

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
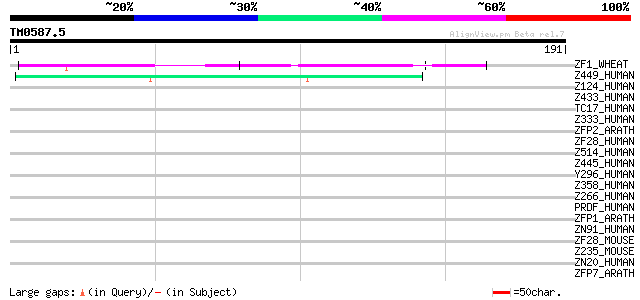
Query= TM0587.5
(191 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
ZF1_WHEAT (Q42430) Zinc-finger protein 1 (WZF1) 66 6e-11
Z449_HUMAN (Q6P9G9) Zinc finger protein 449 (Zinc finger and SCA... 42 9e-04
Z124_HUMAN (Q15973) Zinc finger protein 124 (HZF-16) 42 0.001
Z433_HUMAN (Q8N7K0) Zinc finger protein 433 41 0.001
TC17_HUMAN (O60765) Zinc finger protein 354A (Transcription fact... 40 0.003
Z333_HUMAN (Q96JL9) Zinc finger protein 333 39 0.007
ZFP2_ARATH (Q39261) Zinc finger protein 2 39 0.010
ZF28_HUMAN (Q8NHY6) Zinc finger protein 28 homolog (Zfp-28) (Kru... 39 0.010
Z514_HUMAN (Q96K75) Zinc finger protein 514 39 0.010
Z445_HUMAN (P59923) Zinc finger protein 445 39 0.010
Y296_HUMAN (O15015) Hypothetical zinc finger protein KIAA0296 39 0.010
Z358_HUMAN (Q9NW07) Zinc finger protein 358 38 0.012
Z266_HUMAN (Q14584) Zinc finger protein 266 (Zinc finger protein... 38 0.012
PRDF_HUMAN (P57071) PR-domain zinc finger protein 15 (Zinc finge... 38 0.012
ZFP1_ARATH (Q42485) Zinc finger protein 1 38 0.016
ZN91_HUMAN (Q05481) Zinc finger protein 91 (Zinc finger protein ... 37 0.021
ZF28_MOUSE (P10078) Zinc finger protein 28 (Zfp-28) (mKR5 protein) 37 0.021
Z235_MOUSE (Q61116) Zinc finger protein 235 (Zinc finger protein... 37 0.021
ZN20_HUMAN (P17024) Zinc finger protein 20 (Zinc finger protein ... 37 0.028
ZFP7_ARATH (Q39266) Zinc finger protein 7 37 0.028
>ZF1_WHEAT (Q42430) Zinc-finger protein 1 (WZF1)
Length = 261
Score = 65.9 bits (159), Expect = 6e-11
Identities = 45/142 (31%), Positives = 62/142 (42%), Gaps = 24/142 (16%)
Query: 4 SHSHTQKHVVPSPSS--SPSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKF 61
S Q+ P P S +P +CS CGK F S +AL GH H +
Sbjct: 67 SRGGKQRVQAPQPESFAAPVPAEFKCSVCGKSFSSYQALGGHKTSHRVK----------- 115
Query: 62 RRPSQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSI 121
QP PP A +L + A T +SS +++ H+CSI
Sbjct: 116 ------QPSPPSDAAAAPLVALPAVAAILPSAEPATSSTAASSDG-----ATNRVHRCSI 164
Query: 122 CLRVFSSGQALGGHKRRHWEKG 143
C + F +GQALGGHKR+H++ G
Sbjct: 165 CQKEFPTGQALGGHKRKHYDGG 186
Score = 45.1 bits (105), Expect = 1e-04
Identities = 29/85 (34%), Positives = 41/85 (48%), Gaps = 8/85 (9%)
Query: 80 EEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRH 139
EE +A CLL+L+ + + + E + KCS+C + FSS QALGGHK H
Sbjct: 55 EEENLALCLLMLSRGGKQ--RVQAPQPESFAAPVPAEFKCSVCGKSFSSYQALGGHKTSH 112
Query: 140 WEKGGDLIHHHLPPPAPAPAPVLDL 164
+ PP A AP++ L
Sbjct: 113 ------RVKQPSPPSDAAAAPLVAL 131
>Z449_HUMAN (Q6P9G9) Zinc finger protein 449 (Zinc finger and SCAN
domain containing protein 19)
Length = 518
Score = 42.0 bits (97), Expect = 9e-04
Identities = 37/160 (23%), Positives = 60/160 (37%), Gaps = 20/160 (12%)
Query: 3 NSHSHTQKHVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCH--------PEREWRG 54
NS+ + P SP K +C +CGK F + L GH R H PE R
Sbjct: 301 NSNLEEPLNPKPHKKKSPGEKPHRCPQCGKCFARKSQLTGHQRIHSGEEPHKCPECGKRF 360
Query: 55 INPPPKFRRPSQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTC------------S 102
+ +R + P + ++ + L+ +++ +T S
Sbjct: 361 LRSSDLYRHQRLHTGERPYECTVCKKRFTRRSHLIGHQRTHSEEETYKCLECGKSFCHGS 420
Query: 103 SSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRHWEK 142
S K + + H+C C + FS AL H+R H E+
