

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0544.8
(1753 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
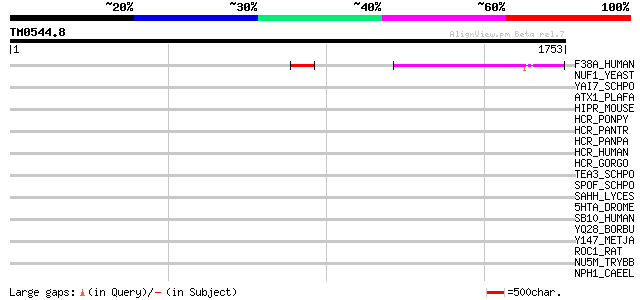
Score E
Sequences producing significant alignments: (bits) Value
F38A_HUMAN (Q92508) Protein FAM38A 216 3e-55
NUF1_YEAST (P32380) NUF1 protein (Spindle poly body spacer prote... 38 0.28
YAI7_SCHPO (Q09894) Hypothetical protein C24B11.07c in chromosome I 37 0.63
ATX1_PLAFA (Q04956) Probable cation-transporting ATPase 1 (EC 3.... 37 0.63
HIPR_MOUSE (Q9JKY5) Huntingtin interacting protein 1 related (Hi... 36 0.82
HCR_PONPY (Q8HZ58) Alpha helical coiled-coil rod protein 36 0.82
HCR_PANTR (Q8HZ60) Alpha helical coiled-coil rod protein 36 0.82
HCR_PANPA (Q8HZ57) Alpha helical coiled-coil rod protein 36 0.82
HCR_HUMAN (Q8TD31) Alpha helical coiled-coil rod protein (Putati... 36 0.82
HCR_GORGO (Q8HZ59) Alpha helical coiled-coil rod protein 36 0.82
TEA3_SCHPO (O14248) Tip elongation aberrant protein 3 (Cell pola... 35 1.4
SPOF_SCHPO (Q10411) Sporulation-specific protein 15 35 1.8
SAHH_LYCES (Q9SWF5) Adenosylhomocysteinase (EC 3.3.1.1) (S-adeno... 35 1.8
5HTA_DROME (P28285) 5-hydroxytryptamine receptor 2A (5-HT recept... 35 1.8
SB10_HUMAN (P48595) Bomapin (Protease inhibitor 10) (Serpin B10) 34 3.1
YQ28_BORBU (O50999) Hypothetical ANK-repeat protein BBB28 34 4.1
Y147_METJA (Q57611) Hypothetical ATP-binding protein MJ0147 34 4.1
ROC1_RAT (Q63644) Rho-associated protein kinase 1 (EC 2.7.1.37) ... 34 4.1
NU5M_TRYBB (P04540) NADH-ubiquinone oxidoreductase chain 5 (EC 1... 34 4.1
NPH1_CAEEL (O17972) Nephrocystin-1 like protein 34 4.1
>F38A_HUMAN (Q92508) Protein FAM38A
Length = 2035
Score = 216 bits (551), Expect = 3e-55
Identities = 155/571 (27%), Positives = 282/571 (49%), Gaps = 41/571 (7%)
Query: 1211 DLYAYIFGADLAVFFLIAIFYESIMKANSEFLEVYQL-EDQFPEDFVLVLMVVFFLIVLD 1269
D+YA +F AD+ F +I + + K ++ L +DQ PE F+++L++ F +V+D
Sbjct: 1473 DVYALMFLADVVDFIIIIFGFWAFGKHSAATDITSSLSDDQVPEAFLVMLLIQFSTMVVD 1532
Query: 1270 RIIYLCSFATAKVIFYLFSLVLFTYSVTKYAWDMDPLRRYSGRLALRAIYFTKAISLVLQ 1329
R +YL K+ F + +LVL + + R ++ + + YF K I L
Sbjct: 1533 RALYLRKTVLGKLAFQV-ALVLAIHLWMFFILPAVTERMFNQNVVAQLWYFVKCIYFALS 1591
Query: 1330 AIQFHFGMPHKSILYRQFLTSSVSRINVMGFRLYRAIPFLYELRCVLDWSCTTTSLTMYD 1389
A Q G P + + FLT + +N+ F+ +R +PFL ELR V+DW T T+L++
