

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0590c.5
(441 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
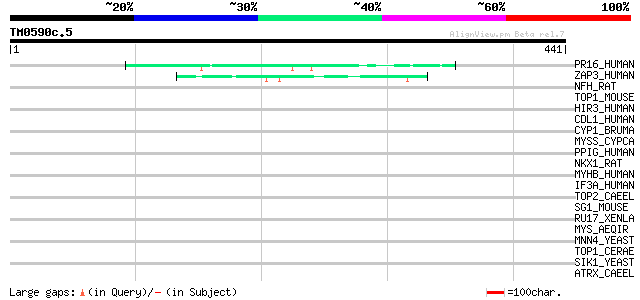
Score E
Sequences producing significant alignments: (bits) Value
PR16_HUMAN (Q92620) Pre-mRNA splicing factor ATP-dependent RNA h... 45 3e-04
ZAP3_HUMAN (P49750) Nuclear protein ZAP3 (ZAP113) 44 6e-04
NFH_RAT (P16884) Neurofilament triplet H protein (200 kDa neurof... 40 0.012
TOP1_MOUSE (Q04750) DNA topoisomerase I (EC 5.99.1.2) 38 0.059
HIR3_HUMAN (Q9BW71) HIRA-interacting protein 3 38 0.059
CDL1_HUMAN (P21127) PITSLRE serine/threonine-protein kinase CDC2... 38 0.059
CYP1_BRUMA (Q27450) Peptidylprolyl isomerase 1 (EC 5.2.1.8) (Pep... 37 0.078
MYSS_CYPCA (Q90339) Myosin heavy chain, fast skeletal muscle 37 0.13
PPIG_HUMAN (Q13427) Peptidyl-prolyl cis-trans isomerase G (EC 5.... 36 0.17
NKX1_RAT (Q9QZM6) Sodium/potassium/calcium exchanger 1 precursor... 36 0.17
MYHB_HUMAN (P35749) Myosin heavy chain, smooth muscle isoform (S... 36 0.17
IF3A_HUMAN (Q14152) Eukaryotic translation initiation factor 3 s... 36 0.17
TOP2_CAEEL (Q23670) Probable DNA topoisomerase II (EC 5.99.1.3) 36 0.23
SG1_MOUSE (P16014) Secretogranin I precursor (SgI) (Chromogranin... 36 0.23
RU17_XENLA (P09406) U1 small nuclear ribonucleoprotein 70 kDa (U... 36 0.23
MYS_AEQIR (P24733) Myosin heavy chain, striated muscle 36 0.23
MNN4_YEAST (P36044) MNN4 protein 36 0.23
TOP1_CERAE (Q7YR26) DNA topoisomerase I (EC 5.99.1.2) 35 0.29
SIK1_YEAST (Q12460) SIK1 protein (Nucleolar protein NOP56) 35 0.29
ATRX_CAEEL (Q9U7E0) Transcriptional regulator ATRX homolog (X-li... 35 0.29
>PR16_HUMAN (Q92620) Pre-mRNA splicing factor ATP-dependent RNA
helicase PRP16 (EC 3.6.1.-) (ATP-dependent RNA helicase
DHX38) (DEAH-box protein 38)
Length = 1227
Score = 45.4 bits (106), Expect = 3e-04
Identities = 59/280 (21%), Positives = 111/280 (39%), Gaps = 41/280 (14%)
Query: 93 NIRQQEGESLKEYMARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMGE- 151
+I + EG L + S E+ A A + LL L + L R+ +
Sbjct: 9 SIHRLEGTDLDCQVGGLICKSKSAASEQHVFKAPAPRPSLLGLDLLASLKRREREEKDDG 68
Query: 152 ---MRARASTYILDEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSV 208
+++ S+Y D E+ +K A+ E GD + + ++KDR PG
Sbjct: 69 EDKKKSKVSSY-KDWEESKDDQKDAEEEGGDQAGQNIRKDRHYRSARVETPSHPGGVSEE 127
Query: 209 FKPTKEQLYPRRDDY------EQRRPWQSKSHRQRE--------ETDMVMNTDVSDMLRG 254
F Q R ++ ++ + W+ + R R+ E D ++ S+ G
Sbjct: 128 FWERSRQRERERREHGVYASSKEEKDWKKEKSRDRDYDRKRDRDERDRSRHSSRSERDGG 187
Query: 255 ASDANLVDEPEAPKYQPRDANPKKWCEFHRSAGHDTDDCWTLQREIDKLIRAGYQGNRQG 314
+ ++ +EPE+P+++P+DA RS + D +GY +R+