Sbjct: 421 SLKRHLKTHTGEKPHRCHNCGKSFSRLTALTLHQRTHTEE 460
>Z124_HUMAN (Q15973) Zinc finger protein 124 (HZF-16)
Length = 296
Score = 41.6 bits (96), Expect = 0.001
Identities = 38/137 (27%), Positives = 52/137 (37%), Gaps = 20/137 (14%)
Query: 5 HSHTQKHVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRGINPPPK-FRR 63
H H + H P C ECGK F +L GH++ H E K FR
Sbjct: 167 HYHERTHTGEKPYV--------CMECGKAFSCLSSLQGHIKAHAGEEPYPCKQCGKAFRY 218
Query: 64 PSQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSS-KSEISCSSSHHHHKCSIC 122
S Q + T I Q+ + + N KG CSSS + + ++C C
Sbjct: 219 ASSLQKH--EKTHIAQKPY--------VCNNCGKGFRCSSSLRDHERTHTGEKPYECQKC 268
Query: 123 LRVFSSGQALGGHKRRH 139
+ FS L HK+ H
Sbjct: 269 GKAFSRASTLWKHKKTH 285
Score = 28.9 bits (63), Expect = 7.6
Identities = 30/136 (22%), Positives = 46/136 (33%), Gaps = 24/136 (17%)
Query: 26 QCSECGKKFWSEKALHGHMRCHP-EREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEV 84
+C ECGK ++L+ H R H E+ + F R S + + T ++ +E
Sbjct: 96 ECMECGKALGFSRSLNRHKRIHTGEKRYECKQCGKAFSRSSHLRDH--ERTHTGEKPYEC 153
Query: 85 AACLLLLANANA-----------KGDTCSSSKSEISCSSSHHHH----------KCSICL 123
C +N K C SC SS H C C
Sbjct: 154 KHCGKAFRYSNCLHYHERTHTGEKPYVCMECGKAFSCLSSLQGHIKAHAGEEPYPCKQCG 213
Query: 124 RVFSSGQALGGHKRRH 139
+ F +L H++ H
Sbjct: 214 KAFRYASSLQKHEKTH 229
>Z433_HUMAN (Q8N7K0) Zinc finger protein 433
Length = 673
Score = 41.2 bits (95), Expect = 0.001
Identities = 34/136 (25%), Positives = 55/136 (40%), Gaps = 18/136 (13%)
Query: 5 HSHTQKHVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHP-EREWRGINPPPKFRR 63
H+H + H P +C +CGK F S + H R H E+ + F+
Sbjct: 270 HAHKRTHTGEKPY--------ECKQCGKAFSSSHSFQIHERTHTGEKPYECKECGKAFKC 321
Query: 64 PSQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICL 123
PS + + T ++ +E C +L+ + + EIS HKC IC
Sbjct: 322 PSSVRRH--ERTHSRKKPYECKHCGKVLSYLTSFQNHLGMHTGEIS-------HKCKICG 372
Query: 124 RVFSSGQALGGHKRRH 139
+ F S +L H++ H
Sbjct: 373 KAFYSPSSLQTHEKTH 388
Score = 30.0 bits (66), Expect = 3.4
Identities = 35/157 (22%), Positives = 49/157 (30%), Gaps = 32/157 (20%)
Query: 5 HSHTQKHVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHP-EREWRGINPPPKFRR 63
H H + H GK +C +CG+ F + H R H E+ + FR
Sbjct: 494 HRHERTHT--------GGKTYECKQCGRSFNCSSSFRYHGRTHTGEKPYECKQCGKAFRS 545
Query: 64 PSQPQPQPPQTTVITQEEHEVAACLLLLANAN-----------AKGDTCSSSKSEISCSS 112
SQ Q T ++ +E C +A+ K C C+S
Sbjct: 546 ASQLQIH--GRTHTGEKPYECKQCGKAFGSASHLQMHGRTHTGEKPYECKQCGKSFGCAS 603
Query: 113 SHHHH----------KCSICLRVFSSGQALGGHKRRH 139
H KC C + F L H R H
Sbjct: 604 RLQMHGRTHTGEKPYKCKQCGKAFGCPSNLRRHGRTH 640
>TC17_HUMAN (O60765) Zinc finger protein 354A (Transcription factor
17) (Zinc finger protein eZNF)
Length = 605
Score = 40.4 bits (93), Expect = 0.003
Identities = 37/118 (31%), Positives = 48/118 (40%), Gaps = 18/118 (15%)
Query: 27 CSECGKKFWSEKALHGHMRCHP-EREWRGINPPPKFRRPSQPQPQPPQTTVITQEEH-EV 84
C ECGK F +L+ H+R H E+ +R F R S I Q+ H E
Sbjct: 272 CKECGKAFTLSTSLYKHLRTHTVEKSYRCKECGKSFSRRS--------GLFIHQKIHAEE 323
Query: 85 AACLLLLANANAKGDTCSSSKSEISCSSSHHHHK---CSICLRVFSSGQALGGHKRRH 139
C N K +CS+S S C H K C+ C F S +L H+R H
Sbjct: 324 NPCKY---NPGRKASSCSTSLS--GCQRIHSRKKSYLCNECGNTFKSSSSLRYHQRIH 376
Score = 37.4 bits (85), Expect = 0.021
Identities = 36/145 (24%), Positives = 50/145 (33%), Gaps = 36/145 (24%)
