSPKFNTEGDAKKRGGAENQAES-TKAPKRRRLTKASSDAGTSKPGAQPTTA 462
A R A S + + A + A S T A + K S S A +A
Sbjct: 233 ATSARDAAASKEAAKSSETNASSSASSAASSATAAGNSAKAAKTSETNARSSETAAGQSA 292
Query: 463 AAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATA--------ASAGV 514
+AA K A +S +AA+ ++S +AG +A AA S+ AT ASA
Sbjct: 293 SAAAGSKTAAASSASAASTSAGQASASATAAGKSAESAASSASTATTKAGEATEQASAAA 352
Query: 515 NATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAAT 574
+ AA +S+T E E+ K+ A + S A + + +AT
Sbjct: 353 RSASAAKTSETNAKASETSAESSKTAAASSASSAASSASSASASKDEATRQASAAKSSAT 412
Query: 575 TSEAAQIEQAPAPEVGSSS 593
T+ E A + + S
Sbjct: 413 TASTKATEAAGSATAAAQS 431
Score = 37.7 bits (86), Expect = 0.14
Identities = 89/376 (23%), Positives = 125/376 (32%), Gaps = 81/376 (21%)
Query: 472 AEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKE 531
A A AA + +A AS+ A AT A A S A + SAG A+ A ++S +
Sbjct: 119 AVAQNTAAAKKSASDASTSAREAATHAADAADSARAASTSAGQAASSAQSASSSAGTAST 178
Query: 532 KENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGS 591
K E KS E S AAT++ AA+ + A
Sbjct: 179 KATEASKSAAA----------------------AESSKSAAATSAGAAKTSETNASASLQ 216
Query: 592 SSYYNMLPNAIEPSEFLLAGLNRDVIEKEVLSQGLNVTKEETLACLLRAGCIFAHTFEKF 651
S+ + + SE + + ++ S N + + A
Sbjct: 217 SAATSASTATTKASEAATSARDAAASKEAAKSSETNASSSASSA---------------- 260
Query: 652 NAANVEAERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQL 711
A++ A A++AK E A + T AG+ A AG K
Sbjct: 261 -ASSATAAGNSAKAAKTSETNARSSE------TAAGQSASAA------AGSKTAAA---- 303
Query: 712 ALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKALA-EQ 770
+S A QA A G K +E A A K A EQ
Sbjct: 304 -----------SSASAASTSAGQASASATAAG-----KSAESAASSASTATTKAGEATEQ 347
Query: 771 EKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIKSLEDELAT 830
+A S + AK TSE KA+ET AE+ T A AS A S
Sbjct: 348 ASAAARSASAAK---------TSETNAKASETSAESSKTAAASSASSAASSASSASASKD 398
Query: 831 EKAKAIEAREQAADIA 846
E + A + +A A
Sbjct: 399 EATRQASAAKSSATTA 414
>FAU_DROME (Q9VGX3) Anoxia up-regulated protein
Length = 619
Score = 61.6 bits (148), Expect = 9e-09
Identities = 92/302 (30%), Positives = 129/302 (42%), Gaps = 43/302 (14%)
Query: 649 EKFNAANVEAERL-KAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGEL 707
E + ++E +R KA+ AK +E A E+R D+L +A K K KI
Sbjct: 323 EALDEVDLEKKRAQKADEAKRREERALKEER-DRLTAEAEKQAAAKAKKAAEEAAKIAAE 381
Query: 708 EDQLA-----LMKEEADELDASLQACKKEKEQAE--KDLIAR--GEALIAKESELAILCA 758
E LA EEA L A+ A +K E+A ++ A+ E K +E A L A
Sbjct: 382 EALLAEAAAQKAAEEAKALKAAEDAAQKAAEEARLAEEAAAQKVAEEAAQKAAEEARL-A 440
Query: 759 ELELVKKALAEQEKKSAESLAL-----AKSDMEAVMQATSEEIKKATETH--AEALATKD 811
E +KA E +K+AE AL A+ EA +A E KA E AE A K
Sbjct: 441 EEAAAQKAAEEAAQKAAEEAALKAAEEARLAEEAAQKAAEEAALKAVEEARAAEEAAQKA 500
Query: 812 AEIA----------SQLAKIKSLEDELATEKAKAIEAREQAADIALDNRERGFYLAKDQA 861
AE A Q + + LE EK E QAA++A R+ LA +
Sbjct: 501 AEEARVAEEARLEEEQRVREQELERLAEIEKESEGELARQAAELAEIARQES-ELAAQEL 559
Query: 862 QHLFPNFDFSAMGVMKE-----------ITAAGLVGPDDPPLIDQNLWTATEEEEEEEEE 910
Q + N + ++ V++E I V P +++L EEEE+EEEE
Sbjct: 560 QAIQKNENETSEPVVEEPVTPVEEQEPIIELGSNVTPTGGNSYEEDL--DAEEEEDEEEE 617
Query: 911 EE 912
EE
Sbjct: 618 EE 619
Score = 52.0 bits (123), Expect = 7e-06
Identities = 60/216 (27%), Positives = 106/216 (48%), Gaps = 19/216 (8%)
Query: 706 ELEDQLALMKEEADELDASLQ----ACKKEKEQ--AEKDLIARGEALIAKESELAILCAE 759
E D++ L K+ A + D + + A K+E+++ AE + A +A A E E A + AE
Sbjct: 323 EALDEVDLEKKRAQKADEAKRREERALKEERDRLTAEAEKQAAAKAKKAAE-EAAKIAAE 381
Query: 760 LELVKKALAEQEKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLA 819
L+ +A A++ + A++L A E Q +EE + A E A+ +A + A+ A++ A
Sbjct: 382 EALLAEAAAQKAAEEAKALKAA----EDAAQKAAEEARLAEEAAAQKVAEEAAQKAAEEA 437
Query: 820 KIKSLEDELATEKAKAIEAREQAADIALDNRERGFYLAKDQAQHLFPNFDFSAMGVMKEI 879
+ L +E A +KA A EA ++AA+ A LA++ AQ A+ +
Sbjct: 438 R---LAEEAAAQKA-AEEAAQKAAEEAALKAAEEARLAEEAAQKAAEEAALKAVEEARAA 493
Query: 880 TAAGLVGPDDPPLIDQNLWTATEEEEEEEEEEEQEK 915
A ++ + ++ A EEE+ E+E E+
Sbjct: 494 EEAAQKAAEEARVAEE----ARLEEEQRVREQELER 525
>AMYH_YEAST (P08640) Glucoamylase S1/S2 precursor (EC 3.2.1.3)
(Glucan 1,4-alpha-glucosidase) (1,4-alpha-D-glucan
glucohydrolase)
Length = 1367
Score = 57.8 bits (138), Expect = 1e-07
Identities = 45/181 (24%), Positives = 83/181 (44%), Gaps = 14/181 (7%)
Query: 428 AENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVA-------AAT 480
+ + ES+ AP T++SS A P + T +++AP + E+S A + T
Sbjct: 462 SSSTTESSSAPVTSSTTESSS-APVPTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTT 520
Query: 481 EPTAVPASSPASAGATAAFAAESSIGATAASAGV--NATKAAASSDTPIGDKEKENETPK 538
E ++ PA +P+S+ ++ A +S ++SA V ++ SS TP+ E+ +
Sbjct: 521 ESSSAPAPTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSTPVTSSTTESSSAP 580
Query: 539 SPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSSYYNML 598
P P S + + P+P+P +S A T ++ E + AP S++ +
Sbjct: 581 VP----TPSSSTTESSSAPVPTPSSSTTESSSAPAPTPSSSTTESSSAPVTSSTTESSSA 636
Query: 599 P 599
P
Sbjct: 637 P 637
Score = 55.1 bits (131), Expect = 9e-07
Identities = 45/178 (25%), Positives = 81/178 (45%), Gaps = 12/178 (6%)
Query: 433 ESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPAS 492
ES+ AP T++SS A P + T +++AP + E+S A T T +S+P +
Sbjct: 377 ESSSAPVTSSTTESSS-APVPTPSSSTTESSSAPVTSSTTESSSAPVTSSTTESSSAPVT 435
Query: 493 AGATAAFAAESSIGATAASAGVNATKAAA---SSDTPIGDKEKENE-----TPKSPPRQ- 543
+ T + +A + T +S+ T +++ SS P+ E+ TP S +
Sbjct: 436 SSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTES 495
Query: 544 -DAPPSPPSTH-DAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSSYYNMLP 599
AP + +T + P+P+P +S A T ++ E + AP S++ + P
Sbjct: 496 SSAPVTSSTTESSSAPVPTPSSSTTESSSAPAPTPSSSTTESSSAPVTSSTTESSSAP 553
Score = 52.0 bits (123), Expect = 7e-06
Identities = 41/186 (22%), Positives = 73/186 (39%), Gaps = 13/186 (6%)
Query: 431 QAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSP 490
++ S P T SS A P + T +++AP + E+S A T T +S+P
Sbjct: 647 ESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVTSSTTESSSAP 706
Query: 491 A---------SAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPP 541
S+ A + S+ +++A ++ SS P+ E+ + P
Sbjct: 707 VPTPSSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVP- 765
Query: 542 RQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSSYYNMLPNA 601
P S + + P+P+P +S T ++ E + AP SS N+ +A
Sbjct: 766 ---TPSSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSVAPVPTPSSSSNITSSA 822
Query: 602 IEPSEF 607
+ F
Sbjct: 823 PSSTPF 828
Score = 51.6 bits (122), Expect = 9e-06
Identities = 43/170 (25%), Positives = 74/170 (43%), Gaps = 14/170 (8%)
Query: 428 AENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPA 487
+ + ES+ AP T++SS A P + T +++AP + E+S A T T +
Sbjct: 333 SSSTTESSSAPVTSSTTESSS-APVPTPSSSTTESSSAPVTSSTTESSSAPVTSSTTESS 391
Query: 488 SSPA---SAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQD 544
S+P S+ T + +A + T +S+ + SS P+ E+ +
Sbjct: 392 SAPVPTPSSSTTESSSAPVTSSTTESSSAPVTSSTTESSSAPVTSSTTESSS-------- 443
Query: 545 APPSPPSTH-DAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSS 593
AP + +T + P+P+P +S A TS + AP P SS+
Sbjct: 444 APVTSSTTESSSAPVPTPSSSTTES-SSAPVTSSTTESSSAPVPTPSSST 492
Score = 50.4 bits (119), Expect = 2e-05
Identities = 44/172 (25%), Positives = 76/172 (43%), Gaps = 15/172 (8%)
Query: 428 AENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVA----AATEPT 483
+ + ES+ AP T++SS TS T +++AP + E+S A + TE +
Sbjct: 399 SSSTTESSSAPVTSSTTESSSAPVTSST----TESSSAPVTSSTTESSSAPVTSSTTESS 454
Query: 484 AVPASSPASAGATAAFAAESSIGATAASAGV--NATKAAASSDTPIGDKEKENETPKSPP 541
+ P +P+S+ ++ A +S ++SA V ++ SS P+ E+ + P
Sbjct: 455 SAPVPTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVP- 513
Query: 542 RQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSS 593
P S + + P P+P +S A TS + AP P SS+
Sbjct: 514 ---TPSSSTTESSSAPAPTPSSSTTES-SSAPVTSSTTESSSAPVPTPSSST 561
Score = 50.4 bits (119), Expect = 2e-05
Identities = 42/178 (23%), Positives = 71/178 (39%), Gaps = 6/178 (3%)
Query: 428 AENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPA 487
+ + ES+ AP T++SS TS + ++A + E+S A T T +
Sbjct: 360 SSSTTESSSAPVTSSTTESSSAPVTSST-TESSSAPVPTPSSSTTESSSAPVTSSTTESS 418
Query: 488 SSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPP 547