Sbjct: 1592 AYQIRCGYPTR--ILGNFLTKKYNHLNLFLFQGFRLVPFLVELRAVMDWVWTDTTLSLSS 1649
Query: 1390 WLKLEDIHASLFLVKCDVDLNRASHQ-QGQKQTKMTKFCNGICLFFVLMCVIWAPMLMYS 1448
W+ +EDI+A++F++KC + + Q +GQK+ K+ K+ G + L+ +IW P+L S
Sbjct: 1650 WMCVEDIYANIFIIKCSRETEKKYPQPKGQKKKKIVKYGMGGLIILFLIAIIWFPLLFMS 1709
Query: 1449 S-GNPTNIANPIKDASARVHIKTLSGRLTLF--ETTLCEIISWETLEARTSLDPKG---- 1501
+ + N D + + + T+ + ++ + E DP+
Sbjct: 1710 LVRSVVGVVNQPIDVTVTLKLGGYEPLFTMSAQQPSIIPFTAQAYEELSRQFDPQPLAMQ 1769
Query: 1502 YLSTYNEKDIQLICCQSDASTLWLVPPIVQARFMKSL---SWNMDLAFSWEFTRDRPRGK 1558
++S Y+ +DI + + LW + P +A+ + L + ++ L F+W F RD +G
Sbjct: 1770 FISQYSPEDIVTAQIEGSSGALWRISPPSRAQMKRELYNGTADITLRFTWNFQRDLAKGG 1829
Query: 1559 EVV----KYELTIQEQDLPKSSEVTEVFNGTSNSFSAF-NIYPRYFRVTGSGDVRSLEQ- 1612
V K+ L + + ++ + GTS+ N++P+Y R + ++Q
Sbjct: 1830 TVEYANEKHMLALAPNSTARR-QLASLLEGTSDQSVVIPNLFPKYIRAPNGPEANPVKQL 1888
Query: 1613 ----SVELVSGDLVLNR-------GNPEWWSFYDLDISDAH-GCGKFPGPMAIIVSEETP 1660
+ + + L R G EWW +++ + C P +I S++
Sbjct: 1889 QPNEEADYLGVRIQLRREQGAGATGFLEWWV---IELQECRTDCNLLP---MVIFSDKVS 1942
Query: 1661 QGIIGDTLSKFSIWGLYITFVLAVGRFIRLQCSDLRMRIPFENLPSCDRLMALCEDIYAA 1720
+G L+ + I GLY++ VL +G+F+R S++ I FE LP DR++ LC+DI+
Sbjct: 1943 PPSLG-FLAGYGIMGLYVSIVLVIGKFVRGFFSEISHSIMFEELPCVDRILKLCQDIFLV 2001
Query: 1721 RAEGELEVEEVLYWTLVKIYRTPHMLLEYTQ 1751
R ELE+EE LY L+ +YR+P ++++T+
Sbjct: 2002 RETRELELEEELYAKLIFLYRSPETMIKWTR 2032
Score = 50.8 bits (120), Expect = 3e-05
Identities = 22/74 (29%), Positives = 47/74 (62%)
Query: 888 LVRHIWSRMRSNNEVVCYCCFILIYLWNFSLLSVVYLAALFLYALCQNTGPSYIFWVLIL 947
L+R ++ + +++E++CY IL ++ S S+V +FL+A+ PS FW+ +
Sbjct: 1189 LLRAVYQCVAAHSELLCYFIIILNHMVTASAGSLVLPVLVFLWAMLSIPRPSKRFWMTAI 1248
Query: 948 IYTEVCILLQYLYQ 961
++TE+ ++++YL+Q
Sbjct: 1249 VFTEIAVVVKYLFQ 1262
Score = 33.5 bits (75), Expect = 5.3
Identities = 54/261 (20%), Positives = 99/261 (37%), Gaps = 35/261 (13%)
Query: 328 LKFWIENMFTLFGLEINMIALLLASFAVLNAISLLYIASLAACVLL-HRLLIKKLWPIFV 386
LK++I F FGLEI + + +N + L+ L A + HR I +LWP +
Sbjct: 497 LKYFINFFFYKFGLEICFLMAVNVIGQRMNFLVTLHGCWLVAILTRRHRQAIARLWPNYC 556
Query: 387 FLFASVVTVEYLAI--------------WMRLTFMNLEIEEHVPCRDCWRVSDIYFSYCR 432
A + +YL W R MN + + + D +R +
Sbjct: 557 LFLALFLLYQYLLCLGMPPALCIDYPWRWSRAVPMNSALIKWLYLPDFFRAPNS------ 610
Query: 433 KCWLGIIVDDPRMLIGYYGVFMFSCFKFRADKSSSLTGLEMYQKIHSQWKSASVLSNLSF 492
