

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0546.13
(1118 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
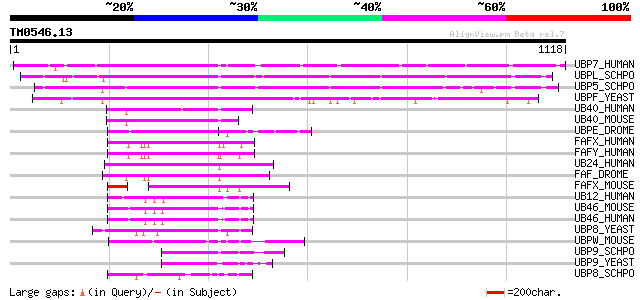
Score E
Sequences producing significant alignments: (bits) Value
UBP7_HUMAN (Q93009) Ubiquitin carboxyl-terminal hydrolase 7 (EC ... 625 e-178
UBPL_SCHPO (Q9UTT1) Ubiquitin carboxyl-terminal hydrolase 21 (EC... 608 e-173
UBP5_SCHPO (Q09879) Probable ubiquitin carboxyl-terminal hydrola... 557 e-158
UBPF_YEAST (P50101) Ubiquitin carboxyl-terminal hydrolase 15 (EC... 485 e-136
UB40_HUMAN (Q9NVE5) Ubiquitin carboxyl-terminal hydrolase 40 (EC... 168 9e-41
UB40_MOUSE (Q8BWR4) Ubiquitin carboxyl-terminal hydrolase 40 (EC... 161 8e-39
UBPE_DROME (Q24574) Ubiquitin carboxyl-terminal hydrolase 64E (E... 160 2e-38
FAFX_HUMAN (Q93008) Probable ubiquitin carboxyl-terminal hydrola... 143 3e-33
FAFY_HUMAN (O00507) Probable ubiquitin carboxyl-terminal hydrola... 141 9e-33
UB24_HUMAN (Q9UPU5) Ubiquitin carboxyl-terminal hydrolase 24 (EC... 139 6e-32
FAF_DROME (P55824) Probable ubiquitin carboxyl-terminal hydrolas... 126 4e-28
FAFX_MOUSE (P70398) Probable ubiquitin carboxyl-terminal hydrola... 124 2e-27
UB12_HUMAN (O75317) Ubiquitin carboxyl-terminal hydrolase 12 (EC... 114 2e-24
UB46_MOUSE (P62069) Ubiquitin carboxyl-terminal hydrolase 46 (EC... 110 2e-23
UB46_HUMAN (P62068) Ubiquitin carboxyl-terminal hydrolase 46 (EC... 110 2e-23
UBP8_YEAST (P50102) Ubiquitin carboxyl-terminal hydrolase 8 (EC ... 105 7e-22
UBPW_MOUSE (Q61068) Ubiquitin carboxyl-terminal hydrolase DUB-1 ... 94 3e-18
UBP9_SCHPO (Q9P7V9) Probable ubiquitin carboxyl-terminal hydrola... 93 5e-18
UBP9_YEAST (P39967) Ubiquitin carboxyl-terminal hydrolase 9 (EC ... 86 6e-16
UBP8_SCHPO (Q09738) Probable ubiquitin carboxyl-terminal hydrola... 82 6e-15
>UBP7_HUMAN (Q93009) Ubiquitin carboxyl-terminal hydrolase 7 (EC
3.1.2.15) (Ubiquitin thiolesterase 7) (Ubiquitin-specific
processing protease 7) (Deubiquitinating enzyme 7)
(Herpesvirus associated ubiquitin-specific protease)
Length = 1102
Score = 625 bits (1612), Expect = e-178
Identities = 412/1139 (36%), Positives = 616/1139 (53%), Gaps = 82/1139 (7%)
Query: 9 IDQQEDEEMLVPHTDLPENNHQPMEV---VAQPEAAPTVESQPVEEPP---QSRFTWRID 62
+ + ED EM TD P Q + VA + T E ++ ++ F + ++
Sbjct: 17 LSEPEDMEMEAGDTDDPPRITQNPVINGNVALSDGHNTAEEDMEDDTSWRSEATFQFTVE 76
Query: 63 NFSRMNVKKLYSEVFVVGGYKWRVLIFPK-------GNNVDYLSMYLDVADSTNLPYGWS 115
FSR++ L FV W++++ P+ +V + +DST+ WS
Sbjct: 77 RFSRLSESVLSPPCFV-RNLPWKIMVMPRFYPDRPHQKSVGFFLQCNAESDSTS----WS 131
Query: 116 RYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGYLLNDTLVV 175
+AQ L ++N ++ + + H F +E+DWGF++FM E+ DP +G++ +D V
Sbjct: 132 CHAQAVLKIINYRDDEKSFSRRISHLFFHKENDWGFSNFMAWSEVTDPEKGFIDDDK--V 189
Query: 176 EAEVLVRRIVDYWT-YDSKKETGYVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTE 234
EV V+ + +DSKK TGYVGLKNQGATCYMNSLLQTL+ RKAVY MPT E
Sbjct: 190 TFEVFVQADAPHGVAWDSKKHTGYVGLKNQGATCYMNSLLQTLFFTNQLRKAVYMMPT-E 248
Query: 235 NDMPAGSIPLALQSLFYKLQYSDTSVATKELTKSFGWDTYDSFMQHDVQELNRVLCEKLE 294
D + S+PLALQ +FY+LQ+SD V TK+LTKSFGW+T DSFMQHDVQEL RVL + +E
Sbjct: 249 GDDSSKSVPLALQRVFYELQHSDKPVGTKKLTKSFGWETLDSFMQHDVQELCRVLLDNVE 308
Query: 295 DKMKGTVVEGTIQKLFEGHHMNYIECINVDYKSTRKESFYDLQLDVKGCHDVYASFDKYV 354
+KMKGT VEGTI KLF G ++YI+C VDY+S R+E +YD+QL +KG +++ SF YV
Sbjct: 309 NKMKGTCVEGTIPKLFRGKMVSYIQCKEVDYRSDRREDYYDIQLSIKGKKNIFESFVDYV 368
Query: 355 EVEPLEGDNKYHAEQYGLQDAKKGVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRYEFP 414
VE L+GDNKY A ++GLQ+A+KGV F+ PPVL LQL RF YD D +KINDR+EFP
Sbjct: 369 AVEQLDGDNKYDAGEHGLQEAEKGVKFLTLPPVLHLQLMRFMYDPQTDQNIKINDRFEFP 428
Query: 415 LELDLDRDDGKYLSPDADRNVRNLYTLHSVLVHSGGVHGGHYYAFIRPTLSDQWYKFDDE 474
+L LD ++L ++ N Y LH+VLVHSG HGGHY ++ P +W KFDD+
Sbjct: 429 EQLPLD----EFLQKTDPKDPAN-YILHAVLVHSGDNHGGHYVVYLNPKGDGKWCKFDDD 483
Query: 475 RVTKEDTKRALEEQYGGEEELPQTNPGFNNTPFKFTKYSNAYMLVYIREADKDKVICNVD 534
V++ + A+E YGG ++ +NAYMLVYIRE+ +V+ V
Sbjct: 484 VVSRCTKEEAIEHNYGGHDD-----------DLSVRHCTNAYMLVYIRESKLSEVLQAVT 532
Query: 535 EKDIAEHLRERLKKEQEEKEHKKKEKAEAHLYTIIKVARNEDLKEQIGKDIYFDLVDHDK 594
+ DI + L ERL++E+ + K+KE+ EAHLY +++ + G D+Y D +K
Sbjct: 533 DHDIPQQLVERLQEEKRIEAQKRKERQEAHLYMQVQIVAEDQFCGHQGNDMY----DEEK 588
Query: 595 VR--SFRVQKQMSFNLFKEEVAKEFGIPVQFQRFWLWAKRQNHTYRPNRPLTPAEEAQSV 652
V+ F+V K S F + +++ G P R W R N T RP A+ +++
Sbjct: 589 VKYTVFKVLKNSSLAEFVQSLSQTMGFPQDQIRLWPMQARSNGTKRPAMLDNEADGNKTM 648
Query: 653 GQVREVSNKVHNAELKLFLEVELGPDLR---PIAPSDKTKDDILLFFKLYDPEKEELRYV 709
++ + N +FLE + P+L P D++LF K+YDP+ L Y
Sbjct: 649 IELSDNEN-----PWTIFLET-VDPELAASGATLPKFDKDHDVMLFLKMYDPKTRSLNYC 702
Query: 710 GRLFVKSTGKPSEILTRLNEMAGYDPDEEIGLYEEIKFEPNVMCEPIDKKLTFRASQLED 769
G ++ + K ++L + + AG+ D + LYEE+K + D L +L D
Sbjct: 703 GHIYTPISCKIRDLLPVMCDRAGFIQDTSLILYEEVKPNLTERIQDYDVSLDKALDELMD 762
Query: 770 GDIICFQK-APAMDSEEHVRYPDVPSYLEYVHNRQVVHFRSLDKPKEDDFCLEMSRLYTY 828
GDII FQK P D+ E P Y +++R V F P + F + +S Y
Sbjct: 763 GDIIVFQKDDPENDNSE---LPTAKEYFRDLYHRVDVIFCDKTIPNDPGFVVTLSNRMNY 819
Query: 829 DDVVEKVAQQLNLDDPSKIRLTPHNCYSQQPKPQPIKYRGVDHLSDMLVHYN-QTSDILY 887
V + VAQ+LN DP ++ Y P P+++ L D+L + + LY
Sbjct: 820 FQVAKTVAQRLN-TDPMLLQFFKSQGYRDGP-GNPLRHNYEGTLRDLLQFFKPRQPKKLY 877
Query: 888 YEILDIPLPELQGLKTLKVAFYHAT-KDEVVSHTIRLPKQSTVGDVLDDLKTKVELSH-P 945
Y+ L + + + + ++ K + ++ ++E + T+ K V D+L++ K VEL
Sbjct: 878 YQQLKMKITDFENRRSFKCIWLNSQFREEEI--TLYPDKHGCVRDLLEECKKAVELGEKA 935
Query: 946 NAELRLLEVFYHKIYKVFPPNEKIETIND-QYWTLRAEEVPEEEKNLG-PHDRLIHVYHF 1003