S+P ++ T + +A + T +S+ + SS P+ S P
Sbjct: 419 SAPVTSSTTESSSAPVTSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAP----VT 474
Query: 548 SPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSSYYNMLPNAIEPS 605
S + + P+P+P +S A TS + AP P SS+ + A PS
Sbjct: 475 SSTTESSSAPVPTPSSSTTESS-SAPVTSSTTESSSAPVPTPSSSTTESSSAPAPTPS 531
Score = 47.8 bits (112), Expect = 1e-04
Identities = 39/183 (21%), Positives = 69/183 (37%), Gaps = 7/183 (3%)
Query: 418 TEGDAKKRGGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVA 477
TE + + ++ S P T SS A P + T +++AP + E+S A
Sbjct: 493 TESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPAPTPSSSTTESSSAPVTSSTTESSSA 552
Query: 478 -------AATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDK 530
+ TE ++ P +S + ++A SS ++SA V ++ + +
Sbjct: 553 PVPTPSSSTTESSSTPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPAP 612
Query: 531 EKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVG 590
+ T +S + S+ P PS S P +S + AP P
Sbjct: 613 TPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVPTPS 672
Query: 591 SSS 593
SS+
Sbjct: 673 SST 675
Score = 47.8 bits (112), Expect = 1e-04
Identities = 39/173 (22%), Positives = 68/173 (38%), Gaps = 12/173 (6%)
Query: 433 ESTKAPKRRRLTKASS---DAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASS 489
ES+ AP T++SS + T++ + P T++ ++ TE ++ P +S
Sbjct: 416 ESSSAPVTSSTTESSSAPVTSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVTS 475
Query: 490 PASAGATAAFAAESSIGATAASAGVNATKAAASS---DTPIGDKEKENETPKSPP----- 541
+ ++A SS ++SA V ++ +SS TP + + P P
Sbjct: 476 STTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPAPTPSSSTT 535
Query: 542 RQDAPPSPPSTHDAGPMPSP-PHQGEKSCPGAATTSEAAQIEQAPAPEVGSSS 593
+ P ST ++ P P P TS + AP P SS+
Sbjct: 536 ESSSAPVTSSTTESSSAPVPTPSSSTTESSSTPVTSSTTESSSAPVPTPSSST 588
Score = 46.2 bits (108), Expect = 4e-04
Identities = 42/202 (20%), Positives = 81/202 (39%), Gaps = 20/202 (9%)
Query: 418 TEGDAKKRGGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPK---GKNVAEA 474
TE + + ++ S P T SS A P + T +++AP + E+
Sbjct: 562 TESSSTPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPAPTPSSSTTES 621
Query: 475 SVA----AATEPTAVPASSPASAGATAAFA-----AESSIGATAASAGVNATKAAASSDT 525
S A + TE ++ P +P+S+ ++ A + S+ +++A ++ SS
Sbjct: 622 SSAPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSA 681
Query: 526 PIGDKEKENE----TPKSPPRQDAPPSPPST----HDAGPMPSPPHQGEKSCPGAATTSE 577
P+ E+ T + AP PS+ + P+P+P +S T
Sbjct: 682 PVTSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVPTPS 741
Query: 578 AAQIEQAPAPEVGSSSYYNMLP 599
++ E + AP S++ + P
Sbjct: 742 SSTTESSSAPVTSSTTESSSAP 763
Score = 46.2 bits (108), Expect = 4e-04
Identities = 41/164 (25%), Positives = 63/164 (38%), Gaps = 7/164 (4%)
Query: 431 QAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTA-VPASS 489
++ S P T SS A P + T +++AP + E+S A P++ SS
Sbjct: 716 ESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESS 775
Query: 490 PASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSP 549
A ++ ESS + + A TP + P S P S
Sbjct: 776 SAPVPTPSSSTTESSSAPVPTPSSSTTESSVAPVPTPSSSSNITSSAPSSTPFSS---ST 832
Query: 550 PSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSS 593
S+ P PS S P +++T+E++ AP P SSS
Sbjct: 833 ESSSVPVPTPSSSTTESSSAPVSSSTTESS---VAPVPTPSSSS 873
Score = 45.8 bits (107), Expect = 5e-04
Identities = 39/165 (23%), Positives = 68/165 (40%), Gaps = 11/165 (6%)
Query: 433 ESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPAS 492
ES+ AP T++SS A P + T +++AP ++ TE ++ P +P+S
Sbjct: 689 ESSSAPVTSSTTESSS-APVPTPSSSTTESSSAP-----VPTPSSSTTESSSAPVPTPSS 742
Query: 493 AGATAAFAAESSIGATAASAGVNATKAAA--SSDTPIGDKEKENETPKSPPRQDAPPSPP 550
+ ++ A +S ++SA V ++ SS P+ S P P S
Sbjct: 743 STTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAP-VPTPSSST 801
Query: 551 STHDAGPMPSPPHQGE--KSCPGAATTSEAAQIEQAPAPEVGSSS 593
+ P+P+P S P + S + + P P SS+
Sbjct: 802 TESSVAPVPTPSSSSNITSSAPSSTPFSSSTESSSVPVPTPSSST 846
Score = 45.4 bits (106), Expect = 7e-04
Identities = 42/186 (22%), Positives = 76/186 (40%), Gaps = 22/186 (11%)
Query: 428 AENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTA--- 484
+ + ES+ AP T++SS A P + T +++ P + E+S A P++
Sbjct: 531 SSSTTESSSAPVTSSTTESSS-APVPTPSSSTTESSSTPVTSSTTESSSAPVPTPSSSTT 589
Query: 485 ------VPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPK 538
VP S ++ +++A A S T +S+ + SS P+
Sbjct: 590 ESSSAPVPTPSSSTTESSSAPAPTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESS 649
Query: 539 SPPRQDAPPSPPSTHDAGPMPSPPHQGEKS--CPGAATTSEAA---------QIEQAPAP 587
S P P S + + P+P+P +S P ++T+E++ + AP P
Sbjct: 650 SAP-VPTPSSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVTSSTTESSSAPVP 708
Query: 588 EVGSSS 593
SS+
Sbjct: 709 TPSSST 714
Score = 44.7 bits (104), Expect = 0.001
Identities = 41/179 (22%), Positives = 70/179 (38%), Gaps = 9/179 (5%)
Query: 433 ESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPAS 492
ES+ AP T++SS A P + T +++AP + E+S +A PT +++ +S
Sbjct: 440 ESSSAPVTSSTTESSS-APVPTPSSSTTESSSAPVTSSTTESS--SAPVPTPSSSTTESS 496
Query: 493 AGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETP---KSPPRQDAPPSP 549
+ + ESS + ++A + TP + + P + AP
Sbjct: 497 SAPVTSSTTESSSAPVPTPSSSTTESSSAPAPTPSSSTTESSSAPVTSSTTESSSAPVPT 556
Query: 550 PSTHDAGPMPSPPHQG---EKSCPGAATTSEAAQIEQAPAPEVGSSSYYNMLPNAIEPS 605
PS+ +P S P +S + AP P SS+ + A PS
Sbjct: 557 PSSSTTESSSTPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPAPTPS 615
Score = 44.3 bits (103), Expect = 0.002
Identities = 50/216 (23%), Positives = 82/216 (37%), Gaps = 14/216 (6%)
Query: 428 AENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPK---GKNVAEASVA----AAT 480
+ + ES+ AP T++SS A P + T +++AP + E+S A + T
Sbjct: 489 SSSTTESSSAPVTSSTTESSS-APVPTPSSSTTESSSAPAPTPSSSTTESSSAPVTSSTT 547
Query: 481 EPTAVPASSPASAGATAAFAAESSIGATAASAGV--NATKAAASSDTPIGDKEKENETPK 538
E ++ P +P+S+ ++ +S ++SA V ++ SS P+
Sbjct: 548 ESSSAPVPTPSSSTTESSSTPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESS 607
Query: 539 SPPRQDAPPSPPSTHDAGPMPSPPHQGE-KSCPGAATTSEAAQIEQAPAPEVGSSSYYNM 597
S P AP ST ++ P E S P +S + AP P SS+ +
Sbjct: 608 SAP---APTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESS 664
Query: 598 LPNAIEPSEFLLAGLNRDVIEKEVLSQGLNVTKEET 633
PS + V S VT T
Sbjct: 665 SAPVPTPSSSTTESSSAPVTSSTTESSSAPVTSSTT 700
Score = 42.7 bits (99), Expect = 0.004
Identities = 40/166 (24%), Positives = 71/166 (42%), Gaps = 17/166 (10%)
Query: 434 STKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASA 493
+T P T SS A P + T +++AP ++ TE ++ P +P+S+
Sbjct: 311 TTPVPTPSSSTTESSSAPVPTPSSSTTESSSAPV--------TSSTTESSSAPVPTPSSS 362
Query: 494 GATAAFAAESSIGATAASAGVNATKAAASS---DTPIGDKEKENETP--KSPPRQDAPPS 548
++ A +S ++SA V ++ +SS TP + + P S + P
Sbjct: 363 TTESSSAPVTSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPV 422
Query: 549 PPSTHDAGPMPSPPHQGEKS-CPGAATTSEAAQIEQAPAPEVGSSS 593
ST ++ P E S P ++T+E++ AP P SS+
Sbjct: 423 TSSTTESSSAPVTSSTTESSSAPVTSSTTESS---SAPVPTPSSST 465
Score = 39.3 bits (90), Expect = 0.049
Identities = 36/168 (21%), Positives = 58/168 (34%), Gaps = 9/168 (5%)
Query: 429 ENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPAS 488
+N TK+ T SS +S + TT++ + + S ++ + T PA+
Sbjct: 202 DNNCGGTKSSTTTSSTSESSTTTSSTSESSTTTSSTSESSTTTSSTSESSTSSSTTAPAT 261
Query: 489 SPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENE---TPKSPPRQDA 545
T + E T S TP K+ T K+
Sbjct: 262 P-----TTTSCTKEKPTPPTTTSCTKEKPTPPHHDTTPCTKKKTTTSKTCTKKTTTPVPT 316
Query: 546 PPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSS 593
P S + + P+P+P +S A TS + AP P SS+
Sbjct: 317 PSSSTTESSSAPVPTPSSSTTES-SSAPVTSSTTESSSAPVPTPSSST 363
Score = 38.5 bits (88), Expect = 0.083
Identities = 36/174 (20%), Positives = 74/174 (41%), Gaps = 14/174 (8%)
Query: 433 ESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPAS 492
++T K++ T + T+ P P+++ V S ++ TE ++ P +S +
Sbjct: 291 DTTPCTKKKTTTSKTCTKKTTTPVPTPSSSTTESSSAPVPTPS-SSTTESSSAPVTSSTT 349
Query: 493 AGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPST 552
++A SS ++SA V ++ +SS P+ E+ + P P S +
Sbjct: 350 ESSSAPVPTPSSSTTESSSAPVTSSTTESSS-APVTSSTTESSSAPVP----TPSSSTTE 404
Query: 553 HDAGPMPSPPHQGEKSCPGAATTSEAAQ-------IEQAPAPEVGSSSYYNMLP 599
+ P+ S + S P ++T+E++ E + AP S++ + P