+I D +L +FS R ++ + G+ + + + V + +
Sbjct: 611 ---TNLISDFLLLLCASQQWQVFSA--ERTEEWQRMAGVNTDRLEPLRGEPNPVPNFIHC 665
Query: 493 ETKGYWTFLDHLRLYGYCHLLDFVLSLILITGTLEYDFLHLGYLGFALVFFRMRLKILKQ 552
++LD L++ + +L VL ++ +TG LGYL +L++
Sbjct: 666 R-----SYLDMLKVAVFRYLFWLVLVVVFVTGATRISIFGLGYLLACFYLLLFGTALLQR 720
Query: 553 GNN----IFRYLRMYNFAVIV 569
++ L +YN VI+
Sbjct: 721 DTRARLVLWDCLILYNVTVII 741
>NUF1_YEAST (P32380) NUF1 protein (Spindle poly body spacer protein
SPC110)
Length = 944
Score = 37.7 bits (86), Expect = 0.28
Identities = 29/120 (24%), Positives = 60/120 (49%), Gaps = 13/120 (10%)
Query: 636 DYVSTYLEKEQIGAILRQQEKKAAWKTAQLRHIRKAEELKHLRSLQVEKMKSEMLNLQNQ 695
D+ E+EQ+ L + E+K KT + + + E ++SL++ + E+ NL N+
Sbjct: 221 DHSGCIEEREQMERKLAELERKL--KTVK-DQVLELENNSDVQSLKLRSKEDELKNLMNE 277
Query: 696 LHSMSTDANYSNASLKNDGLRERRNSFLENEFRKQALDTNTESIGTLDVNQSLLSEKSES 755
L+ + ++A + L+ F +NE RK+ + N I + +++ L +++ES
Sbjct: 278 LNELKSNAEEKDTQLE----------FKKNELRKRTNELNELKIKSDEMDLQLKQKQNES 327
>YAI7_SCHPO (Q09894) Hypothetical protein C24B11.07c in chromosome I
Length = 561
Score = 36.6 bits (83), Expect = 0.63
Identities = 20/58 (34%), Positives = 26/58 (44%)
Query: 814 SQAQSLGNMAVNNLVNYLKIEREGQEPTDDSSEDEEYYEIENQNSGAEPLEPTFSTNS 871
S +QSLGN ++ + TDDS E E Y+ + N N G EP S S
Sbjct: 481 SSSQSLGNEVLSRPNSNSNSAESRNGETDDSGESETYFTLMNNNKGVEPRASVASETS 538
>ATX1_PLAFA (Q04956) Probable cation-transporting ATPase 1 (EC
3.6.3.-)
Length = 1956
Score = 36.6 bits (83), Expect = 0.63
Identities = 50/222 (22%), Positives = 95/222 (42%), Gaps = 8/222 (3%)
Query: 694 NQLH-SMSTDANYSNASLKNDGLRERRNSFLENEFRKQALDTNT--ESIGTLDVNQSLLS 750
N LH + ST ++Y S D E N + K+ +D N+ E +G N+ L+
Sbjct: 610 NDLHINDSTCSSYLLNSETKDAYCEYYNIDHLCDINKKNMDINSKNELMGKYSKNE-LMG 668
Query: 751 EKSESPLVPEYMKHPMGSPHGIVEVKERTKNNDFLDLEIRNRYKLPGKRNALVSAVHFIG 810
+ ++ L+ +Y K+ + + E+ + N+ + +N +N L+ I
Sbjct: 669 KTIKNELMGKYSKNELMGKYSKNELMGKYSKNELMGKYSKNELMGKYSKNELMGKT--IK 726
Query: 811 SGISQAQSLGNMAVNNLVNYLKIEREGQEPTDDSSEDEEYYEIENQNSGAEPLEPTFSTN 870
+ + ++ +M +N NY +D+ EY+ I NS P E S N
Sbjct: 727 NQVGVDTNIYHMNCDNDYNYDYPCDYNCNNCNDTYHRLEYHNINKDNSFNIPPEKNKSYN 786
Query: 871 SINEHTGSDTSCLQIGILVRHIWSRMRSNNEVVCYCCFILIY 912
+I+EH + L + H S++ NN+++ IL++
Sbjct: 787 NISEHIKINYPLLFEALACCHTLSKV--NNKIMGDVLEILMF 826
>HIPR_MOUSE (Q9JKY5) Huntingtin interacting protein 1 related
