

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0541.11
(269 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
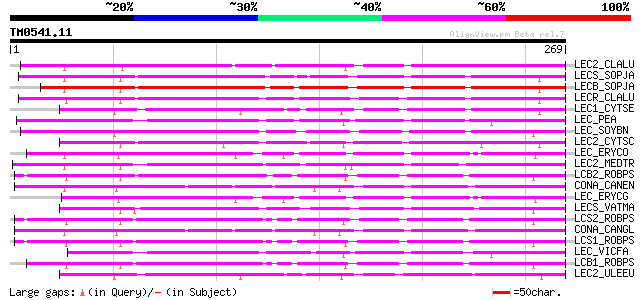
Score E
Sequences producing significant alignments: (bits) Value
LEC2_CLALU (Q39529) Agglutinin II precursor (ClAII) (LecClAII) 201 2e-51
LECS_SOPJA (P93535) Seed lectin precursor (LECSJASG) 179 6e-45
LECB_SOPJA (P93538) Bark lectin precursor (LECSJABG) (Fragment) 177 2e-44
LECR_CLALU (Q39527) Lectin-related protein precursor (CLLRP) (LR... 169 5e-42
LEC1_CYTSE (P22970) Anti-H(O) lectin I (CSA-I) 167 2e-41
LEC_PEA (P02867) Lectin precursor 166 7e-41
LEC_SOYBN (P05046) Lectin precursor (Agglutinin) (SBA) 162 6e-40
LEC2_CYTSC (P29257) 2-acetamido-2-deoxy-D-galactose-binding seed... 162 1e-39
LEC_ERYCO (P16404) Lectin precursor (ECorL) 161 1e-39
LEC2_MEDTR (Q01807) Truncated lectin 2 precursor 161 2e-39
LCB2_ROBPS (Q42372) Bark agglutinin I, polypeptide B precursor (... 161 2e-39
CONA_CANEN (P02866) Concanavalin A precursor (Con A) 161 2e-39
LEC_ERYCG (P83410) Lectin (ECL) 160 2e-39
LECS_VATMA (P81371) Seed lectin (VML) 160 2e-39
LCS2_ROBPS (Q41161) Seed agglutinin II precursor (RPSAII) (LECRP... 160 3e-39
CONA_CANGL (P14894) Concanavalin A precursor (Con A) 159 5e-39
LCS1_ROBPS (Q41162) Seed agglutinin I precursor (RPSAI) (LECRPAS1) 155 7e-38
LEC_VICFA (P02871) Favin (Lectin) 154 2e-37
LCB1_ROBPS (Q41159) Bark agglutinin I, polypeptide A precursor (... 154 3e-37
LEC2_ULEEU (P22973) Anti-H(O) lectin II (UEA-II) 153 5e-37
>LEC2_CLALU (Q39529) Agglutinin II precursor (ClAII) (LecClAII)
Length = 290
Score = 201 bits (511), Expect = 2e-51
Identities = 116/268 (43%), Positives = 158/268 (58%), Gaps = 13/268 (4%)
Query: 6 SISLLTTLLPFILLITTVKS-DSFSFNFPRFESDTKTILSDGDANTTGG--VLQLTKKDL 62
S+ LL + F++L+ V S DS SF F F D + ++ GDA + G LQLTK D
Sbjct: 16 SLPLLAFITLFLMLLNRVNSSDSLSFTFDNFRPDQRDLILQGDAKISSGGDSLQLTKTDT 75
Query: 63 YGIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDF 122
G P + S G+ ++ +HL D+ T ++A F T F+FV++ + P GDG F+IA +
Sbjct: 76 SGKPVRGSVGRALYYTPLHLWDSSTNRLASFQTTFTFVLSSPTNNP-GDGIAFFIAPPET 134
Query: 123 DFPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNV-WDPSYPSNSPHVGIDINSIR 181
P GG LGLF P NA N S NQ+VAVEFD+F N WDPS+ H+GID+N+I+
Sbjct: 135 TIPPG-SSGGLLGLFSPDNALNNSLNQIVAVEFDTFVNNNWDPSHR----HIGIDVNTIK 189
Query: 182 SVATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSD 241
S AT W + G + A ISY S +K LSV YPN+ + V+Y+VDL L +