Sbjct: 405 SSSAPVTSSTTESS-SAPVTSSTTESSSAPVTSSTTESSSAPVTSSTTESSSAP 457
Score = 37.0 bits (84), Expect = 0.24
Identities = 28/149 (18%), Positives = 56/149 (36%), Gaps = 10/149 (6%)
Query: 460 TTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKA 519
T ++ + + + ++ +E + +S+ S+ T++ + S+ +T A A T
Sbjct: 208 TKSSTTTSSTSESSTTTSSTSESSTTTSSTSESSTTTSSTSESSTSSSTTAPATPTTTSC 267
Query: 520 AASSDTPIGDKEKENETPKSPPRQDAPPSPP---------STHDAGPMPSPPHQGEKSCP 570
TP E P +PP D P + P+P+P +S
Sbjct: 268 TKEKPTPPTTTSCTKEKP-TPPHHDTTPCTKKKTTTSKTCTKKTTTPVPTPSSSTTESSS 326
Query: 571 GAATTSEAAQIEQAPAPEVGSSSYYNMLP 599
T ++ E + AP S++ + P
Sbjct: 327 APVPTPSSSTTESSSAPVTSSTTESSSAP 355
>DYNA_MOUSE (O08788) Dynactin 1 (150 kDa dynein-associated
polypeptide) (DP-150) (DAP-150) (p150-glued)
Length = 1281
Score = 56.2 bits (134), Expect = 4e-07
Identities = 91/388 (23%), Positives = 153/388 (38%), Gaps = 51/388 (13%)
Query: 493 AGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETP-KSPPRQDAPPSPPS 551
A T+ +SS G +A AA + G K K+ T K+ R+ P P S
Sbjct: 101 ADTTSPETPDSSASKVLKREGADA---AAKTSKLRGLKPKKAPTARKTTTRRPKPTRPAS 157
Query: 552 THDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAP-------APEVGSSSYYNMLPNAIEP 604
T AGP S G S G ++SE + Q P P + S LP+ +
Sbjct: 158 TGVAGPSSSLGPSGSASA-GELSSSEPSTPAQTPLAAPIIPTPALTSPGAAPPLPSPSKE 216
Query: 605 SEFLLAGLNRDVIEKEVLSQGLNVTKEETLACLLRAGCIFAHTFEKFNAANVEAERLKAE 664
E GL V + E + L + + E A L ++ ++ E+++
Sbjct: 217 EE----GLRAQVRDLEEKLETLRLKRSEDKAKL-----------KELEKHKIQLEQVQEW 261
Query: 665 SAKHQEAAAAWEKRFDKLATQAGK----DKVYADKMIGTA-GIKIGELEDQLA------- 712
+K QE A ++R + +A + + Y ++M TA I++ L+ ++A
Sbjct: 262 KSKMQEQQADLQRRLKEARKEAKEALEAKERYMEEMADTADAIEMATLDKEMAEERAESL 321
Query: 713 -----LMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKAL 767
+KE DEL L+ K E E+ D A L E + A L L ++
Sbjct: 322 QQEVEALKERVDELTTDLEILKAEIEEKGSDGAASSYQLKQLEEQNARLKDALVRMRDLS 381
Query: 768 AEQEKKSAESLALAK---SDMEAVMQ---ATSEEIKKATETHAEALATKDAEIASQLAKI 821
+ ++++ + L + ++E V Q EE+ +A T E DA + ++ +
Sbjct: 382 SSEKQEHVKLQKLMEKKNQELEVVRQQRERLQEELSQAESTIDELKEQVDAALGAE-EMV 440
Query: 822 KSLEDELATEKAKAIEAREQAADIALDN 849
+ L D + K E RE D+ N
Sbjct: 441 EMLTDRNLNLEEKVRELRETVGDLEAMN 468
>RRB1_CANFA (Q28298) Ribosome-binding protein 1 (180 kDa ribosome
receptor) (RRp)
Length = 1534
Score = 55.1 bits (131), Expect = 9e-07
Identities = 108/547 (19%), Positives = 190/547 (33%), Gaps = 81/547 (14%)
Query: 423 KKRGGAENQAESTK-APKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATE 481
KK GA NQ + + AP + + + + + G GA A +A A
Sbjct: 630 KKAEGAPNQGKKAEGAPNQGKKAEGAQNQGKKAEGAPNQGKKADLVANQGTKAEGVAGQG 689
Query: 482 PTAVPASSPASAGATAAFAAESSIGAT----AASAGVNATKAAASSDTPIGDKEKENETP 537
A A + G + S G+ A N +K A S+ PI K +
Sbjct: 690 KKAEGAPNQGKKGEGTPNQGKKSEGSPNQGKKVDASANQSKRAESA--PIQGKNADMVQS 747
Query: 538 KSPPRQDAPPSPPSTHDAGPMPSPP----------------------HQGEKSCPGAATT 575
+ P+Q+AP S P PP ++GE +
Sbjct: 748 QEAPKQEAPAKKKSGSKKKGEPGPPDSDSPLYLPYKTLVSTVGSMVFNEGEAQRLIEILS 807
Query: 576 SEAAQIEQAPAPEVGSSSYYNMLPNAIEPSEFLLAGLNRDVIEKEVLSQGLNVTKEETLA 635
+A I+ +L +E E LLA D V L +E A
Sbjct: 808 EKAGVIQDTWHKATQKGDPVAILKRQLEEKEKLLATEQEDAA---VAKSKLREVNKELAA 864
Query: 636 CLLRAGCIFAHTFEKFNAANVEAERLKAESAKHQEAAA-------AWEKRFDKLATQAGK 688
EK AA EA+ K A+ QE A ++ + ++ GK
Sbjct: 865 -------------EKAKAAAGEAKVKKQLVAREQEITAVQARIEASYREHVKEVQQLQGK 911
Query: 689 DKVYADKMIGTAGIKIGELEDQLALMKEEADELDASLQA-------------CKKEKEQA 735
+ +++ ++ L+ + +++++ ++ + +++ K KE
Sbjct: 912 IRTLQEQLENGPNTQLARLQQENSILRDALNQATSQVESKQNTELAKLRQELSKVSKELV 971
Query: 736 EKDLIARGEALIAKESELAILCAELELVKKALAEQEKKSAESLALAKSDMEAVMQATSEE 795
EK AR E K E E ++++ + +E + A L + E TS
Sbjct: 972 EKSEAARQEEQQRKALETKTAALEKQVLQLQASHKESEEALQKRLDEVSRELCRSQTSHA 1031
Query: 796 IKKATETHAEALATKDAEIASQL----AKIKSLEDELATEKAKAIEAREQAADIALDNRE 851
+A A+ + AE+ S+L A++KS +EL+ + EAR + + + R
Sbjct: 1032 SLRADAEKAQEQQQQMAELHSKLQSSEAEVKSKSEELSGLHGQLKEARAENSQLMERIRS 1091
Query: 852 RGFYLAKDQAQHLFPNFDFSAMGVMKEITAAGLVGPDDPPLIDQNLWTATEEEEEEEEEE 911
L QA+ ++ A+ ++ +W +E E +E
Sbjct: 1092 IEALLEAGQARDT------------QDAQASRAEHQARLKELESQVWCLEKEATELKEAV 1139
Query: 912 EQEKENN 918
EQ+K N
Sbjct: 1140 EQQKVKN 1146
Score = 35.4 bits (80), Expect = 0.70
Identities = 46/194 (23%), Positives = 92/194 (46%), Gaps = 14/194 (7%)
Query: 649 EKFNAANVEAERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELE 708
E+ + + + + +AE+++ E + E + A QA +D A ++ ELE
Sbjct: 1066 EELSGLHGQLKEARAENSQLMERIRSIEALLE--AGQA-RDTQDAQASRAEHQARLKELE 1122
Query: 709 DQLALMKEEADELDASLQACKKE----KEQAEKDLIARGEALIAKESELAILCAELELVK 764
Q+ +++EA EL +++ K + +E+ K + A A A E +L L E +
Sbjct: 1123 SQVWCLEKEATELKEAVEQQKVKNNDLREKNWKAMEALASAERACEEKLRSLTQAKEESE 1182
Query: 765 KALAEQEKKSAES-LALAKSDMEAVMQATSEEIKKATETHAEALATKDA------EIASQ 817
K L+ E ++ E+ LAL + + Q+ +E +++ E E L + A ++AS+
Sbjct: 1183 KQLSLTEAQTKEALLALLPALSSSAPQSYTEWLQELREKGPELLKQRPADTDPSSDLASK 1242
Query: 818 LAKIKSLEDELATE 831
L + + ++ L E
Sbjct: 1243 LREAEETQNNLQAE 1256
>DYNA_RAT (P28023) Dynactin 1 (150 kDa dynein-associated
polypeptide) (DP-150) (DAP-150) (p150-glued)
Length = 1280
Score = 55.1 bits (131), Expect = 9e-07
Identities = 86/355 (24%), Positives = 141/355 (39%), Gaps = 35/355 (9%)
Query: 519 AAASSDTPIGDKEKENETP-KSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSE 577
AAA + G K K+ T K+ R+ P P ST AGP S G S G ++SE
Sbjct: 124 AAAKTSKLRGLKPKKAPTARKTTTRRPKPTRPASTGVAGPSSSLGPSGSASA-GELSSSE 182
Query: 578 AAQIEQAP-------APEVGSSSYYNMLPNAIEPSEFLLAGLNRDVIEKEVLSQGLNVTK 630
+ Q P P + S LP+ + E GL V + E + L + +
Sbjct: 183 PSTPAQTPLAAPIIPTPALTSPGAAPPLPSPSKEEE----GLRDQVRDLEEKLETLRLKR 238
Query: 631 EETLACLLRAGCIFAHTFE---------KFNAANVEAERLKAESAKHQEAAAAWEKRFDK 681
E A L + H + K + +R E+ + +EA A E+ ++
Sbjct: 239 SEDKAKLKE---LEKHKIQLEQVQEWKSKMQEQQADLQRRLKEAKEAKEALEAKERYMEE 295
Query: 682 LATQAGK-DKVYADKMIGTAGIKIGELEDQLALMKEEADELDASLQACKKEKEQAEKDLI 740
+A A + DK + A + L+ ++ +KE DEL L+ K E E+ D
Sbjct: 296 MADTADAIEMATLDKEM--AEERAESLQQEVEALKERVDELTTDLEILKAEIEEKGSDGA 353
Query: 741 ARGEALIAKESELAILCAELELVKKALAEQEKKSAESLALAK---SDMEAVMQ---ATSE 794
A L E + A L L ++ + ++++ + L + ++E V Q E
Sbjct: 354 ASSYQLKQLEEQNARLKDALVRMRDLSSSEKQEHVKLQKLMEKKNQELEVVRQQRERLQE 413
Query: 795 EIKKATETHAEALATKDAEIASQLAKIKSLEDELATEKAKAIEAREQAADIALDN 849
E+ +A T E DA + ++ ++ L D + K E RE D+ N
Sbjct: 414 ELSQAESTIDELKEQVDAALGAE-EMVEMLTDRNLNLEEKVRELRETVGDLEAMN 467
>KINL_LEICH (P46865) Kinesin-like protein K39 (Fragment)
Length = 955
Score = 54.3 bits (129), Expect = 1e-06
Identities = 69/275 (25%), Positives = 116/275 (42%), Gaps = 38/275 (13%)
Query: 590 GSSSYYNMLPNAIEPSEFLL----AGLNRDVIEKEVLSQGLNVTKEETLACLLRAGCIFA 645
G+ + Y++ P F+L G ++ + V LN EETL+ L A +
Sbjct: 334 GAKAQYSVAPFRDSKLTFILKDSLGGNSKTFMIATVSPSALNY--EETLSTLRYA----S 387
Query: 646 HTFEKFNAANVEAERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIG 705
+ N A V E + + E+ D AG D Y +
Sbjct: 388 RARDIVNVAQVN------EDPRARRIRELEEQMEDMRQAMAGGDPAY-----------VS 430
Query: 706 ELEDQLALMKEEADELDASLQACKKEKE--QAEKDLIARGEALIAKESELAILCAELELV 763
EL+ +LAL++ EA + A LQA ++E+E Q ++ L+ E A++SEL A L+
Sbjct: 431 ELKKKLALLESEAQKRAADLQALEREREHNQVQERLLRATE---AEKSELESRAAALQEE 487
Query: 764 KKALAEQEKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIKS 823
A Q K K + + +E+ K ++ KDAEIAS+ K++S
Sbjct: 488 MTATRRQADKMQALNLRLKEEQARKERELLKEMAKKDAALSKVRRRKDAEIASEREKLES 547
Query: 824 LEDELATEKAK------AIEAREQAADIALDNRER 852
+L E+ + A++ ++ AL++ ER
Sbjct: 548 TVAQLEREQREREVALDALQTHQRKLQEALESSER 582
Score = 46.6 bits (109), Expect = 3e-04
Identities = 46/187 (24%), Positives = 82/187 (43%), Gaps = 18/187 (9%)
Query: 670 EAAAAWEKRFDKLATQAGKDKVYADKMIGTAGI----------KIGELEDQLALMKEEAD 719
E A WE + A A +D+ A ++ A ++ LE QL +E A
Sbjct: 658 ELATEWEDALRERAL-AERDEAAAAELDAAASTSQNARESACERLTSLEQQLRESEERAA 716
Query: 720 ELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKALAEQEKKSAESLA 779
EL + L+A K AE+D R E +L E E LA Q + +A +
Sbjct: 717 ELASQLEATAAAKSSAEQD---RENTRATLEQQL----RESEARAAELASQLEATAAAKM 769
Query: 780 LAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIKSLEDELATEKAKAIEAR 839
A+ D E ++++ + E AE + ++ A++++ + E AT + + ++
Sbjct: 770 SAEQDRENTRATLEQQLRDSEERAAELASQLESTTAAKMSAEQDRESTRATLEQQLRDSE 829
Query: 840 EQAADIA 846