Query: 5 HSHTQKHVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRP 64
+ H + H V K +C ECGK F L H + H E NP K
Sbjct: 286 YKHLRTHTVE--------KSYRCKECGKSFSRRSGLFIHQKIHAEENPCKYNPGRKASSC 337
Query: 65 SQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHH------- 117
S ++ C + ++ K C+ + SSS +H
Sbjct: 338 ST----------------SLSGCQRI--HSRKKSYLCNECGNTFKSSSSLRYHQRIHTGE 379
Query: 118 ---KCSICLRVFSSGQALGGHKRRH 139
KCS C R FS +L H+R H
Sbjct: 380 KPFKCSECGRAFSQSASLIQHERIH 404
Score = 30.8 bits (68), Expect = 2.0
Identities = 27/121 (22%), Positives = 42/121 (34%), Gaps = 22/121 (18%)
Query: 26 QCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEVA 85
+C+ECGK F S L+ H H ++ N K +T+I E
Sbjct: 411 RCNECGKGFTSISRLNRHRIIHTGEKYYNCNECGK--------ALSSHSTLIIHER---- 458
Query: 86 ACLLLLANANAKGDTCSSSKSEISCSSSHHH-------HKCSICLRVFSSGQALGGHKRR 138
+ K C + + S H +KC+ C + F +L H+R
Sbjct: 459 ---IHTGEKPCKCKVCGKAFRQSSALIQHQRMHTGERPYKCNECGKTFRCNSSLSNHQRI 515
Query: 139 H 139
H
Sbjct: 516 H 516
>Z333_HUMAN (Q96JL9) Zinc finger protein 333
Length = 665
Score = 38.9 bits (89), Expect = 0.007
Identities = 36/138 (26%), Positives = 53/138 (38%), Gaps = 14/138 (10%)
Query: 3 NSHSHTQKHVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRGINPPPK-F 61
N S + HV P +CS+CGK F +L H+R H + N K F
Sbjct: 460 NQPSSLRSHVRTHTGEKPF----ECSQCGKAFREHSSLKTHLRTHTREKPYECNQCGKPF 515
Query: 62 RRPSQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSI 121
R + T ++ +E A C +L+ S+ KS + + + C
Sbjct: 516 RTSTHLNVHKRIHT--GEKLYECATCGQVLSR-------LSTLKSHMRTHTGEKPYVCQE 566
Query: 122 CLRVFSSGQALGGHKRRH 139
C R FS +L H R H
Sbjct: 567 CGRAFSEPSSLRKHARTH 584
Score = 30.0 bits (66), Expect = 3.4
Identities = 28/115 (24%), Positives = 46/115 (39%), Gaps = 10/115 (8%)
Query: 26 QCSECGKKFWSEKALHGHMRCHP-EREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEV 84
+CS+CGK F L H R H E+ + + F +PS + T ++ E
Sbjct: 423 ECSQCGKTFTRNFNLILHQRNHTGEKPYECKDCGKAFNQPSSLRSH--VRTHTGEKPFEC 480
Query: 85 AACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRH 139
+ C SS K+ + + ++C+ C + F + L HKR H
Sbjct: 481 SQCGKAFREH-------SSLKTHLRTHTREKPYECNQCGKPFRTSTHLNVHKRIH 528
Score = 28.9 bits (63), Expect = 7.6
Identities = 13/41 (31%), Positives = 21/41 (50%)
Query: 11 HVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHPERE 51
H++ + +G+ QC++C K F +L H R H RE
Sbjct: 604 HLIVHVRTHSAGRPYQCNQCEKAFRHSSSLTVHKRTHVGRE 644
>ZFP2_ARATH (Q39261) Zinc finger protein 2
Length = 150
Score = 38.5 bits (88), Expect = 0.010
Identities = 19/52 (36%), Positives = 26/52 (49%)
Query: 88 LLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRH 139
L L+ ++ + SSS + SC C+ C R F S QALGGH+ H
Sbjct: 23 LELVLEPSSMSSSSSSSTNSSSCLEQPRVFSCNYCQRKFYSSQALGGHQNAH 74
Score = 29.3 bits (64), Expect = 5.8
Identities = 17/42 (40%), Positives = 20/42 (47%), Gaps = 9/42 (21%)
Query: 15 SPSSSPSGKWSQCSE---------CGKKFWSEKALHGHMRCH 47
S SSS S S C E C +KF+S +AL GH H
Sbjct: 33 SSSSSSSTNSSSCLEQPRVFSCNYCQRKFYSSQALGGHQNAH 74
>ZF28_HUMAN (Q8NHY6) Zinc finger protein 28 homolog (Zfp-28)
(Kruppel-like zinc finger factor X6)
Length = 868
Score = 38.5 bits (88), Expect = 0.010
Identities = 37/142 (26%), Positives = 54/142 (37%), Gaps = 19/142 (13%)
Query: 4 SHSHTQK-HVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHP-EREWRGINPPPKF 61
S + TQK H+ + K +C ECGK F L H R H E+ ++ + F