Sbjct: 190 SSATVRWQRE---NGSLATAQISYNSDTKKLSVVSSYPNTQANEDYTVSYDVDLKTELPE 246
Query: 242 WVLVGFTAATGSVSETHDILSWAFNSFL 269
WV VGF+ +TG + H+ILSW FNS L
Sbjct: 247 WVRVGFSGSTGGYVQNHNILSWTFNSNL 274
>LECS_SOPJA (P93535) Seed lectin precursor (LECSJASG)
Length = 292
Score = 179 bits (454), Expect = 6e-45
Identities = 110/270 (40%), Positives = 160/270 (58%), Gaps = 19/270 (7%)
Query: 5 FSISLLTTLLPFILLITTVKS-DSFSFNFPRFESDTKTILSDGDANTTG-GVLQLTKKDL 62
FS+ L ++ F+LL+ V S + SF+FP+F S+ + +L GDA + G LQLT +
Sbjct: 17 FSVVLAISITFFLLLLNKVNSAEILSFSFPKFASNQEDLLLQGDALVSSKGELQLTTVE- 75
Query: 63 YGIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDF 122
G+P NS G+ ++ VH+ D TG+VA F T FSFVV + DG F++A +
Sbjct: 76 NGVPIWNSTGRALYYAPVHIWDKSTGRVASFATSFSFVVKAPVASKSADGIAFFLAPPNN 135
Query: 123 DFPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDINSIRS 182
GG LGLF + +N S+ Q++AV+FD+ N WDP N+ H+GID+NSI S
Sbjct: 136 QI--QGPGGGHLGLFH-SSGYN-SSYQIIAVDFDTHINAWDP----NTRHIGIDVNSINS 187
Query: 183 VATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDW 242
T W GE+ LISY++A++ L+V + YP+S + ++ VDL ++L +W
Sbjct: 188 TKTVTWGWQ---NGEVANVLISYQAATETLTVSLTYPSS--QTSYILSAAVDLKSILPEW 242
Query: 243 VLVGFTAATGSVS---ETHDILSWAFNSFL 269
V VGFTAATG + ETHD+LSW+F S L
Sbjct: 243 VRVGFTAATGLTTQYVETHDVLSWSFTSTL 272
>LECB_SOPJA (P93538) Bark lectin precursor (LECSJABG) (Fragment)
Length = 270
Score = 177 bits (449), Expect = 2e-44
Identities = 105/259 (40%), Positives = 159/259 (60%), Gaps = 19/259 (7%)
Query: 16 FILLITTVKS-DSFSFNFPRFESDTKTILSDGDANTTG-GVLQLTKKDLYGIPSKNSFGQ 73
F+LL+ V S + SF+FP+F S+ + +L GDA + G LQLT + G+P NS G+
Sbjct: 6 FLLLLNKVNSAEILSFSFPKFVSNQEDLLLQGDALVSSEGELQLTTVE-NGVPVWNSTGR 64
Query: 74 TTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFDFPEDPKEGGF 133
++ VH+ DN TG+VA F T FSFVV + DG F++A L+ GG
Sbjct: 65 ALYYAPVHIWDNSTGRVASFATSFSFVVKAPVASKSADGIAFFLAPLNNQI--HGAGGGL 122
Query: 134 LGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDINSIRSVATAPWPIDLV 193
GLF+ ++ +S+ Q+VAVEFD+ TN WDP N+ H+GID+NS++S T W +
Sbjct: 123 YGLFN--SSSYSSSYQIVAVEFDTHTNAWDP----NTRHIGIDVNSVKSTKTVTWGWE-- 174
Query: 194 PQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDWVLVGFTAATGS 253
GE+ LI+Y++A+++L+V + YP++ + ++ VDL ++L +WV VGFTA TG
Sbjct: 175 -NGEVANVLITYQAATEMLTVSLTYPSN--QTSYILSAAVDLKSILPEWVRVGFTATTGL 231
Query: 254 VS---ETHDILSWAFNSFL 269
+ ET+D+LSW+F S L
Sbjct: 232 TTQYVETNDVLSWSFTSTL 250