+ +LRLLE+ +KI V +E +E ++ T R EE+P ++ ++ ++ L+ V HF
Sbjct: 936 SGKLRLLEIVSYKIIGVHQEDELLECLSPATSRTFRIEEIPLDQVDIDKENEMLVTVAHF 995
Query: 1004 TKDTSQNQMQIQNFGEPFFLVIREGETLTEIKVRIQKKLQVPDDEFEKWKFAFFALGRPE 1063
K+ FG PF L I +GE E+ RIQ L + + EFEK+KFA GR +
Sbjct: 996 HKEV------FGTFGIPFLLRIHQGEHFREVMKRIQSLLDIQEKEFEKFKFAIVMTGRHQ 1049
Query: 1064 YLQDSDIVSNRFQRRDVYGAWEQ---YLGLEHTDNAPKRSYAVNQNRHTF-EKPVKIYN 1118
Y+ + + N G +LGL+H + APKRS R+T+ EK +KI+N
Sbjct: 1050 YINEDEYEVNLKDFEPQPGNMSHPRPWLGLDHFNKAPKRS------RYTYLEKAIKIHN 1102
>UBPL_SCHPO (Q9UTT1) Ubiquitin carboxyl-terminal hydrolase 21 (EC
3.1.2.15) (Ubiquitin thiolesterase 21)
(Ubiquitin-specific processing protease 21)
(Deubiquitinating enzyme 21)
Length = 1129
Score = 608 bits (1567), Expect = e-173
Identities = 407/1130 (36%), Positives = 612/1130 (54%), Gaps = 97/1130 (8%)
Query: 23 DLPENNHQPMEVVAQPEAAPT-VESQP-VEEPPQSRFTWRIDNFSRMNVKKLYSEVFVVG 80
+L E++ +P+ E + V +P +EE + ++W + NFS + K YS +F G
Sbjct: 18 ELEESSQEPLRADNYEEIYNSLVHHEPDLEEAAHASYSWVVKNFSTLE-DKTYSPLFKAG 76
Query: 81 GYKWRVLIFPKG-NNVDYLSMYLDVADS----------TNLPYG--------------WS 115
WR+++FPKG N +Y S++L+ L G +S
Sbjct: 77 HTTWRIVLFPKGCNQTEYASVFLEYLPQCKVEAIRKYEAELAAGKTPTIDPEIVNDETYS 136
Query: 116 RYAQFSLAVVNQIQNKYTVRKDTQH-QFNARESDWGFTSFMPLGELYDPSRGY----LLN 170
AQF+L++ N +Q+ ++ +T H +F + DWGFT F+ L ++ P+ + L N
Sbjct: 137 CCAQFALSLSN-VQDPTVMQINTSHHRFRSEVKDWGFTRFVDLRKIAVPTPEFPVPFLEN 195
Query: 171 DTLVVEAEVLVRRIVD--------YWTYDSKKETGYVGLKNQGATCYMNSLLQTLYHIPY 222
D + + V VR + D + Y+SKKETGYVGLKNQGATCYMNSLLQ+L+
Sbjct: 196 DEICIS--VTVRVLQDPTGVLWHSFVNYNSKKETGYVGLKNQGATCYMNSLLQSLFFTNI 253
Query: 223 FRKAVYHMPTTENDMPAGSIPLALQSLFYKLQYSDTSVATKELTKSFGWDTYDSFMQHDV 282
FRK VY +PT +ND S+ ALQ +FY L+ V+T ELT+SFGW+++DSFMQHD+
Sbjct: 254 FRKTVYKIPT-DNDDSRDSVAYALQRVFYNLEKQREPVSTTELTRSFGWNSFDSFMQHDI 312
Query: 283 QELNRVLCEKLEDKMKGTVVEGTIQKLFEGHHMNYIECINVDYKSTRKESFYDLQLDVKG 342
QE NRVL + LE KMKGT VE + +F G +Y++CI+V+Y+S+R E F+D+QL+VKG
Sbjct: 313 QEFNRVLQDNLEKKMKGTEVENALNDIFVGKMKSYVKCIDVNYESSRVEDFWDIQLNVKG 372
Query: 343 CHDVYASFDKYVEVEPLEGDNKYHAEQYGLQDAKKGVLFIDFPPVLQLQLKRFEYDFMRD 402
+ SF ++VE L GDNKY+AE +GLQDA KG++F P VLQLQLKRF+YD +RD
Sbjct: 373 MDTLEDSFRDAIQVETLTGDNKYYAEGHGLQDAHKGIIFESLPNVLQLQLKRFDYDMLRD 432
Query: 403 TMVKINDRYEFPLELDLDRDDGKYLSPDADRNVRNLYTLHSVLVHSGGVHGGHYYAFIRP 462
MVKINDR+EFPLE+DL+ YLS AD++ ++Y LH VLVH G +HGGHYYA I+P
Sbjct: 433 MMVKINDRHEFPLEIDLE----PYLSETADKSESHVYVLHGVLVHGGDLHGGHYYALIKP 488
Query: 463 TLSDQWYKFDDERVTKEDTKRALEEQYGGEEELPQTNPGFNNTPFKFTKYSNAYMLVYIR 522
W+KFDD+RVT+ K LE+ YGGE G+N PFK ++ NAYMLVY R
Sbjct: 489 EKDSNWFKFDDDRVTRATIKEVLEDNYGGEP--AGRAKGYNGNPFK--RFMNAYMLVYFR 544
Query: 523 EADKDKVICNVDEKDIAEHLRERLKKEQEEKEHKKKEKAEAHLYTIIKVARNEDLKEQIG 582
++ D ++ V +D+ H+R L +E E K E+ E +Y ++V + K+ G
Sbjct: 545 KSRLDHILSPVTAEDVPFHVRNTLDEEHRVVERKLLEREEQQIYRRVRVLTTDGFKKYHG 604
Query: 583 KDIY-FDLVDHDKVR-SFRVQKQMSFNLFKEEVAKEFGIPVQFQRFWLWAKRQNHTYRPN 640
D+ F D D V + ++++ + ++ +A R WL RQN T R +
Sbjct: 605 FDMTDFSASDDDPVLITTKIKRNANIWDLQKHLAGLLNRDTSGIRIWLMTNRQNRTVRVD 664
Query: 641 RPLTPAEEAQSVGQVREVSNKVHNAELKLFLEV--ELGPDLRPIAPSDKTKDDILLFFKL 698
PL ++ V Q+ ++ + + ++++++E E L +D D +F K+
Sbjct: 665 LPLD--KKTILVDQICDMHIR-KDMDMRVYVEFLSEHNQLLADFGATDDNDFDTYIFLKI 721
Query: 699 YDPEKEELRYVGRLFVKSTGKPSEILTRLNEMAGYDPDEEIGLYEEIKFEPNVMCEPIDK 758
+D E +++ + L V S + + E + D I YEEIK M + +D
Sbjct: 722 FDYETQQISGLADLHVSKNSPISSLSEWIREHLKWSSDVPITYYEEIK---TGMVDVLDP 778
Query: 759 KLTFRASQLEDGDIICFQKAPAMDSEEHVRYP--DVPSYLEYVHNRQVVHF--RSLDKPK 814
+F S+++ GDIICF+K DS +P +++ +R V+ F R D
Sbjct: 779 NASFEKSEIQVGDIICFEKKLVHDSSSDTSHPYKSALDLYDFMAHRVVITFEPRYSDDTN 838
Query: 815 EDDFCLEMSRLYTYDDVVEKVAQQLNLDDPSKIRLTPHNCYSQQPKP---QPIKYRGVDH 871
F L ++ Y D+ VA +LN+ DP+ ++ T + S+ P+ P K+ +
Sbjct: 839 NGVFDLVLTTHTNYTDMARAVANKLNV-DPNYLQFTMAHLPSRTPRSVIRNPSKFTLQNA 897
Query: 872 LSDMLVHYNQTSDILYYEILDIPLPELQGLKTLKVAFYHATKDEVVSHTIRLPKQSTVGD 931
+ H NQ + +++YE+LDI L EL+ + ++V F + K+ TV D
Sbjct: 898 IPSTYSH-NQ-NVVMFYEVLDITLSELERKQLIRVHFLSNGISHETQMEFYVDKEGTVED 955
Query: 932 VLDDLKTKVELSHPNA-ELRLLEVFYHKIYKVFPPNEKIETINDQYWTLRAEEVPEEEK- 989
+L + KV L+ +A LRL EV+ H+I K P + I +N ++ T E P+EE+
Sbjct: 956 ILRQVTQKVPLNAEDASRLRLYEVYNHRILKSHLPTDGIYDLN-EFSTAYVEVTPKEEQM 1014
Query: 990 NLGPHDRL-IHVYHFTKDTSQNQMQIQNFGEPFFLVIREGETLTEIKVRIQKKLQVPDDE 1048
L D + I V HF KD S+ PF+ V+ GETL ++K R+QK+L D +
Sbjct: 1015 QLKTDDAVSIVVQHFFKDLSRLH------DIPFYFVLLRGETLKDLKKRLQKRLGYNDTQ 1068
Query: 1049 FEKWKFAFF---ALGRPEYLQDSDIVSNRFQRRDVYGAWE---QYLGLEH 1092
F K K A + G+P YL D D V +YG E LGL+H
Sbjct: 1069 FSKVKLAVLQAQSFGKPYYLTDDDEV--------LYGELEPQSHILGLDH 1110
>UBP5_SCHPO (Q09879) Probable ubiquitin carboxyl-terminal hydrolase 5
(EC 3.1.2.15) (Ubiquitin thiolesterase 5)
(Ubiquitin-specific processing protease 5)
(Deubiquitinating enzyme 5)
Length = 1108
Score = 557 bits (1435), Expect = e-158
Identities = 384/1091 (35%), Positives = 577/1091 (52%), Gaps = 79/1091 (7%)
Query: 50 EEPPQSRFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGN--NVDYLSMYLDV--A 105
EE RFTW I ++ ++ ++ S F VG ++++ FP+G + + S++L+ +
Sbjct: 51 EESGFQRFTWHIKSWHELD-RRAVSPQFAVGSRQFKITYFPQGTLQSAGFTSIFLEYIPS 109
Query: 106 DSTNLPYGWSRYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELY---D 162
+ L + QF+ + N + +V +F+ DWGFT F L +L
Sbjct: 110 EEEKLSNKYGCCCQFAFVISNPRKPSLSVANSAHCRFSPEIVDWGFTQFAELKKLLCRQA 169
Query: 163 PSRGYLLND-TLVVEAEVLVRRIV------DYWTYDSKKETGYVGLKNQGATCYMNSLLQ 215