Sbjct: 651 SKAFTQKAHLAQHQKTHTGEKPYECKECGKAFSQTTHLIQHQRVHTGEKPYKCMECGKAF 710
Query: 62 RRPSQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHH----H 117
S Q Q +E C +KS + C H +
Sbjct: 711 GDNSSCTQH--QRLHTGQRPYECIEC-----------GKAFKTKSSLICHRRSHTGEKPY 757
Query: 118 KCSICLRVFSSGQALGGHKRRH 139
+CS+C + FS Q+L H+R H
Sbjct: 758 ECSVCGKAFSHRQSLSVHQRIH 779
Score = 28.9 bits (63), Expect = 7.6
Identities = 10/22 (45%), Positives = 13/22 (58%)
Query: 26 QCSECGKKFWSEKALHGHMRCH 47
+C+ECGK F + H RCH
Sbjct: 449 KCNECGKAFSDGSSFARHQRCH 470
>Z514_HUMAN (Q96K75) Zinc finger protein 514
Length = 400
Score = 38.5 bits (88), Expect = 0.010
Identities = 35/129 (27%), Positives = 51/129 (39%), Gaps = 16/129 (12%)
Query: 15 SPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHP-EREWRGINPPPKFRRPSQPQPQPPQ 73
S + P K +C+ECGK F + L H RCH E+ + + F S Q
Sbjct: 194 SQRTHPEKKSCKCNECGKSFHFQSELRRHQRCHTGEKPYECSDCGRAFGHISSLIKH--Q 251
Query: 74 TTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSH---HHHKCSICLRVFSSGQ 130
T ++ +E + C G S S S + H +KC+ C R F
Sbjct: 252 RTHTGEKPYECSEC----------GRAFSQSSSLVLHYRFHTGEKPYKCNECGRAFGHTS 301
Query: 131 ALGGHKRRH 139
+L H+R H
Sbjct: 302 SLIKHQRTH 310
Score = 33.1 bits (74), Expect = 0.40
Identities = 34/118 (28%), Positives = 46/118 (38%), Gaps = 16/118 (13%)
Query: 26 QCSECGKKFWSEKALHGHMRCHP-EREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEV 84
+CSECG+ F +L H R H E+ ++ F S Q T ++ +E
Sbjct: 261 ECSECGRAFSQSSSLVLHYRFHTGEKPYKCNECGRAFGHTSSLIKH--QRTHTGEKPYEC 318
Query: 85 AACLLLLANANAKGDTCSSSKSEISCSSSH---HHHKCSICLRVFSSGQALGGHKRRH 139
C G T S S S I H +KC+ C R FS +L H R H
Sbjct: 319 REC----------GRTFSQSSSLIVHYRFHTGEKPYKCNKCGRAFSQSSSLTQHYRFH 366
>Z445_HUMAN (P59923) Zinc finger protein 445
Length = 1031
Score = 38.5 bits (88), Expect = 0.010
Identities = 32/117 (27%), Positives = 41/117 (34%), Gaps = 16/117 (13%)
Query: 26 QCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEVA 85
+C CGK+F L H R H +PP R S + + EE +
Sbjct: 897 KCQWCGKEFIGRHTLSSHQRKHTRAAQAERSPPA---RSSSQDTKLRLQKLKPSEEMPLE 953
Query: 86 ACLLLLANANAKGDTCSSSKSEISC---SSSHHHHKCSICLRVFSSGQALGGHKRRH 139
C + CS S S HKCSIC + F+ L HKR H
Sbjct: 954 DCK----------EACSQSSRLTGLQDISIGKKCHKCSICGKTFNKSSQLISHKRFH 1000
Score = 30.0 bits (66), Expect = 3.4
Identities = 31/136 (22%), Positives = 50/136 (35%), Gaps = 14/136 (10%)
Query: 7 HTQKHVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQ 66
H + +V+ K +C++C K F HMR H E ++ + + R S
Sbjct: 607 HCKSYVLEHQRIHTQEKPYKCTKCRKTFRWRSNFTRHMRLHEEEKFYKQDECREGFRQS- 665
Query: 67 PQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHK---CSICL 123
P PQ ++ C G T + K+ + H K CS C
Sbjct: 666 PDCSQPQGAPAVEKTFLCQQC----------GKTFTRKKTLVDHQRIHTGEKPYQCSDCG 715
Query: 124 RVFSSGQALGGHKRRH 139
+ F+ A HK++H
Sbjct: 716 KDFAYRSAFIVHKKKH 731
>Y296_HUMAN (O15015) Hypothetical zinc finger protein KIAA0296
Length = 1829
Score = 38.5 bits (88), Expect = 0.010
Identities = 34/118 (28%), Positives = 44/118 (36%), Gaps = 28/118 (23%)
Query: 26 QCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEVA 85
+CSECGK F K L H R H ER RG K R +P
Sbjct: 1259 RCSECGKAFRLRKQLASHQRVHMER--RGGGGTRKATREDRP--------------FRCG 1302
Query: 86 ACLLLLANANAKGDTCSSSKSEISCSSSHH--HHKCSICLRVFSSGQALGGHKRRHWE 141
C G T + S ++ SH + C C + +S+ AL H+R H E