Sbjct: 188 SERSSRRNEPESPRHRPKDAATPS-----RSTWEEED--------------SGYGSSRRS 228
Query: 315 QWRNGGDQNKAHKREEERADTKGKKKQESAAIATKGADDT 354
QW + R+ ER+ + ++ ++ K +DDT
Sbjct: 229 QWES--PSPTPSYRDSERSHRLSTRDRD-RSVRGKYSDDT 265
>ZAP3_HUMAN (P49750) Nuclear protein ZAP3 (ZAP113)
Length = 1822
Score = 44.3 bits (103), Expect = 6e-04
Identities = 47/214 (21%), Positives = 76/214 (34%), Gaps = 36/214 (16%)
Query: 133 LPGGLNSKLTRKPARSMGEMRARASTYILDEEDEAFKRKRAKLEKGDTSPKRVKKDRSGE 192
+PGGL R A S R R ++ R+ E+ P+R G
Sbjct: 898 VPGGLQGSQDRGAAGS----RERGPPRRAGSQERGPLRRAGSRER--IPPRRAGSRERGP 951
Query: 193 DKGDGKQQRP------GKGKSVFKPTK----EQLYPRRDDYEQRRPWQSKSHRQREETDM 242
+G G ++R G+ + F+P E++YP D R PW R EE +
Sbjct: 952 PRGPGSRERGLGRSDFGRDRGPFRPEPGDGGEKMYPYHRDEPPRAPWNHGEERGHEEFPL 1011
Query: 243 VMNTDVSDMLRGASDANLVDEPEAPKYQPRDANPKKWCEFHRSAGHDTDDCWTLQREIDK 302
D + +D+ + +Y W E R DT + + +
Sbjct: 1012 -------DGRNAPMERERLDDWDRERY---------WRECERDYQDDTLELYNREDRFSA 1055
Query: 303 LIRAGYQGNRQG----QWRNGGDQNKAHKREEER 332
+ G+R+G W D ++ + RE ER
Sbjct: 1056 PPSRSHDGDRRGPWWDDWERDQDMDEDYNREMER 1089
>NFH_RAT (P16884) Neurofilament triplet H protein (200 kDa
neurofilament protein) (Neurofilament heavy polypeptide)
(NF-H) (Fragment)
Length = 831
Score = 40.0 bits (92), Expect = 0.012
Identities = 33/131 (25%), Positives = 57/131 (43%), Gaps = 22/131 (16%)
Query: 141 LTRKPARSMGEMRARASTYILDEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKG-DGKQ 199
LT KP S GE + ++EA ++K A E+ + VK++ ++K D K
Sbjct: 689 LTEKPKDSPGEAK----------KEEAKEKKAAAPEEETPAKLGVKEEAKPKEKAEDAKA 738
Query: 200 QRPGKGKSVFKPTKEQL--YPRRDDYEQRRPWQSKSHRQREETDMVMNTDVSDMLRGASD 257
+ P K KP KE++ P + D ++ + +SK ++ + + + D
Sbjct: 739 KEPSKPSEKEKPKKEEVPAAPEKKDTKEEKTTESKKREEKPKMEAKAKEE---------D 789
Query: 258 ANLVDEPEAPK 268
L EP PK
Sbjct: 790 KGLPQEPSKPK 800
>TOP1_MOUSE (Q04750) DNA topoisomerase I (EC 5.99.1.2)
Length = 767
Score = 37.7 bits (86), Expect = 0.059
Identities = 20/76 (26%), Positives = 42/76 (54%), Gaps = 8/76 (10%)
Query: 162 DEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRP---GKGKSVFKPTKEQLYP 218
DE+D +K K+ K T + +K R E++ DGK ++P K K V +P ++ P
Sbjct: 143 DEDDADYKPKKIK-----TEDIKKEKKRKSEEEEDGKLKKPKNKDKDKKVAEPDNKKKKP 197
Query: 219 RRDDYEQRRPWQSKSH 234
++++ ++ + W+ + +
Sbjct: 198 KKEEEQKWKWWEEERY 213
>HIR3_HUMAN (Q9BW71) HIRA-interacting protein 3
Length = 556
Score = 37.7 bits (86), Expect = 0.059
Identities = 66/290 (22%), Positives = 110/290 (37%), Gaps = 46/290 (15%)
Query: 107 ARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMGEMRARASTYILDEEDE 166
A S + E EE A + + G +K + K + E A EE+