>LECR_CLALU (Q39527) Lectin-related protein precursor (CLLRP)
(LRPCL) (Fragment)
Length = 290
Score = 169 bits (429), Expect = 5e-42
Identities = 106/270 (39%), Positives = 158/270 (58%), Gaps = 20/270 (7%)
Query: 5 FSISLLTTLLPFILLITTVKSD-SFSFNFPRFESDTKTILSDGDANTTG-GVLQLTKKDL 62
FS+ L ++ ++LL+ V S+ + SF F +F S+ +L GDA + G LQLT+ +
Sbjct: 16 FSVVLAISITFYLLLLNKVNSEEALSFTFTKFVSNQDELLLQGDALVSSKGELQLTRVE- 74
Query: 63 YGIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDF 122
G P +S G+ + VH+ D+ TG VA F T F+FVV DG F++A D
Sbjct: 75 NGQPIPHSVGRALYSDPVHIWDSSTGSVASFVTSFTFVVEAPNENKTADGIAFFLAPPD- 133
Query: 123 DFPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDINSIRS 182
+ GGFLGLF+ ++ S+NQ++AVEFD+F+N WDP+ + H+GID+NSI S
Sbjct: 134 --TQVQSLGGFLGLFN--SSVYNSSNQILAVEFDTFSNSWDPT----ARHIGIDVNSIES 185
Query: 183 VATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDW 242
TA W GE+ LI+Y + ++ L + YP+S + ++ VDL ++L +W
Sbjct: 186 TRTATWG---WRNGEVAIVLITYVAPAETLIASLTYPSS--QTSYILSAAVDLKSILPEW 240
Query: 243 VLVGFTAATGSVS---ETHDILSWAFNSFL 269
V VGF+AATG + ETHD+LSW+F S L
Sbjct: 241 VRVGFSAATGRSAGYVETHDVLSWSFTSTL 270
>LEC1_CYTSE (P22970) Anti-H(O) lectin I (CSA-I)
Length = 244
Score = 167 bits (423), Expect = 2e-41
Identities = 101/253 (39%), Positives = 148/253 (57%), Gaps = 23/253 (9%)
Query: 25 SDSFSFNFPRFESDTKTILSDGDAN-TTGGVLQLTKKDLYGIPSKNSFGQTTFFGAVHLS 83
+D SFNF +F + IL G+A+ +T GVLQ+TK P+ S G+ + VH+
Sbjct: 2 NDHLSFNFDKFVPNQNNILFQGEASVSTTGVLQVTKVSK---PATRSIGRALYAAPVHIW 58
Query: 84 DNRTGKVADFTTEFSFVVNPKGSEPHG-DGFTFYIAGLDFDFPEDPKEGGFLGLFDPKNA 142
D+ TG+VA F T FSFVV + + +G DG TF++A + P G F GLF+ +
Sbjct: 59 DSTTGRVASFETSFSFVVKDEPEKSNGVDGLTFFLAPANSQIPSGSSAGLF-GLFNSSD- 116
Query: 143 FNTSANQVVAVEFDSFT----NVWDPSYPSNSPHVGIDINSIRSVATAPWPIDLVPQGEI 198
N S+NQ++AVEFD++ N WDP + H+G+D+NSI+S+ T W GE+
Sbjct: 117 -NKSSNQIIAVEFDTYFGKTYNPWDPDFK----HIGVDVNSIKSIKTVKWD---WRNGEV 168
Query: 199 GKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDWVLVGFTAATGSVS--E 256
+I+YR+ +K L+V + YP+ + V VDL A+L +WV VGF+A G+ + E
Sbjct: 169 ANVVITYRAPTKSLTVSLSYPSD--QTSNIVTASVDLKAILPEWVSVGFSAGVGNAAEFE 226
Query: 257 THDILSWAFNSFL 269
THD+LSW F S L
Sbjct: 227 THDVLSWYFTSNL 239
>LEC_PEA (P02867) Lectin precursor
Length = 275
Score = 166 bits (419), Expect = 7e-41
Identities = 103/271 (38%), Positives = 151/271 (55%), Gaps = 22/271 (8%)
Query: 4 SFSISLLTTLLPFILLITTVKSDSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKKDLY 63
SF L+ LL IL +++ SF +F D + ++ GD TT L LTK