Sbjct: 1303 QC----------GRTYRHAGSLLNHRRSHETGQYSCPTCPKTYSNRMALKDHQRLHSE 1350
Score = 37.7 bits (86), Expect = 0.016
Identities = 33/119 (27%), Positives = 50/119 (41%), Gaps = 13/119 (10%)
Query: 27 CSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEVAA 86
CS C K+ ++ AL H+R H R +G+ + +PS P P P T +E E
Sbjct: 403 CSLCSKQLFNAAALKNHVRAH-HRPRQGVG---ENGQPSVP-PAPLLLAETTHKEEEDPT 457
Query: 87 CLLLLANANAKGDTCSSSKSEISCSSSHHH------HKCSICLRVFSSGQALGGHKRRH 139
L + K C + +H H ++CS+C R + + AL H R H
Sbjct: 458 --TTLDHRPYKCSECGRAYRHRGSLVNHRHSHRTGEYQCSLCPRKYPNLMALRNHVRVH 514
Score = 34.7 bits (78), Expect = 0.14
Identities = 20/52 (38%), Positives = 23/52 (43%), Gaps = 4/52 (7%)
Query: 27 CSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQTTVIT 78
C+ CGK F + AL HMR H R PP RP + P TV T
Sbjct: 77 CTTCGKDFSNPMALKSHMRTHAPEGRRRHRPP----RPKEATPHLQGETVST 124
Score = 32.3 bits (72), Expect = 0.69
Identities = 32/136 (23%), Positives = 50/136 (36%), Gaps = 21/136 (15%)
Query: 26 QCSECGKKFWSEKALHGHMRCHPEREWRGIN-----PPPKFRRPSQPQPQPPQTTVITQE 80
QCS C +K+ + AL H+R H + R + P + P P + T +
Sbjct: 493 QCSLCPRKYPNLMALRNHVRVHCKAARRSADIGAEGAPSHLKVELPPDPVEAEAAPHTDQ 552
Query: 81 EHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRH- 139
+H + A T ++ K+ H CSIC +F ++L H H
Sbjct: 553 DH------VCKHEEEATDITPAADKTAA--------HICSICGLLFEDAESLERHGLTHG 598
Query: 140 -WEKGGDLIHHHLPPP 154
EK + PP
Sbjct: 599 AGEKENSRTETTMSPP 614
Score = 31.6 bits (70), Expect = 1.2
Identities = 28/85 (32%), Positives = 33/85 (37%), Gaps = 15/85 (17%)
Query: 6 SHTQKHVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRGINP--PPKFRR 63
+H++ H P K CS CGK F + L GH R H RE P P FRR
Sbjct: 1721 NHSRTHTDP--------KRHCCSICGKAFRTAARLEGHGRVHAPREGPFTCPHCPRHFRR 1772
Query: 64 -----PSQPQPQPPQTTVITQEEHE 83
Q Q Q T + HE
Sbjct: 1773 RISFVQHQQQHQEEWTVAGSGRGHE 1797
Score = 30.8 bits (68), Expect = 2.0
Identities = 29/129 (22%), Positives = 51/129 (39%), Gaps = 16/129 (12%)
Query: 27 CSECGKKFWSEKALHGHMRCHPE-REWRGINPPPKFRRP------------SQPQPQPPQ 73
C+ C K+F + AL H R H + R + + P FR P + +P+ +
Sbjct: 268 CAICFKEFSNLMALKNHSRLHAQYRPYHCPHCPRVFRLPRELLEHQQSHEGERQEPRWEE 327
Query: 74 TTVITQEEHEVAACLLLLANA---NAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQ 130
+ T H + L +A N + +S + E S + +C C R +
Sbjct: 328 KGMPTTNGHTDESSQDQLPSAQMLNGSAELSTSGELEDSGLEEYRPFRCGDCGRTYRHAG 387
Query: 131 ALGGHKRRH 139
+L H++ H
Sbjct: 388 SLINHRKSH 396
Score = 30.4 bits (67), Expect = 2.6
Identities = 16/50 (32%), Positives = 21/50 (42%)
Query: 103 SSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRHWEKGGDLIHHHLP 152
S K+ + H CSIC + F + L GH R H + G H P
Sbjct: 1718 SLKNHSRTHTDPKRHCCSICGKAFRTAARLEGHGRVHAPREGPFTCPHCP 1767
Score = 29.3 bits (64), Expect = 5.8
Identities = 16/46 (34%), Positives = 20/46 (42%), Gaps = 4/46 (8%)
Query: 3 NSHSHTQKHVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHP 48
NS SH+ V + +G CS+CG F L H CHP
Sbjct: 803 NSSSHSANAV----TGWQAGAAHTCSDCGHSFPHATGLLSHRPCHP 844
>Z358_HUMAN (Q9NW07) Zinc finger protein 358
Length = 481
Score = 38.1 bits (87), Expect = 0.012
Identities = 36/133 (27%), Positives = 51/133 (38%), Gaps = 12/133 (9%)
Query: 8 TQKHVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHP-EREWRGINPPPKFRRPSQ 66
T V+P+P+S P + C +CG+ F L H R H E+ +R + F +
Sbjct: 49 TSPAVLPAPASPP--RPFSCPDCGRAFRRSSGLSQHRRTHSGEKPYRCPDCGKSFSHGAT 106