(Hip1-related)
Length = 1068
Score = 36.2 bits (82), Expect = 0.82
Identities = 52/214 (24%), Positives = 87/214 (40%), Gaps = 22/214 (10%)
Query: 652 RQQEKKAAWKTAQLRHIRKAEELKHLRSLQVEKMKSEMLNLQNQLHSMSTDANYSNASLK 711
R+Q++KA QLRH EL L++LQ+E +++ L + + + +T+A YS
Sbjct: 399 RKQKQKALVDNEQLRH-----ELAQLKALQLEGARNQGLREEAERKASATEARYSK---- 449
Query: 712 NDGLRERRNSFLENEFRKQALDTNTESIGTLDVNQSLLSEKSESPLVPEYMKHPMGSPHG 771
L+E+ + + + L N ++ L V Q + E V E + M
Sbjct: 450 ---LKEKHSELINT--HAELLRKNADTAKQLTVTQ---QSQEEVARVKEQLAFQMEQAKR 501
Query: 772 IVEVK--ERTKNNDFLDLEIRNRYKLPGKRNALVSAVHFIGSGISQAQSLGNMAVNNLVN 829
E+K E++ + L E+ R + +S GS +S N L
Sbjct: 502 ESEMKMEEQSDQLEKLKRELAARAGELARAQEALSRTEQSGSELSSRLDTLNAEKEALSG 561
Query: 830 YLKIEREGQEPTDDS--SEDEEYYEIENQNSGAE 861
++ +RE + S E EE E Q S E
Sbjct: 562 VVR-QREAELLAAQSLVREKEEALSQEQQRSSQE 594
>HCR_PONPY (Q8HZ58) Alpha helical coiled-coil rod protein
Length = 782
Score = 36.2 bits (82), Expect = 0.82
Identities = 34/111 (30%), Positives = 52/111 (46%), Gaps = 16/111 (14%)
Query: 652 RQQEKKAAWKTAQL--RHIRKAEELKHLRSLQVEKMKSEMLNLQNQLHSMSTDANYSNAS 709
R++ KA Q+ R ++ E + LR LQ E K E L +L + D N A+
Sbjct: 627 RREHAKAVVSLRQIQRRAAQEKERSQELRRLQEEARKEEGQRLARRLQELERDKNLMLAT 686
Query: 710 LKNDGLRERRNSFLENEFRKQALDTNTESIGTLDVNQSLLSEKSESPLVPE 760
L+ +GL R +++Q L T S+ LD +S++S SP PE
Sbjct: 687 LQQEGLLSR--------YKQQRLLTVLPSL--LDKKKSVVS----SPRPPE 723
>HCR_PANTR (Q8HZ60) Alpha helical coiled-coil rod protein
Length = 782
Score = 36.2 bits (82), Expect = 0.82
Identities = 34/111 (30%), Positives = 52/111 (46%), Gaps = 16/111 (14%)
Query: 652 RQQEKKAAWKTAQL--RHIRKAEELKHLRSLQVEKMKSEMLNLQNQLHSMSTDANYSNAS 709
R++ KA Q+ R ++ E + LR LQ E K E L +L + D N A+
Sbjct: 627 RREHAKAVVSLRQIQRRAAQEKERSQELRRLQEEARKEEGQRLARRLQELERDKNLMLAT 686
Query: 710 LKNDGLRERRNSFLENEFRKQALDTNTESIGTLDVNQSLLSEKSESPLVPE 760
L+ +GL R +++Q L T S+ LD +S++S SP PE
Sbjct: 687 LQQEGLLSR--------YKQQRLLTVLPSL--LDKKKSVVS----SPRPPE 723
>HCR_PANPA (Q8HZ57) Alpha helical coiled-coil rod protein
Length = 782
Score = 36.2 bits (82), Expect = 0.82
Identities = 34/111 (30%), Positives = 52/111 (46%), Gaps = 16/111 (14%)
Query: 652 RQQEKKAAWKTAQL--RHIRKAEELKHLRSLQVEKMKSEMLNLQNQLHSMSTDANYSNAS 709
R++ KA Q+ R ++ E + LR LQ E K E L +L + D N A+
Sbjct: 627 RREHAKAVVSLRQIQRRAAQEKERSQELRRLQEEARKEEGQRLARRLQELERDKNLMLAT 686