Sbjct: 10 SFYAIFLSILLTTILFFKVNSTETTSFLITKFSPDQQNLIFQGDGYTTKEKLTLTKA--- 66
Query: 64 GIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFD 123
KN+ G+ + +H+ D TG VA+F T F+FV+N S DGFTF+IA +D
Sbjct: 67 ---VKNTVGRALYSSPIHIWDRETGNVANFVTSFTFVINAPNSYNVADGFTFFIAPVD-- 121
Query: 124 FPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTN-VWDPSYPSNSPHVGIDINSIRS 182
+ GG+LG+F+ +A Q VAVEFD+F N WDPS + H+GID+NSI+S
Sbjct: 122 -TKPQTGGGYLGVFN--SAEYDKTTQTVAVEFDTFYNAAWDPS--NRDRHIGIDVNSIKS 176
Query: 183 VATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYE----VDLGAV 238
V T W + GE +I++ +A+ VL+V + YPNS ++ E +Y V L V
Sbjct: 177 VNTKSWKLQ---NGEEANVVIAFNAATNVLTVSLTYPNS-LEEENVTSYTLSDVVSLKDV 232
Query: 239 LSDWVLVGFTAATGSVSETHDILSWAFNSFL 269
+ +WV +GF+A TG+ H++LSW+F+S L
Sbjct: 233 VPEWVRIGFSATTGAEYAAHEVLSWSFHSEL 263
>LEC_SOYBN (P05046) Lectin precursor (Agglutinin) (SBA)
Length = 285
Score = 162 bits (411), Expect = 6e-40
Identities = 97/267 (36%), Positives = 149/267 (55%), Gaps = 19/267 (7%)
Query: 6 SISLLTTLLPFILLITTVKSDSFSFNFPRFESDTKTILSDGDAN-TTGGVLQLTKKDLYG 64
S+SL TL+ +L +++ SF++ +F ++ GDA T+ G LQL K D G
Sbjct: 14 SLSLTLTLVLVLLTSKANSAETVSFSWNKFVPKQPNMILQGDAIVTSSGKLQLNKVDENG 73
Query: 65 IPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFDF 124
P +S G+ + +H+ D TG VA F F+F ++ DG F++A +D
Sbjct: 74 TPKPSSLGRALYSTPIHIWDKETGSVASFAASFNFTFYAPDTKRLADGLAFFLAPID--- 130
Query: 125 PEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDINSIRSVA 184
+ G+LGLF+ N S +QVVAVEFD+F N WDP +PH+GI++NSIRS+
Sbjct: 131 TKPQTHAGYLGLFNE----NESGDQVVAVEFDTFRNSWDPP----NPHIGINVNSIRSIK 182
Query: 185 TAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDWVL 244
T W + ++ K LI+Y +++ +L +VYP+ + ++ VDL L +WV
Sbjct: 183 TTSWDL---ANNKVAKVLITYDASTSLLVASLVYPSQ--RTSNILSDVVDLKTSLPEWVR 237
Query: 245 VGFTAATG--SVSETHDILSWAFNSFL 269
+GF+AATG E+HD+LSW+F S L
Sbjct: 238 IGFSAATGLDIPGESHDVLSWSFASNL 264
>LEC2_CYTSC (P29257) 2-acetamido-2-deoxy-D-galactose-binding seed
lectin II (CSII)
Length = 248
Score = 162 bits (409), Expect = 1e-39
Identities = 99/254 (38%), Positives = 146/254 (56%), Gaps = 20/254 (7%)
Query: 25 SDSFSFNFPRFESDTKTILSDGDANTTG-GVLQLTKKDLYGIPSKNSFGQTTFFGAVHLS 83
S+ SF+F +F++D K ++ DA T G LQLT + G P+ S G+ + +H+
Sbjct: 1 SEELSFSFTKFKTDQKNLILQRDALITPTGKLQLTTVE-NGKPAAYSLGRALYSTPIHIW 59
Query: 84 DNRTGKVADFTTEFSFVVN--PKGSEPHGDGFTFYIAGLDFDFPEDPKEGGFLGLFDPKN 141
D TG A F T FSFV++ P S DG F++A D + GG+LGLF+ +