Sbjct: 185 ASVSRKQAREESEESEAEPVQRTAKKVEGNKGTK-SLKESEQESEEEILAQKKEQREEEV 243
Query: 167 AFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSVFKPTKEQLYPRRDDYEQR 226
+ K EKGD P+ RS + +++R K KS K +L D E++
Sbjct: 244 EEEEKEEDEEKGDWKPRT----RSNGRRKSAREERSCKQKSQAK----RLLGDSDSEEEQ 295
Query: 227 RPWQSK---SHRQREETDMVMNTDVSDMLRGAS-DANLVDEPEAPKYQPRDANPKKWCEF 282
+ S S R RE + D + + G + DE ++ K +P +K
Sbjct: 296 KEAASSGDDSGRDREPPVQRKSEDRTQLKGGKRLSGSSEDEEDSGKGEPTAKGSRKMARL 355
Query: 283 HRSAGHDTDDCWTLQREIDKLIRAGYQGNRQGQWRNGGDQNKAHKREEERADTKGKKKQE 342
++G ++D L+RE+ AG GG Q + R +++ KG+ +
Sbjct: 356 GSTSGEESD----LEREVSDS-EAG-----------GGPQGERKNRSSKKSSRKGRTRSS 399
Query: 343 SAAIATKGADDTFARHSGPPVGTINTIAGGFGGGGDTHAA---RKRHVRA 389
S++ +D + G G G G+ H A KR++RA
Sbjct: 400 SSS-----SDGSPEAKGG---------KAGSGRRGEDHPAVMRLKRYIRA 435
>CDL1_HUMAN (P21127) PITSLRE serine/threonine-protein kinase CDC2L1
(EC 2.7.1.37) (Galactosyltransferase associated protein
kinase p58/GTA) (Cell division cycle 2-like 1) (CLK-1)
(CDK11) (p58 CLK-1)
Length = 795
Score = 37.7 bits (86), Expect = 0.059
Identities = 48/272 (17%), Positives = 108/272 (39%), Gaps = 20/272 (7%)
Query: 82 KNQPVTINDLYNIRQQEGESLKEYMARYSAASVKVED--EEPRACALAFKNGLLPGGLNS 139
+N P D R +E +SL + + KV +E R ++ GG ++
Sbjct: 63 RNSPYRREDSMEDRGEEDDSLAIKPPQQMSRKEKVHHRKDEKRKEKRRHRSHSAEGGKHA 122
Query: 140 KLTRKPARSMGEMRARASTYILDEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQ 199
++ K R R EE + +R+ + ++ + + + +++R ++ + K+
Sbjct: 123 RVKEKEREHERRKRHR-------EEQDKARREWERQKRREMAREHSRRERDRLEQLERKR 175
Query: 200 QRPGKGKSVFKPTKEQLYPRRDDYEQRRPWQSKSHRQREETDMVMNTDVSDMLRGASDAN 259
+R K + K +EQ R E+R+ + ++ R+ M D SD ++ + +
Sbjct: 176 ERERKMREQQKEQREQKERERRAEERRK--EREARREVSAHHRTMREDYSDKVKASHWSR 233
Query: 260 LVDEPEAPKYQ------PRDANP---KKWCEFHRSAGHDTDDCWTLQREIDKLIRAGYQG 310
P +++ P +A P +K + + D LQ D +
Sbjct: 234 SPPRPPRERFELGDGRKPGEARPARAQKPAQLKEEKMEERDLLSDLQDISDSERKTSSAE 293
Query: 311 NRQGQWRNGGDQNKAHKREEERADTKGKKKQE 342
+ + +G ++ + + EEE + ++ +E
Sbjct: 294 SSSAESGSGSEEEEEEEEEEEEEGSTSEESEE 325
Score = 32.0 bits (71), Expect = 3.3
Identities = 42/209 (20%), Positives = 82/209 (39%), Gaps = 19/209 (9%)
Query: 149 MGEMRARASTYILDEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSV 208
MG+ + LDE + KR++ + EK + KR+K + K D ++ + +
Sbjct: 1 MGDEKDSWKVKTLDEILQEKKRRKEQEEKAEI--KRLKNSDDRDSKRDSLEEGELRDHRM 58
Query: 209 FKPTKEQLYPRRDDYEQR---------RPWQSKS-----HRQREETDMVMNTDVSDMLRG 254
+ Y R D E R +P Q S H +++E S G
Sbjct: 59 EITIRNSPYRREDSMEDRGEEDDSLAIKPPQQMSRKEKVHHRKDEKRKEKRRHRSHSAEG 118
Query: 255 ASDANLVDEPEAPKYQPR--DANPKKWCEFHRSAGHDTDDCWTLQREIDKLIRAGYQGNR 312
A + ++ + + R + K E+ R + + +RE D+L + + R