Query: 710 LKNDGLRERRNSFLENEFRKQALDTNTESIGTLDVNQSLLSEKSESPLVPE 760
L+ +GL R +++Q L T S+ LD +S++S SP PE
Sbjct: 687 LQQEGLLSR--------YKQQRLLTVLPSL--LDKKKSVVS----SPRPPE 723
>HCR_HUMAN (Q8TD31) Alpha helical coiled-coil rod protein (Putative
gene 8 protein) (Pg8)
Length = 782
Score = 36.2 bits (82), Expect = 0.82
Identities = 34/111 (30%), Positives = 52/111 (46%), Gaps = 16/111 (14%)
Query: 652 RQQEKKAAWKTAQL--RHIRKAEELKHLRSLQVEKMKSEMLNLQNQLHSMSTDANYSNAS 709
R++ KA Q+ R ++ E + LR LQ E K E L +L + D N A+
Sbjct: 627 RREHAKAVVSLRQIQRRAAQEKERSQELRRLQEEARKEEGQRLARRLQELERDKNLMLAT 686
Query: 710 LKNDGLRERRNSFLENEFRKQALDTNTESIGTLDVNQSLLSEKSESPLVPE 760
L+ +GL R +++Q L T S+ LD +S++S SP PE
Sbjct: 687 LQQEGLLSR--------YKQQRLLTVLPSL--LDKKKSVVS----SPRPPE 723
>HCR_GORGO (Q8HZ59) Alpha helical coiled-coil rod protein
Length = 782
Score = 36.2 bits (82), Expect = 0.82
Identities = 34/111 (30%), Positives = 52/111 (46%), Gaps = 16/111 (14%)
Query: 652 RQQEKKAAWKTAQL--RHIRKAEELKHLRSLQVEKMKSEMLNLQNQLHSMSTDANYSNAS 709
R++ KA Q+ R ++ E + LR LQ E K E L +L + D N A+
Sbjct: 627 RREHAKAVVSLRQIQRRAAQEKERSQELRRLQEEARKEEGQRLARRLQELERDKNLMLAT 686
Query: 710 LKNDGLRERRNSFLENEFRKQALDTNTESIGTLDVNQSLLSEKSESPLVPE 760
L+ +GL R +++Q L T S+ LD +S++S SP PE
Sbjct: 687 LQQEGLLSR--------YKQQRLLTVLPSL--LDKKKSVVS----SPRPPE 723
>TEA3_SCHPO (O14248) Tip elongation aberrant protein 3 (Cell
polarity protein tea3)
Length = 1125
Score = 35.4 bits (80), Expect = 1.4
Identities = 55/243 (22%), Positives = 110/243 (44%), Gaps = 26/243 (10%)
Query: 518 SLILITGTLEYDFLHLGYLGFALVFFRMRLKI---LKQGNNIFRYLRMYNFAVIVLSLAY 574
SLI G LEY+ G L A +++ ++ LKQ + L+ N + V
Sbjct: 650 SLIEKVGKLEYEVQ--GTLEEATSYYQKNTELQQLLKQNESASELLKSRNEKLCVDYDKL 707
Query: 575 QSPFLGDFSEI----------KSGFIEYINELVGFQK----YDYGFRITSRSALVEIIIY 620
+S F D S+I +S ++ ELV ++ +YG + L+E+
Sbjct: 708 RSVFEEDSSKILSLQKENENLQSQILQISEELVDYRSRCEALEYG-NYELETKLIEMHDR 766
Query: 621 MLVSLQSYMFSFPEFDYVSTYLEKEQIGAILRQQEKKAAWKTAQLRHIRKAEELKHLRSL 680
+ + S D +T + + ++ R++++K+ T Q + + E +++R +
Sbjct: 767 VEMQTNVIEASASALDVSNTAILSFE-DSLRRERDEKS---TLQQKCLNLQYEYENVR-I 821
Query: 681 QVEKMKSEMLNLQNQLHSMSTDANYSNASLKNDGLRERRNSFLENEFRKQALDTNTESIG 740
++E ++S L L++ L +DA YS A +++ GL + +S EN+ + + + I
Sbjct: 822 ELENLQSRALELESALEQSVSDAKYSKAIMQS-GLSKLLSSINENKDNLKEFSKSKQKIS 880
Query: 741 TLD 743