E+AA++A
Sbjct: 830 ERAAELA 836
Score = 41.6 bits (96), Expect = 0.010
Identities = 67/290 (23%), Positives = 118/290 (40%), Gaps = 31/290 (10%)
Query: 579 AQIEQAPAPEVGSSSYYNMLPNAIEPSEFLLAGLNRDVIEKEVLSQGLNVTKEETLACLL 638
A+ ++A A E+ +++ + NA E + L L + + E E + L E T A
Sbjct: 673 AERDEAAAAELDAAASTSQ--NARESACERLTSLEQQLRESEERAAELASQLEATAAAKS 730
Query: 639 RAGCIFAHTFEKFNAANVEAERLKAESAKHQEAAAA--WEKRFDKLATQAGKDKVYAD-- 694
A +T E+E AE A EA AA D+ T+A ++ D
Sbjct: 731 SAEQDRENTRATLEQQLRESEARAAELASQLEATAAAKMSAEQDRENTRATLEQQLRDSE 790
Query: 695 --------KMIGTAGIKIGE----------LEDQLALMKEEADELDASLQACKKEKEQAE 736
++ T K+ LE QL +E A EL + L++ K AE
Sbjct: 791 ERAAELASQLESTTAAKMSAEQDRESTRATLEQQLRDSEERAAELASQLESTTAAKMSAE 850
Query: 737 KDLIARGEALIAKESELAILCAELELVKKALAEQEKKSAESLALAKSDMEAVMQATSEEI 796
+D R E +L E E LA Q + + + A+ D E+ +++
Sbjct: 851 QD---RESTRATLEQQL----RESEERAAELASQLESTTAAKMSAEQDRESTRATLEQQL 903
Query: 797 KKATETHAEALATKDAEIASQLAKIKSLEDELATEKAKAIEAREQAADIA 846
+ + E AE + +A A++ + + E+ A + + ++ E+AA++A
Sbjct: 904 RDSEERAAELASQLEATAAAKSSAEQDRENTRAALEQQLRDSEERAAELA 953
>MST1_DROHY (Q08695) Axoneme-associated protein mst101(1)
Length = 344
Score = 53.9 bits (128), Expect = 2e-06
Identities = 65/203 (32%), Positives = 92/203 (45%), Gaps = 31/203 (15%)
Query: 658 AERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKEE 717
AE+ K A +E AA +K K A A K+K A+K K E A ++E
Sbjct: 72 AEKKKCAEAAKKEKEAAEKK---KCAEAAKKEKEAAEKK------KCAEA----AKKEKE 118
Query: 718 ADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKALAE-------- 769
A E +A KKEKE AEK A A KE+ CAE +K AE
Sbjct: 119 AAEKKKCAEAAKKEKEAAEKKKCAEA-AKKEKEAAEKKKCAEAAKKEKEAAEKKKCAEAA 177
Query: 770 QEKKSAESLALAKSDMEAV-----MQATSEEIKKATETHAEALATKDAEIA-SQLAKIKS 823
Q+KK AE LAK + EA +A +E + A + E A K+ E A + + ++
Sbjct: 178 QKKKCAE---LAKKEQEAAEKKKCAEAAKKEKEAAEKKKCEERAKKEKEAAEKKKCEERA 234
Query: 824 LEDELATEKAKAIEAREQAADIA 846
+++ A EK K EA ++ + A
Sbjct: 235 KKEKEAAEKKKCAEAAKKEKEAA 257
Score = 46.2 bits (108), Expect = 4e-04
Identities = 44/148 (29%), Positives = 67/148 (44%), Gaps = 12/148 (8%)
Query: 710 QLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKALAE 769
+ A ++EA E +A KKEKE AEK A A KE+ CAE +K AE
Sbjct: 63 EAAKKEKEAAEKKKCAEAAKKEKEAAEKKKCAEA-AKKEKEAAEKKKCAEAAKKEKEAAE 121
Query: 770 ---------QEKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAK 820
+EK++AE A++ + A ++ +A + EA K A+Q K
Sbjct: 122 KKKCAEAAKKEKEAAEKKKCAEAAKKEKEAAEKKKCAEAAKKEKEAAEKKKCAEAAQKKK 181
Query: 821 IKSL--EDELATEKAKAIEAREQAADIA 846
L +++ A EK K EA ++ + A
Sbjct: 182 CAELAKKEQEAAEKKKCAEAAKKEKEAA 209
Score = 45.1 bits (105), Expect = 9e-04
Identities = 56/195 (28%), Positives = 84/195 (42%), Gaps = 20/195 (10%)
Query: 658 AERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKEE 717
AE+ K A +E AA +K K A A K+K A+K + + + LA ++E
Sbjct: 136 AEKKKCAEAAKKEKEAAEKK---KCAEAAKKEKEAAEKKKCAEAAQKKKCAE-LAKKEQE 191
Query: 718 ADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKALAEQEKKSAES 777
A E +A KKEKE AEK + E KE E A KK E+ KK E+
Sbjct: 192 AAEKKKCAEAAKKEKEAAEK---KKCEERAKKEKEAA--------EKKKCEERAKKEKEA 240
Query: 778 LALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIKSLEDELATEKAKAIE 837
K A + + E KK E A K AE+A + ++ E + +KA
Sbjct: 241 AEKKKCAEAAKKEKEAAEKKKCAEA---AQKKKCAELAKKAK--EAAEKKKCAKKAGEKG 295
Query: 838 AREQAADIALDNRER 852
+++ +D N ++
Sbjct: 296 SKQSGSDKGKKNGKK 310
>M24_STRPY (P12379) M protein, serotype 24 precursor
Length = 539
Score = 53.9 bits (128), Expect = 2e-06
Identities = 106/449 (23%), Positives = 172/449 (37%), Gaps = 54/449 (12%)
Query: 379 LEGADLSRTPKYLEDMNFTNAELLKARERRMARFSPKFNTEGDAKKRGGAENQAESTKAP 438
L+ +DLS K L+D N E L +AK++ +++ S KA
Sbjct: 71 LKNSDLSFNNKALKDHNDELTEELS-----------------NAKEKLRKNDKSLSEKAS 113
Query: 439 KRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNV-AEASVAAATEPTAVPASSPASAGATA 497
K + L +D + GA + A + K K + AE + AA + A A +TA
Sbjct: 114 KIQELEARKADLEKALEGAMNFSTADSAKIKTLEAEKAALAARKADLEKALEGAMNFSTA 173
Query: 498 AFAAESSIGATAASAGVN------ATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPS 551
A ++ A A+ A + A + T K K E K+ +
Sbjct: 174 DSAKIKTLEAEKAALEARQAELEKALEGAMNFSTADSAKIKTLEAEKAALAARKADLEKA 233
Query: 552 THDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSSYYNMLPNAIEPSEFLLAG 611
A + K+ EA Q E A E G+ ++ I+ E A
Sbjct: 234 LEGAMNFSTADSAKIKTLEAEKAALEARQAELEKALE-GAMNFSTADSAKIKTLEAEKAA 292
Query: 612 LNRDVIEKEVLSQGLNVTKEETLACLLRAGCIFAHTFEKFNAANVEAERLKAESAKHQEA 671
L + + E SQ LN ++ LR + +K +EAE K E
Sbjct: 293 LEAEKADLEHQSQVLNANRQS-----LRRDLDASREAKK----QLEAEHQKLEEQNKISE 343
Query: 672 AAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKEEADELDASLQACKKE 731
A+ R D A++ K ++ A+ +LE+Q + + L L A ++
Sbjct: 344 ASRQSLRRDLDASREAKKQLEAEHQ---------KLEEQNKISEASRQSLRRDLDASREA 394
Query: 732 KEQAEKDLIARGEALIAKESELAILCAELELVKKALAEQEKKSAESLALAKSDMEAVMQA 791
K+Q EK AL S+L A LE + K L E +K + + A ++ +EA +A
Sbjct: 395 KKQVEK-------ALEEANSKL----AALEKLNKELEESKKLTEKEKAELQAKLEAEAKA 443
Query: 792 TSEEIKKATETHAEALATKDAEIASQLAK 820
E++ K E A+ A K ++ + AK
Sbjct: 444 LKEKLAKQAEELAKLRAGKASDSQTPDAK 472
>RL22_DROME (P50887) 60S ribosomal protein L22
Length = 299
Score = 53.5 bits (127), Expect = 2e-06
Identities = 52/172 (30%), Positives = 74/172 (42%), Gaps = 12/172 (6%)
Query: 422 AKKRGGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATE 481
AK G A + A K+ A++ KP A+ AAA KNV +AS AA
Sbjct: 5 AKTNKGDTKTAAAKPAEKKAAPAAAAAKGKVEKPKAEAAKPAAA-AAKNVKKASEAAKDV 63
Query: 482 PTAVPASSPASAGATAAFAAESSIGA-----TAASAGVNATKAAASSDTPIGDKEKENET 536
A A+ PA+A AA A +S A AA+ +A AAA + +K T
Sbjct: 64 KAAAAAAKPAAAKPAAAKPAAASKDAGKKAPAAAAPKKDAKAAAAPAPAKAAPAKKAAST 123
Query: 537 P-KSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAP 587
P +PP + A P+ + A P+P + P A + + + APAP
Sbjct: 124 PAAAPPAKKAAPAKAAA-PAAAAPAP----AAAAPAVAKPAPKPKAKAAPAP 170
Score = 40.4 bits (93), Expect = 0.022
Identities = 38/131 (29%), Positives = 52/131 (39%), Gaps = 14/131 (10%)
Query: 421 DAKKRGGAENQAESTKAPKRRRLTK--ASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAA 478
+ KK A ++ A + K A+ A SK + AAAAPK A A+ A
Sbjct: 52 NVKKASEAAKDVKAAAAAAKPAAAKPAAAKPAAASKDAGKKAPAAAAPKKDAKAAAAPAP 111
Query: 479 ATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPK 538
A A A+S +A A AA + A AA+A A A A + P
Sbjct: 112 AKAAPAKKAASTPAAAPPAKKAAPAKAAAPAAAAPAPAAAAPAVA------------KPA 159
Query: 539 SPPRQDAPPSP 549
P+ A P+P
Sbjct: 160 PKPKAKAAPAP 170
Score = 40.4 bits (93), Expect = 0.022
Identities = 49/167 (29%), Positives = 66/167 (39%), Gaps = 25/167 (14%)
Query: 450 AGTSKPGAQPT-TAAAAPKGKNVAEASVAA---ATEPTAVPASSPASAGATAAFAAESSI 505
A T+K T TAAA P K A A+ AA +P A A A+A A+E++
Sbjct: 2 APTAKTNKGDTKTAAAKPAEKKAAPAAAAAKGKVEKPKAEAAKPAAAAAKNVKKASEAAK 61
Query: 506 GATAASAGVNATKAA--------ASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDA-- 555
AA+A A AA A++ G K PK + A P+P A
Sbjct: 62 DVKAAAAA--AKPAAAKPAAAKPAAASKDAGKKAPAAAAPKKDAKAAAAPAPAKAAPAKK 119
Query: 556 ---GPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSSYYNMLP 599
P +PP +K+ P A AA APAP + + P
Sbjct: 120 AASTPAAAPP--AKKAAPAKAAAPAAA----APAPAAAAPAVAKPAP 160
>ALGP_PSEAE (P15276) Transcriptional regulatory protein algP
(Alginate regulatory protein algR3)
Length = 352
Score = 53.5 bits (127), Expect = 2e-06
Identities = 58/214 (27%), Positives = 86/214 (40%), Gaps = 15/214 (7%)
Query: 403 KARERRMARFSPKFNTEGDAK---KRGGAENQAESTKAPKRRRLTKASSDAGTSKPGAQP 459
KA E R A+ + K + AK K A+ A+ P + K ++ +KP A+P
Sbjct: 128 KALESRKAKPATKPAAKAAAKPAVKTVAAKPAAKPAAKPAAKPAAKPAAKTAAAKPAAKP 187
Query: 460 TTAAAA-PKGKNVAEASVAA-----ATEPTAVPASSPASAGATAAFAAESSIGATAASAG 513
T AA P K A+ + A A +P A PA+ PA+ A A AA+ + A
Sbjct: 188 TAKPAAKPAAKPAAKTAAAKPAAKPAAKPVAKPAAKPAAKTAAAKPAAKPAAKPVAKPTA 247
Query: 514 VNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSP---PHQGEKSCP 570
A K AA+ + + P + A P + A P P P +
Sbjct: 248 KPAAKTAAAKPAAKPAAKPAAKPAAKPVAKSAAAKPAAKPAAKPAAKPAAKPAAKPVAAK 307
Query: 571 GAAT---TSEAAQIEQAPAPEVGSSSYYNMLPNA 601
AAT T+ AA+ P+ +SS + P A
Sbjct: 308 PAATKPATAPAAKPAATPSAPAAASSAASATPAA 341
Score = 48.5 bits (114), Expect = 8e-05
Identities = 40/156 (25%), Positives = 65/156 (41%), Gaps = 6/156 (3%)
Query: 422 AKKRGGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAA-PKGKNVAEASVAA-A 479