Sbjct: 119 GKHARVKEKEREHERRKRHREEQDKARREWERQKRREMAREHS-RRERDRLEQLERKRER 177
Query: 313 QGQWRNGGDQNKAHKREEERADTKGKKKQ 341
+ + R + + K E RA+ + K+++
Sbjct: 178 ERKMREQQKEQREQKERERRAEERRKERE 206
>CYP1_BRUMA (Q27450) Peptidylprolyl isomerase 1 (EC 5.2.1.8)
(Peptidylprolyl cis-trans isomerase 1) (PPIase 1)
(Cyclophilin) (BmCYP-1)
Length = 843
Score = 37.4 bits (85), Expect = 0.078
Identities = 51/268 (19%), Positives = 108/268 (40%), Gaps = 29/268 (10%)
Query: 86 VTINDLYNIRQQEGESLKEYMARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKP 145
VT++ +IR + E+++ ++ + A E++ + + K G G ++
Sbjct: 472 VTLDSAEDIRDSDDEAIRIHLLK---AKKMAEEKTKQEAKMLEKTGDKEGRDQKTISEAK 528
Query: 146 ARSMGEMRARASTYILDEEDEAFKRKRAK--LEKGDTSPKRVKKDRSGEDKGDGKQQRPG 203
+ E + + DE ++ K+ K + + DT K+ +K S E +G K R
Sbjct: 529 QKDSAEKDRQHREHKNDELEKRAIEKQDKDQIVERDTGSKQRRKSDSKEHRG--KTDRKH 586
Query: 204 KGKSVFKPTKEQLYPRR-DDYEQRRPWQSKSHRQREETDMVMNTDVSDMLRGASDANLVD 262
+ KS+ + + + DD +++ KS E+T + N+VD
Sbjct: 587 RSKSIEEDGRRSTSREKLDDLKRKETSGQKSQADSEQTV-------------EAKTNVVD 633
Query: 263 EPEAPKYQPRDANPKKWCEFHRSAGHDTDDCWTLQREIDKLIRAGYQGNRQGQWRNGGDQ 322
+ K+ ++ ++ + L+ E K Q N + + GG++
Sbjct: 634 SNSDNSKMSVNGKLKEVSSTNKE--NEVSEQKDLKAESTKSEEIKQQVNEVSRKQKGGEK 691
Query: 323 NKAHKREE------ERADTKGKKKQESA 344
K HKR E R+ + G++++ S+
Sbjct: 692 PKEHKRNERSRSRRRRSRSNGRRRRSSS 719
>MYSS_CYPCA (Q90339) Myosin heavy chain, fast skeletal muscle
Length = 1935
Score = 36.6 bits (83), Expect = 0.13
Identities = 41/164 (25%), Positives = 72/164 (43%), Gaps = 30/164 (18%)
Query: 87 TINDLYNIRQQEGESLKEYMARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPA 146
+I DL N +QQ E +K+ S K+EDE+ L ++L +K
Sbjct: 1066 SIMDLENEKQQSDEKIKKKDFEISQLLSKIEDEQ---------------SLGAQLQKK-- 1108
Query: 147 RSMGEMRARASTYILDEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRPGKGK 206
+ E++AR L+EE EA + RAK+EK +R R E+ + ++ G
Sbjct: 1109 --IKELQARIEE--LEEEIEAERAARAKVEK-----QRADLSRELEEISERLEEAGGATA 1159
Query: 207 SVFKPTKEQLYPRRDDYEQRRPWQSKSHRQREETDMVMNTDVSD 250
+ + K+ R ++++ R +S Q E T + + +D
Sbjct: 1160 AQIEMNKK----REAEFQKMRRDLEESTLQHEATAAALRKEQAD 1199
>PPIG_HUMAN (Q13427) Peptidyl-prolyl cis-trans isomerase G (EC
5.2.1.8) (Peptidyl-prolyl isomerase G) (PPIase G)
(Rotamase G) (Cyclophilin G) (Clk associating
RS-cyclophilin) (CARS-cyclophilin) (CARS-Cyp) (SRcyp)
(CASP10)
Length = 754
Score = 36.2 bits (82), Expect = 0.17
Identities = 50/243 (20%), Positives = 96/243 (38%), Gaps = 19/243 (7%)
Query: 126 LAFKNGLLPGGLNSKLTRKPARSMGEMRARASTYILDEEDEAFKRKRAKLEKGDTSPKRV 185
L ++G G + R P+RS R R S E + +R ++ G+ + +
Sbjct: 325 LVTRSGRKIKGRGPRRYRTPSRSRSRDRFRRSETPPHWRQEMQRAQRMRVSSGE---RWI 381