Query: 67 PQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVF 126
T ++ AAC A G + K S S HH C +C + F
Sbjct: 107 LAQHRGIHT--GARPYQCAAC------GKAFGWRSTLLKHRSSHSGEKPHH-CPVCGKAF 157
Query: 127 SSGQALGGHKRRH 139
G L H R H
Sbjct: 158 GHGSLLAQHLRTH 170
Score = 33.9 bits (76), Expect = 0.24
Identities = 30/114 (26%), Positives = 45/114 (39%), Gaps = 10/114 (8%)
Query: 27 CSECGKKFWSEKALHGHMRCH-PEREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEVA 85
C +CGK F AL H R H ER +R + F + S Q T + +
Sbjct: 206 CPQCGKAFGQSSALLQHQRTHTAERPYRCPHCGKAFGQSSNLQHHLRIHT--GERPYACP 263
Query: 86 ACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRH 139
C + A G + S+ + S ++C +C + F +L HKR H
Sbjct: 264 HC------SKAFGQS-SALLQHLHVHSGERPYRCQLCGKAFGQASSLTKHKRVH 310
>Z266_HUMAN (Q14584) Zinc finger protein 266 (Zinc finger protein
HZF1) (Fragment)
Length = 200
Score = 38.1 bits (87), Expect = 0.012
Identities = 30/126 (23%), Positives = 45/126 (34%), Gaps = 32/126 (25%)
Query: 26 QCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEVA 85
+C ECGK F +L+ HMR H ++ P T + + +
Sbjct: 88 ECLECGKAFTHSSSLNNHMRTHSAKK--------------------PFTCMECGKAFKFP 127
Query: 86 AC--LLLLANANAKGDTCSSSKSEISCSSSHHHH----------KCSICLRVFSSGQALG 133
C L + + K C S S+S H +C C + FSS +
Sbjct: 128 TCVNLHMRIHTGEKPYKCKQCGKSFSYSNSFQLHERTHTGEKPYECKECGKAFSSSSSFR 187
Query: 134 GHKRRH 139
H+RRH
Sbjct: 188 NHERRH 193
Score = 29.3 bits (64), Expect = 5.8
Identities = 29/119 (24%), Positives = 42/119 (34%), Gaps = 12/119 (10%)
Query: 26 QCSECGKKFWSEKALHGHMRCHP-EREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEV 84
+C +CGK F L H R H ER + F R S+ T ++ E
Sbjct: 4 KCKDCGKAFTQNSDLTKHARTHSGERPYECKECGKAFARSSRLSEH--TRTHTGEKPFEC 61
Query: 85 AACLLLLANANAKGDTCSSSKS-EISCSSSHHHHKCSICLRVFSSGQALGGHKRRHWEK 142
C K SS+ S + + +C C + F+ +L H R H K
Sbjct: 62 VKC--------GKAFAISSNLSGHLRIHTGEKPFECLECGKAFTHSSSLNNHMRTHSAK 112
>PRDF_HUMAN (P57071) PR-domain zinc finger protein 15 (Zinc finger
protein 298)
Length = 1507
Score = 38.1 bits (87), Expect = 0.012
Identities = 27/117 (23%), Positives = 45/117 (38%), Gaps = 4/117 (3%)
Query: 26 QCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEVA 85
QC+ C K F + L H+R H ++ ++ F R + Q +EV
Sbjct: 739 QCNICSKIFQNSSNLSRHVRSHGDKLFKCEECAKLFSRKESLK----QHVSYKHSRNEVD 794
Query: 86 ACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRHWEK 142
K S+ +C + +C +C R FS+ L HK++H +K
Sbjct: 795 GEYRYRCGTCEKTFRIESALEFHNCRTDDKTFQCEMCFRFFSTNSNLSKHKKKHGDK 851
>ZFP1_ARATH (Q42485) Zinc finger protein 1
Length = 228
Score = 37.7 bits (86), Expect = 0.016
Identities = 23/71 (32%), Positives = 31/71 (43%), Gaps = 11/71 (15%)
Query: 80 EEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHK-----------CSICLRVFSS 128
++ E+ LL A + S S S+SHH H+ C+ C R F S
Sbjct: 20 QDLELGLTLLSRGTATSSELNLIDSFKTSSSSTSHHQHQQEQLADPRVFSCNYCQRKFYS 79
Query: 129 GQALGGHKRRH 139
QALGGH+ H
Sbjct: 80 SQALGGHQNAH 90
Score = 28.9 bits (63), Expect = 7.6
Identities = 10/21 (47%), Positives = 14/21 (66%)
Query: 27 CSECGKKFWSEKALHGHMRCH 47
C+ C +KF+S +AL GH H
Sbjct: 70 CNYCQRKFYSSQALGGHQNAH 90
>ZN91_HUMAN (Q05481) Zinc finger protein 91 (Zinc finger protein
HTF10) (HPF7)
Length = 1191
Score = 37.4 bits (85), Expect = 0.021
Identities = 30/121 (24%), Positives = 45/121 (36%), Gaps = 22/121 (18%)
