

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0541.15
(322 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
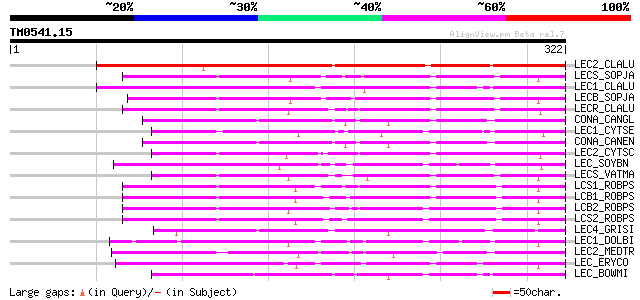
Score E
Sequences producing significant alignments: (bits) Value
LEC2_CLALU (Q39529) Agglutinin II precursor (ClAII) (LecClAII) 213 5e-55
LECS_SOPJA (P93535) Seed lectin precursor (LECSJASG) 183 4e-46
LEC1_CLALU (Q39528) Agglutinin I precursor (ClAI) (LecClAI) 183 6e-46
LECB_SOPJA (P93538) Bark lectin precursor (LECSJABG) (Fragment) 181 2e-45
LECR_CLALU (Q39527) Lectin-related protein precursor (CLLRP) (LR... 175 1e-43
CONA_CANGL (P14894) Concanavalin A precursor (Con A) 171 2e-42
LEC1_CYTSE (P22970) Anti-H(O) lectin I (CSA-I) 169 9e-42
CONA_CANEN (P02866) Concanavalin A precursor (Con A) 168 2e-41
LEC2_CYTSC (P29257) 2-acetamido-2-deoxy-D-galactose-binding seed... 166 6e-41
LEC_SOYBN (P05046) Lectin precursor (Agglutinin) (SBA) 166 9e-41
LECS_VATMA (P81371) Seed lectin (VML) 165 2e-40
LCS1_ROBPS (Q41162) Seed agglutinin I precursor (RPSAI) (LECRPAS1) 162 1e-39
LCB1_ROBPS (Q41159) Bark agglutinin I, polypeptide A precursor (... 160 3e-39
LCB2_ROBPS (Q42372) Bark agglutinin I, polypeptide B precursor (... 160 5e-39
LCS2_ROBPS (Q41161) Seed agglutinin II precursor (RPSAII) (LECRP... 159 1e-38
LEC4_GRISI (P24146) Lectin IV (GS4) 153 6e-37
LEC1_DOLBI (P05045) Seed lectin subunit I precursor (SL) [Contai... 150 4e-36
LEC2_MEDTR (Q01807) Truncated lectin 2 precursor 150 5e-36
LEC_ERYCO (P16404) Lectin precursor (ECorL) 149 7e-36
LEC_BOWMI (P42088) Lectin (Agglutinin) (BMA) 149 9e-36
>LEC2_CLALU (Q39529) Agglutinin II precursor (ClAII) (LecClAII)
Length = 290
Score = 213 bits (542), Expect = 5e-55
Identities = 120/276 (43%), Positives = 170/276 (61%), Gaps = 9/276 (3%)
Query: 51 SQLNYQPTMANITSELLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTN 110
S N T I+ LLA + L+L+ SDSLSF+F F PD R ++L GDA ++
Sbjct: 4 SNTNLLQTKKPISLPLLAFITLFLMLLNRVNSSDSLSFTFDNFRPDQRDLILQGDAKISS 63
Query: 111 G--TLQLTKIDQYGNPSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNHSQPH-GD 167
G +LQLTK D G P S G + +Y +HL D T R+A F+T FTFV++ + GD
Sbjct: 64 GGDSLQLTKTDTSGKPVRGSVGRALYYTPLHLWDSSTNRLASFQTTFTFVLSSPTNNPGD 123
Query: 168 GFTFFLASLDFEFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSF-SNSFDPNFPHIG 226
G FF+A + P G GG LGLF + A N S N IVAVEFD+F +N++DP+ HIG
Sbjct: 124 GIAFFIAPPETTIPPGSSGG-LLGLFSPDNALNNSLNQIVAVEFDTFVNNNWDPSHRHIG 182
Query: 227 IDINTIRSAVTIPWSVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPI 286
ID+NTI+S+ T+ W + G++ A+ISY+S ++ L+VV YP Q + + Y +
Sbjct: 183 IDVNTIKSSATVRWQ---RENGSLATAQISYNSDTKKLSVVSSYPNTQANEDY-TVSYDV 238
Query: 287 DIESYLSEWVVIGFSATTGEFAETHSILSWSFNSTL 322
D+++ L EWV +GFS +TG + + H+ILSW+FNS L
Sbjct: 239 DLKTELPEWVRVGFSGSTGGYVQNHNILSWTFNSNL 274
>LECS_SOPJA (P93535) Seed lectin precursor (LECSJASG)
Length = 292
Score = 183 bits (465), Expect = 4e-46
Identities = 112/263 (42%), Positives = 157/263 (59%), Gaps = 17/263 (6%)
Query: 66 LLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTN-GTLQLTKIDQYGNP 124
L ++ F LLL+ ++ LSFSFP F + +LL GDA ++ G LQLT ++ G P