Query: 186 KKDRSGEDKGDGKQQRPGKGKSVFKPTKEQLYPRRDDYEQRRPWQSKSHRQR-EETDMVM 244
K D+S ++ Q+ P + K K T + + + + + K H+ +E D+
Sbjct: 382 KGDKSELNEIKENQRSPVRVKER-KITDHRNVSESPNRKNEKEKKVKDHKSNSKERDIRR 440
Query: 245 NTDVSDMLRG-----ASDANLVDEPEAPKYQPRDANPKKWCEFH---RSAGHDTDDCWTL 296
N++ D + A + E K + RD+ + E RS G D ++
Sbjct: 441 NSEKDDKYKNKVKKRAKSKSRSKSKEKSKSKERDSKHNRNEEKRMRSRSKGRDHENVKEK 500
Query: 297 QREIDKLIRAGYQGNRQGQWRNGGDQNKAHKREEERADTKGKKKQESAAIATKGADDTFA 356
+++ D +G Q + R+ + + E +K K+K A ++ D T
Sbjct: 501 EKQSDS------KGKDQERSRSKEKSKQLESKSNEHDHSKSKEKDRRAQSRSRECDITKG 554
Query: 357 RHS 359
+HS
Sbjct: 555 KHS 557
>NKX1_RAT (Q9QZM6) Sodium/potassium/calcium exchanger 1 precursor
(Na(+)/K(+)/Ca(2+)-exchange protein 1) (Retinal rod
Na-Ca+K exchanger)
Length = 1181
Score = 36.2 bits (82), Expect = 0.17
Identities = 39/194 (20%), Positives = 72/194 (37%), Gaps = 23/194 (11%)
Query: 162 DEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQ-----QRPGKGKSVFKPTKEQL 216
D E E + +G+T + + ++ GE + +GK+ + +GK V + +
Sbjct: 788 DHEGETEAEGKEVEHEGETEAEGTEDEQEGETEAEGKEVEQEGETEAEGKEVEHEVETEA 847
Query: 217 YPRRDDYEQRRPWQSKSHRQREETDMVMNTDVSDMLR--------GASDANLVDEPEAPK 268
+ ++E + K ET+ N + G ++A DE E
Sbjct: 848 ERKETNHEGETEAEGKEADHEGETEAEGNVEHQGETEAEGKVEHEGETEAGEKDEHEGQS 907
Query: 269 YQPRDANPKKWCEFHRSAGHDTDDCWTLQREIDKLIRAGYQGNRQGQWRNGGDQNKAHKR 328
D K E A + D C T Q G +G G +GGD +
Sbjct: 908 ETQADDTEVKDGEGEAEANAE-DQCETAQ---------GEKGADGGGGSDGGDSEEEEDE 957
Query: 329 EEERADTKGKKKQE 342
E+E + + ++++E
Sbjct: 958 EDEEEEEEEEEEEE 971
Score = 33.1 bits (74), Expect = 1.5
Identities = 38/196 (19%), Positives = 78/196 (39%), Gaps = 9/196 (4%)
Query: 162 DEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSVFKPTKE---QLYP 218
+E + + + K E+G+T +R + + E + GK+++ G+ +S K +E +
Sbjct: 725 EEGERETEAEGKKDEEGETEAERKEDGQEEETETKGKEKQEGETESEGKDEQEGETEAEG 784
Query: 219 RRDDYEQRRPWQSKSHRQREETDMVMNTDVSDMLRGASDANLVDEPEAPKYQPRDANPKK 278
+ D+E + K ET+ D + A + E E + + ++ +
Sbjct: 785 KEADHEGETEAEGKEVEHEGETEAEGTEDEQEGETEAEGKEVEQEGET-EAEGKEVEHEV 843
Query: 279 WCEFHRSAGHDTDDCWTLQREIDKLIRAGYQGNRQGQWRNGGDQNKAHKREEERADTKGK 338
E R + + +E D +GN + Q + +A + E +T+
Sbjct: 844 ETEAERKETNHEGETEAEGKEADHEGETEAEGNVEHQ-----GETEAEGKVEHEGETEAG 898
Query: 339 KKQESAAIATKGADDT 354
+K E + ADDT
Sbjct: 899 EKDEHEGQSETQADDT 914
>MYHB_HUMAN (P35749) Myosin heavy chain, smooth muscle isoform (SMMHC)
Length = 1972
Score = 36.2 bits (82), Expect = 0.17
Identities = 29/106 (27%), Positives = 47/106 (43%), Gaps = 4/106 (3%)
Query: 142 TRKPARSMGEMRARASTYI--LDEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDK--GDG 197