P ++ D L++ A V + + + YDSK TGYVGLKNQGATCYMNSLLQ
Sbjct: 170 PDVPPIVEDGALLLTAYVRILKDPTGVLWHSFNDYDSKIATGYVGLKNQGATCYMNSLLQ 229
Query: 216 TLYHIPYFRKAVYHMPTTENDMPAG--SIPLALQSLFYKLQYSDTSVATKELTKSFGWDT 273
+LY I FR+ VY +PT D P G SI ALQ FY LQ+ + V+T ELTKSFGWD+
Sbjct: 230 SLYIIHAFRRIVYQIPT---DSPQGKDSIAYALQRCFYNLQFMNEPVSTTELTKSFGWDS 286
Query: 274 YDSFMQHDVQELNRVLCEKLEDKMKGTVVEGTIQKLFEGHHMNYIECINVDYKSTRKESF 333
DSFMQHDVQE NRVL + LE M+ T VE + LF G +YI C+NV+++S R E +
Sbjct: 287 LDSFMQHDVQEFNRVLQDNLERSMRDTKVENALTNLFVGKMKSYIACVNVNFESARSEDY 346
Query: 334 YDLQLDVKGCHDVYASFDKYVEVEPLEGDNKYHAEQYGLQDAKKGVLFIDFPPVLQLQLK 393
+D+QL+VKG ++ SF Y++VE LEGDN Y A+ YG Q+AKKGV+F FPP+L LQLK
Sbjct: 347 WDIQLNVKGMKNLEDSFRSYIQVETLEGDNCYFADTYGFQEAKKGVIFESFPPILHLQLK 406
Query: 394 RFEYDFMRDTMVKINDRYEFPLELDLDRDDGKYLSPDADRNVRNLYTLHSVLVHSGGVHG 453
RFEYDF RD M+KINDRYEFPLE D +LSP+AD++ Y L+ VLVHSG +H
Sbjct: 407 RFEYDFERDMMIKINDRYEFPLEFDAK----AFLSPEADQSQNCEYVLYGVLVHSGDLHN 462
Query: 454 GHYYAFIRPTLSDQWYKFDDERVTKEDTKRALEEQYGGEEELPQTNPGFNNTPFKFTKYS 513
GHYYA ++ WYK+DD RVT+ + LEE YGG+ + +P F +P K ++
Sbjct: 463 GHYYALLKTEKDGPWYKYDDTRVTRATLREVLEENYGGDYIM---HPPF-RSPVKLKRFM 518
Query: 514 NAYMLVYIREADKDKVICNVDEKDIAEHLRERLKKEQEEKEHKKKEKAEAHLYTIIKVAR 573
+AYML+Y+R+ D+++ V +I EHL+E L + E ++KE+ E+HLYT +++
Sbjct: 519 SAYMLLYLRKDKLDELMNPVSADEIPEHLKEALNPSIQLAELRRKERLESHLYTKVQLIT 578
Query: 574 NEDLKEQIGKDI--YFDLVDHDKVRSFRVQKQMSFNLFKEEVAKEFGIPVQFQRFWLWAK 631
E E DI + + + + FR++K+ F+ F VA++ G P + RFW K
Sbjct: 579 PEFYSEHHEFDIADFGNAYKEETIPQFRIKKEAKFSEFIPIVAEKLGYPQECMRFWYVVK 638
Query: 632 RQNHTYRPNRPLTPAEEAQSVGQVREVSNKVHNAELKLFLEVELGPDLRPIAPSDKTKD- 690
R N T R P+ E ++ +V+ V N L+L+LE+ +L T +
Sbjct: 639 RHNCTVRVESPVN--ELNSTMEEVKNVWNS-QGEILRLYLEITPENELSSSLTHQNTGEW 695
Query: 691 DILLFFKLYDPEKEELRYVGRLFVKSTGKPSEILTRLNEMAGYDPDEEIGLYEEIKFEPN 750
+ +F K +D + +E+ G L V + + I L E A + + +YEEIK P
Sbjct: 696 NAFIFVKYFDRKSQEISGCGTLHVNKSDEIRSICPLLCERANLPKNTPLNIYEEIK--PG 753
Query: 751 VMCEPIDKKLTFRASQLEDGDIICFQKAPAMDSEEHVRYPDVPSYL---EYVHNRQVVHF 807
M + + + TF S+L GDIICF+ E+ + S L +++ N+ +V F
Sbjct: 754 -MVDFLRLEKTFTQSELSTGDIICFEPCRPSALEDDIVNSGFDSALKLYDFLSNKVLVLF 812
Query: 808 RS--LDKPKEDDFCLEMSRLYTYDDVVEKVAQQLNLDDPSKIRLTPHN--CYSQ---QPK 860
R +D+ +F + + R YDD+ ++ Q+L + IRLT N YS P
Sbjct: 813 RPRFIDQDSIIEFEMLLDRRIKYDDLCIELGQKLGI-GADHIRLTTCNPLTYSAGMVVPN 871
Query: 861 PQPIKYRGVDHLSDMLVHYNQTSDILYYEILDIPLPELQGLKTLKVAFYHATKDEVVSHT 920
I + + S+ S++++YE +D+ L +L + +++ + D + +
Sbjct: 872 DSNITLYEILYSSE----EEMPSNVIFYETMDVSLSDLDRKRLVRLRW---LVDGLANIE 924
Query: 921 IRLPKQSTVGDVLDDLKTKVELSHPNAEL-----RLLEVFYHKIYKVFPPNEKIETINDQ 975
+ + GD+ +DL V P+++L R+ EVF + ++ I TIN
Sbjct: 925 LVEAYINKSGDI-NDLFGAVCERFPDSDLRKKKVRVYEVFESRYHRDLSLRTLIRTINPA 983
Query: 976 YWTLRAEEVPEEEKNLGPHDRLIHVYHFTKDTSQNQMQIQNFGEPFFLVIREGETLTEIK 1035
TL E VP ++ L P ++++ V+HF KD ++ G PF VI+ E + K
Sbjct: 984 A-TLVGEVVPLDQLQLYPEEKIVQVHHFHKDIARIH------GIPFSFVIKPQEKFIDTK 1036
Query: 1036 VRIQKKLQVPDDEFEKWKFAF--FALGRPEYLQDSDIVSNRFQRRDVYGAWEQYLGLEHT 1093
+R+ + Q P+ F KF F R YL D DI Y E+ G
Sbjct: 1037 LRLAARTQYPESIFSVIKFCVVDFDNNRVVYLNDEDI---------TYDVVEKLNGTLAL 1087
Query: 1094 DNAPKRSYAVN 1104
D A K S N
Sbjct: 1088 DRAKKDSKKPN 1098
>UBPF_YEAST (P50101) Ubiquitin carboxyl-terminal hydrolase 15 (EC
3.1.2.15) (Ubiquitin thiolesterase 15)
(Ubiquitin-specific processing protease 15)
(Deubiquitinating enzyme 15)
Length = 1230
Score = 485 bits (1249), Expect = e-136
Identities = 361/1155 (31%), Positives = 570/1155 (49%), Gaps = 158/1155 (13%)
Query: 47 QPVEEPPQSRFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNNVDYLSMYLD--- 103
+ VE + FTW I +++ + K S F +G ++W +L+FP+GN+ +++YL+
Sbjct: 32 EEVETLLEDSFTWNIPDWNELTNPKYNSPRFRIGDFEWDILLFPQGNHNKGVAVYLEPHP 91
Query: 104 -------VADSTNLPYGWSRYAQFSLAVVNQIQNKYTVR--KDTQHQFNARESDWGFTSF 154
+ + W AQF++ + ++ N T+ + H+FNA ++DWGF +
Sbjct: 92 EEKLDETTGEMVPVDPDWYCCAQFAIGI-SRPGNGDTINLINKSHHRFNALDTDWGFANL 150
Query: 155 MPLGELYDPSRG----YLLNDTLVVEAEVLVRRIV------DYWTYDSKKETGYVGLKNQ 204
+ L L PS+G +L TL + A V + + ++ YDSKK TGYVG +NQ
Sbjct: 151 IDLNNLKHPSKGRPLSFLNEGTLNITAYVRILKDPTGVLWHNFLNYDSKKVTGYVGFRNQ 210
Query: 205 GATCYMNSLLQTLYHIPYFRKAVYHMPTTENDMPAGSIPLALQSLFYKLQYSDTSVATKE 264
GATCY+NSLLQ+ + YFRK VY +PT E++ P S+PLALQ FY+LQ SD + T E
Sbjct: 211 GATCYLNSLLQSYFFTKYFRKLVYEIPT-EHESPNNSVPLALQRAFYQLQVSDIPLDTLE 269
Query: 265 LTKSFGWDTYDSFMQHDVQELNRVLCEKLEDKMKGTVVEGTIQKLFEGHHMNYIECINVD 324
LT+SFGWDT +SF QHDVQELNR+L ++LE+ MKGT VEG + ++F G +YI+CINVD
Sbjct: 270 LTRSFGWDTAESFTQHDVQELNRILMDRLENNMKGTPVEGKLNEIFVGKMKSYIKCINVD 329
Query: 325 YKSTRKESFYDLQLDVKGCHDVYASFDKYVEVEPLEGDNKYHAEQYGLQDAKKGVLFIDF 384
Y+S R E F+DLQL+VK ++ SFD Y+E+E + G+N+Y A+ YGLQDA+KGV+F F
Sbjct: 330 YESARVEDFWDLQLNVKNFKNLQESFDNYIEMELMNGENQYAAQDYGLQDAQKGVIFESF 389
Query: 385 PPVLQLQLKRFEYDFMRDTMVKINDRYEFPLELDL----DRDDGKYL--SPDADRNVRNL 438
PPVL LQLKRFEYDF D MVK+ND+YEFP +DL D+D K S + D+N +
Sbjct: 390 PPVLHLQLKRFEYDFNYDQMVKVNDKYEFPETIDLSPFVDKDVLKKTLDSENKDKN-PYV 448
Query: 439 YTLHSVLVHSGGVHGGHYYAFIRPTLSDQWYKFDDERVTKEDTKRALEEQYGGE---EEL 495
Y LH VLVHSG + GHYY I+P + DQWY+FDDERV + K+ +E +G + +E
Sbjct: 449 YNLHGVLVHSGDISTGHYYTLIKPGVEDQWYRFDDERVWRVTKKQVFQENFGCDRLPDEK 508
Query: 496 PQTNPGFNNTPFKFTKYSNAYMLVYIREADKDKVICNVDEKDIAEHLRERLKKEQEEKEH 555
+T + ++++AYMLVYIR+ ++ ++ V E D+ +H+ R+++E +E+E
Sbjct: 509 VRTMTRGEYQNYIIQRHTSAYMLVYIRQEQEEDLLRPVLESDVPKHVITRVREEIKERET 568
Query: 556 KKKEKAEAHLYTIIKVARNEDLKEQIGKDIYFDLVDHDKVRSFRVQ------KQMSFNLF 609
K+KE EAHLY +++ +KE I + FD HD R F + +Q++ +