Sbjct: 60 DKSTGDEASFATFFSFVISDAPNPSTAATDGLAFFLAPAD---TQPQSAGGYLGLFEKDS 116
Query: 142 AFNTSANQVVAVEFDSFTN-VWDPSYPSNSPHVGIDINSIRSVATAPWPIDLVPQGEIGK 200
++N S+NQ+VAVEFD++ N WD P +PH+GID+N+I+S + W G +
Sbjct: 117 SYN-SSNQIVAVEFDTYYNSAWD---PQTNPHIGIDVNTIKSKKVSSWGF---KNGNVAT 169
Query: 201 ALISYRSASKVLSVFVVYPNSPVKIET--FVAYEVDLGAVLSDWVLVGFTAATGSVS--- 255
LI+Y+ +SK L +VYP+ +T ++ VDL A + +WV +GF+A TG
Sbjct: 170 VLITYQPSSKSLVASLVYPSGQTSDKTSYIISANVDLKATVPEWVRIGFSATTGQTDNYI 229
Query: 256 ETHDILSWAFNSFL 269
ETHDILSW+F S L
Sbjct: 230 ETHDILSWSFKSKL 243
>LEC_ERYCO (P16404) Lectin precursor (ECorL)
Length = 281
Score = 161 bits (408), Expect = 1e-39
Identities = 104/268 (38%), Positives = 155/268 (57%), Gaps = 21/268 (7%)
Query: 9 LLTTLLPFILLITTVKS-DSFSFNFPRFESDTKTILSDGDANTT-GGVLQLTKKDLYGIP 66
L +L F+L++ V S ++ SF+F FE + G A T GVLQLTK + G+P
Sbjct: 10 LALSLTLFLLILNKVNSVETISFSFSEFEPGNDNLTLQGAALITQSGVLQLTKINQNGMP 69
Query: 67 SKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEP-HGDGFTFYIAGLDFDFP 125
+ +S G+T + VH+ D TG VA F T FSF + + P DG F++
Sbjct: 70 AWDSTGRTLYAKPVHIWDMTTGTVASFETRFSFSIEQPYTRPLPADGLVFFMG----PTK 125
Query: 126 EDPKEG-GFLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDINSIRSVA 184
P +G G+LG+F+ N+ ++ Q + VEFD+F+N WD P PH+GID+NSIRS+
Sbjct: 126 SKPAQGYGYLGIFN--NSKQDNSYQTLGVEFDTFSNPWD---PPQVPHIGIDVNSIRSIK 180
Query: 185 TAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDWVL 244
T P+ +D G++ +I Y ++SK+L +VYP+S I T +A VD+ VL +WV
Sbjct: 181 TQPFQLD---NGQVANVVIKYDASSKILHAVLVYPSSGA-IYT-IAEIVDVKQVLPEWVD 235
Query: 245 VGFTAATGS---VSETHDILSWAFNSFL 269
VG + ATG+ +ETHD+ SW+F + L
Sbjct: 236 VGLSGATGAQRDAAETHDVYSWSFQASL 263
>LEC2_MEDTR (Q01807) Truncated lectin 2 precursor
Length = 280
Score = 161 bits (407), Expect = 2e-39
Identities = 101/274 (36%), Positives = 155/274 (55%), Gaps = 21/274 (7%)
Query: 2 ANSFSISLLTTLLPFILLITTVKS-DSFSFNFPRFESDTKTILSDGDANTTG-GVLQLTK 59
+++FS L +L FILL+ V S ++ SF+ +F D K ++ GDA T G L+L+K
Sbjct: 3 SSNFSCILSISLTFFILLLNKVNSAETTSFSITKFVPDQKNLIFQGDAKTASTGKLELSK 62
Query: 60 KDLYGIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAG 119
KNS G+ + +H+ D++TG VA+F T F+F + + DG F+IA
Sbjct: 63 A------VKNSIGRALYSAPIHIWDSKTGSVANFQTTFTFTITAPNTYNVADGLAFFIAP 116
Query: 120 LDFDFPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNV-WDP---SYPSNSPHVGI 175
+D P+ GG+LG+FD K + Q VAVE D+F N WDP + S H+GI
Sbjct: 117 IDTK-PKSIHHGGYLGVFDSKT--YKKSIQTVAVEIDTFYNAQWDPNPGNISSTGRHIGI 173