TR + EMR + + + L E+ E FKR +A L+K + ++ D +GE + G
Sbjct: 1187 TRSHEAQVQEMRQKHAQAVEELTEQLEQFKRAKANLDKNKQTLEKENADLAGELRVLGQA 1246
Query: 198 KQQRPGKGKSVFKPTKEQLYPRRDDYEQRRPWQSKSHRQREETDMV 243
KQ+ K K + +E D R K H+ + E + V
Sbjct: 1247 KQEVEHKKKKLEAQVQELQSKCSDGERARAELNDKVHKLQNEVESV 1292
>IF3A_HUMAN (Q14152) Eukaryotic translation initiation factor 3
subunit 10 (eIF-3 theta) (eIF3 p167) (eIF3 p180) (eIF3
p185) (eIF3a)
Length = 1382
Score = 36.2 bits (82), Expect = 0.17
Identities = 42/220 (19%), Positives = 86/220 (39%), Gaps = 26/220 (11%)
Query: 154 ARASTY---ILDEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSVFK 210
AR S Y + E+ + + +LE+ K ++ +K + +Q+R
Sbjct: 767 ARQSVYEEKLKQFEERLAEERHNRLEERKRQRKEERRITYYREKEEEEQRR--------- 817
Query: 211 PTKEQLYPRRDDYEQRRPWQSKSHRQREETDMVMNTDVSDMLRGASDANLVDEPEAPKYQ 270
+EQ+ R++ E R + RE + V + + + + + + + +
Sbjct: 818 -AEEQMLKEREERE-RAERAKREEELREYQERVKKLEEVERKKRQRELEIEERERRREEE 875
Query: 271 PRDANPKKWCEFHRSAGHDTDDCWTLQREIDKLIRAGYQGNRQGQWRNGG--DQNKAHKR 328
R + + R D++ W E D R +G + +WR G D++++H+R
Sbjct: 876 RRLGDSSLSRKDSRWGDRDSEGTWRKGPEADSEWR---RGPPEKEWRRGEGRDEDRSHRR 932
Query: 329 EEER-------ADTKGKKKQESAAIATKGADDTFARHSGP 361
+EER D + + + + +G DD GP
Sbjct: 933 DEERPRRLGDDEDREPSLRPDDDRVPRRGMDDDRGPRRGP 972
>TOP2_CAEEL (Q23670) Probable DNA topoisomerase II (EC 5.99.1.3)
Length = 1520
Score = 35.8 bits (81), Expect = 0.23
Identities = 30/135 (22%), Positives = 59/135 (43%), Gaps = 13/135 (9%)
Query: 96 QQEGESLKEYMARYSAASVKVEDEE------PRACALAFKNGLL-----PGGLNSKLTRK 144
++EG+ +K++M+ + + K E + + F G+ G + K
Sbjct: 1312 KKEGQDIKKFMSPAAPKTAKKEKSDGFNSDMSEESDVEFDEGIDFDSDDDGVEREDVVSK 1371
Query: 145 PARSMGEMRARASTYILDEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRPGK 204
P G+ A+A L ++DE +K A +K +PK+ K + S GD ++ K
Sbjct: 1372 PKPRTGKGAAKAEVIDLSDDDEVPAKKPAPAKK--AAPKKKKSEFSDLSGGDSDEEAEKK 1429
Query: 205 GKSVFKPTKEQLYPR 219
+ KP+ ++ P+
Sbjct: 1430 PSTSKKPSPKKAAPK 1444
>SG1_MOUSE (P16014) Secretogranin I precursor (SgI) (Chromogranin B)
(CgB)
Length = 677
Score = 35.8 bits (81), Expect = 0.23
Identities = 24/111 (21%), Positives = 55/111 (48%), Gaps = 7/111 (6%)
Query: 130 NGLLPGGLNSKLTRKPARSMGEMRARASTYILDEEDEAFKRKRAKLEKGDTSPKRVK--- 186
+G P ++K + A+ E ARA + ++ E++ R+++ E G+ + ++ K
Sbjct: 190 SGEKPNTFSNKRSEASAKKKDESVARADAHSMELEEKTHSREQSSQESGEETRRQEKPQE 249
Query: 187 ---KDRSGEDKGDGKQQRPG-KGKSVFKPTKEQLYPRRDDYEQRRPWQSKS 233
+D+S E+ +G++ G +G+ + + ++ RR + R +KS
Sbjct: 250 LTDQDQSQEESQEGEEGEEGEEGEEGEEDSASEVTKRRPRHHHGRSGSNKS 300
>RU17_XENLA (P09406) U1 small nuclear ribonucleoprotein 70 kDa (U1
snRNP 70 kDa) (snRNP70) (U1-70K)