L+
Sbjct: 881 YLE 883
>SPOF_SCHPO (Q10411) Sporulation-specific protein 15
Length = 1957
Score = 35.0 bits (79), Expect = 1.8
Identities = 36/156 (23%), Positives = 70/156 (44%), Gaps = 17/156 (10%)
Query: 642 LEKEQIGAILRQQEKKAAWKTAQL--RHIRKAEELKHLRSLQVEKMKSEMLNLQNQLHSM 699
LE+ I +Q++ + K Q +RK+EE L+ + ++ + NL+ + ++
Sbjct: 670 LEESNKSLIKKQEDVDSLEKNIQTLKEDLRKSEEALRFSKLEAKNLREVIDNLKGKHETL 729
Query: 700 STDANYSNASLKNDGLRERRNSFLENEFRKQALDTN--TESIGTLDVNQSLLSEKSESPL 757
N ++SL + + N+ L +E K + D T ++ TL S ++S + L
Sbjct: 730 EAQRNDLHSSLSD---AKNTNAILSSELTKSSEDVKRLTANVETL-TQDSKAMKQSFTSL 785
Query: 758 VPEY-----MKHPMGSPHGIVEVKERTKNNDFLDLE 788
V Y + H + H V +++NN L+ E
Sbjct: 786 VNSYQSISNLYHELRDDH----VNMQSQNNTLLESE 817
>SAHH_LYCES (Q9SWF5) Adenosylhomocysteinase (EC 3.3.1.1)
(S-adenosyl-L-homocysteine hydrolase) (AdoHcyase)
Length = 485
Score = 35.0 bits (79), Expect = 1.8
Identities = 26/96 (27%), Positives = 44/96 (45%), Gaps = 15/96 (15%)
Query: 1598 YFRVTGSGDVRSLEQSVELVSGDLVLNRGNPEWWSFYDLDISDAHGCGKFPGPMAIIVSE 1657
+ TG+ D+ ++ ++ + +V N G+ +D +I D HG FPG I +
Sbjct: 321 FVTTTGNKDIIMVDHMRKMKNNAIVCNIGH------FDNEI-DMHGLETFPGVKRITIKP 373
Query: 1658 ETPQGIIGDTLSKFSIWGLYITFVLAVGRFIRLQCS 1693
+T + + DT S VLA GR + L C+
Sbjct: 374 QTDRWVFPDTNSGI--------IVLAEGRLMNLGCA 401
>5HTA_DROME (P28285) 5-hydroxytryptamine receptor 2A (5-HT receptor)
(Serotonin receptor 2A)
Length = 834
Score = 35.0 bits (79), Expect = 1.8
Identities = 23/92 (25%), Positives = 39/92 (42%), Gaps = 3/92 (3%)
Query: 800 NALVSAVHFIGSGISQAQSLG--NMAVNNLVNYLKIEREGQEPTDDSSEDEEYYEIENQN 857
N + + H + Q +S + AVN + + E +GQ P ED E E +++
Sbjct: 517 NQIATVSHLVALAKQQGKSTAKSSAAVNGMAPSGRQEDDGQRPEHGEQEDREELEDQDEQ 576
Query: 858 SGAEPLEPTFSTNSINEHTGSDTSCLQIGILV 889
G +P T +T + + D C G+ V
Sbjct: 577 VGPQPTTATSATTAAGTNESED-QCKANGVEV 607
>SB10_HUMAN (P48595) Bomapin (Protease inhibitor 10) (Serpin B10)
Length = 397
Score = 34.3 bits (77), Expect = 3.1
Identities = 25/111 (22%), Positives = 46/111 (40%), Gaps = 6/111 (5%)
Query: 655 EKKAAWKTAQLRHIRKAEELKHLRSLQVEKMKSEMLNLQNQLHSMSTDANYSNASLKNDG 714
EK W +A + + + + HL ++E + +L++ L SM +S + G
Sbjct: 271 EKLNEWTSADMMELYEVQ--LHLPKFKLE----DSYDLKSTLSSMGMSDAFSQSKADFSG 324
Query: 715 LRERRNSFLENEFRKQALDTNTESIGTLDVNQSLLSEKSESPLVPEYMKHP 765
+ RN FL N F K ++ N + + S + + P + HP