Sbjct: 21 LAISITFFLLLLNKVNSAEILSFSFPKFASNQEDLLLQGDALVSSKGELQLTTVEN-GVP 79
Query: 125 SPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNH--SQPHGDGFTFFLASLDFEFPE 182
+S+G + +Y VH+ DK TGRVA F T F+FVV + DG FFLA + +
Sbjct: 80 IWNSTGRALYYAPVHIWDKSTGRVASFATSFSFVVKAPVASKSADGIAFFLAPPNNQIQ- 138
Query: 183 GGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNSFDPNFPHIGIDINTIRSAVTIPWSV 242
G GGG LGLF + YN+S I+AV+FD+ N++DPN HIGID+N+I S T+ W
Sbjct: 139 -GPGGGHLGLF-HSSGYNSSYQ-IIAVDFDTHINAWDPNTRHIGIDVNSINSTKTVTWGW 195
Query: 243 NDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSA 302
+ G + N ISY + + LTV + YP Q + + A +D++S L EWV +GF+A
Sbjct: 196 QN---GEVANVLISYQAATETLTVSLTYPSSQTSYILSA---AVDLKSILPEWVRVGFTA 249
Query: 303 TTG---EFAETHSILSWSFNSTL 322
TG ++ ETH +LSWSF STL
Sbjct: 250 ATGLTTQYVETHDVLSWSFTSTL 272
>LEC1_CLALU (Q39528) Agglutinin I precursor (ClAI) (LecClAI)
Length = 293
Score = 183 bits (464), Expect = 6e-46
Identities = 114/284 (40%), Positives = 164/284 (57%), Gaps = 22/284 (7%)
Query: 51 SQLNYQPTMANITSELLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDAN-TT 109
S N+ T ++ LLA + L+L+ SDSLSF+F F P+ ++ DA+ ++
Sbjct: 4 SNTNFLETKKPLSLPLLAFITIYLMLLHRVNSSDSLSFTFNNFPPNSEDLIFQKDASISS 63
Query: 110 NGTLQLTKIDQYGNPSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTF-VVNHSQPHGDG 168
N TL+LT+I G P+ S G + +Y V L DK TGR+A F+T F+F + + +Q GDG
Sbjct: 64 NETLELTRISSSGQPATSSVGRALYYTPVRLWDKSTGRLASFKTTFSFAITSPTQDPGDG 123
Query: 169 FTFFLASLDFEFPEGGGGGGFLGLFDEETAYNTSKN---------SIVAVEFDSFSN-SF 218
F FF+A D G GGG LGLF+ N+S N IVAVEFD++ N
Sbjct: 124 FAFFIAPPD---TTPGYGGGLLGLFNGFNLRNSSNNGVAVNNQSAQIVAVEFDTYINGQC 180
Query: 219 DPNFPHIGIDINTIRSAVTIPWSVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTN 278
DP + H+GID+N+I S W + G A+ISY+ S+ LT V YP N+T
Sbjct: 181 DPKYRHVGIDVNSITSLAYTQWQWQN---GVKATAQISYNPASQKLTAVTSYP---NSTP 234
Query: 279 IDAIKYPIDIESYLSEWVVIGFSATTGEFAETHSILSWSFNSTL 322
+ + ID+++ L EWV +GFSA+TG+ E +SIL+WSF+S+L
Sbjct: 235 L-TVSLDIDLQTVLPEWVRVGFSASTGQNVERNSILAWSFSSSL 277
>LECB_SOPJA (P93538) Bark lectin precursor (LECSJABG) (Fragment)
Length = 270
Score = 181 bits (459), Expect = 2e-45
Identities = 108/260 (41%), Positives = 157/260 (59%), Gaps = 17/260 (6%)
Query: 69 TLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTN-GTLQLTKIDQYGNPSPH 127
++ F LLL+ ++ LSFSFP F + +LL GDA ++ G LQLT ++ G P +
Sbjct: 2 SITFFLLLLNKVNSAEILSFSFPKFVSNQEDLLLQGDALVSSEGELQLTTVEN-GVPVWN 60
Query: 128 SSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNH--SQPHGDGFTFFLASLDFEFPEGGG 185
S+G + +Y VH+ D TGRVA F T F+FVV + DG FFLA L+ + G
Sbjct: 61 STGRALYYAPVHIWDNSTGRVASFATSFSFVVKAPVASKSADGIAFFLAPLNNQIH--GA 118
Query: 186 GGGFLGLFDEETAYNTSKNSIVAVEFDSFSNSFDPNFPHIGIDINTIRSAVTIPWSVNDQ 245
GGG GLF+ + +S IVAVEFD+ +N++DPN HIGID+N+++S T+ W +
Sbjct: 119 GGGLYGLFNSSSY--SSSYQIVAVEFDTHTNAWDPNTRHIGIDVNSVKSTKTVTWGWEN- 175
Query: 246 PQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSATTG 305
G + N I+Y + + LTV + YP Q + + A +D++S L EWV +GF+ATTG
Sbjct: 176 --GEVANVLITYQAATEMLTVSLTYPSNQTSYILSA---AVDLKSILPEWVRVGFTATTG 230
Query: 306 ---EFAETHSILSWSFNSTL 322