Length = 471
Score = 35.8 bits (81), Expect = 0.23
Identities = 42/192 (21%), Positives = 77/192 (39%), Gaps = 17/192 (8%)
Query: 162 DEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSVFKPTKEQLYPRRD 221
+ E E +R+R++ + +R + R E++ +++ K K K K++ RR
Sbjct: 239 EREKEPRERRRSRSRER----RRKSRSREKEERKRTREKSKDKDKEKDKDNKDRDRKRRS 294
Query: 222 DYEQRRPWQSKSHRQREE--------TDMVMNTDVSDMLRGASDANLVDEPEAPKYQ--- 270
+R+ + + ++EE D D G L EPE +
Sbjct: 295 RSRERKRERDRDREKKEERVEAEVPEADDAPQDDAQIGDLGIDGIELKQEPEEKSRERDR 354
Query: 271 PRDANPKKWCEFHRSAGHDTD-DCWTLQREIDKLIRAGYQGNRQGQWRNGGDQNKAHKRE 329
RD + +K E R D D D R+ D+ +R+ +++K HKRE
Sbjct: 355 ERDRDREKG-EKDRDKDRDRDRDRRRSHRDRDREKDRDRDRDRRRDRDRDRERDKDHKRE 413
Query: 330 EERADTKGKKKQ 341
+R D K+++
Sbjct: 414 RDRGDRSEKREE 425
>MYS_AEQIR (P24733) Myosin heavy chain, striated muscle
Length = 1938
Score = 35.8 bits (81), Expect = 0.23
Identities = 39/144 (27%), Positives = 67/144 (46%), Gaps = 30/144 (20%)
Query: 82 KNQPVTINDLYNIRQQEGESLKEYMARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKL 141
K+ + DL ++++ E+++ A S+ + K+EDE+ L S+L
Sbjct: 1057 KSTQENVEDLERVKRELEENVRRKEAEISSLNSKLEDEQ---------------NLVSQL 1101
Query: 142 TRKPARSMGEMRARASTYILDEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQR 201
RK + E++AR L+EE EA + RAK+EK +R + +R E+ G+ +
Sbjct: 1102 QRK----IKELQARIEE--LEEELEAERNARAKVEK-----QRAELNRELEELGERLDEA 1150
Query: 202 PGKGKSVFKPTK----EQLYPRRD 221
G + + K E L RRD
Sbjct: 1151 GGATSAQIELNKKREAELLKIRRD 1174
>MNN4_YEAST (P36044) MNN4 protein
Length = 1178
Score = 35.8 bits (81), Expect = 0.23
Identities = 22/114 (19%), Positives = 54/114 (47%), Gaps = 1/114 (0%)
Query: 160 ILDEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSVFKPTKEQLYPR 219
+L+E K+K+ + EK + KK + E+K +++ K + K KE+ +
Sbjct: 1033 LLEERKRREKKKKEEEEKKKKEEEE-KKKKEEEEKKKKEEEEKKKKEEEEKKKKEEEEKK 1091
Query: 220 RDDYEQRRPWQSKSHRQREETDMVMNTDVSDMLRGASDANLVDEPEAPKYQPRD 273
+ + E+++ + + +++EE + + N D + + +E E K + ++
Sbjct: 1092 KQEEEEKKKKEEEEKKKQEEGEKMKNEDEENKKNEDEEKKKNEEEEKKKQEEKN 1145
>TOP1_CERAE (Q7YR26) DNA topoisomerase I (EC 5.99.1.2)
Length = 767
Score = 35.4 bits (80), Expect = 0.29
Identities = 20/76 (26%), Positives = 42/76 (54%), Gaps = 8/76 (10%)
Query: 162 DEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRP---GKGKSVFKPTKEQLYP 218
DE+D +K K+ K T + +K R E++ DGK ++P K K V +P ++ P
Sbjct: 143 DEDDADYKPKKIK-----TEDIKKEKKRKLEEEEDGKLRKPKNKDKDKKVPEPDNKKKKP 197
Query: 219 RRDDYEQRRPWQSKSH 234
++++ ++ + W+ + +
Sbjct: 198 KKEEEQKWKWWEEERY 213
Score = 31.6 bits (70), Expect = 4.3
Identities = 25/125 (20%), Positives = 52/125 (41%), Gaps = 7/125 (5%)