A K A+ A+ P + K ++ +KP A+P A P K A+ + A A
Sbjct: 200 AAKTAAAKPAAKPAAKPVAKPAAKPAAKTAAAKPAAKPAAKPVAKPTAKPAAKTAAAKPA 259
Query: 480 TEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKS 539
+P A PA+ PA+ + AA+ + A A A K AA P+ K + +
Sbjct: 260 AKPAAKPAAKPAAKPVAKSAAAKPAAKPAAKPAAKPAAKPAAK---PVAAKPAATKPATA 316
Query: 540 P-PRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAAT 574
P + A PS P+ + +P + P +A+
Sbjct: 317 PAAKPAATPSAPAAASSAASATPAAGSNGAAPTSAS 352
Score = 47.0 bits (110), Expect = 2e-04
Identities = 36/108 (33%), Positives = 51/108 (46%), Gaps = 4/108 (3%)
Query: 418 TEGDAKKRGGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPTT-AAAAPKGKNVAE--A 474
T A K A+ A+ P + K + + +KP A+P AA P K A+ A
Sbjct: 246 TAKPAAKTAAAKPAAKPAAKPAAKPAAKPVAKSAAAKPAAKPAAKPAAKPAAKPAAKPVA 305
Query: 475 SVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAAS 522
+ AAT+P PA+ PA A +A AA S+ AT A+ A +AS
Sbjct: 306 AKPAATKPATAPAAKPA-ATPSAPAAASSAASATPAAGSNGAAPTSAS 352
Score = 34.7 bits (78), Expect = 1.2
Identities = 51/225 (22%), Positives = 74/225 (32%), Gaps = 25/225 (11%)
Query: 379 LEGADLSRTPKYLEDMNFTNAELLKARERRMARFSPKFNTEGDAKKRGGAENQAESTK-- 436
LEGA + L D A+L K R + + DA K G + QA++ +
Sbjct: 26 LEGA----CKQALVDSEKLLAKLEKQRGKAQEKLHKARTKLQDAAKAGKTKAQAKARETI 81
Query: 437 --------------APKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEP 482
A R + D S AQ GK + AT+P
Sbjct: 82 SDLEEALDTLKARQADTRTYIVGLKRDVQESLKLAQGVGKVKEAAGKALESRKAKPATKP 141
Query: 483 TAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPR 542
A A+ PA A AA+ + A A A K AA+ + + P
Sbjct: 142 AAKAAAKPAVKTVAAKPAAKPAAKPAAKPAAKPAAKTAAAKPAAKPTAKPAAKPAAKPAA 201
Query: 543 QDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAP 587
+ A P + A P+ P + P A T + + A P
Sbjct: 202 KTAAAKPAAKPAAKPVAKPAAK-----PAAKTAAAKPAAKPAAKP 241
>XP2_XENLA (P17437) Skin secretory protein xP2 precursor (APEG
protein)
Length = 439
Score = 52.4 bits (124), Expect = 6e-06
Identities = 50/177 (28%), Positives = 65/177 (36%), Gaps = 10/177 (5%)
Query: 419 EGDAKKRGGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVA------ 472
EG+A AE +A + + A +A P A A +G+ A
Sbjct: 125 EGEAPAPAPAEGEAPAPAPAEGEAPAPAEGEAPAPAPAEVEAPAPAPAEGEAPAPAPAEG 184
Query: 473 EASVAAATEPTA-VPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKE 531
EA A E A PA + A A A AE A A + G A A + P
Sbjct: 185 EAPAPAPAEGEAPAPAPAEGEAPAPAPAPAEGEAPAPAPAEGEAPAPAPAEGEAP-APAP 243
Query: 532 KENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIE-QAPAP 587
E E P P + P+P P P+P +GE P A A E +APAP
Sbjct: 244 AEGEAPAPAPAEGEAPAPAPAEGEAPAPAPA-EGEAPAPAPAEGEAPAPAEGEAPAP 299
Score = 49.7 bits (117), Expect = 4e-05
Identities = 52/177 (29%), Positives = 66/177 (36%), Gaps = 9/177 (5%)
Query: 419 EGDAKKRGGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAA 478
EG+A AE +A + + A ++ P P A EA A
Sbjct: 173 EGEAPAPAPAEGEAPAPAPAEGEAPAPAPAEGEAPAPAPAPAEGEAPAPAPAEGEAPAPA 232
Query: 479 ATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPK 538
E A PA +PA A A AE A A + G A A + P E E P
Sbjct: 233 PAEGEA-PAPAPAEGEAPAPAPAEGEAPAPAPAEGEAPAPAPAEGEAP-APAPAEGEAP- 289
Query: 539 SPPRQDAP-PSPPSTHDAGPMPS---PPHQGEKSCPGAATTSEAAQIEQAPAP-EVG 590
+P +AP P+P P P+ P E P AA +E APAP EVG
Sbjct: 290 APAEGEAPAPAPAEGEAPAPAPAEGGAPSPAEGGAP-AAAPAEGGAPAPAPAPVEVG 345
Score = 49.3 bits (116), Expect = 5e-05
Identities = 51/183 (27%), Positives = 67/183 (35%), Gaps = 10/183 (5%)
Query: 419 EGDAKKRGGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASV-- 476
+G A AE A + + A ++ G P A A +G+ A A
Sbjct: 77 DGGAPAPAPAEGGAPAPAPAEGGAPAPAPAEGGAPAPAEGGAPAPAPAEGEAPAPAPAEG 136
Query: 477 -AAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENE 535
A A P A +PA A A AE A A + G A A + P E E
Sbjct: 137 EAPAPAPAEGEAPAPAEGEAPAPAPAEVEAPAPAPAEGEAPAPAPAEGEAP-APAPAEGE 195
Query: 536 TPKSPPRQDAPPSPPSTHDAGPMPSP-PHQGEKSCP----GAATTSEAAQIE-QAPAPEV 589
P P + P+P G P+P P +GE P G A A+ E APAP
Sbjct: 196 APAPAPAEGEAPAPAPAPAEGEAPAPAPAEGEAPAPAPAEGEAPAPAPAEGEAPAPAPAE 255
Query: 590 GSS 592
G +
Sbjct: 256 GEA 258
Score = 47.0 bits (110), Expect = 2e-04
Identities = 53/182 (29%), Positives = 66/182 (36%), Gaps = 24/182 (13%)
Query: 419 EGDAKKRGGAENQAESTKAPKRRRLTKASSDAGTSKPG-AQPTTAAAAPKGKNVAEASVA 477
E A GGA A + A A +D G P A+ A AP AE A
Sbjct: 53 EAPAPAEGGAPAPAPAEGAEP------APADGGAPAPAPAEGGAPAPAP-----AEGG-A 100
Query: 478 AATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEK----- 532
A P A +PA GA A AE A A + G A A + P + +
Sbjct: 101 PAPAPAEGGAPAPAEGGAPAPAPAEGEAPAPAPAEGEAPAPAPAEGEAPAPAEGEAPAPA 160
Query: 533 --ENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVG 590
E E P P + P+P P P+P +GE P A E APAP G
Sbjct: 161 PAEVEAPAPAPAEGEAPAPAPAEGEAPAPAPA-EGEAPAPAPA---EGEAPAPAPAPAEG 216
Query: 591 SS 592
+
Sbjct: 217 EA 218
Score = 45.4 bits (106), Expect = 7e-04
Identities = 41/148 (27%), Positives = 52/148 (34%), Gaps = 12/148 (8%)
Query: 446 ASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTA---VPASSPASAGATAAFAAE 502
A ++ G P A A +G A A A A PA +PA GA A AE
Sbjct: 38 APAEGGAPAPAPAEGEAPAPAEGGAPAPAPAEGAEPAPADGGAPAPAPAEGGAPAPAPAE 97
Query: 503 SSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPP 562
A A + G A + P E E P P + P+P P P+
Sbjct: 98 GGAPAPAPAEGGAPAPAEGGAPAPA---PAEGEAPAPAPAEGEAPAPAPAEGEAPAPA-- 152
Query: 563 HQGEKSCPGAATTSEAAQIE---QAPAP 587
+GE P A A +APAP
Sbjct: 153 -EGEAPAPAPAEVEAPAPAPAEGEAPAP 179
>GOA4_HUMAN (Q13439) Golgi autoantigen, golgin subfamily A member 4
(Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (72.1
protein)
Length = 2230
Score = 52.4 bits (124), Expect = 6e-06
Identities = 38/138 (27%), Positives = 67/138 (48%), Gaps = 14/138 (10%)
Query: 703 KIGELEDQLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELEL 762
++ E E+++ ++ + + A L+ KKE E + ++ E L A E L E E
Sbjct: 1579 RVKEAEEKILTLENQVYSMKAELETKKKELEHVNLSVKSKEEELKALEDRL-----ESES 1633
Query: 763 VKKALAEQEKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIK 822
K LAE ++K+ + +A K + + M+ E+ KK TE+H L TK E +
Sbjct: 1634 AAK-LAELKRKAEQKIAAIKKQLLSQMEEKEEQYKKGTESHLSELNTKLQE--------R 1684
Query: 823 SLEDELATEKAKAIEARE 840
E + EK K++E+ +
Sbjct: 1685 EREVHILEEKLKSVESSQ 1702
Score = 39.3 bits (90), Expect = 0.049
Identities = 47/213 (22%), Positives = 92/213 (43%), Gaps = 20/213 (9%)
Query: 657 EAERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKE 716
+ E L+ + K + AA + +K + A K + K+ E+++Q+ +++
Sbjct: 397 QGEELREQKEKSERAAF---EELEKALSTAQKTEEARRKLKA-------EMDEQIKTIEK 446
Query: 717 EADELDASLQACKKEKEQAEKDLIARGE----ALIAKESELAILCAELELVKKALA-EQE 771
++E SLQ +Q D++ + A + K E + E EL KK E+E
Sbjct: 447 TSEEERISLQQELSRVKQEVVDVMKKSSEEQIAKLQKLHEKELARKEQELTKKLQTRERE 506
Query: 772 KKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIKSLEDELATE 831
+ +AL KS E ++ + E+ ++ + E K A + K++ L+ E T
Sbjct: 507 FQEQMKVALEKSQSE-YLKISQEKEQQESLALEELELQKKAILTESENKLRDLQQEAETY 565
Query: 832 KAKAIEAREQAADIALDNRERGFYLAKDQAQHL 864
+ + +E +N+ + +KD A HL
Sbjct: 566 RTRILELESSLEKSLQENKNQ----SKDLAVHL 594
Score = 36.2 bits (82), Expect = 0.41
Identities = 39/155 (25%), Positives = 66/155 (42%), Gaps = 7/155 (4%)
Query: 701 GIKIGELEDQLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAEL 760
G + LE +K + + L + + KEQ L + EAL + +L EL
Sbjct: 275 GTSVKTLETLQQRVKRQENLLKRCKETIQSHKEQCTL-LTSEKEAL---QEQLDERLQEL 330
Query: 761 ELVKKALAEQEKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAK 820
E +K ++ K L AK+ +E + Q I + E L K+ EIA ++
Sbjct: 331 EKIKDLHMAEKTKLITQLRDAKNLIEQLEQDKGMVIAETKRQMHETLEMKEEEIAQLRSR 390
Query: 821 IKSLE---DELATEKAKAIEAREQAADIALDNRER 852
IK + +EL +K K+ A + + AL ++
Sbjct: 391 IKQMTTQGEELREQKEKSERAAFEELEKALSTAQK 425
Score = 35.8 bits (81), Expect = 0.54
Identities = 56/243 (23%), Positives = 100/243 (41%), Gaps = 19/243 (7%)
Query: 612 LNRDVIEKEVLSQGLNVTKEETLACLLRAGCIFAHTFEKFNAANVEAERLKAESAKHQEA 671
L R++ EKE L Q + KEET++ C + K A + ++ QE
Sbjct: 1741 LQRNLTEKEKLLQRVGQEKEETVSSHFEMRCQYQERLIKLEHAEAKQHEDQSMIGHLQEE 1800
Query: 672 AAAWEKRFDKLATQ-----AGKDKVYA----DKMIGTAGIKIGELEDQLALMKEEADELD 722
K++ + Q GK+ + A + + + E E +++++ ELD
Sbjct: 1801 LEEKNKKYSLIVAQHVEKEGGKNNIQAKQNLENVFDDVQKTLQEKELTCQILEQKIKELD 1860
Query: 723 ASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKALAEQEKKSAESLALAK 782
+ L K+ ++L ++ E L A + ++ EL+++ E+KS L K
Sbjct: 1861 SCLVRQKEVHRVEMEELTSKYEKLQALQ-QMDGRNKPTELLEE---NTEEKSKSHLVQPK 1916
Query: 783 --SDMEAVMQATSEEIKKA-TETHAEALATKDAEIASQLAKI-KSLEDELATEKAKAIEA 838
S+MEA Q E K A E + L + + L + K + EL K + +
Sbjct: 1917 LLSNMEA--QHNDLEFKLAGAEREKQKLGKEIVRLQKDLRMLRKEHQQELEILKKEYDQE 1974
Query: 839 REQ 841
RE+
Sbjct: 1975 REE 1977
Score = 35.8 bits (81), Expect = 0.54
Identities = 46/213 (21%), Positives = 90/213 (41%), Gaps = 32/213 (15%)
Query: 657 EAERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKE 716
++E LK K Q+ + A E+ ++ ++ + K+ +L+ + +
Sbjct: 519 QSEYLKISQEKEQQESLALEEL-----------ELQKKAILTESENKLRDLQQEAETYRT 567
Query: 717 EADELDASLQACKKEKEQAEKDLIARGEALIAKES-ELAILC----AELELVKKALAEQE 771
EL++SL+ +E + KDL EA K + E+ ++ ELE +K +Q+
Sbjct: 568 RILELESSLEKSLQENKNQSKDLAVHLEAEKNKHNKEITVMVEKHKTELESLKH---QQD 624
Query: 772 KKSAESLALAKSDMEAVMQATSEEIKKATET-----------HAEALATKDAE-IASQLA 819
E L + K + M+ E+ ++ ET H E + K E + +
Sbjct: 625 ALWTEKLQVLKQQYQTEMEKLREKCEQEKETLLKDKEIIFQAHIEEMNEKTLEKLDVKQT 684
Query: 820 KIKSLEDELATEKAKAIEAREQAADIALDNRER 852
+++SL EL +E KA E+ + D ++
Sbjct: 685 ELESLSSEL-SEVLKARHKLEEELSVLKDQTDK 716
Score = 34.3 bits (77), Expect = 1.6
Identities = 30/137 (21%), Positives = 62/137 (44%), Gaps = 3/137 (2%)
Query: 703 KIGELEDQLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELEL 762
++ E E+Q+ MK + + L +A +KE ++ + + + EL+ + L
Sbjct: 1298 QLEEKENQIKSMKADIESLVTEKEALQKEGGNQQQAASEKESCITQLKKELSENINAVTL 1357
Query: 763 VKKALAEQEKKSAESLALAKSDMEAVMQ--ATSEEIKKATETHAEALATKDAEIASQLAK 820
+K+ L E +K SL+ +D+ +Q + E + A + + + E+ Q+
Sbjct: 1358 MKEELKE-KKVEISSLSKQLTDLNVQLQNSISLSEKEAAISSLRKQYDEEKCELLDQVQD 1416
Query: 821 IKSLEDELATEKAKAIE 837
+ D L+ EK A+E
Sbjct: 1417 LSFKVDTLSKEKISALE 1433
>CALD_CHICK (P12957) Caldesmon (CDM)
Length = 771
Score = 52.4 bits (124), Expect = 6e-06
Identities = 64/221 (28%), Positives = 104/221 (46%), Gaps = 32/221 (14%)
Query: 657 EAERLKAESAKHQEAAAAWEKRFDK--------LATQAGKDKVYADKMIGTAGIKIGE-- 706
E E+ K + + A EK DK LA A D T + GE
Sbjct: 186 EEEKPKEVPTEENQVDVAVEKSTDKEEVVETKTLAVNAEND---------TNAMLEGEQS 236
Query: 707 LEDQLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKA 766
+ D KEEA++ L+A +KE+ +AE++ A E A+E + A E ++A
Sbjct: 237 ITDAADKEKEEAEKEREKLEAEEKERLKAEEEKKAAEEKQKAEEEKKAA-----EERERA 291
Query: 767 LAEQEKKSAESLALAKSDMEAVM-----QATSEEIKKATETHAEALATKDAEIASQLAKI 821
AE+EK++AE AK++ E +A +EE +KA E A+ A ++ + A + AK
Sbjct: 292 KAEEEKRAAEERERAKAEEERKAAEERERAKAEEERKAAEERAK--AEEERKAAEERAKA 349
Query: 822 KSLEDELATEKAKAIEAREQAADIALDNRERGFYLAKDQAQ 862
+ E + A E+AKA + R+ A + E A+++A+
Sbjct: 350 EE-ERKAAEERAKAEKERKAAEERERAKAEEEKRAAEEKAR 389
Score = 51.2 bits (121), Expect = 1e-05
Identities = 52/188 (27%), Positives = 94/188 (49%), Gaps = 18/188 (9%)
Query: 658 AERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKEE 717
A++ K E+ K +E A EK +A ++K A++ + E E + A +E
Sbjct: 241 ADKEKEEAEKEREKLEAEEKE----RLKAEEEKKAAEEK------QKAEEEKKAAEERER 290
Query: 718 ADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKALAEQEKKSAES 777
A + A ++E+ +AE++ A E AK E E +A AE+E+K+AE
Sbjct: 291 AKAEEEKRAAEERERAKAEEERKAAEERERAKAEEERKAAEE-----RAKAEEERKAAEE 345
Query: 778 LALAKSDMEAVMQ-ATSEEIKKATETHAEALATKDAEIASQLAKIKS--LEDELATEKAK 834
A A+ + +A + A +E+ +KA E A A ++ A + A++++ L+++ E+ K
Sbjct: 346 RAKAEEERKAAEERAKAEKERKAAEERERAKAEEEKRAAEEKARLEAEKLKEKKKMEEKK 405
Query: 835 AIEAREQA 842
A E + QA
Sbjct: 406 AQEEKAQA 413
Score = 50.1 bits (118), Expect = 3e-05
Identities = 60/191 (31%), Positives = 91/191 (47%), Gaps = 30/191 (15%)
Query: 657 EAERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKE 716
E ERLKAE +E AA EK+ + +A +++ A E E + A +E
Sbjct: 259 EKERLKAE----EEKKAAEEKQKAEEEKKAAEERERAK----------AEEEKRAAEERE 304
Query: 717 EADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKALAEQEKKSAE 776
A + A ++E+ +AE++ A E A+E A AE ++A AE+E+K+AE
Sbjct: 305 RAKAEEERKAAEERERAKAEEERKAAEERAKAEEERKA---AE----ERAKAEEERKAAE 357
Query: 777 SLALAKSDMEAVMQ---ATSEEIKKATETHAEALATKDAEIASQLAKIKSLEDELATEKA 833
A A+ + +A + A +EE K+A E A A K E K K E + EKA
Sbjct: 358 ERAKAEKERKAAEERERAKAEEEKRAAEEKARLEAEKLKE------KKKMEEKKAQEEKA 411
Query: 834 KAIEAREQAAD 844
+A R+Q D
Sbjct: 412 QANLLRKQEED 422
Score = 42.4 bits (98), Expect = 0.006
Identities = 66/284 (23%), Positives = 117/284 (40%), Gaps = 19/284 (6%)
Query: 649 EKFNAANVEAERLKAE----SAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKI 704
E+ A E ER KAE +A+ +E A A E+R K A + + K ++ K
Sbjct: 279 EEEKKAAEERERAKAEEEKRAAEERERAKAEEER--KAAEERERAKAEEERKAAEERAKA 336
Query: 705 GELEDQLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVK 764
E + +E A+ + K EKE+ + R +A K + E E +K
Sbjct: 337 EEERKAAEERAKAEEERKAAEERAKAEKERKAAEERERAKAEEEKRAAEEKARLEAEKLK 396
Query: 765 KALAEQEKKSAESLALA------KSDMEAVMQATSEEIKKATETHAEALATKDAEIASQL 818
+ +EKK+ E A A + D EA ++A E + + + ++ KD + +
Sbjct: 397 EKKKMEEKKAQEEKAQANLLRKQEEDKEAKVEAKKESLPEKLQPTSKKDQVKDNKDKEKA 456
Query: 819 AK--IKSLED-ELATEKAKAIEAREQAADIALDNRERGFYLAKDQAQHLFPNFDFSAMGV 875
K +KS+ D + + KA + L + E F + + N + +
Sbjct: 457 PKEEMKSVWDRKRGVPEQKAQNGERELTTPKLKSTENAFGRSNLKGA---ANAE-AGSEK 512
Query: 876 MKEITAAGLVGPDDPPLIDQNLWTATEEEEEEEEEEEQEKENNE 919
+KE V D+ + EEEE+++++EE E++ E
Sbjct: 513 LKEKQQEAAVELDELKKRREERRKILEEEEQKKKQEEAERKIRE 556
Score = 38.9 bits (89), Expect = 0.063
Identities = 89/440 (20%), Positives = 152/440 (34%), Gaps = 56/440 (12%)
Query: 445 KASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESS 504
K D+ KP PT +N + +V +T+ V E+
Sbjct: 180 KEEKDSEEEKPKEVPTE-------ENQVDVAVEKSTDKEEV---------------VETK 217
Query: 505 IGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQ 564
A A NA S T DKEKE + + + +
Sbjct: 218 TLAVNAENDTNAMLEGEQSITDAADKEKEEAEKEREKLEAEEKERLKAEEEKKAAEEKQK 277
Query: 565 GEKSCPGAATTSEAAQIEQAPAPEVGSSSYYNMLPNAIEPSEFLLAGLNRDVIEKEVLSQ 624
E+ A A E+ A E + A E E A R E+ ++
Sbjct: 278 AEEEKKAAEERERAKAEEEKRAAEERERAKAEEERKAAEERERAKAEEERKAAEERAKAE 337
Query: 625 GLNVTKEETL-ACLLRAGCIFAHTFEKFNAANVEAERLKAESAKHQEAAAAWEK---RFD 680
EE A R EK A E ER KAE +E AA EK +
Sbjct: 338 EERKAAEERAKAEEERKAAEERAKAEKERKAAEERERAKAE----EEKRAAEEKARLEAE 393
Query: 681 KLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKEEADELDASLQACKK--------EK 732
KL + ++ A + A + + ED+ A ++ + + L LQ K +K
Sbjct: 394 KLKEKKKMEEKKAQEEKAQANLLRKQEEDKEAKVEAKKESLPEKLQPTSKKDQVKDNKDK 453
Query: 733 EQAEKDLIA------RG-EALIAKESELAILCAELELVKKALAEQEKKSAESLALAKSDM 785
E+A K+ + RG A+ E + +L+ + A K A + +
Sbjct: 454 EKAPKEEMKSVWDRKRGVPEQKAQNGERELTTPKLKSTENAFGRSNLKGAANAEAGSEKL 513
Query: 786 EAVMQATS---EEIKKATETHAEALATKDAEIASQLA--------KIKSLEDELATEKAK 834
+ Q + +E+KK E + L ++ + + A + K +++E+ +A+
Sbjct: 514 KEKQQEAAVELDELKKRREERRKILEEEEQKKKQEEAERKIREEEEKKRMKEEIERRRAE 573
Query: 835 AIEAREQAADIALDNRERGF 854
A E R++ + + ++ F
Sbjct: 574 AAEKRQKVPEDGVSEEKKPF 593
>RRB1_HUMAN (Q9P2E9) Ribosome-binding protein 1 (Ribosome receptor
protein) (180 kDa ribosome receptor homolog) (ES/130
related protein)
Length = 1410
Score = 52.0 bits (123), Expect = 7e-06
Identities = 90/452 (19%), Positives = 173/452 (37%), Gaps = 64/452 (14%)
Query: 419 EGDAKKRGGAENQAESTKAPKRRRLTKASSDA--GTSKPGAQPTTAAAAPKGKNVA---- 472
+ +K GA+NQ + T+ + ++ ++ + G P + A +G V
Sbjct: 506 QNQGQKGEGAQNQGKKTEGAQGKKAERSPNQGKKGEGAPIQGKKADSVANQGTKVEGITN 565
Query: 473 EASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEK 532
+ A + A + G A AA +AS T A S + P K++
Sbjct: 566 QGKKAEGSPSEGKKAEGSPNQGKKADAAANQGKKTESASVQGRNTDVAQSPEAP---KQE 622
Query: 533 ENETPKSPPRQDAPPSPPSTHDAGPMPSPP------------HQGEKSCPGAATTSEAAQ 580
KS ++ P PP GP+ P ++GE + +A
Sbjct: 623 APAKKKSGSKKKGEPGPPDAD--GPLYLPYKTLVSTVGSMVFNEGEAQRLIEILSEKAGI 680
Query: 581 IEQAPAPEVGSSSYYNMLPNAIEPSEFLLAGLNRDVIEKEVLSQGLNVTKEETLACLLRA 640
I+ +L +E E LLA D V L +E A
Sbjct: 681 IQDTWHKATQKGDPVAILKRQLEEKEKLLATEQEDAA---VAKSKLRELNKEMAA----- 732
Query: 641 GCIFAHTFEKFNAANVEAERLKAESAKHQEAAA-------AWEKRFDKLATQAGKDKVYA 693
EK AA EA+ K A+ QE A ++ + ++ GK +
Sbjct: 733 --------EKAKAAAGEAKVKKQLVAREQEITAVQARMQASYREHVKEVQQLQGKIRTLQ 784
Query: 694 DKMIGTAGIKIGELEDQLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESEL 753
+++ ++ L+ + +++++ ++ + +++ K+ AE + + + ++KE