++ ET+ +LSWSF STL
Sbjct: 231 LTTQYVETNDVLSWSFTSTL 250
>LECR_CLALU (Q39527) Lectin-related protein precursor (CLLRP)
(LRPCL) (Fragment)
Length = 290
Score = 175 bits (444), Expect = 1e-43
Identities = 111/263 (42%), Positives = 151/263 (57%), Gaps = 18/263 (6%)
Query: 66 LLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTN-GTLQLTKIDQYGNP 124
L ++ F LLL+ ++LSF+F F + +LL GDA ++ G LQLT+++ G P
Sbjct: 20 LAISITFYLLLLNKVNSEEALSFTFTKFVSNQDELLLQGDALVSSKGELQLTRVEN-GQP 78
Query: 125 SPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVN--HSQPHGDGFTFFLASLDFEFPE 182
PHS G + + VH+ D TG VA F T FTFVV + DG FFLA D +
Sbjct: 79 IPHSVGRALYSDPVHIWDSSTGSVASFVTSFTFVVEAPNENKTADGIAFFLAPPDTQVQS 138
Query: 183 GGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNSFDPNFPHIGIDINTIRSAVTIPWSV 242
GG FLGLF+ + YN+S N I+AVEFD+FSNS+DP HIGID+N+I S T W
Sbjct: 139 LGG---FLGLFNS-SVYNSS-NQILAVEFDTFSNSWDPTARHIGIDVNSIESTRTATWGW 193
Query: 243 NDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSA 302
+ G + I+Y + + L + YP Q + + A +D++S L EWV +GFSA
Sbjct: 194 RN---GEVAIVLITYVAPAETLIASLTYPSSQTSYILSA---AVDLKSILPEWVRVGFSA 247
Query: 303 TTGE---FAETHSILSWSFNSTL 322
TG + ETH +LSWSF STL
Sbjct: 248 ATGRSAGYVETHDVLSWSFTSTL 270
>CONA_CANGL (P14894) Concanavalin A precursor (Con A)
Length = 290
Score = 171 bits (434), Expect = 2e-42
Identities = 102/261 (39%), Positives = 152/261 (58%), Gaps = 27/261 (10%)
Query: 78 TTTVKSDSLSFSFPTFDPDVRVILLNGDANT-TNGTLQLTKIDQYGNPSPHSSGISAFYG 136
++T ++++L F F F D + ++L GDA T T+G L+LT++ G+P S G + FY
Sbjct: 29 SSTHETNALHFMFNQFSKDQKDLILQGDATTGTDGNLELTRVSSNGSPQGSSVGRALFYA 88
Query: 137 AVHLSDKKTGRVADFRTEFTFVVNHSQPH-GDGFTFFLASLDFEFPEGGGGGGFLGLF-- 193
VH+ + + VA F FTF++ H DG FF++++D P G G LGLF
Sbjct: 89 PVHIWES-SAVVASFDATFTFLIKSPDSHPADGIAFFISNIDSSIPSGSTGR-LLGLFPD 146
Query: 194 ----------DEETAYNTSKNSIVAVEFDSFSNSF--DPNFPHIGIDINTIRSAVTIPWS 241
D AYN ++IVAVE D++ N+ DPN+PHIGIDI ++RS T W+
Sbjct: 147 ANVIRNSTTIDFNAAYNA--DTIVAVELDTYPNTDIGDPNYPHIGIDIKSVRSKKTAKWN 204
Query: 242 VNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFS 301
+ + G + A I Y+SV + L+ VV YP G + T + Y +D+++ L EWV +G S
Sbjct: 205 MQN---GKVGTAHIIYNSVGKRLSAVVSYPNGDSAT----VSYDVDLDNVLPEWVRVGLS 257
Query: 302 ATTGEFAETHSILSWSFNSTL 322
A+TG + ET++ILSWSF S L
Sbjct: 258 ASTGLYKETNTILSWSFTSKL 278
>LEC1_CYTSE (P22970) Anti-H(O) lectin I (CSA-I)
Length = 244
Score = 169 bits (428), Expect = 9e-42
Identities = 104/250 (41%), Positives = 147/250 (58%), Gaps = 22/250 (8%)
Query: 83 SDSLSFSFPTFDPDVRVILLNGDAN-TTNGTLQLTKIDQYGNPSPHSSGISAFYGAVHLS 141
+D LSF+F F P+ IL G+A+ +T G LQ+TK+ + P+ S G + + VH+
Sbjct: 2 NDHLSFNFDKFVPNQNNILFQGEASVSTTGVLQVTKVSK---PATRSIGRALYAAPVHIW 58
Query: 142 DKKTGRVADFRTEFTFVVNHSQPHG---DGFTFFLASLDFEFPEGGGGGGFLGLFDEETA 198
D TGRVA F T F+FVV DG TFFLA + + P G G F GLF+ ++
Sbjct: 59 DSTTGRVASFETSFSFVVKDEPEKSNGVDGLTFFLAPANSQIPSGSSAGLF-GLFN--SS 115
Query: 199 YNTSKNSIVAVEFDSF----SNSFDPNFPHIGIDINTIRSAVTIPWSVNDQPQGAIQNAK 254
N S N I+AVEFD++ N +DP+F HIG+D+N+I+S T+ W D G + N
Sbjct: 116 DNKSSNQIIAVEFDTYFGKTYNPWDPDFKHIGVDVNSIKSIKTVKW---DWRNGEVANVV 172