Sbjct: 569 KEKEIREAHLYVTLRL---HSIKEFIHYE-GFDYFAHDGFRLFAEELNDSGLQQINLKVL 624
Query: 610 K--------EEVAKEFGIPVQFQ-RFWLWAKRQNHTYRPNRPLT------PAEEAQSVGQ 654
+ + + IP + ++W R+N T R +P+ +EA +
Sbjct: 625 RTTKLSDIFASIKETMNIPQERDVKYWKMDYRRNSTLRLTQPINFESVNITLQEALKKEK 684
Query: 655 VREVS---------------------------------------NKVHNAEL--KLFLEV 673
R + NK+ A L K L+
Sbjct: 685 KRTMQTQYGEEGVASTEEDDKALLETVSFLDLFIEEPYLELQFLNKLKEASLISKAQLDD 744
Query: 674 ELGPDLRPIAPSDKTKDDI-----------LLFFKLYDPEKEELRYVGRLFVKSTGKPSE 722
EL +R P + TK I LLF K YDP ++L G V + S+
Sbjct: 745 ELISTIRTNLP-ELTKGGIEPVFATDNKSNLLFVKSYDPHTQKLLGFGHFAVNQLQQLSD 803
Query: 723 ILTRLNEMAGYDPDEEIGLYEEIKFEPNVMCEPIDKKLTFRASQLEDGDIICFQKAPAMD 782
I + + +E++ YEE+ +P + E I K T + ++ GDI+ F+ A+
Sbjct: 804 ISAIIED--SISSNEKLTFYEEV--QPGTINE-IYMKETIYDADIDTGDIVSFEVPGAVL 858
Query: 783 SEEHVRYPDVPSYLEYVHNRQVVHFRSLDKPKE---------DDFCLEMSRLYTYDDVVE 833
+ Y + + Y+ R + F D E + F +S YDD+
Sbjct: 859 PDTFPVYATIKDFYSYLRYRVKLKFSKFDGSSEEYGVSNEIPESFEFWISAYAPYDDLAR 918
Query: 834 KVAQQLNLDDPSKIRLTPHNCYSQQPKPQPIKYRGVDHLSDMLVHYNQTSDI--LYYEIL 891
V++ ++ P +++ YS + L+D L+ I +E+L
Sbjct: 919 MVSKYAHV-KPEYLKIIA--LYSN----GRFVLKSTSLLNDYLLKDFNCDQIPPFAFEVL 971
Query: 892 DIPLPELQGLKTLKVAFYHATKDEVVSHTIRLPKQSTVGDVLDDLKTKVELSHPNAELRL 951
+PL EL+ L+ +K+ + + + T L+ ++ K+ + E L
Sbjct: 972 SVPLKELERLRPIKLYWLKNSYIHYQCFEFEVANDYTESQFLEKVQHKIGFTDEEKENIL 1031
Query: 952 LEVFYHKIYKVFPPNEKIETINDQYWTLRAEEVPEEEKNLGPHDRLIHVY-----HFTKD 1006
L + ++ ++ ++ L +PEE K +RL +V F D
Sbjct: 1032 LWTNTNFQFQGLLSDQNTFKDVSKHSLLFGRILPEESKLFKELNRLENVQTSSLEDFMDD 1091
Query: 1007 TSQNQMQIQNFGEPFFLVIREGETLTEIKVRIQKKLQ--------------VPDDEFEKW 1052
+ + + + + I E + V +Q+ + +PD+ F K
Sbjct: 1092 ENATDRPMDD-EQDLGMAIEHSEDMKGRIVVVQQYFKDLENRHGISFLFNLIPDETFPKT 1150
Query: 1053 K---FAFFALGRPEY 1064
K A F LG+ E+
Sbjct: 1151 KDRLHAKFGLGQKEF 1165
>UB40_HUMAN (Q9NVE5) Ubiquitin carboxyl-terminal hydrolase 40 (EC
3.1.2.15) (Ubiquitin thiolesterase 40)
(Ubiquitin-specific processing protease 40)
(Deubiquitinating enzyme 40) (Fragment)
Length = 1142
Score = 168 bits (425), Expect = 9e-41
Identities = 110/307 (35%), Positives = 156/307 (49%), Gaps = 20/307 (6%)
Query: 196 TGYVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTE------NDMPAGS---IPLAL 246
T G++NQG TCY+NSLLQTL+ P FR+A++ + E D P IPL L
Sbjct: 38 TNLSGIRNQGGTCYLNSLLQTLHFTPEFREALFSLGPEELGLFEDKDKPDAKVRIIPLQL 97
Query: 247 QSLFYKLQYSDTSVA-TKELTKSFGWDTYDSFMQHDVQELNRVLCEKLEDKMKGTVVEGT 305
Q LF +L D A T +LT SFGW + + QHDVQELNR+L LE + GT
Sbjct: 98 QRLFAQLLLLDQEAASTADLTDSFGWTSNEEMRQHDVQELNRILFSALETSLVGTSGHDL 157
Query: 306 IQKLFEGHHMNYIECINVDYKSTRKESFYDLQLDVK---GCHDVYASFDKYVEVEPLEGD 362
I +L+ G +N I C S R+E F DL + VK G D A ++ YVE E + D
Sbjct: 158 IYRLYHGTIVNQIVCKECKNVSERQEDFLDLTVAVKNVSGLED--ALWNMYVEEEVFDCD 215
Query: 363 NKYHAEQYG-LQDAKKGVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRYEFPLELDLDR 421
N YH L A K PP L + L RF +DF++ K Y FPL ++L
Sbjct: 216 NLYHCGTCDRLVKAAKSAKLRKLPPFLTVSLLRFNFDFVKCERYKETSCYTFPLRINLK- 274
Query: 422 DDGKYLSPDADRNVRNLYTLHSVLVHSGGVHGGHYYAFIRPTLSDQWYKFDDERVTKEDT 481
+ ++ +Y L SV++H GG +GGHY+ +I+ ++F +E+ +
Sbjct: 275 ---PFCEQSELDDLEYIYDLFSVIIHKGGCYGGHYHVYIKDVDHLGNWQFQEEKSKPDVN 331
Query: 482 KRALEEQ 488
+ L+ +
Sbjct: 332 LKDLQSE 338
>UB40_MOUSE (Q8BWR4) Ubiquitin carboxyl-terminal hydrolase 40 (EC
3.1.2.15) (Ubiquitin thiolesterase 40)
(Ubiquitin-specific processing protease 40)
(Deubiquitinating enzyme 40) (Fragment)
Length = 1140
Score = 161 bits (408), Expect = 8e-39
Identities = 103/278 (37%), Positives = 141/278 (50%), Gaps = 16/278 (5%)
Query: 196 TGYVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTE------NDMPAGS---IPLAL 246
T G++NQG TCY++SLLQTL+ P FR+A++ + E D P IPL L
Sbjct: 38 TNLSGIRNQGGTCYLSSLLQTLHFTPEFREALFSLGPEELGSLEDKDKPDAKVRIIPLQL 97
Query: 247 QSLFYKLQYSDTSVA-TKELTKSFGWDTYDSFMQHDVQELNRVLCEKLEDKMKGTVVEGT 305
Q LF +L D A T +LT SFGW + QHDVQELNR+L LE + GT
Sbjct: 98 QRLFAQLLLLDQEAASTIDLTDSFGWTNDEEMRQHDVQELNRILFSALETSLVGTSGHDL 157
Query: 306 IQKLFEGHHMNYIECINVDYKSTRKESFYDLQLDVKGCHDVYASF-DKYVEVEPLEGDNK 364
I +L+ G +N I C S R+E F DL + VK + + YVE E + DN
Sbjct: 158 IHRLYHGTIVNQIVCKECKNISERQEDFLDLTVAVKNVSGLEDELCNMYVEEEIFDYDNL 217
Query: 365 YHAEQYG-LQDAKKGVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRYEFPLELDLDRDD 423
YH L A K PP L + L RF +DF++ K Y FPL ++L
Sbjct: 218 YHCGTCDRLVKAAKSAKLRKLPPFLTISLLRFNFDFVKCERYKDTSCYTFPLRINLK--- 274
Query: 424 GKYLSPDADRNVRNLYTLHSVLVHSGGVHGGHYYAFIR 461
+ ++ +Y L SV++H GG +GGHY+ +I+
Sbjct: 275 -PFCEQSELDDMEYMYDLFSVIIHKGGCYGGHYHVYIK 311
>UBPE_DROME (Q24574) Ubiquitin carboxyl-terminal hydrolase 64E (EC
3.1.2.15) (Ubiquitin thiolesterase 64E)
(Ubiquitin-specific processing protease 64E)
(Deubiquitinating enzyme 64E)
Length = 898
Score = 160 bits (405), Expect = 2e-38
Identities = 96/233 (41%), Positives = 139/233 (59%), Gaps = 11/233 (4%)
Query: 197 GYVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDMPAGSIPLALQSLFYKLQYS 256
GYVGL NQ TCY+NSLLQ L+ P FR A+Y +ND A LQ LF LQ S
Sbjct: 53 GYVGLVNQAMTCYLNSLLQALFMTPEFRNALYRWEF-DNDNEAKK-SYQLQKLFLNLQTS 110
Query: 257 D-TSVATKELTKSFGWDTYDSFMQHDVQELNRVLCEKLEDKMKGTVVEGTIQKLFEGHHM 315
+V T +LT+SFGWD+ +++ QHD+QEL RV+ + LE K K T I L+EG
Sbjct: 111 PKAAVETTDLTRSFGWDSTEAWQQHDIQELCRVMFDALEHKFKNTKQANLISNLYEGKMN 170
Query: 316 NYIECINVD-YKSTRKESFYDLQLDVK--GCHDVYASFDK----YVEVEPLEGDNKYHAE 368
Y++C+ + K+ +++F D+ L V+ G Y S ++ +V+ E L+G+N+Y E