Sbjct: 325 MSSARNLFLSNVFHKAFVEINEQGTEAAAGSGSEIDIRIRVPSIEFNANHP 375
>YQ28_BORBU (O50999) Hypothetical ANK-repeat protein BBB28
Length = 414
Score = 33.9 bits (76), Expect = 4.1
Identities = 32/108 (29%), Positives = 51/108 (46%), Gaps = 12/108 (11%)
Query: 515 FVLSLILITGTLEYDFLHLGYLGFALVFFRMRLKILKQGNNIF--RYLRMYNFAVIVLSL 572
+VLS I + +L +G+L F + +R KI + N++ YL + +I +L
Sbjct: 4 YVLSKIFLYS----GYLVIGFLSFTIFNKNLRNKIRDKLKNLYFLYYLTFFTLFIISSNL 59
Query: 573 AY---QSPFLGDFSEIKSGFIEY--INELVGFQKYDYGFRITSRSALV 615
+Y + L DF+ + F E INE FQKY F + R L+
Sbjct: 60 SYYFTEKQLLEDFNIFEKEFFEIHKINEQF-FQKYLLNFPVPIRMELM 106
>Y147_METJA (Q57611) Hypothetical ATP-binding protein MJ0147
Length = 371
Score = 33.9 bits (76), Expect = 4.1
Identities = 46/278 (16%), Positives = 115/278 (40%), Gaps = 18/278 (6%)
Query: 591 EYINELVGFQKYDYGFRITSRSALVEIIIYMLVSLQSYMFSFPEFDYVSTYLEK-----E 645
E+I + +K D+ +I +S ++ +I L PE ++ + EK +
Sbjct: 72 EFIEAIFTTKKDDFFEKIKDKSEVLNLITKGAKILTG--IPIPEVEFDKLFEEKINDAFQ 129
Query: 646 QIGAILRQQEKKAAWKTAQLRHIRKAEELK-HLRSLQVEKMKSEMLNLQNQLH-----SM 699
+ +IL + +K L ++ +++ + + ++++ +++L + H +
Sbjct: 130 YLNSILLEVKKSGKQPVLILDELQMIKDVVLNWQKYLLKELFQFLVSLTKEQHLCHVFCL 189
Query: 700 STDANYSNASLKNDGLRERRNSFLENEFRKQALDTNTESIGTLDVNQSLLSEKSESPLVP 759
S+D+ + L R L ++F K+ + + +N+ L E E L+
Sbjct: 190 SSDSLFIEYVYSAGELEGRAKYLLVDDFDKETALKFMDFLAKEILNKKLSDEDKE--LIY 247
Query: 760 EYMKHPMGSPHGIVEVKERTKNNDFLDLEIRNRYKLPGKRNALVSAVHFIGSGISQAQSL 819
Y+ + +++ K D L+L ++ + K + + +I + +
Sbjct: 248 NYVGGKPVYIYSVIDEMRYRKLEDILNLMLKEETQ---KLKYFLKELDYIKPKVELKDEI 304
Query: 820 GNMAVNNLVNYLKIEREGQEPTDDSSEDEEYYEIENQN 857
+ ++++N LK+ +E E +DD + Y + +N
Sbjct: 305 IEIKKDDIINALKLFKENYEVSDDDIPEPVYIYLVKKN 342
>ROC1_RAT (Q63644) Rho-associated protein kinase 1 (EC 2.7.1.37)
(Rho-associated, coiled-coil containing protein kinase
1) (p160 ROCK-1) (p160ROCK) (p150 RhoA-binding kinase
ROK beta)
Length = 1369
Score = 33.9 bits (76), Expect = 4.1
Identities = 55/223 (24%), Positives = 95/223 (41%), Gaps = 41/223 (18%)
Query: 653 QQEKKAAWKTAQL-RHIRKAEELKHLRS---LQVEKMKSEMLNLQNQLHSMSTDANYSNA 708
Q + A K QL + + +A +L S +++ K +EM +QL S++ + N
Sbjct: 531 QSSQLANEKLTQLQKQLEEANDLLRTESDTAVRLRKSHTEMSKSVSQLESLNRELQERNR 590
Query: 709 SLKNDGLRERRNSF-----LENEFRKQALDTNTESIGTLDVNQSLLSEKSESPLVPEYMK 763
L+N + ++ + LE E R + D +E IG L + L E+ +++K