Query: 255 ISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSATTGEFA--ETHS 312
I+Y + ++ LTV + YP Q T+NI + +D+++ L EWV +GFSA G A ETH
Sbjct: 173 ITYRAPTKSLTVSLSYPSDQ-TSNI--VTASVDLKAILPEWVSVGFSAGVGNAAEFETHD 229
Query: 313 ILSWSFNSTL 322
+LSW F S L
Sbjct: 230 VLSWYFTSNL 239
>CONA_CANEN (P02866) Concanavalin A precursor (Con A)
Length = 290
Score = 168 bits (425), Expect = 2e-41
Identities = 100/261 (38%), Positives = 151/261 (57%), Gaps = 27/261 (10%)
Query: 78 TTTVKSDSLSFSFPTFDPDVRVILLNGDANT-TNGTLQLTKIDQYGNPSPHSSGISAFYG 136
++T ++++L F F F D + ++L GDA T T+G L+LT++ G+P S G + FY
Sbjct: 29 SSTHETNALHFMFNQFSKDQKDLILQGDATTGTDGNLELTRVSSNGSPQGSSVGRALFYA 88
Query: 137 AVHLSDKKTGRVADFRTEFTFVVNHSQPH-GDGFTFFLASLDFEFPEGGGGGGFLGLF-- 193
VH+ + + VA F FTF++ H DG FF++++D P G G LGLF
Sbjct: 89 PVHIWES-SAVVASFEATFTFLIKSPDSHPADGIAFFISNIDSSIPSGSTGR-LLGLFPD 146
Query: 194 ----------DEETAYNTSKNSIVAVEFDSFSNSF--DPNFPHIGIDINTIRSAVTIPWS 241
D AYN ++IVAVE D++ N+ DP++PHIGIDI ++RS T W+
Sbjct: 147 ANVIRNSTTIDFNAAYNA--DTIVAVELDTYPNTDIGDPSYPHIGIDIKSVRSKKTAKWN 204
Query: 242 VNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFS 301
+ + G + A I Y+SV + L+ VV YP + T + Y +D+++ L EWV +G S
Sbjct: 205 MQN---GKVGTAHIIYNSVDKRLSAVVSYPNADSAT----VSYDVDLDNVLPEWVRVGLS 257
Query: 302 ATTGEFAETHSILSWSFNSTL 322
A+TG + ET++ILSWSF S L
Sbjct: 258 ASTGLYKETNTILSWSFTSKL 278
>LEC2_CYTSC (P29257) 2-acetamido-2-deoxy-D-galactose-binding seed
lectin II (CSII)
Length = 248
Score = 166 bits (421), Expect = 6e-41
Identities = 107/251 (42%), Positives = 148/251 (58%), Gaps = 19/251 (7%)
Query: 83 SDSLSFSFPTFDPDVRVILLNGDANTT-NGTLQLTKIDQYGNPSPHSSGISAFYGAVHLS 141
S+ LSFSF F D + ++L DA T G LQLT ++ G P+ +S G + + +H+
Sbjct: 1 SEELSFSFTKFKTDQKNLILQRDALITPTGKLQLTTVEN-GKPAAYSLGRALYSTPIHIW 59
Query: 142 DKKTGRVADFRTEFTFVV----NHSQPHGDGFTFFLASLDFEFPEGGGGGGFLGLFDEET 197
DK TG A F T F+FV+ N S DG FFLA D + P+ GG+LGLF++++
Sbjct: 60 DKSTGDEASFATFFSFVISDAPNPSTAATDGLAFFLAPADTQ-PQ--SAGGYLGLFEKDS 116
Query: 198 AYNTSKNSIVAVEFDSFSNS-FDPNF-PHIGIDINTIRSAVTIPWSVNDQPQGAIQNAKI 255
+YN+S N IVAVEFD++ NS +DP PHIGID+NTI+S W + G + I
Sbjct: 117 SYNSS-NQIVAVEFDTYYNSAWDPQTNPHIGIDVNTIKSKKVSSWGFKN---GNVATVLI 172
Query: 256 SYSSVSRDLTVVVHYPPGQNTTNID-AIKYPIDIESYLSEWVVIGFSATTGE---FAETH 311
+Y S+ L + YP GQ + I +D+++ + EWV IGFSATTG+ + ETH
Sbjct: 173 TYQPSSKSLVASLVYPSGQTSDKTSYIISANVDLKATVPEWVRIGFSATTGQTDNYIETH 232
Query: 312 SILSWSFNSTL 322
ILSWSF S L
Sbjct: 233 DILSWSFKSKL 243
>LEC_SOYBN (P05046) Lectin precursor (Agglutinin) (SBA)
Length = 285
Score = 166 bits (419), Expect = 9e-41
Identities = 110/268 (41%), Positives = 154/268 (57%), Gaps = 19/268 (7%)
Query: 61 NITSELLATLIFSLLLITTTVKS-DSLSFSFPTFDPDVRVILLNGDAN-TTNGTLQLTKI 118
N+ L TL L+L+T+ S +++SFS+ F P ++L GDA T++G LQL K+
Sbjct: 10 NVVVSLSLTLTLVLVLLTSKANSAETVSFSWNKFVPKQPNMILQGDAIVTSSGKLQLNKV 69
Query: 119 DQYGNPSPHSSGISAFYGAVHLSDKKTGRVADFRTEF--TFVVNHSQPHGDGFTFFLASL 176
D+ G P P S G + + +H+ DK+TG VA F F TF ++ DG FFLA +
Sbjct: 70 DENGTPKPSSLGRALYSTPIHIWDKETGSVASFAASFNFTFYAPDTKRLADGLAFFLAPI 129
Query: 177 DFEFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNSFDPNFPHIGIDINTIRSAV 236