Sbjct: 171 AYVKCLGCNTEKNDCEDTFLDIPLPVRPFGSSSAYGSIEEALRAFVQPETLDGNNQYLCE 230
Query: 369 QYGLQ-DAKKGVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRYEFPLELDLD 420
+ + DA KG+ F FP +L L LKRF++D+ +K+NDR FP L+L+
Sbjct: 231 KCKKKCDAHKGLHFKSFPYILTLHLKRFDFDYQTMHRIKLNDRVTFPQTLNLN 283
Score = 77.4 bits (189), Expect = 2e-13
Identities = 54/195 (27%), Positives = 101/195 (51%), Gaps = 23/195 (11%)
Query: 420 DRDDGKYLSPDADRNVRN-----LYTLHSVLVHSGGVHGGHYYAFIRPTLSDQWYKFDDE 474
D D+G +S ++ + LY L ++++HSG GGHYYA+I+ +++W+ F+D+
Sbjct: 339 DEDEGIDMSSSTSKSAKQGSGPYLYELFAIMIHSGSASGGHYYAYIKDFDNNEWFCFNDQ 398
Query: 475 RVTKEDTKRALEEQYGGEEELPQTNPGFNNTPFKFTKYSNAYMLVYIR-EADKDKVICNV 533
VT T+ ++ +GG P + +T +NAYML+Y + +A +++++ V
Sbjct: 399 NVT-SITQEDIQRSFGG--------PNGSYYSSAYTSSTNAYMLMYRQVDAKRNELVAKV 449
Query: 534 DEKDIAEHLRERLKKEQEEKEHKKKEKAEAHLYTIIKVARNEDLKEQIGKDIYFDLVDHD 593
+D EH++ L K E+E + + H+ T+ +A + K + +YF
Sbjct: 450 --RDFPEHIKTLLPKLHSEEE-TRVSRLGRHI-TVTDLALPDLYKPR----VYFYNPSLK 501
Query: 594 KVRSFRVQKQMSFNL 608
K++ RV SFN+
Sbjct: 502 KMKITRVYVSQSFNI 516
>FAFX_HUMAN (Q93008) Probable ubiquitin carboxyl-terminal hydrolase
FAF-X (EC 3.1.2.15) (Ubiquitin thiolesterase FAF-X)
(Ubiquitin-specific processing protease FAF-X)
(Deubiquitinating enzyme FAF-X) (Fat facets protein
related, X-linked) (Ubiquitin-specif
Length = 2547
Score = 143 bits (360), Expect = 3e-33
Identities = 107/370 (28%), Positives = 166/370 (43%), Gaps = 74/370 (20%)
Query: 197 GYVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDMP-----------AGSIPLA 245
G+VGLKN GATCYMNS++Q LY IP R + + T +D+ ++
Sbjct: 1548 GFVGLKNAGATCYMNSVIQQLYMIPSIRNGILAIEGTGSDVDDDMSGDEKQDNESNVDPR 1607
Query: 246 LQSLFYKLQYSDTSVATK-------------ELTKSFG---------------WDTYDSF 277
Y Q+ D +K L FG W + +
Sbjct: 1608 DDVFGYPQQFEDKPALSKTEDRKEYNIGVLRHLQVIFGHLAASRLQYYVPRGFWKQFRLW 1667
Query: 278 -------MQHDVQELNRVLCEKLEDKMKGTVVEGTIQKLFEGHHMNYIECINVDYKSTRK 330
QHD E L + L++ +K + K+ G + C ++ +
Sbjct: 1668 GEPVNLREQHDALEFFNSLVDSLDEALKALGHPAMLSKVLGGSFADQKICQGCPHRYECE 1727
Query: 331 ESFYDLQLDVKGCHDVYASFDKYVEVEPLEGDNKYHAEQYGLQ-DAKKGVLFIDFPPVLQ 389
ESF L +D++ ++ S ++YV+ + LEG N YH E+ + D K +L PPVL
Sbjct: 1728 ESFTTLNVDIRNHQNLLDSLEQYVKGDLLEGANAYHCEKCNKKVDTVKRLLIKKLPPVLA 1787
Query: 390 LQLKRFEYDFMRDTMVKINDRYEFPLELDLD--------RDDGKYLSP-----------D 430
+QLKRF+YD+ R+ +K ND +EFP ELD++ + +G ++P +
Sbjct: 1788 IQLKRFDYDWERECAIKFNDYFEFPRELDMEPYTVAGVAKLEGDNVNPESQLIQQSEQSE 1847
Query: 431 ADRNVRNLYTLHSVLVHSGGVHGGHYYAFIRPTLS-----DQWYKFDDERVTK---EDTK 482
++ Y L VLVHSG GGHYY++I ++WYKFDD VT+ +D +
Sbjct: 1848 SETAGSTKYRLVGVLVHSGQASGGHYYSYIIQRNGGDGERNRWYKFDDGDVTECKMDDDE 1907
Query: 483 RALEEQYGGE 492
+ +GGE
Sbjct: 1908 EMKNQCFGGE 1917
>FAFY_HUMAN (O00507) Probable ubiquitin carboxyl-terminal hydrolase
FAF-Y (EC 3.1.2.15) (Ubiquitin thiolesterase FAF-Y)
(Ubiquitin-specific processing protease FAF-Y)
(Deubiquitinating enzyme FAF-Y) (Fat facets protein
related, Y-linked) (Ubiquitin-specif
Length = 2555
Score = 141 bits (356), Expect = 9e-33
Identities = 108/370 (29%), Positives = 164/370 (44%), Gaps = 74/370 (20%)
Query: 197 GYVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDM-----------PAGSIPLA 245
G+VGLKN GATCYMNS++Q LY IP R ++ + T +D+ ++
Sbjct: 1557 GFVGLKNAGATCYMNSVIQQLYMIPSIRNSILAIEGTGSDLHDDMFGDEKQDSESNVDPR 1616
Query: 246 LQSLFYKLQYSDTSVATK-------------ELTKSFG---------------WDTYDSF 277
Y Q+ D +K L FG W + +
Sbjct: 1617 DDVFGYPHQFEDKPALSKTEDRKEYNIGVLRHLQVIFGHLAASQLQYYVPRGFWKQFRLW 1676
Query: 278 -------MQHDVQELNRVLCEKLEDKMKGTVVEGTIQKLFEGHHMNYIECINVDYKSTRK 330
QHD E L + L++ +K + K+ G + C ++ +
Sbjct: 1677 GEPVNLREQHDALEFFNSLVDSLDEALKALGHPAILSKVLGGSFADQKICQGCPHRFECE 1736
Query: 331 ESFYDLQLDVKGCHDVYASFDKYVEVEPLEGDNKYHAEQYGLQ-DAKKGVLFIDFPPVLQ 389
ESF L +D++ ++ S ++Y++ + LEG N YH E+ + D K +L P VL
Sbjct: 1737 ESFTTLNVDIRNHQNLLDSLEQYIKGDLLEGANAYHCEKCDKKVDTVKRLLIKKLPRVLA 1796
Query: 390 LQLKRFEYDFMRDTMVKINDRYEFPLELD-----------LDRDD--------GKYLSPD 430
+QLKRF+YD+ R+ +K ND +EFP ELD L+RD+ + D
Sbjct: 1797 IQLKRFDYDWERECAIKFNDYFEFPRELDMGPYTVAGVANLERDNVNSENELIEQKEQSD 1856
Query: 431 ADRNVRNLYTLHSVLVHSGGVHGGHYYAFI-----RPTLSDQWYKFDDERVTK---EDTK 482
+ Y L VLVHSG GGHYY++I + +D WYKFDD VT+ +D +
Sbjct: 1857 NETAGGTKYRLVGVLVHSGQASGGHYYSYIIQRNGKDDQTDHWYKFDDGDVTECKMDDDE 1916
Query: 483 RALEEQYGGE 492
+ +GGE
Sbjct: 1917 EMKNQCFGGE 1926
>UB24_HUMAN (Q9UPU5) Ubiquitin carboxyl-terminal hydrolase 24 (EC
3.1.2.15) (Ubiquitin thiolesterase 24)
(Ubiquitin-specific processing protease 24)
(Deubiquitinating enzyme 24) (Fragment)
Length = 977
Score = 139 bits (349), Expect = 6e-32
Identities = 107/371 (28%), Positives = 169/371 (44%), Gaps = 33/371 (8%)
Query: 191 DSKKETGYVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDMPAGSIPLALQSLF 250
DS+ +G+VGL+N GATCYMN++ Q LY P +++ + + D P S+ +QSLF
Sbjct: 38 DSRSSSGFVGLRNGGATCYMNAVFQQLYMQPGLPESLLSVDD-DTDNPDDSVFYQVQSLF 96
Query: 251 YKLQYSDTSVATKE----LTKSFGWDTYDSFMQHDVQELNRVLCEKLEDKMKGTVVEGTI 306
L S E + K + + Y Q D E L +++++ +K +
Sbjct: 97 GHLMESKLQYYVPENFWKIFKMWNKELYVR-EQQDAYEFFTSLIDQMDEYLKKMGRDQIF 155
Query: 307 QKLFEGHHMNYIECINVDYKSTRKESFYDLQLDVKGCHDVYASFDKYVEVEPLEGDNKYH 366
+ F+G + + C + ++ R+E+F L L V C + S D++V E LEG N Y+
Sbjct: 156 KNTFQGIYSDQKICKDCPHRYEREEAFMALNLGVTSCQSLEISLDQFVRGEVLEGSNAYY 215
Query: 367 AEQ-YGLQDAKKGVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRYEFPLELDLDR---- 421
E+ + K P VL + L RF +D+ +K +++ FP L+++
Sbjct: 216 CEKCKEKRITVKRTCIKSLPSVLVIHLMRFGFDWESGRSIKYDEQIRFPWMLNMEPYTVS 275