Query: 26 QCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEVA 85
+C ECGK F S L+GH R H + P K + Q++ +T+ +
Sbjct: 1051 KCEECGKAFISSSTLNGHKRIHTREK------PYKCEECGKAF---SQSSTLTRHKR--- 1098
Query: 86 ACLLLLANANAKGDTCSSSKSEISCSSSH-------HHHKCSICLRVFSSGQALGGHKRR 138
L K C + E S + H +KC C + F+ L HK+
Sbjct: 1099 ---LHTGEKPYKCGECGKAFKESSALTKHKIIHTGEKPYKCEKCCKAFNQSSILTNHKKI 1155
Query: 139 H 139
H
Sbjct: 1156 H 1156
Score = 32.3 bits (72), Expect = 0.69
Identities = 32/124 (25%), Positives = 47/124 (37%), Gaps = 28/124 (22%)
Query: 26 QCSECGKKFWSEKALHGHMRCHP-EREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEV 84
+C ECGK F L H R H E+ ++ F R S+ +T +
Sbjct: 995 KCEECGKAFSQSSTLTRHTRMHTGEKPYKCEECGKAFNRSSK----------LTTHK--- 1041
Query: 85 AACLLLLANANAKGDTCSSSKSEISCSSSHHH---------HKCSICLRVFSSGQALGGH 135
++ K + C K+ IS S+ + H +KC C + FS L H
Sbjct: 1042 ---IIHTGEKPYKCEEC--GKAFISSSTLNGHKRIHTREKPYKCEECGKAFSQSSTLTRH 1096
Query: 136 KRRH 139
KR H
Sbjct: 1097 KRLH 1100
Score = 29.3 bits (64), Expect = 5.8
Identities = 30/116 (25%), Positives = 42/116 (35%), Gaps = 12/116 (10%)
Query: 26 QCSECGKKFWSEKALHGHMRCHP-EREWRGINPPPKFRRPSQPQPQPPQTTVITQEE-HE 83
+C ECGK F L H R H E+ ++ F SQ + T E+ ++
Sbjct: 911 KCEECGKAFSQPSHLTTHKRMHTGEKPYKCEECGKAF---SQSSTLTTHKIIHTGEKPYK 967
Query: 84 VAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRH 139
C A K T + K + +KC C + FS L H R H
Sbjct: 968 CEEC----GKAFRKSSTLTEHK---IIHTGEKPYKCEECGKAFSQSSTLTRHTRMH 1016
>ZF28_MOUSE (P10078) Zinc finger protein 28 (Zfp-28) (mKR5 protein)
Length = 825
Score = 37.4 bits (85), Expect = 0.021
Identities = 37/137 (27%), Positives = 52/137 (37%), Gaps = 17/137 (12%)
Query: 8 TQK-HVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHP-EREWRGINPPPKFRRPS 65
TQK H+ + K +C ECGK F L H R H E+ ++ + F S
Sbjct: 612 TQKAHLAQHQKTHTGEKPYECKECGKAFSQTTHLIQHQRVHTGEKPYKCLECGKAFGDNS 671
Query: 66 QPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEIS---CSSSHHHHKCSIC 122
T Q +E C G T + S I C + ++CS C
Sbjct: 672 SCTQHRRLHT--GQRPYECVEC----------GKTFKTKSSLICHRRCHTGEKPYECSAC 719
Query: 123 LRVFSSGQALGGHKRRH 139
+ FS Q+L H+R H
Sbjct: 720 GKAFSHRQSLSVHQRIH 736
>Z235_MOUSE (Q61116) Zinc finger protein 235 (Zinc finger protein
93) (Zfp-93)
Length = 645
Score = 37.4 bits (85), Expect = 0.021
Identities = 42/147 (28%), Positives = 54/147 (36%), Gaps = 26/147 (17%)
Query: 16 PSSSPSGKWSQCSECGKKFWSEKALHGHMRCHP-EREWRGINPPPKFRRPS--------Q 66
PS P K C ECGK F AL H R H E+ +R + F R S
Sbjct: 276 PSVHPGRKRYWCHECGKGFRQSSALQTHQRVHTGEKPYRCDSCGKGFSRSSDLNIHRRVH 335
Query: 67 PQPQPPQTTV----ITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHH----- 117
+P + V TQ H A + + K C SCSS+ H H
Sbjct: 336 TGEKPYKCEVCGKGFTQWAHLQAHERI---HTGEKPYKCGDCGKRFSCSSNLHTHQRVHT 392
Query: 118 -----KCSICLRVFSSGQALGGHKRRH 139
+C+ C + FS L H+R H
Sbjct: 393 EEKPYECNECGKRFSLSGNLDIHQRVH 419
Score = 30.4 bits (67), Expect = 2.6
Identities = 29/115 (25%), Positives = 44/115 (38%), Gaps = 12/115 (10%)
Query: 27 CSECGKKFWSEKALHGHMRCHP-EREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEVA 85
CS CGK F H R H E+ +R +F P Q+ ++ ++
Sbjct: 455 CSVCGKNFSRSSHFLDHQRIHTGEKPYRCEVCGKRF--PWSLSLHSHQSVHTGKKPYKCG 512
Query: 86 ACLLLLANANAKG-DTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRH 139
C KG SS ++ S + KC++C + FS L H+R H