Query: 176 DINSIRSVATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDL 235
D+NSI+S++T PW ++ + I + A+ VLSV V Y P+ +++ V L
Sbjct: 174 DVNSIKSISTVPWSLE---NNKKANVAIGFNGATNVLSVDVEY---PLIRHYTLSHVVPL 227
Query: 236 GAVLSDWVLVGFTAATGSVSETHDILSWAFNSFL 269
V+ +WV +GF+++TG+ HDILSW+F+S L
Sbjct: 228 KDVVPEWVRIGFSSSTGAEYSAHDILSWSFDSKL 261
>LCB2_ROBPS (Q42372) Bark agglutinin I, polypeptide B precursor
(RPbAI) (LECRPA2)
Length = 286
Score = 161 bits (407), Expect = 2e-39
Identities = 110/273 (40%), Positives = 153/273 (55%), Gaps = 21/273 (7%)
Query: 3 NSFSISLLTTLLPFILLITTVKSD-SFSFNFPRFESDTKTILSDGDANTTG-GVLQLTKK 60
NSF + LL+ F+LL+ V S S SF+FP+F+ ++ DA T GVLQLT
Sbjct: 10 NSFLL-LLSISFFFLLLLNKVNSTGSLSFSFPKFKHSQPDLIFQSDALVTSKGVLQLTTV 68
Query: 61 DLYGIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGL 120
+ G P +S G+ + + D+ TG VA F T FSF++ DG F++A +
Sbjct: 69 N-DGRPVYDSIGRVLYAAPFQIWDSTTGNVASFVTSFSFIIKAPNEGKTADGLVFFLAPV 127
Query: 121 DFDFPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNV-WDPSYPSNSPHVGIDINS 179
P K GG LGLF K+ +NQ+VAVEFD+F NV WDP N H+GID+NS
Sbjct: 128 GSTQP--LKGGGLLGLF--KDESYNKSNQIVAVEFDTFRNVAWDP----NGIHMGIDVNS 179
Query: 180 IRSVATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVL 239
I+SV T W GE+ ISY +++K L+ +VYP+ ++ ++ VDL VL
Sbjct: 180 IQSVRTVRWD---WANGEVANVFISYEASTKSLTASLVYPS--LEKSFILSAIVDLKKVL 234
Query: 240 SDWVLVGFTAATG---SVSETHDILSWAFNSFL 269
+WV VGFTA TG +T+D+LSW+F S L
Sbjct: 235 PEWVRVGFTATTGLSEDYVQTNDVLSWSFESNL 267
>CONA_CANEN (P02866) Concanavalin A precursor (Con A)
Length = 290
Score = 161 bits (407), Expect = 2e-39
Identities = 105/285 (36%), Positives = 152/285 (52%), Gaps = 31/285 (10%)
Query: 3 NSFSISLLTTLLPFILLITTVKS-----DSFSFNFPRFESDTKTILSDGDANT-TGGVLQ 56
+S + + T + F++++ V S ++ F F +F D K ++ GDA T T G L+
Sbjct: 7 SSLFLPIFTFITMFLMVVNKVSSSTHETNALHFMFNQFSKDQKDLILQGDATTGTDGNLE 66
Query: 57 LTKKDLYGIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFY 116
LT+ G P +S G+ F+ VH+ ++ + VA F F+F++ S P DG F+
Sbjct: 67 LTRVSSNGSPQGSSVGRALFYAPVHIWES-SAVVASFEATFTFLIKSPDSHP-ADGIAFF 124
Query: 117 IAGLDFDFPEDPKEGGFLGLFDPKNAFNTS----------ANQVVAVEFDSF--TNVWDP 164
I+ +D P G LGLF N S A+ +VAVE D++ T++ DP
Sbjct: 125 ISNIDSSIPSG-STGRLLGLFPDANVIRNSTTIDFNAAYNADTIVAVELDTYPNTDIGDP 183
Query: 165 SYPSNSPHVGIDINSIRSVATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVK 224
SYP H+GIDI S+RS TA W + G++G A I Y S K LS V YPN+
Sbjct: 184 SYP----HIGIDIKSVRSKKTAKWNMQ---NGKVGTAHIIYNSVDKRLSAVVSYPNAD-- 234