Query: 422 ------------------DDGKYLSPDADRNVRNLYTLHSVLVHSGGVHGGHYYAFI--- 460
D G SP + Y L V+VHSG H GHYY+FI
Sbjct: 276 GMARQDSSSEVGENGRSVDQGGGGSPRKKVALTENYELVGVIVHSGQAHAGHYYSFIKDR 335
Query: 461 RPTLSDQWYKFDDERVTKED-TKRALEEQYGGEEELPQTNPGFNNTPFKFTKYSNAYMLV 519
R +WYKF+D + + D LE + G E P+ N +Y NAYML
Sbjct: 336 RGCGKGKWYKFNDTVIEEFDLNDETLEYECFGGEYRPKVYDQTNPYTDVRRRYWNAYMLF 395
Query: 520 YIREADKDKVI 530
Y R +D++ +
Sbjct: 396 YQRVSDQNSPV 406
>FAF_DROME (P55824) Probable ubiquitin carboxyl-terminal hydrolase FAF
(EC 3.1.2.15) (Ubiquitin thiolesterase FAF)
(Ubiquitin-specific processing protease FAF)
(Deubiquitinating enzyme FAF) (Fat facets protein)
Length = 2778
Score = 126 bits (316), Expect = 4e-28
Identities = 113/409 (27%), Positives = 177/409 (42%), Gaps = 74/409 (18%)
Query: 188 WTY----DSKKETGYVGLKNQGATCYMNSLLQTLYHIPYFRKAVY--HMPTT-------- 233
W Y ++ G+ GLKN GATCYMNS+LQ LY +P R + H T
Sbjct: 1653 WDYLPPVGARPTKGFCGLKNAGATCYMNSVLQQLYMVPAVRVGILRAHGAATTDGEDFSG 1712
Query: 234 ENDMPAGSIPLAL----QSLFYKLQYSDTSV--ATKELTKSF------------------ 269
++D+ G + AL S L S +++ ++ K++
Sbjct: 1713 DSDLTGGGLGSALFSGPASALVSLPSSSSTIEDGLHDVRKNYHVVILKHVQAIFAHLGHS 1772
Query: 270 --------GWDTYDSFM--------QHDVQELNRVLCEKLEDKMKGTVVEGTIQKLFEGH 313
G T+ + Q D E L E L++ +K + G
Sbjct: 1773 ALQYYVPRGLWTHFKLLGEPVNLREQQDAVEFFMSLLESLDEGLKALGQPQLMNATLGGS 1832
Query: 314 HMNYIECINVDYKSTRKESFYDLQLDVKGCHDVYASFDKYVEVEPLEGDNKYHAEQYGLQ 373
+ C ++ +++E F +D++ + S ++YV+ E LEG + YH ++ +
Sbjct: 1833 FSDQKICQECPHRYSKEEPFSVFSVDIRNHSSLTESLEQYVKGELLEGADAYHCDKCDKK 1892
Query: 374 DAK-KGVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRYEFPLELDLD--------RDDG 424
K V PPVL +QLKRFEYD+ R +K ND +EFP LD++ + +G
Sbjct: 1893 VVTVKRVCVKKLPPVLAIQLKRFEYDYERVCAIKFNDYFEFPRILDMEPYTVSGLAKLEG 1952
Query: 425 KY--LSPDADRNVRNL-YTLHSVLVHSGGVHGGHYYAFI---RPTLSD-QWYKFDDERVT 477
+ + + NV Y L ++VHSG GGHY+++I P QWYKFDD VT
Sbjct: 1953 EVVEVGDNCQTNVETTKYELTGIVVHSGQASGGHYFSYILSKNPANGKCQWYKFDDGEVT 2012
Query: 478 K---EDTKRALEEQYGGEEELPQTNPGFNNTPFKFTK-YSNAYMLVYIR 522
+ + + E +GGE + ++ K + NAYML Y R
Sbjct: 2013 ECKMHEDEEMKAECFGGEYMGETYDNNLKRMQYRRQKRWWNAYMLFYTR 2061
>FAFX_MOUSE (P70398) Probable ubiquitin carboxyl-terminal hydrolase
FAF-X (EC 3.1.2.15) (Ubiquitin thiolesterase FAF-X)
(Ubiquitin-specific processing protease FAF-X)
(Deubiquitinating enzyme FAF-X) (Fat facets protein
related, X-linked) (Ubiquitin-specif
Length = 2559
Score = 124 bits (310), Expect = 2e-27
Identities = 92/318 (28%), Positives = 151/318 (46%), Gaps = 32/318 (10%)
Query: 279 QHDVQELNRVLCEKLEDKMKGTVVEGTIQKLFEGHHMNYIECINVDYKSTRKESFYDLQL 338
QHD + L + L++ +K + K+ G + C ++ +ESF L +
Sbjct: 1683 QHDALKSFNSLVDSLDEALKALGHPAMLSKVLGGSFADQKICQGCPHRYECEESFTTLNV 1742
Query: 339 DVKGCHDVYASFDKYVEVEPLEGDNKYHAEQYGLQ-DAKKGVLFIDFPPVLQLQLKRFEY 397
D++ ++ S ++YV+ + LEG N YH E+ + D K +L PPVL +QLKRF+Y
Sbjct: 1743 DIRNHQNLLDSLEQYVKGDLLEGANAYHCEKCNKKVDTVKRLLIKKLPPVLAIQLKRFDY 1802
Query: 398 DFMRDTMVKINDRYEFPLELDLD--------RDDGKYLSPDADRNVRN-----------L 438
D+ R+ +K ND +EFP ELD++ + +G ++P++ +N
Sbjct: 1803 DWERECAIKFNDYFEFPRELDMEPYTVAGVAKLEGDNVNPESQLIQQNEQSESEKAGSTK 1862
Query: 439 YTLHSVLVHSGGVHGGHYYAFI-----RPTLSDQWYKFDDERVTK---EDTKRALEEQYG 490
Y L VLVHSG GGHYY++I ++WYKFDD VT+ +D + + +G
Sbjct: 1863 YRLVGVLVHSGQASGGHYYSYIIQRNGGDGEKNRWYKFDDGDVTECKMDDDEEMKTQCFG 1922
Query: 491 GEEELPQTNPGFNNTPFKFTK-YSNAYMLVYIRE---ADKDKVICNVDEKDIAEHLRERL 546
GE + ++ K + NAY+L Y R D+VI + E I + +
Sbjct: 1923 GEYMGEVLDHMMKRMSYRRQKRWWNAYILFYERMDTIGHDDEVIRYISEIAITTRPHQIV 1982
Query: 547 KKEQEEKEHKKKEKAEAH 564
E+ +K+ H
Sbjct: 1983 MPSAIERSVRKQNVQFMH 2000
Score = 50.1 bits (118), Expect = 3e-05
Identities = 22/41 (53%), Positives = 29/41 (70%)
Query: 197 GYVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDM 237
G+VGLKN GATCYMNS++Q LY IP R + + T +D+
Sbjct: 1555 GFVGLKNAGATCYMNSVIQQLYMIPSIRNGILAIEGTGSDV 1595
>UB12_HUMAN (O75317) Ubiquitin carboxyl-terminal hydrolase 12 (EC
3.1.2.15) (Ubiquitin thiolesterase 12)
(Ubiquitin-specific processing protease 12)
(Deubiquitinating enzyme 12) (Ubiquitin hydrolyzing
enzyme 1)
Length = 355
Score = 114 bits (284), Expect = 2e-24
Identities = 96/323 (29%), Positives = 140/323 (42%), Gaps = 41/323 (12%)
Query: 198 YVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDMPAGSIPLALQSLFYKLQYSD 257
Y GL N G TCY NS+LQ LY FR+ V + S+ L LF+ +
Sbjct: 23 YFGLVNFGNTCYCNSVLQALYFCRPFREKV--LAYKSQPRKKESLLTCLADLFHSIATQK 80
Query: 258 TSVATKELTKSFGW-----DTYDSFMQHDVQELNRVLC---------EKLEDKMKGTVVE 303
V K + +D++MQ D E L E+ ++K G +
Sbjct: 81 KKVGVIPPKKFITRLRKENELFDNYMQQDAHEFLNYLLNTIADILQEERKQEKQNGRLPN 140
Query: 304 GTIQ--------------KLFEGHHMNYIECINVDYKSTRKESFYDLQLDVKGCHDVYAS 349
G I ++F+G N C+ + S++ E F DL +DV+ +
Sbjct: 141 GNIDNENNNSTPDPTWVHEIFQGTLTNETRCLTCETISSKDEDFLDLSVDVEQNTSITHC 200
Query: 350 FDKYVEVEPLEGDNKYHAEQY-GLQDAKKGVLFIDFPPVLQLQLKRFEYDFMRDTMVKIN 408
+ E L + KY+ E+ Q+A K + P +L L LKRF+Y K++
Sbjct: 201 LRGFSNTETLCSEYKYYCEECRSKQEAHKRMKVKKLPMILALHLKRFKYMDQLHRYTKLS 260
Query: 409 DRYEFPLELDLDRDDGKYLSPDADRNVRNLYTLHSVLVHSG-GVHGGHYYAFIRPTLSDQ 467
R FPLEL L G +PD +Y L +V+VH G G + GHY A ++ D
Sbjct: 261 YRVVFPLELRLFNTSGDATNPD------RMYDLVAVVVHCGSGPNRGHYIAIVKS--HDF 312
Query: 468 WYKFDDERVTKEDTKRALEEQYG 490
W FDD+ V K D +A+EE YG
Sbjct: 313 WLLFDDDIVEKIDA-QAIEEFYG 334
>UB46_MOUSE (P62069) Ubiquitin carboxyl-terminal hydrolase 46 (EC
3.1.2.15) (Ubiquitin thiolesterase 46)
(Ubiquitin-specific processing protease 46)
(Deubiquitinating enzyme 46)
Length = 366