Query: 162 DEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSVFKPTKEQLYP--- 218
D+ + K KR + + + ++KK++ + + + F P KE + P
Sbjct: 81 DKHKDRDKEKRKEEKVRASGDAKIKKEKENGFSSPPQIKDEPEDDGYFVPPKEDIKPLKR 140
Query: 219 -RRDDYEQRRPWQSKSHRQREETDMVMNTDVSDMLRGASDANL---VDEPEAPKYQPRDA 274
R +D +P + K+ ++E + + LR + + V EP+ K +P+
Sbjct: 141 PRDEDDADYKPKKIKTEDIKKEKKRKLEEEEDGKLRKPKNKDKDKKVPEPDNKKKKPKKE 200
Query: 275 NPKKW 279
+KW
Sbjct: 201 EEQKW 205
>SIK1_YEAST (Q12460) SIK1 protein (Nucleolar protein NOP56)
Length = 504
Score = 35.4 bits (80), Expect = 0.29
Identities = 22/66 (33%), Positives = 38/66 (57%), Gaps = 4/66 (6%)
Query: 144 KPARSMGEMRARASTYI--LDEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQR 201
KPA + E + + S+ L+++DE K K+ K K + K+ KKD+ ++K D K+++
Sbjct: 439 KPAAEVEETKEKESSKKRKLEDDDEEKKEKKEKKSKKEKKEKKEKKDK--KEKKDKKEKK 496
Query: 202 PGKGKS 207
K KS
Sbjct: 497 DKKKKS 502
>ATRX_CAEEL (Q9U7E0) Transcriptional regulator ATRX homolog
(X-linked nuclear protein-1)
Length = 1359
Score = 35.4 bits (80), Expect = 0.29
Identities = 45/228 (19%), Positives = 86/228 (36%), Gaps = 41/228 (17%)
Query: 143 RKPARSMGEMRARASTYILDEEDEAFK------RKRAKLE------------KGDTSPKR 184
++PA+ + +A +S D+E+E+ + RKRAK E K S K+
Sbjct: 53 KRPAK---KRKASSSEEDDDDEEESPRKSSKKSRKRAKSESESDESDEEEDRKKSKSKKK 109
Query: 185 V----------KKDRSGEDKGDGKQQRPGKGKSVFKPTKEQLYPRRDDYEQRRPWQSKSH 234
V K+ S + D ++R K K K TK+Q + + KS
Sbjct: 110 VDQKKKEKSKKKRTTSSSEDEDSDEEREQKSKKKSKKTKKQTSSESSEESEEERKVKKSK 169
Query: 235 RQREETDMVMNTDVSDMLRGASDANLVDEPEAPKYQPRDANPKKWCEFHRSAGHDTDDCW 294
+ +E++ + + A + DE E P + + KK S D +
Sbjct: 170 KNKEKS----------VKKRAETSEESDEDEKPSKKSKKGLKKKAKSESESESEDEKEVK 219
Query: 295 TLQREIDKLIRAGYQGNRQGQWRNGGDQNKAHKREEERADTKGKKKQE 342
+++ K+++ + + + ++ K K E + K +E
Sbjct: 220 KSKKKSKKVVKKESESEDEAPEKKKTEKRKRSKTSSEESSESEKSDEE 267
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.315 0.132 0.387
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 55,071,098
Number of Sequences: 164201
Number of extensions: 2485479
Number of successful extensions: 8027
Number of sequences better than 10.0: 116
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 109
Number of HSP's that attempted gapping in prelim test: 7810
Number of HSP's gapped (non-prelim): 254
length of query: 441
length of database: 59,974,054
effective HSP length: 113
effective length of query: 328
effective length of database: 41,419,341
effective search space: 13585543848
effective search space used: 13585543848
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.5 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0590c.5