Sbjct: 513 EC--------GKGFSHASSLQAHHSVHTGEKPFKCNVCQKQFSKTSNLQAHQRVH 559
>ZN20_HUMAN (P17024) Zinc finger protein 20 (Zinc finger protein
KOX13) (DKFZp572P0920)
Length = 532
Score = 37.0 bits (84), Expect = 0.028
Identities = 38/156 (24%), Positives = 54/156 (34%), Gaps = 28/156 (17%)
Query: 6 SHTQKHVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRGINPPPK-FRRP 64
SH QKH P +C +CGK F L H + H E + G K FR
Sbjct: 324 SHLQKHGRTHTGEKPY----ECRQCGKAFRCTSDLQRHEKTHTEDKPYGCKQCGKGFRCA 379
Query: 65 SQPQPQPPQTTVITQEEHEVAACLLLL-----------ANANAKGDTCSSSKSEISCSSS 113
SQ Q + T ++ HE C + + K C SS
Sbjct: 380 SQLQIH--ERTHSGEKPHECKECGKVFKYFSSLRIHERTHTGEKPHECKQCGKAFRYFSS 437
Query: 114 HHHH----------KCSICLRVFSSGQALGGHKRRH 139
H H +C +C + F+ ++ H+R H
Sbjct: 438 LHIHERTHTGDKPYECKVCGKAFTCSSSIRYHERTH 473
Score = 31.6 bits (70), Expect = 1.2
Identities = 36/138 (26%), Positives = 48/138 (34%), Gaps = 16/138 (11%)
Query: 4 SHSHTQKHVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHP-EREWRGINPPPKFR 62
S S Q H + P +C +CGK F L H R H E+ + FR
Sbjct: 294 SFSSIQYHKMTHTGEKPY----ECKQCGKAFRCGSHLQKHGRTHTGEKPYECRQCGKAFR 349
Query: 63 RPSQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSS-KSEISCSSSHHHHKCSI 121
S Q T E + C KG C+S + S H+C
Sbjct: 350 CTSDLQRHEK-----THTEDKPYGC-----KQCGKGFRCASQLQIHERTHSGEKPHECKE 399
Query: 122 CLRVFSSGQALGGHKRRH 139
C +VF +L H+R H
Sbjct: 400 CGKVFKYFSSLRIHERTH 417
Score = 29.3 bits (64), Expect = 5.8
Identities = 34/146 (23%), Positives = 47/146 (31%), Gaps = 32/146 (21%)
Query: 4 SHSHTQKHVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRR 63
SHS Q+H V P +C CGK F+ H R H G+ P ++
Sbjct: 182 SHSCIQRHRVMHSGDGPY----KCKFCGKAFYFLNLCLIHERIHT-----GVKP---YKC 229
Query: 64 PSQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHK----- 118
+ TT+ E + D C + S S HK
Sbjct: 230 KQCGKAFTRSTTLPVHER----------THTGVNADECKECGNAFSFPSEIRRHKRSHTG 279
Query: 119 -----CSICLRVFSSGQALGGHKRRH 139
C C +VF S ++ HK H
Sbjct: 280 EKPYECKQCGKVFISFSSIQYHKMTH 305
>ZFP7_ARATH (Q39266) Zinc finger protein 7
Length = 209
Score = 37.0 bits (84), Expect = 0.028
Identities = 20/64 (31%), Positives = 25/64 (38%), Gaps = 10/64 (15%)
Query: 99 DTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRH----------WEKGGDLIH 148
DT + C ++ C+ C R F S QALGGH+ H G H
Sbjct: 41 DTFNDDTKSTKCEANPRVFSCNYCRRKFYSSQALGGHQNAHKRERTMAKRAMHMGRMFGH 100
Query: 149 HHLP 152
HH P
Sbjct: 101 HHRP 104
Score = 28.9 bits (63), Expect = 7.6
Identities = 10/21 (47%), Positives = 14/21 (66%)
Query: 27 CSECGKKFWSEKALHGHMRCH 47
C+ C +KF+S +AL GH H
Sbjct: 61 CNYCRRKFYSSQALGGHQNAH 81
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.132 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,330,311
Number of Sequences: 164201
Number of extensions: 1382612
Number of successful extensions: 14097
Number of sequences better than 10.0: 427
Number of HSP's better than 10.0 without gapping: 258
Number of HSP's successfully gapped in prelim test: 171
Number of HSP's that attempted gapping in prelim test: 9737
Number of HSP's gapped (non-prelim): 3641
length of query: 191
length of database: 59,974,054
effective HSP length: 104
effective length of query: 87
effective length of database: 42,897,150
effective search space: 3732052050
effective search space used: 3732052050
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0587.5