Score = 110 bits (275), Expect = 2e-23
Identities = 96/323 (29%), Positives = 140/323 (42%), Gaps = 41/323 (12%)
Query: 198 YVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDMPAGSIPLALQSLFYKLQYSD 257
Y GL N G TCY NS+LQ LY FR+ V + ++ L LF+ +
Sbjct: 34 YFGLVNFGNTCYCNSVLQALYFCRPFRENVLAYKAQQKKKE--NLLTCLADLFHSIATQK 91
Query: 258 TSVATKELTKSFGW-----DTYDSFMQHDVQELNRVLC---------EKLEDKMKGTVVE 303
V K D +D++MQ D E L EK ++K G +
Sbjct: 92 KKVGVIPPKKFISRLRKENDLFDNYMQQDAHEFLNYLLNTIADILQEEKKQEKQNGKLKN 151
Query: 304 GT--------------IQKLFEGHHMNYIECINVDYKSTRKESFYDLQLDVKGCHDVYAS 349
G + ++F+G N C+N + S++ E F DL +DV+ +
Sbjct: 152 GNMNEPAENNKPELTWVHEIFQGTLTNETRCLNCETVSSKDEDFLDLSVDVEQNTSITHC 211
Query: 350 FDKYVEVEPLEGDNKYHAEQ-YGLQDAKKGVLFIDFPPVLQLQLKRFEYDFMRDTMVKIN 408
+ E L + KY+ E Q+A+K + P +L L LKRF+Y K++
Sbjct: 212 LRDFSNTETLCSEQKYYCETCCSKQEAQKRMRVKKLPMILALHLKRFKYMEQLHRYTKLS 271
Query: 409 DRYEFPLELDLDRDDGKYLSPDADRNVRNLYTLHSVLVHSG-GVHGGHYYAFIRPTLSDQ 467
R FPLEL L S DA N+ +Y L +V+VH G G + GHY ++
Sbjct: 272 YRVVFPLELRLFN-----TSSDA-VNLDRMYDLVAVVVHCGSGPNRGHYITIVKS--HGF 323
Query: 468 WYKFDDERVTKEDTKRALEEQYG 490
W FDD+ V K D +A+EE YG
Sbjct: 324 WLLFDDDIVEKIDA-QAIEEFYG 345
>UB46_HUMAN (P62068) Ubiquitin carboxyl-terminal hydrolase 46 (EC
3.1.2.15) (Ubiquitin thiolesterase 46)
(Ubiquitin-specific processing protease 46)
(Deubiquitinating enzyme 46)
Length = 366
Score = 110 bits (275), Expect = 2e-23
Identities = 96/323 (29%), Positives = 140/323 (42%), Gaps = 41/323 (12%)
Query: 198 YVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDMPAGSIPLALQSLFYKLQYSD 257
Y GL N G TCY NS+LQ LY FR+ V + ++ L LF+ +
Sbjct: 34 YFGLVNFGNTCYCNSVLQALYFCRPFRENVLAYKAQQKKKE--NLLTCLADLFHSIATQK 91
Query: 258 TSVATKELTKSFGW-----DTYDSFMQHDVQELNRVLC---------EKLEDKMKGTVVE 303
V K D +D++MQ D E L EK ++K G +
Sbjct: 92 KKVGVIPPKKFISRLRKENDLFDNYMQQDAHEFLNYLLNTIADILQEEKKQEKQNGKLKN 151
Query: 304 GT--------------IQKLFEGHHMNYIECINVDYKSTRKESFYDLQLDVKGCHDVYAS 349
G + ++F+G N C+N + S++ E F DL +DV+ +
Sbjct: 152 GNMNEPAENNKPELTWVHEIFQGTLTNETRCLNCETVSSKDEDFLDLSVDVEQNTSITHC 211
Query: 350 FDKYVEVEPLEGDNKYHAEQ-YGLQDAKKGVLFIDFPPVLQLQLKRFEYDFMRDTMVKIN 408
+ E L + KY+ E Q+A+K + P +L L LKRF+Y K++
Sbjct: 212 LRDFSNTETLCSEQKYYCETCCSKQEAQKRMRVKKLPMILALHLKRFKYMEQLHRYTKLS 271
Query: 409 DRYEFPLELDLDRDDGKYLSPDADRNVRNLYTLHSVLVHSG-GVHGGHYYAFIRPTLSDQ 467
R FPLEL L S DA N+ +Y L +V+VH G G + GHY ++
Sbjct: 272 YRVVFPLELRLFN-----TSSDA-VNLDRMYDLVAVVVHCGSGPNRGHYITIVKS--HGF 323
Query: 468 WYKFDDERVTKEDTKRALEEQYG 490
W FDD+ V K D +A+EE YG
Sbjct: 324 WLLFDDDIVEKIDA-QAIEEFYG 345
>UBP8_YEAST (P50102) Ubiquitin carboxyl-terminal hydrolase 8 (EC
3.1.2.15) (Ubiquitin thiolesterase 8)
(Ubiquitin-specific processing protease 8)
(Deubiquitinating enzyme 8)
Length = 471
Score = 105 bits (262), Expect = 7e-22
Identities = 98/362 (27%), Positives = 159/362 (43%), Gaps = 51/362 (14%)
Query: 168 LLNDTLVVEA--EVLVRRIVDYWTYDSKKETGYVGLKNQGATCYMNSLLQTLYHIPYFRK 225
L+ND ++ + +V + +V ++ G GL N G+TC+M+S+LQ L H PYF +
Sbjct: 108 LINDAILAKYWDDVCTKTMVP----SMERRDGLSGLINMGSTCFMSSILQCLIHNPYFIR 163
Query: 226 ---AVYHMPTTENDMPAGSIPLALQSLFYKL-------QYSDTSVATKELTK-----SFG 270
+ H + P AL + ++L Q S +S +T T +
Sbjct: 164 HSMSQIHSNNCKVRSPDKCFSCALDKIVHELYGALNTKQASSSSTSTNRQTGFIYLLTCA 223
Query: 271 WDTYDS---FMQHDVQELNRVLCEKLED-------------KMKGTVVEGTIQKLFEGHH 314
W + + Q D E + + ++ + E + +FEG
Sbjct: 224 WKINQNLAGYSQQDAHEFWQFIINQIHQSYVLDLPNAKEVSRANNKQCECIVHTVFEGSL 283
Query: 315 MNYIECINVDYKS-TRKESFYDLQLDVKGCHDVYASFDKYVEVEPLEGDNKYHAEQYGLQ 373
+ I C S T + F DL LD+K +Y D + + E L+ N + E Q
Sbjct: 284 ESSIVCPGCQNNSKTTIDPFLDLSLDIKDKKKLYECLDSFHKKEQLKDFNYHCGECNSTQ 343
Query: 374 DAKKGVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRYEFPLELDLDRDDGKYLS-PDAD 432
DA K + P VL LQLKRFE+ + + K++D EFP L++ Y S + D
Sbjct: 344 DAIKQLGIHKLPSVLVLQLKRFEH-LLNGSNRKLDDFIEFPTYLNMK----NYCSTKEKD 398
Query: 433 RNVRN------LYTLHSVLVHSGGVHGGHYYAFIRPTLSDQWYKFDDERVTKEDTKRALE 486
++ N +Y L ++ H G V+ GHY AF + + QW+KF+D V+ + L+
Sbjct: 399 KHSENGKVPDIIYELIGIVSHKGTVNEGHYIAFCKIS-GGQWFKFNDSMVSSISQEEVLK 457
Query: 487 EQ 488
EQ
Sbjct: 458 EQ 459
>UBPW_MOUSE (Q61068) Ubiquitin carboxyl-terminal hydrolase DUB-1 (EC
3.1.2.15) (Ubiquitin thiolesterase DUB-1)
(Ubiquitin-specific processing protease DUB-1)
(Deubiquitinating enzyme 1)
Length = 526
Score = 93.6 bits (231), Expect = 3e-18
Identities = 106/408 (25%), Positives = 172/408 (41%), Gaps = 51/408 (12%)
Query: 200 GLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDM-PAGSIPLALQSLFYK-LQYSD 257
GL+N G +CY+N+ LQ L H P + ++ P G A+++L + L +S
Sbjct: 52 GLQNTGNSCYLNAALQCLTHTPPLADYMLSQEHSQTCCSPEGCKLCAMEALVTQSLLHSH 111
Query: 258 TSVATKE---LTKSFGWDTYDS---FMQHDVQELNRVLCEKLEDKMKGTVVEGT-IQKLF 310
+ K LT +F + F+ ++ ++ C ++ + K T + + I +F
Sbjct: 112 SGDVMKPSHILTSAFHKHQQEDAHEFLMFTLETMHES-CLQVHRQSKPTSEDSSPIHDIF 170
Query: 311 EGHHMNYIECINVDYKSTRKESFYDLQLDVKGCHDVYASFDKYVEVEPLEGDNKYHAEQY 370
G + I+C+ S + F D+ LD+ V + + E L GDN Y+ +
Sbjct: 171 GGWWRSQIKCLLCQGTSDTYDRFLDIPLDISSAQSVKQALWDTEKSEELCGDNAYYCGKC 230
Query: 371 GLQDAKKGVLFIDFPP-VLQLQLKRFEYDFMRDTMVKINDRYEFPLELDLDRDDGKYLSP 429
+ L + P VL + L RF T K++ + +P LDL YLS
Sbjct: 231 RQKMPASKTLHVHIAPKVLMVVLNRFS----AFTGNKLDRKVSYPEFLDLK----PYLSE 282
Query: 430 DADRNVRNLYTLHSVLVHSGGV-HGGHYYAFIRPTLSDQWYKFDDERVTKEDTKRALEEQ 488
+ Y L++VLVH G H GHY+ ++ +WYK DD +VT+ D L E
Sbjct: 283 PTGGPLP--YALYAVLVHDGATSHSGHYFCCVKAG-HGKWYKMDDTKVTRCDVTSVLNE- 338