D + P+ G +LGLF+E N S + +VAVEFD+F NS+DP PHIGI++N+IRS
Sbjct: 130 DTK-PQTHAG--YLGLFNE----NESGDQVVAVEFDTFRNSWDPPNPHIGINVNSIRSIK 182
Query: 237 TIPWSVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWV 296
T W D + I+Y + S L V P Q T+NI + +D+++ L EWV
Sbjct: 183 TTSW---DLANNKVAKVLITYDA-STSLLVASLVYPSQRTSNI--LSDVVDLKTSLPEWV 236
Query: 297 VIGFSATTGEF--AETHSILSWSFNSTL 322
IGFSA TG E+H +LSWSF S L
Sbjct: 237 RIGFSAATGLDIPGESHDVLSWSFASNL 264
>LECS_VATMA (P81371) Seed lectin (VML)
Length = 240
Score = 165 bits (417), Expect = 2e-40
Identities = 105/248 (42%), Positives = 143/248 (57%), Gaps = 24/248 (9%)
Query: 83 SDSLSFSFPTFDPDVRVILLNGDANTTN-GTLQLTKIDQYGNPSPHSSGISAFYGAVHLS 141
S+ +SFSF F+P+ + I+L GDA T+ G LQLTK+ G P HS G + + +H+
Sbjct: 1 SEVVSFSFTKFNPNPKDIILQGDALVTSKGKLQLTKVKD-GKPVDHSLGRALYAAPIHIW 59
Query: 142 DKKTGRVADFRTEFTFVVN--HSQPHGDGFTFFLASLDFEFPEGGGGGGFLGLFDEETAY 199
D T RVA F T F+FVV DG FFLA D + + GG FLGLF
Sbjct: 60 DDSTDRVASFATSFSFVVEAPDESKTADGIAFFLAPPDTQPQKDGG---FLGLF------ 110
Query: 200 NTSKNSI--VAVEFDSFSNSFDPNFPHIGIDINTIRSAVTIPWSVNDQPQGAIQNAKISY 257
N S SI VAVEFD+FSN++DP+ HIGI++N+I S + W + G + N ISY
Sbjct: 111 NDSNKSIQTVAVEFDTFSNTWDPSARHIGINVNSIESMKYVKWGWEN---GKVANVYISY 167
Query: 258 SSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSATTG---EFAETHSIL 314
+ ++ LT + YP + + A +D++S L EWV +GFSAT+G + ETH +L
Sbjct: 168 EASTKTLTASLTYPSNATSYIVSA---NVDLKSALPEWVRVGFSATSGLSRDHVETHDVL 224
Query: 315 SWSFNSTL 322
WSF STL
Sbjct: 225 DWSFTSTL 232
>LCS1_ROBPS (Q41162) Seed agglutinin I precursor (RPSAI) (LECRPAS1)
Length = 285
Score = 162 bits (409), Expect = 1e-39
Identities = 107/264 (40%), Positives = 149/264 (55%), Gaps = 19/264 (7%)
Query: 66 LLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDA-NTTNGTLQLTKIDQYGNP 124
LL+ F LLL+ + SLSFSFP F P+ ++ DA T+ G LQLT + G P
Sbjct: 15 LLSISFFFLLLLNKVNSTGSLSFSFPKFAPNQPYLIFQRDALVTSTGVLQLTNVVN-GVP 73
Query: 125 SPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNHSQP--HGDGFTFFLASLDFEFPE 182
S G + + + D TG VA F T F+F++ P DG FFLA +D +
Sbjct: 74 PRRSIGRALYAAPFQIWDNTTGNVASFVTSFSFIIQAPNPATTADGLAFFLAPVD---TQ 130
Query: 183 GGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSN-SFDPNFPHIGIDINTIRSAVTIPWS 241
G GG LG+F ++ +YN S N IVAVEFD+FSN FDP H+GI++N+I S T+PW+
Sbjct: 131 PGDLGGMLGIF-KDGSYNKS-NQIVAVEFDTFSNIHFDPKGRHMGINVNSIVSVKTVPWN 188
Query: 242 VNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFS 301
+ G + N ISY + ++ L + YP + + I AI +D++ L EWV GFS
Sbjct: 189 WTN---GEVANVFISYEASTKSLNASLVYPSLETSFIIHAI---VDVKDVLPEWVRFGFS 242
Query: 302 ATTG---EFAETHSILSWSFNSTL 322
ATTG + +T+ +LSWSF S L
Sbjct: 243 ATTGIDTGYVQTNDVLSWSFESNL 266
>LCB1_ROBPS (Q41159) Bark agglutinin I, polypeptide A precursor
(RPbAI) (LECRPA1)
Length = 285
Score = 160 bits (406), Expect = 3e-39
Identities = 105/264 (39%), Positives = 147/264 (54%), Gaps = 19/264 (7%)
Query: 66 LLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDA-NTTNGTLQLTKIDQYGNP 124
LL+ F LLL+ + SLSFSFP F P+ ++ DA T+ G LQLT + G P
Sbjct: 15 LLSISFFFLLLLNKVNSTGSLSFSFPKFAPNQPYLIFQRDALVTSTGVLQLTNVVN-GVP 73