Sbjct: 591 MLENSKSQADKDYYQLQAVLEAERRDRGHD--SEMIGDLQARITSLQEE------VKHLK 642
Query: 764 HPMGSPHG-------IVEVKERTKNNDFLDLEIRNRYKLPGKRNALVSAV--HFI----- 809
H + G ++ E+ KNN LEI YKL + L V H +
Sbjct: 643 HNLERVEGERKEAQDMLNHSEKEKNN----LEIDLNYKLKSIQQRLEQEVNEHKVTKARL 698
Query: 810 ---GSGISQAQSLGNMAVNNLVNYLKIEREGQEPTDDSSEDEE 849
I +A+S +A+ + LK ERE +E ++ + E
Sbjct: 699 TDKHQSIEEAKS---VAMCEMEKKLKEEREAREKAENRVVETE 738
>NU5M_TRYBB (P04540) NADH-ubiquinone oxidoreductase chain 5 (EC
1.6.5.3)
Length = 590
Score = 33.9 bits (76), Expect = 4.1
Identities = 25/90 (27%), Positives = 42/90 (45%), Gaps = 9/90 (10%)
Query: 321 KTALKVSLKFWIENMFTLFGLEIN--------MIALLLASFAVLNAISLLYIASL-AACV 371
K L + L WI F L L ++A LL FA+LN+ S+ ++ +
Sbjct: 336 KATLFIVLGIWIHIFFGLQDLRCYFFMYFCGCVLARLLLIFAILNSCSIWFLCGFYCKDM 395
Query: 372 LLHRLLIKKLWPIFVFLFASVVTVEYLAIW 401
LL L++ + I FLF S++ + + I+
Sbjct: 396 LLALLMLLSFYNIIEFLFISIIFIFFTMIY 425
>NPH1_CAEEL (O17972) Nephrocystin-1 like protein
Length = 682
Score = 33.9 bits (76), Expect = 4.1
Identities = 41/166 (24%), Positives = 69/166 (40%), Gaps = 22/166 (13%)
Query: 622 LVSLQSYMFSFPEFDYVSTYLEKEQ--IGAILRQQEKKAAWKTAQLRHIR------KAEE 673
++SLQ + FP+F+Y LEKEQ A R +K + Q H + + E+
Sbjct: 7 ILSLQDAINRFPQFEYQINRLEKEQKDTEAASRASFRKHFVRQCQELHRQLDDHRNRIEK 66
Query: 674 LKHLRSLQVEKMKSEMLNLQNQLHSMSTDANYSNASLKNDGLRE----------RRNSFL 723
K + + E ++ L+ +L ++S + + S+ D E RR S +
Sbjct: 67 AKTDETYKKENALEQLDKLKQRLTALSPEKEQLSFSVSVDSQSEEEKPKMAAIGRRKSTM 126
Query: 724 ENEFRKQALDTNTESIGTLDVNQSLLSEKSESPLVPEYMKHPMGSP 769
N+ + D ++E I T DV L + S P +H P
Sbjct: 127 YNDDESEDSDNDSEIIET-DVQ---LDDPLPSQPQPPQQQHQQPQP 168
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.325 0.139 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 197,265,436
Number of Sequences: 164201
Number of extensions: 8280007
Number of successful extensions: 26549
Number of sequences better than 10.0: 35
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 27
Number of HSP's that attempted gapping in prelim test: 26496
Number of HSP's gapped (non-prelim): 75
length of query: 1753
length of database: 59,974,054
effective HSP length: 124
effective length of query: 1629
effective length of database: 39,613,130
effective search space: 64529788770
effective search space used: 64529788770
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0544.8