Query: 489 YGGEEELPQTNPGFNNTPFKFTKYSNAYMLVYIREADKDKVICNVDEKDIAEHL-RERLK 547
NAY+L Y+++A+ +V ++ E I E L E
Sbjct: 339 -------------------------NAYVLFYVQQANLKQVSIDMPEGRINEVLDPEYQL 373
Query: 548 KEQEEKEHKKKEKAEAHLYTIIKVARNEDLKE-QIGKDIYFDLVDHDK 594
K+ K+HKKK L + +KE +GK V+H K
Sbjct: 374 KKSRRKKHKKKSPFTEDLGEPCENRDKRAIKETSLGKGKVLQEVNHKK 421
>UBP9_SCHPO (Q9P7V9) Probable ubiquitin carboxyl-terminal hydrolase
9 (EC 3.1.2.15) (Ubiquitin thiolesterase 9)
(Ubiquitin-specific processing protease 9)
(Deubiquitinating enzyme 9)
Length = 585
Score = 92.8 bits (229), Expect = 5e-18
Identities = 72/251 (28%), Positives = 119/251 (46%), Gaps = 29/251 (11%)
Query: 306 IQKLFEGHHMNYIECINVDYKSTRKESFYDLQLDVKGCHDVYASFDKYVEVEPLEGDNKY 365
+ LFEG + +C+ + ++R ESF DL +D++ V + + E L NK+
Sbjct: 230 VHSLFEGTLTSETKCLTCENITSRDESFLDLSIDIENHTSVTSCLRSFSASEMLSSKNKF 289
Query: 366 HAEQ-YGLQDAKKGVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRYEFPLELDLDRDDG 424
H + LQ+A+K + P +L L LKRF+Y+ ++ K+ F E+ L
Sbjct: 290 HCDVCKSLQEAEKRMKIKKLPKILSLHLKRFKYNETQEGHDKLFYTIVFTNEMRL----- 344
Query: 425 KYLSPDADRNVRNLYTLHSVLVH-SGGVHGGHYYAFIRPTLSDQWYKFDDERVTKEDTKR 483
+ + + N +Y L SV+VH GG H GHY + +R T + W FDDE VT + +
Sbjct: 345 -FTTTEDAENAERMYYLSSVIVHVGGGPHRGHYVSIVR-TKTYGWVLFDDENVTPVN-EN 401
Query: 484 ALEEQYGGEEELPQTNPGFNNTPFKFTKYSNAYMLVYIREADKDKVICNVDEKDIAE--H 541
L+ +G + PG + AY+L Y ++D + VD K+ +
Sbjct: 402 YLQRFFGDQ-------PG----------QATAYVLFYTAADEEDDDVSEVDTKESIKPMS 444
Query: 542 LRERLKKEQEE 552
+ +LK+E E
Sbjct: 445 IPSQLKQESVE 455
Score = 40.4 bits (93), Expect = 0.027
Identities = 23/71 (32%), Positives = 37/71 (51%), Gaps = 7/71 (9%)
Query: 198 YVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDMPAGSIPLALQSLFYKLQYSD 257
+ GL N G TCY++S+L +LYH+ FR ++ N P S P +S+ K + +
Sbjct: 40 FYGLTNYGNTCYVSSVLVSLYHLKPFRDSL-------NSYPLPSAPPNFKSVCTKTNHPE 92
Query: 258 TSVATKELTKS 268
+S + KS
Sbjct: 93 SSSSRHSKKKS 103
>UBP9_YEAST (P39967) Ubiquitin carboxyl-terminal hydrolase 9 (EC
3.1.2.15) (Ubiquitin thiolesterase 9)
(Ubiquitin-specific processing protease 9)
(Deubiquitinating enzyme 9)
Length = 754
Score = 85.9 bits (211), Expect = 6e-16
Identities = 71/227 (31%), Positives = 106/227 (46%), Gaps = 32/227 (14%)
Query: 306 IQKLFEGHHMNYIECINVDYKSTRKESFYDLQLDVKGCH--DVYASFDKYVEVEPLEGDN 363
I LF+G N I+C+ D ++R E F D ++V+G D+ Y + E L G N
Sbjct: 473 ITDLFKGTLTNRIKCLTCDNITSRDEPFLDFPIEVQGDEETDIQKMLKSYHQREMLNGVN 532
Query: 364 KYHAEQ-YGLQDAKKGVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRYEFPLELDLDRD 422
K++ + YGLQ+A++ V P +L L LKRF+Y + + +K+ ++ +PL LD
Sbjct: 533 KFYCNKCYGLQEAERMVGLKQLPHILSLHLKRFKYSEEQKSNIKLFNKILYPLTLD---- 588
Query: 423 DGKYLSPDADRNVRNLYTLHSVLVHSG-GVHGGHYYAFIRPTLSDQWYKFDDERVTKEDT 481
+S + +V Y L V++H G G GHY R W +DDE T E
Sbjct: 589 ----VSSTFNTSVYKKYELSGVVIHMGSGPQHGHYVCICR-NEKFGWLLYDDE--TVESI 641
Query: 482 KRALEEQYGGEEELPQTNPGFNNTPFKFTKYSNAYMLVYIREADKDK 528
K Q+ G +PG T AY+L Y +E DK
Sbjct: 642 KEETVLQFTG-------HPGDQTT---------AYVLFY-KETQADK 671
Score = 42.4 bits (98), Expect = 0.007
Identities = 30/73 (41%), Positives = 36/73 (49%), Gaps = 6/73 (8%)
Query: 161 YDPSRGYLLNDTLVVEAEVLVRRIVDYWTYD--SKKETGYVGLKNQGATCYMNSLLQTLY 218
YD S Y + E+ +L I D Y S K GY +N G TCY NS+LQ LY
Sbjct: 98 YDSSTTYE-DRAFDTESSILFTTITDLMPYGDGSNKVFGY---ENFGNTCYCNSVLQCLY 153
Query: 219 HIPYFRKAVYHMP 231
+IP FR V P
Sbjct: 154 NIPEFRCNVLRYP 166
>UBP8_SCHPO (Q09738) Probable ubiquitin carboxyl-terminal hydrolase
8 (EC 3.1.2.15) (Ubiquitin thiolesterase 8)
(Ubiquitin-specific processing protease 8)
(Deubiquitinating enzyme 8)
Length = 449
Score = 82.4 bits (202), Expect = 6e-15
Identities = 80/312 (25%), Positives = 135/312 (42%), Gaps = 48/312 (15%)
Query: 197 GYVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDMPAGS-IPLALQSLFYKLQ- 254
G G++N GATC+M+ +LQ++ H P R + T D + + A+ +F +
Sbjct: 143 GLRGIQNLGATCFMSVILQSILHNPLVRNLFFSGFHTSTDCKRPTCMTCAIDDMFSSIYN 202
Query: 255 -------YSDTSVATK--ELTKSFGWDTYDSFMQHDVQELNRVLCEKLE-DKMKGTVVEG 304
Y T+V +L+KS + Q D E L +++ + GT +
Sbjct: 203 SKNKSTFYGPTAVLNLMWKLSKSL-----CGYSQQDGHEFFVYLLDQMHTESGGGTSMPC 257
Query: 305 T--IQKLFEGHHMNYIECINVDYKSTRKESFYDLQLDV-----KGCHDVYASFDKYVEVE 357
T I ++F G N + C++ + + D+ LD+ +GC + + S +K
Sbjct: 258 TCPIHRIFSGSLKNVVTCLDCKKERVAVDPLMDISLDINEPTLQGCLERFVSKEKV---- 313
Query: 358 PLEGDNKYHAEQYGLQDAKKGVLFIDFPPVLQLQLKRFEY-DFMRDTMVKINDRYEFPLE 416
+Y G ++A K ++F PP + +QLKRFE +F T KI+ + +P
Sbjct: 314 ------QYSCHSCGSKNAIKQLVFDKLPPTICMQLKRFEQNNFAMST--KIDKQVSYPAF 365
Query: 417 LDLDRDDGKYLSPDADRNVRNLYTLHSVLVHSGGVHGGHYYAFIRPTLSDQWYKFDDERV 476
L R + D D Y L+SV+ H G + GHY A+ +QW+ DD +
Sbjct: 366 L---RMRYNFNQDDVD------YQLYSVVCHKGTLDTGHYIAY--TYYQNQWFLLDDTTI 414
Query: 477 TKEDTKRALEEQ 488
+ L Q
Sbjct: 415 VEVKESEVLNSQ 426
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 141,688,384
Number of Sequences: 164201
Number of extensions: 6679396
Number of successful extensions: 21893
Number of sequences better than 10.0: 145
Number of HSP's better than 10.0 without gapping: 86
Number of HSP's successfully gapped in prelim test: 59
Number of HSP's that attempted gapping in prelim test: 21214
Number of HSP's gapped (non-prelim): 418
length of query: 1118
length of database: 59,974,054
effective HSP length: 121
effective length of query: 997
effective length of database: 40,105,733
effective search space: 39985415801
effective search space used: 39985415801
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 71 (32.0 bits)
Lotus: description of TM0546.13