Query: 125 SPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNHSQP--HGDGFTFFLASLDFEFPE 182
S S G + + + D TG VA F T F+F++ P DG FFLA +D + +
Sbjct: 74 SGKSLGRALYAAPFQIWDSTTGNVASFVTSFSFIIQAPNPTTTADGLAFFLAPVDTQPLD 133
Query: 183 GGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSN-SFDPNFPHIGIDINTIRSAVTIPWS 241
GG LG+F + Y N IVAVEFD+FSN FDP H+GI++N+I S T+PW+
Sbjct: 134 ---VGGMLGIFKD--GYFNKSNQIVAVEFDTFSNIHFDPKGRHMGINVNSIVSIKTVPWN 188
Query: 242 VNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFS 301
+ G + N ISY + ++ LT + YP + + + AI +D++ L EWV GFS
Sbjct: 189 WTN---GEVANVFISYEASTKSLTASLVYPSLETSFIVHAI---VDVKDVLPEWVRFGFS 242
Query: 302 ATTG---EFAETHSILSWSFNSTL 322
ATTG + +T+ +LSWSF S L
Sbjct: 243 ATTGIDKGYVQTNDVLSWSFESNL 266
>LCB2_ROBPS (Q42372) Bark agglutinin I, polypeptide B precursor
(RPbAI) (LECRPA2)
Length = 286
Score = 160 bits (404), Expect = 5e-39
Identities = 106/264 (40%), Positives = 149/264 (56%), Gaps = 18/264 (6%)
Query: 66 LLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTN-GTLQLTKIDQYGNP 124
LL+ F LLL+ + SLSFSFP F ++ DA T+ G LQLT ++ G P
Sbjct: 15 LLSISFFFLLLLNKVNSTGSLSFSFPKFKHSQPDLIFQSDALVTSKGVLQLTTVND-GRP 73
Query: 125 SPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVN--HSQPHGDGFTFFLASLDFEFPE 182
S G + + D TG VA F T F+F++ + DG FFLA + P
Sbjct: 74 VYDSIGRVLYAAPFQIWDSTTGNVASFVTSFSFIIKAPNEGKTADGLVFFLAPVGSTQPL 133
Query: 183 GGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSN-SFDPNFPHIGIDINTIRSAVTIPWS 241
GGG LGLF +E+ YN S N IVAVEFD+F N ++DPN H+GID+N+I+S T+ W
Sbjct: 134 KGGG--LLGLFKDES-YNKS-NQIVAVEFDTFRNVAWDPNGIHMGIDVNSIQSVRTVRW- 188
Query: 242 VNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFS 301
D G + N ISY + ++ LT + YP + + + AI +D++ L EWV +GF+
Sbjct: 189 --DWANGEVANVFISYEASTKSLTASLVYPSLEKSFILSAI---VDLKKVLPEWVRVGFT 243
Query: 302 ATTG---EFAETHSILSWSFNSTL 322
ATTG ++ +T+ +LSWSF S L
Sbjct: 244 ATTGLSEDYVQTNDVLSWSFESNL 267
>LCS2_ROBPS (Q41161) Seed agglutinin II precursor (RPSAII)
(LECRPAS2)
Length = 285
Score = 159 bits (401), Expect = 1e-38
Identities = 105/264 (39%), Positives = 146/264 (54%), Gaps = 19/264 (7%)
Query: 66 LLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDA-NTTNGTLQLTKIDQYGNP 124
LL+ F LLL+ + SLSFSFP F P+ ++ DA T+ G LQLT + G P
Sbjct: 15 LLSISFFFLLLLNKVNSTGSLSFSFPKFAPNQPYLIFQRDALVTSTGVLQLTNVVN-GVP 73
Query: 125 SPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNHSQP--HGDGFTFFLASLDFEFPE 182
S S G + + + D TG VA F T F+F++ P DG FFLA +D + +
Sbjct: 74 SRKSLGRALYAAPFQIWDSTTGNVASFVTSFSFIIQAPNPATTADGLAFFLAPVDTQPLD 133
Query: 183 GGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSN-SFDPNFPHIGIDINTIRSAVTIPWS 241
GG LG+F + Y N IVAVEFD+FSN +DP H+GI++N+I S T+PW
Sbjct: 134 ---LGGMLGIF--KNGYFNKSNQIVAVEFDTFSNRHWDPTGRHMGINVNSIVSVKTVPW- 187
Query: 242 VNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFS 301
+ G + N ISY + ++ LT + YP + + I AI +D++ L EWV GFS
Sbjct: 188 --NWANGEVANVFISYEASTKSLTASLVYPSLETSFIIHAI---VDVKDVLPEWVRFGFS 242
Query: 302 ATTG---EFAETHSILSWSFNSTL 322
ATTG + +T+ +LSWSF S L
Sbjct: 243 ATTGIDTGYVQTNDVLSWSFESNL 266
>LEC4_GRISI (P24146) Lectin IV (GS4)
Length = 243