Sbjct: 785 EQLENGPNTQLARLQQENSILRDALNQATSQVES----KQNAELAKLRQELSKVSKE--- 837
Query: 754 AILCAELELVKKALAEQEKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAE 813
L + E V++ EQ++K+ E+ A A +QA+ E + EAL + E
Sbjct: 838 --LVEKSEAVRQD--EQQRKALEAKAAAFEKQVLQLQASHRESE-------EALQKRLDE 886
Query: 814 IASQLAKIKSLEDELATEKAKAIEAREQAADI 845
++ +L +S L + KA E ++Q A++
Sbjct: 887 VSRELCHTQSSHASLRADAEKAQEQQQQMAEL 918
Score = 40.8 bits (94), Expect = 0.017
Identities = 45/203 (22%), Positives = 89/203 (43%), Gaps = 23/203 (11%)
Query: 649 EKFNAANVEAERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELE 708
EK A + ++ KA AK AAA+EK+ +L + + K ++ E+
Sbjct: 840 EKSEAVRQDEQQRKALEAK----AAAFEKQVLQLQASHRESEEALQK-------RLDEVS 888
Query: 709 DQLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKALA 768
+L ASL+A ++ ++ ++ + L + E+E+ C EL + L
Sbjct: 889 RELC----HTQSSHASLRADAEKAQEQQQQMAELHSKLQSSEAEVRSKCEELSGLHGQLQ 944
Query: 769 EQEKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIKSLEDEL 828
E ++++ +S +EA+++A A+ + AE Q ++K LE ++
Sbjct: 945 EARAENSQLTERIRS-IEALLEA-------GQARDAQDVQASQAEADQQQTRLKELESQV 996
Query: 829 ATEKAKAIEAREQAADIALDNRE 851
+ + +AIE RE + N +
Sbjct: 997 SGLEKEAIELREAVEQQKVKNND 1019
Score = 32.7 bits (73), Expect = 4.5
Identities = 39/194 (20%), Positives = 87/194 (44%), Gaps = 11/194 (5%)
Query: 649 EKFNAANVEAERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELE 708
E+ + + + + +AE+++ E + E + + +D + ++ ELE
Sbjct: 934 EELSGLHGQLQEARAENSQLTERIRSIEALLEAGQARDAQDVQASQAEADQQQTRLKELE 993
Query: 709 DQLALMKEEADELDASLQACKKE----KEQAEKDLIARGEALIAKESELAILCAELELVK 764
Q++ +++EA EL +++ K + +E+ K + A A A + +L L E +
Sbjct: 994 SQVSGLEKEAIELREAVEQQKVKNNDLREKNWKAMEALATAEQACKEKLLSLTQAKEESE 1053
Query: 765 KALAEQEKKSAESLALAKSDMEAVMQAT----SEEIKKATET---HAEALATKDAEIASQ 817
K L E ++ E+L ++ + Q +++K+ T H A A +++AS+
Sbjct: 1054 KQLCLIEAQTMEALLALLPELSVLAQQNYTEWLQDLKEKGPTLLKHPPAPAEPSSDLASK 1113
Query: 818 LAKIKSLEDELATE 831
L + + + L E
Sbjct: 1114 LREAEETQSTLQAE 1127
>A180_HUMAN (O60641) Clathrin coat assembly protein AP180 (Clathrin
coat associated protein AP180) (91 kDa
synaptosomal-associated protein)
Length = 907
Score = 52.0 bits (123), Expect = 7e-06
Identities = 55/188 (29%), Positives = 81/188 (42%), Gaps = 20/188 (10%)
Query: 422 AKKRGGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPT--TAAAAPK----GKNVAEAS 475
A G A +E AP A+ DA + P+ +A AAP+ E S
Sbjct: 445 AASPGEAPAASEGAAAPATPTPVAAALDACSGNDPFAPSEGSAEAAPELDLFAMKPPETS 504
Query: 476 VAAATEPTA-----VPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTP---- 526
V T PTA VPA++P+ A A AA AA ++ ATAA+ T AA ++ P
Sbjct: 505 VPVVT-PTASTAPPVPATAPSPAPAVAA-AAAATTAATAAATTTTTTSAATATTAPPALD 562
Query: 527 -IGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAP 585
GD + + P+ DA PS SPP QG P ++ T++ ++
Sbjct: 563 IFGDLFESTPEVAAAPKPDAAPS-IDLFSTDAFSSPP-QGASPVPESSLTADLLSVDAFA 620
Query: 586 APEVGSSS 593
AP +++
Sbjct: 621 APSPATTA 628
>TMPB_TREPH (P29720) Treponemal membrane protein B precursor
(Antigen tmpB)
Length = 384
Score = 51.6 bits (122), Expect = 9e-06
Identities = 57/192 (29%), Positives = 91/192 (46%), Gaps = 13/192 (6%)
Query: 680 DKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKEEADELDASLQACKKE---KEQAE 736
DK+ +A DK A+K + +D+ A KE+A + A+ KE KE+A
Sbjct: 146 DKVIAKAAADKAAAEKAAKEKAAREKSAKDKAA--KEKAAKEKAAKDKAAKEKAAKEKAA 203
Query: 737 KDLIARGEALIAKESELAILCAELELVKKALAEQE--KKSAESLALAKSDMEAVMQATSE 794
KD A+ +A AKE + A+ + K A++E +K+AE A K+ EA + +E
Sbjct: 204 KDKAAKEKA--AKEKAAREMAAKEKAAKDKAAKEEAARKAAEEAAARKAAEEAAARKAAE 261
Query: 795 EIKKATETHAEALATKDAEIASQLAKIKSLEDELATEKAKAIEAREQAADIALDNRERGF 854
E +A AE A + A A + A K+ E+ L EK + + E +RE +
Sbjct: 262 E--EAARIAAEEEAARKA--AEEEAARKAAEEALYNEKGEKVLPSEYKVLTWKLDRECFW 317
Query: 855 YLAKDQAQHLFP 866
+AK+ A + P
Sbjct: 318 NIAKNPAVYNDP 329
Score = 46.6 bits (109), Expect = 3e-04
Identities = 44/128 (34%), Positives = 60/128 (46%), Gaps = 14/128 (10%)
Query: 729 KKEKEQAEKDLIARGEALIA-------KESELAILCAELELVKKALAEQEKK-----SAE 776
K +K + LIA GEA+ A K ++A+ CAE L E E K +A+
Sbjct: 97 KADKNYPTEYLIA-GEAIKAGGIAFDNKNYDVAVTCAEKALESLKTVEPEDKVIAKAAAD 155
Query: 777 SLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIKSLEDELATEK-AKA 835
A K+ E + S + K A E A+ A KD + AK K+ +D+ A EK AK
Sbjct: 156 KAAAEKAAKEKAAREKSAKDKAAKEKAAKEKAAKDKAAKEKAAKEKAAKDKAAKEKAAKE 215
Query: 836 IEAREQAA 843
ARE AA
Sbjct: 216 KAAREMAA 223
>DYNA_HUMAN (Q14203) Dynactin 1 (150 kDa dynein-associated
polypeptide) (DP-150) (DAP-150) (p150-glued) (p135)
Length = 1278
Score = 51.6 bits (122), Expect = 9e-06
Identities = 89/388 (22%), Positives = 151/388 (37%), Gaps = 51/388 (13%)
Query: 493 AGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETP-KSPPRQDAPPSPPS 551
A T+ +SS G + T A + G K K+ T K+ R+ P P S
Sbjct: 101 ADTTSPETPDSSASKVLKREGTDTT---AKTSKLRGLKPKKAPTARKTTTRRPKPTRPAS 157
Query: 552 THDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAP-------APEVGSSSYYNMLPNAIEP 604
T AG S G S G ++SE + Q P P + S LP+ +
Sbjct: 158 TGVAGASSSLGPSGSASA-GELSSSEPSTPAQTPLAAPIIPTPVLTSPGAVPPLPSPSKE 216
Query: 605 SEFLLAGLNRDVIEKEVLSQGLNVTKEETLACLLRAGCIFAHTFEKFNAANVEAERLKAE 664
E GL V + E + L + + E A L ++ ++ E+++
Sbjct: 217 EE----GLRAQVRDLEEKLETLRLKRAEDKAKL-----------KELEKHKIQLEQVQEW 261
Query: 665 SAKHQEAAAAWEKRFDKLATQAGK----DKVYADKMIGTA-GIKIGELEDQLA------- 712
+K QE A ++R + +A + + Y ++M TA I++ L+ ++A
Sbjct: 262 KSKMQEQQADLQRRLKEARKEAKEALEAKERYMEEMADTADAIEMATLDKEMAEERAESL 321
Query: 713 -----LMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKAL 767
+KE DEL L+ K E E+ D A L E + A L L ++
Sbjct: 322 QQEVEALKERVDELTTDLEILKAEIEEKGSDGAASSYQLKQLEEQNARLKDALVRMRDLS 381
Query: 768 AEQEKKSAESLALAK---SDMEAVMQ---ATSEEIKKATETHAEALATKDAEIASQLAKI 821
+ ++++ + L + ++E V Q EE+ +A T E DA + ++ +
Sbjct: 382 SSEKQEHVKLQKLMEKKNQELEVVRQQRERLQEELSQAESTIDELKEQVDAALGAE-EMV 440
Query: 822 KSLEDELATEKAKAIEAREQAADIALDN 849
+ L D + K E RE D+ N
Sbjct: 441 EMLTDRNLNLEEKVRELRETVGDLEAMN 468
>DYNA_CHICK (P35458) Dynactin 1 (150 kDa dynein-associated
polypeptide) (DP-150) (DAP-150) (p150-glued)
Length = 1224
Score = 51.6 bits (122), Expect = 9e-06
Identities = 90/379 (23%), Positives = 156/379 (40%), Gaps = 70/379 (18%)
Query: 501 AESSIGATAASAGVNATK----AAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAG 556
A+++ T SA + K AA G K K+ T + P P S PS+ AG
Sbjct: 102 ADTTSPETPESAALKVPKRHSRXAAKGSKLRGAKPKKT-TARRPKPTRTPTSAPSSGTAG 160
Query: 557 PMPSPPHQGEKSCPGAATTSEAAQIEQAP--APEVGSSSYYN----MLPNAIEPSEFLLA 610
P S G G ++SE + Q P AP + S S + M+P+ + E L +
Sbjct: 161 PSGSASASG-----GEMSSSEPSTPAQTPLVAPVIPSPSLTSPVAPMVPSPTKEEENLRS 215
Query: 611 GLNRDVIEKEVLSQGLNVTKEETLACLLRAGCIFAHTFEKFNAANVEAERLKAESAKHQE 670
+ RD+ EK + L + + E A L EK+ ++ E+++ +K QE
Sbjct: 216 QV-RDLEEK---LETLKIKRNEDKAKLKE--------LEKYK---IQLEQVQEWKSKMQE 260
Query: 671 AAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKEEADELDASLQACKK 730
A ++R K A + KD LE + M+E AD DA ++
Sbjct: 261 QQADLQRRL-KEAKKEAKDA----------------LEAKERYMEEMADTADA-IEMATL 302
Query: 731 EKEQAEKDLIARGEALIAKESELAILCAELELVK-----------------KALAEQEKK 773
+KE AE+ + + + + + ++ L +LE++K K L EQ +
Sbjct: 303 DKEMAEERAESLQQEVDSLKEKVEYLTMDLEILKHEIEEKGSDGAASSYQVKQLEEQNAR 362
Query: 774 SAESLALAKSDMEAVMQATSEEIKKATE---THAEALATKDAEIASQLAKIKSLEDELAT 830
E+L + D+ A + +++K E T E+L + ++ ++ + + DEL
Sbjct: 363 LKEALVRMR-DLSASEKQEHVKLQKQMEKKNTELESLRQQREKLQEEVKQAEKTVDELKE 421
Query: 831 EKAKAIEAREQAADIALDN 849
+ A+ A E + N
Sbjct: 422 QVDAALGAEEMVETLTERN 440
Score = 45.1 bits (105), Expect = 9e-04
Identities = 86/445 (19%), Positives = 172/445 (38%), Gaps = 39/445 (8%)
Query: 414 PKFNTEGDAK--KRGGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNV 471
PK ++ AK K GA+ + + + PK R ++ +GT A P G
Sbjct: 118 PKRHSRXAAKGSKLRGAKPKKTTARRPKPTRTPTSAPSSGT-----------AGPSGSAS 166
Query: 472 AEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKE 531
A +++EP S+PA A S+ + A + TK + + + D E
Sbjct: 167 ASGGEMSSSEP-----STPAQTPLVAPVIPSPSLTSPVAPMVPSPTKEEENLRSQVRDLE 221
Query: 532 KENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGS 591
++ ET K +D + K A + + A +
Sbjct: 222 EKLETLKIKRNEDKAKLKELEKYKIQLEQVQEWKSKMQEQQADLQRRLKEAKKEAKDALE 281
Query: 592 SSYYNMLPNAIEPSEFLLAGLNRDVIEK--EVLSQGLNVTKEETLACLLRAGCIFAHTFE 649