Score = 153 bits (386), Expect = 6e-37
Identities = 101/246 (41%), Positives = 140/246 (56%), Gaps = 16/246 (6%)
Query: 84 DSLSFSFPTFDP----DVRVILLNGDANTTNGTLQLTKIDQYGNPSPHSSGISAFYGAVH 139
++++F++P F + I GDA G LQLTK D GNP S+G +++ V
Sbjct: 2 NTVNFTYPDFWSYSLKNGTEITFLGDATRIPGALQLTKTDANGNPVRSSAGQASYSEPVF 61
Query: 140 LSDKKTGRVADFRTEFTFVV-NHSQPHGDGFTFFLASLDFEFPEGGGGGGFLGLFDEETA 198
L D TG+ A F T FTF++ N+ P DG FFLA +D + GG FLGLF ETA
Sbjct: 62 LWDS-TGKAASFYTSFTFLLKNYGAPTADGLAFFLAPVDSSVKDYGG---FLGLFRHETA 117
Query: 199 YNTSKNSIVAVEFDSFSNSF--DPNFPHIGIDINTIRSAVTIPWSVNDQPQGAIQNAKIS 256
+ SKN +VAVEFD++ N DP +PHIGID+N+I S T W +D +I A I+
Sbjct: 118 ADPSKNQVVAVEFDTWINKDWNDPPYPHIGIDVNSIVSVATTRWENDDAYGSSIATAHIT 177
Query: 257 YSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSATTGEFAETHSILSW 316
Y + S+ LTV++ Y G++ + + +D+ L + V IGFSA G + E ILSW
Sbjct: 178 YDARSKILTVLLSYEHGRDY----ILSHVVDLAKVLPQKVRIGFSAGVG-YDEVTYILSW 232
Query: 317 SFNSTL 322
F STL
Sbjct: 233 HFFSTL 238
>LEC1_DOLBI (P05045) Seed lectin subunit I precursor (SL) [Contains:
Seed lectin subunit II]
Length = 275
Score = 150 bits (379), Expect = 4e-36
Identities = 104/270 (38%), Positives = 150/270 (55%), Gaps = 21/270 (7%)
Query: 59 MANITSELLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTNGTLQLTKI 118
MA+ T ++ +L LLL+T ++ SFSF F+ +L GDA ++G LQLTK+
Sbjct: 1 MASSTVSVVLSLF--LLLLTQANSANIQSFSFKNFNSPS--FILQGDATVSSGKLQLTKV 56
Query: 119 DQYGNPSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVN--HSQPHGDGFTFFLASL 176
+ G P+P S G + + + + DK TG VA + T FT ++ DG F L +
Sbjct: 57 KENGIPTPSSLGRAFYSSPIQIYDKSTGAVASWATSFTVKISAPSKASFADGIAFALVPV 116
Query: 177 DFEFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNS-FDPNFPHIGIDINTIRSA 235
E GG+LG+FD + YN S + VAVEFD+FSNS +DP+ HIGID+N+I+S
Sbjct: 117 G---SEPRRNGGYLGVFDSD-VYNNSAQT-VAVEFDTFSNSGWDPSMKHIGIDVNSIKSI 171
Query: 236 VTIPWSVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEW 295
T+ W D G I+Y++ + L V P + T+ I + +DI + L E+
Sbjct: 172 ATVSW---DLANGENAEILITYNAAT-SLLVASLVHPSRRTSYI--LSERVDITNELPEY 225
Query: 296 VVIGFSATTG---EFAETHSILSWSFNSTL 322
V +GFSATTG + ETH +LSWSF S L
Sbjct: 226 VSVGFSATTGLSEGYIETHDVLSWSFASKL 255
>LEC2_MEDTR (Q01807) Truncated lectin 2 precursor
Length = 280
Score = 150 bits (378), Expect = 5e-36
Identities = 98/274 (35%), Positives = 155/274 (55%), Gaps = 27/274 (9%)
Query: 60 ANITSELLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTN-GTLQLTKI 118
+N + L +L F +LL+ +++ SFS F PD + ++ GDA T + G L+L+K
Sbjct: 4 SNFSCILSISLTFFILLLNKVNSAETTSFSITKFVPDQKNLIFQGDAKTASTGKLELSKA 63
Query: 119 DQYGNPSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNHSQPHG--DGFTFFLASL 176
+ +S G + + +H+ D KTG VA+F+T FTF + + DG FF+A +
Sbjct: 64 VK------NSIGRALYSAPIHIWDSKTGSVANFQTTFTFTITAPNTYNVADGLAFFIAPI 117
Query: 177 DFEFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNS-FDPN-------FPHIGID 228
D + P+ GG+LG+FD +T Y S + VAVE D+F N+ +DPN HIGID
Sbjct: 118 DTK-PKSIHHGGYLGVFDSKT-YKKSIQT-VAVEIDTFYNAQWDPNPGNISSTGRHIGID 174
Query: 229 INTIRSAVTIPWSVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDI 288
+N+I+S T+PWS+ + + N I ++ + L+V V YP ++ T + + + +
Sbjct: 175 VNSIKSISTVPWSLENNKKA---NVAIGFNGATNVLSVDVEYPLIRHYT----LSHVVPL 227
Query: 289 ESYLSEWVVIGFSATTGEFAETHSILSWSFNSTL 322
+ + EWV IGFS++TG H ILSWSF+S L
Sbjct: 228 KDVVPEWVRIGFSSSTGAEYSAHDILSWSFDSKL 261
>LEC_ERYCO (P16404) Lectin precursor (ECorL)
Length = 281
Score = 149 bits (377), Expect = 7e-36
Identities = 100/271 (36%), Positives = 149/271 (54%), Gaps = 23/271 (8%)
Query: 62 ITSELLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTT-NGTLQLTKIDQ 120
+ S L +L LL++ +++SFSF F+P + L G A T +G LQLTKI+Q
Sbjct: 6 LCSVLALSLTLFLLILNKVNSVETISFSFSEFEPGNDNLTLQGAALITQSGVLQLTKINQ 65
Query: 121 YGNPSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNHSQPH-----GDGFTFFLAS 175
G P+ S+G + + VH+ D TG VA F T F+F + QP+ DG FF+
Sbjct: 66 NGMPAWDSTGRTLYAKPVHIWDMTTGTVASFETRFSFSI--EQPYTRPLPADGLVFFMGP 123
Query: 176 LDFEFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNSFD-PNFPHIGIDINTIRS 234
+ G G+LG+F+ N+ + + VEFD+FSN +D P PHIGID+N+IRS
Sbjct: 124 TK---SKPAQGYGYLGIFNNSKQDNSYQT--LGVEFDTFSNPWDPPQVPHIGIDVNSIRS 178
Query: 235 AVTIPWSVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSE 294
T P+ +++ G + N I Y + S+ L V+ YP ++ I I +D++ L E
Sbjct: 179 IKTQPFQLDN---GQVANVVIKYDASSKILHAVLVYP---SSGAIYTIAEIVDVKQVLPE 232
Query: 295 WVVIGFSATTG---EFAETHSILSWSFNSTL 322
WV +G S TG + AETH + SWSF ++L
Sbjct: 233 WVDVGLSGATGAQRDAAETHDVYSWSFQASL 263
>LEC_BOWMI (P42088) Lectin (Agglutinin) (BMA)
Length = 240
Score = 149 bits (376), Expect = 9e-36
Identities = 94/244 (38%), Positives = 141/244 (57%), Gaps = 14/244 (5%)
Query: 83 SDSLSFSFPTFDPDVRVILLNGDANT-TNGTLQLTKIDQYGNPSPHSSGISAFYGAVHLS 141
++S+ F+F F+ + ++ GDA+ +N LQLTK+D GNP S G + + + L
Sbjct: 1 ANSVCFTFTDFESGQQDLIFQGDASVGSNKALQLTKVDSKGNPQGGSVGRALYTAPIRLW 60
Query: 142 DKKTGRVADFRTEFTFVVNH-SQPHGDGFTFFLASLDFEFPEGGGGGGFLGLFDEETAYN 200
+ + VA F T FTF ++ S TFF+AS D + P G GG LGLF
Sbjct: 61 -QSSSLVASFETTFTFSISQGSSTPAAALTFFIASPDTKIPSGSGGR-LLGLFGSSNNAG 118
Query: 201 TSKNSIVAVEFDSFSNSF--DPNFPHIGIDINTIRSAVTIPWSVNDQPQGAIQNAKISYS 258
S N +VAVEFD++ N+ DPN+ HIGID+N+IRS W D G A ISY+
Sbjct: 119 -SDNGVVAVEFDTYPNTDIGDPNYRHIGIDVNSIRSKAASKW---DWQNGKTATAHISYN 174
Query: 259 SVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSATTGEFAETHSILSWSF 318
S S+ L+VV YP N++ + + + +++ + V +GFSATTG++ +T++IL+WSF
Sbjct: 175 SASKRLSVVSSYP---NSSPV-VVSFDVELNNVGPPDVRVGFSATTGQYTQTNNILAWSF 230
Query: 319 NSTL 322
S+L
Sbjct: 231 RSSL 234
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.135 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,805,514
Number of Sequences: 164201
Number of extensions: 1675168
Number of successful extensions: 4462
Number of sequences better than 10.0: 83
Number of HSP's better than 10.0 without gapping: 70
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 4080
Number of HSP's gapped (non-prelim): 93
length of query: 322
length of database: 59,974,054
effective HSP length: 110
effective length of query: 212
effective length of database: 41,911,944
effective search space: 8885332128
effective search space used: 8885332128
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0541.15