+ M A +A L++++ E+ E L Q ++ KE+ + L I H E
Sbjct: 282 AKERYMEEMADTADAIEMATLDKEMAEERAESLQQEVDSLKEK-VEYLTMDLEILKHEIE 340
Query: 650 KFN----AANVEAERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIG 705
+ A++ + ++L+ ++A+ +EA R L+ ++ V K + ++
Sbjct: 341 EKGSDGAASSYQVKQLEEQNARLKEALV----RMRDLSASEKQEHVKLQKQMEKKNTELE 396
Query: 706 ELEDQLALMKEEADELDASLQACKKE------KEQAEKDLIARGEALIAKESELAILCAE 759
L Q ++EE + + ++ K++ E+ + L R L K EL +
Sbjct: 397 SLRQQREKLQEEVKQAEKTVDELKEQVDAALGAEEMVETLTERNLDLEEKVRELRETVGD 456
Query: 760 LELVKKALAEQEKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQ-- 817
LE + + E ++ + E+ + ++ + A E +K E E +A I
Sbjct: 457 LEAMNEMNDELQENARETELELREQLD-LAAARVREAEKRVEAAQETVADYQQTIKKYRE 515
Query: 818 -LAKIKSLEDELATEKAKAIEAREQ 841
A ++ + EL +++ + E ++Q
Sbjct: 516 LTAHLQDVNRELMSQQEASAEKQQQ 540
>NFH_MOUSE (P19246) Neurofilament triplet H protein (200 kDa
neurofilament protein) (Neurofilament heavy polypeptide)
(NF-H)
Length = 1087
Score = 51.2 bits (121), Expect = 1e-05
Identities = 93/436 (21%), Positives = 160/436 (36%), Gaps = 46/436 (10%)
Query: 403 KARERRMARFSPKFNTEGDAKKRGGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTA 462
+A+ A+ + + G+AK ++ AE+ + + +A S A PG + A
Sbjct: 661 EAKSPAEAKSPAEVKSPGEAKSPAEPKSPAEAKSPAEVKSPAEAKSPAEVKSPGEAKSPA 720
Query: 463 AAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAAS 522
A + + A+V + E + P + + A A + A+S I + K A
Sbjct: 721 AVKSPAEAKSPAAVKSPGEAKS-PGEAKSPAEAKSPAEAKSPIEVKSPEKAKTPVKEGAK 779
Query: 523 SDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIE 582
S + K E KSP ++D P P SP +G K A +
Sbjct: 780 SPA----EAKSPEKAKSPVKEDIKP-PAEAKSPEKAKSPVKEGAKPPEKAKPLDVKSPEA 834
Query: 583 QAPAPEVGSSSYYNMLPNAIEPSEFLLAGLNRDV----IEKEVLSQGLNVTKEETLACLL 638
Q P E + +P I P E + + E+ S+ + KEE +
Sbjct: 835 QTPVQEEAT------VPTDIRPPEQVKSPAKEKAKSPEKEEAKTSEKVAPKKEEVKS--- 885
Query: 639 RAGCIFAHTFEKFNAANVEAERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIG 698
++ A ++++ E A E + D+ +A K KV K
Sbjct: 886 --------PVKEEVKAKEPPKKVEEEKTLPTPKTEAKESKKDEAPKEAPKPKVEEKKETP 937
Query: 699 TAGIKIGELEDQLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCA 758
T K E + KEEA E ++ + +E+ L + EA +++E A
Sbjct: 938 TEKPKDSTAEAK----KEEAGEKKKAVAS----EEETPAKLGVKEEAKPKEKTETTKTEA 989
Query: 759 ELELVKKALAEQEKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQL 818
E A++ K E+ K +M A + + +K TE+ K E
Sbjct: 990 E-----DTKAKEPSKPTETEKPKKEEMPAAPEKKDTKEEKTTESR------KPEEKPKME 1038
Query: 819 AKIKSLEDELATEKAK 834
AK+K + L+ E +K
Sbjct: 1039 AKVKEDDKSLSKEPSK 1054
Score = 48.9 bits (115), Expect = 6e-05
Identities = 98/440 (22%), Positives = 167/440 (37%), Gaps = 31/440 (7%)
Query: 421 DAKKRGGAENQAESTKAPKRRRLTKASSDAGTSKPGA--QPTTAAAAPKGKNVAEASVAA 478
+AK A++ AE+ + + +A S A P P T + + K+ +EA A
Sbjct: 595 EAKSPAEAKSPAEAKSPAEAKSPAEAKSPAEAKSPAEAKSPATVKSPGEAKSPSEAKSPA 654
Query: 479 ATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPK 538
+ A A SPA A + A + + A K+ A +P + K K
Sbjct: 655 EAKSPA-EAKSPAEAKSPAEVKSPGEAKSPAEPKSPAEAKSPAEVKSPA--EAKSPAEVK 711
Query: 539 SPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPG-AATTSEAAQIEQAPAPEVGSSSYYNM 597
SP +P + S +A + GE PG A + +EA +A +P S
Sbjct: 712 SPGEAKSPAAVKSPAEAKSPAAVKSPGEAKSPGEAKSPAEAKSPAEAKSPIEVKSPEKAK 771
Query: 598 LP---NAIEPSEFLLAGLNRDVIEKEVLSQGLNVTKEETLACLLRAGCIFAHTFEKFNAA 654
P A P+E + + KE + E ++ G A EK
Sbjct: 772 TPVKEGAKSPAEAKSPEKAKSPV-KEDIKPPAEAKSPEKAKSPVKEG---AKPPEKAKPL 827
Query: 655 NVEAERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYA-DKMIGTAGIKIGELEDQL-A 712
+V++ +A++ +EA + R + K+K + +K K+ ++++ +
Sbjct: 828 DVKSP--EAQTPVQEEATVPTDIRPPEQVKSPAKEKAKSPEKEEAKTSEKVAPKKEEVKS 885
Query: 713 LMKEEAD--------ELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVK 764
+KEE E + +L K E ++++KD A EA K E E
Sbjct: 886 PVKEEVKAKEPPKKVEEEKTLPTPKTEAKESKKD-EAPKEAPKPKVEEKKETPTEKPKDS 944
Query: 765 KALAEQEKKSAESLALAKSDMEAVMQATSEEIK-----KATETHAEALATKDAEIASQLA 819
A A++E+ + A+A + EE K + T+T AE K+ ++
Sbjct: 945 TAEAKKEEAGEKKKAVASEEETPAKLGVKEEAKPKEKTETTKTEAEDTKAKEPSKPTETE 1004
Query: 820 KIKSLEDELATEKAKAIEAR 839
K K E A EK E +
Sbjct: 1005 KPKKEEMPAAPEKKDTKEEK 1024
Score = 43.1 bits (100), Expect = 0.003
Identities = 108/543 (19%), Positives = 197/543 (35%), Gaps = 58/543 (10%)
Query: 403 KARERRMARFSPKFNTEGDAKKRGGAEN--------------QAESTKAPKRRRLTKASS 448
K+R + A+ + + G+AK A++ +A+S PK K+ +
Sbjct: 511 KSRVKEEAKSPGEAKSPGEAKSPAEAKSPGEAKSPGEAKSPGEAKSPAEPKSPAEPKSPA 570
Query: 449 DAGTSKPGAQPTTAAAAPKGKNVAEA-------SVAAATEPTAVPASSPASAGATAAFAA 501
+A + P T + + K+ +EA S A A P + + A + A A A
Sbjct: 571 EAKSPAEPKSPATVKSPGEAKSPSEAKSPAEAKSPAEAKSPAEAKSPAEAKSPAEAKSPA 630
Query: 502 ESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSP 561
E+ AT S G +A + S+ + K KSP +P S +A P
Sbjct: 631 EAKSPATVKSPG----EAKSPSEAKSPAEAKSPAEAKSPAEAKSPAEVKSPGEAKSPAEP 686
Query: 562 PHQGEKSCPG-AATTSEAAQIEQAPAP-EVGSSSYYNMLPNAIEPSEFLLAGLNRDVIEK 619
E P + +EA + +P E S + A P+ G
Sbjct: 687 KSPAEAKSPAEVKSPAEAKSPAEVKSPGEAKSPAAVKSPAEAKSPAAVKSPG-------- 738
Query: 620 EVLSQGLNVTKEETLACLLRAGCIFAHTFEKFNAANVEAERLKAESAKHQEAAAAWEKRF 679
E S G + E + I + EK E + AE+ ++A + ++
Sbjct: 739 EAKSPGEAKSPAEAKSPAEAKSPIEVKSPEKAKTPVKEGAKSPAEAKSPEKAKSPVKEDI 798
Query: 680 DKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKEEAD---ELDASLQACKKEKEQAE 736
A +K K G K E L + EA + +A++ + EQ +
Sbjct: 799 KPPAEAKSPEKA---KSPVKEGAKPPEKAKPLDVKSPEAQTPVQEEATVPTDIRPPEQVK 855
Query: 737 KDLIARGEALIAKESELAILCAELELVKKALAEQEKKSAESLALAKSDMEAVMQATSEEI 796
+ ++ +E++ + A + K+ ++E K+ E K + E + E
Sbjct: 856 SPAKEKAKSPEKEEAKTSEKVAPKKEEVKSPVKEEVKAKE--PPKKVEEEKTLPTPKTEA 913
Query: 797 KKATETHAEALATKDAEIASQLAKIKSLEDELATEKAKAIEAREQAADIALDNRERGFYL 856
K++ + A A K + + +D +T +AK EA E+ +A +
Sbjct: 914 KESKKDEAPKEAPKPKVEEKKETPTEKPKD--STAEAKKEEAGEKKKAVASEEETPAKLG 971
Query: 857 AKDQAQHLFPNFDFSAMGVMKEITAAGLVGPDDPPLIDQNLWTATEEEEEEEEEEEQEKE 916
K++A+ KE T +D + + T TE+ ++EE EK+
Sbjct: 972 VKEEAK-------------PKEKTETTKTEAEDTKAKEPSKPTETEKPKKEEMPAAPEKK 1018
Query: 917 NNE 919
+ +
Sbjct: 1019 DTK 1021
Score = 40.0 bits (92), Expect = 0.028
Identities = 36/138 (26%), Positives = 52/138 (37%), Gaps = 2/138 (1%)
Query: 457 AQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIG-ATAASAGVN 515
AQ A +G+ E +AAAT P A A+SP + S G A + +
Sbjct: 473 AQGQEGEEAEEGEEKEEEELAAATSPPAEEAASPEKETKSRVKEEAKSPGEAKSPGEAKS 532
Query: 516 ATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPG-AAT 574
+A + + + K KSP +P P S +A P PG A +
Sbjct: 533 PAEAKSPGEAKSPGEAKSPGEAKSPAEPKSPAEPKSPAEAKSPAEPKSPATVKSPGEAKS 592
Query: 575 TSEAAQIEQAPAPEVGSS 592
SEA +A +P S
Sbjct: 593 PSEAKSPAEAKSPAEAKS 610
Score = 38.9 bits (89), Expect = 0.063
Identities = 41/191 (21%), Positives = 71/191 (36%), Gaps = 8/191 (4%)
Query: 403 KARERRMARFSPKFNTEGDAKKRGGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTA 462
K E A SP +K + + E+ + + +A S A PG +
Sbjct: 487 KEEEELAAATSPPAEEAASPEKETKSRVKEEAKSPGEAKSPGEAKSPAEAKSPGEAKSPG 546
Query: 463 AAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAAS 522
A G+ + A + EP SPA A + A + +++ + + + K+ A
Sbjct: 547 EAKSPGEAKSPAEPKSPAEP-----KSPAEAKSPAEPKSPATVKSPGEAKSPSEAKSPAE 601
Query: 523 SDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPG-AATTSEAAQI 581
+ +P + K KSP +P S +A + GE P A + +EA
Sbjct: 602 AKSPA--EAKSPAEAKSPAEAKSPAEAKSPAEAKSPATVKSPGEAKSPSEAKSPAEAKSP 659
Query: 582 EQAPAPEVGSS 592
+A +P S
Sbjct: 660 AEAKSPAEAKS 670
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.312 0.128 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 108,478,868
Number of Sequences: 164201
Number of extensions: 4880922
Number of successful extensions: 46449
Number of sequences better than 10.0: 1695
Number of HSP's better than 10.0 without gapping: 310
Number of HSP's successfully gapped in prelim test: 1447
Number of HSP's that attempted gapping in prelim test: 34786
Number of HSP's gapped (non-prelim): 7722
length of query: 919
length of database: 59,974,054
effective HSP length: 119
effective length of query: 800
effective length of database: 40,434,135
effective search space: 32347308000
effective search space used: 32347308000
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 71 (32.0 bits)
Lotus: description of TM0590c.8