Query: 225 IETFVAYEVDLGAVLSDWVLVGFTAATGSVSETHDILSWAFNSFL 269
V+Y+VDL VL +WV VG +A+TG ET+ ILSW+F S L
Sbjct: 235 -SATVSYDVDLDNVLPEWVRVGLSASTGLYKETNTILSWSFTSKL 278
>LEC_ERYCG (P83410) Lectin (ECL)
Length = 239
Score = 160 bits (406), Expect = 2e-39
Identities = 101/250 (40%), Positives = 147/250 (58%), Gaps = 20/250 (8%)
Query: 26 DSFSFNFPRFESDTKTILSDGDANTT-GGVLQLTKKDLYGIPSKNSFGQTTFFGAVHLSD 84
++ SF+F FE + G A T GVLQLTK + G+P+ +S G+T + VH+ D
Sbjct: 2 ETISFSFSEFEPGNDNLTLQGAALITQSGVLQLTKINQNGMPAWDSTGRTLYTKPVHMWD 61
Query: 85 NRTGKVADFTTEFSFVVNPKGSEP-HGDGFTFYIAGLDFDFPEDPKEG-GFLGLFDPKNA 142
+ TG VA F T FSF + + P DG F++ P +G G+LG+F+ N+
Sbjct: 62 STTGTVASFETRFSFSIEQPYTRPLPADGLVFFMG----PTKSKPAQGYGYLGVFN--NS 115
Query: 143 FNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDINSIRSVATAPWPIDLVPQGEIGKAL 202
++ Q +AVEFD+F+N WD P PH+GID+NSIRS+ T P+ +D G++ +
Sbjct: 116 KQDNSYQTLAVEFDTFSNPWD---PPQVPHIGIDVNSIRSIKTQPFQLD---NGQVANVV 169
Query: 203 ISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDWVLVGFTAATGS---VSETHD 259
I Y + SK+L V +VYP+S I T +A VD+ VL DWV VG + ATG+ +ETHD
Sbjct: 170 IKYDAPSKILHVVLVYPSSGA-IYT-IAEIVDVKQVLPDWVDVGLSGATGAQRDAAETHD 227
Query: 260 ILSWAFNSFL 269
+ SW+F + L
Sbjct: 228 VYSWSFQASL 237
>LECS_VATMA (P81371) Seed lectin (VML)
Length = 240
Score = 160 bits (406), Expect = 2e-39
Identities = 99/250 (39%), Positives = 141/250 (55%), Gaps = 23/250 (9%)
Query: 25 SDSFSFNFPRFESDTKTILSDGDANTTG-GVLQLTK-KDLYGIPSKNSFGQTTFFGAVHL 82
S+ SF+F +F + K I+ GDA T G LQLTK KD G P +S G+ + +H+
Sbjct: 1 SEVVSFSFTKFNPNPKDIILQGDALVTSKGKLQLTKVKD--GKPVDHSLGRALYAAPIHI 58
Query: 83 SDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFDFPEDPKEGGFLGLFDPKNA 142
D+ T +VA F T FSFVV DG F++A D + K+GGFLGLF+ N
Sbjct: 59 WDDSTDRVASFATSFSFVVEAPDESKTADGIAFFLAPPD---TQPQKDGGFLGLFNDSN- 114
Query: 143 FNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDINSIRSVATAPWPIDLVPQGEIGKAL 202
+ Q VAVEFD+F+N WDPS + H+GI++NSI S+ W + G++
Sbjct: 115 ---KSIQTVAVEFDTFSNTWDPS----ARHIGINVNSIESMKYVKWGWE---NGKVANVY 164
Query: 203 ISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDWVLVGFTAATG---SVSETHD 259
ISY +++K L+ + YP++ V+ VDL + L +WV VGF+A +G ETHD
Sbjct: 165 ISYEASTKTLTASLTYPSNATSY--IVSANVDLKSALPEWVRVGFSATSGLSRDHVETHD 222
Query: 260 ILSWAFNSFL 269
+L W+F S L
Sbjct: 223 VLDWSFTSTL 232
>LCS2_ROBPS (Q41161) Seed agglutinin II precursor (RPSAII)
(LECRPAS2)
Length = 285
Score = 160 bits (405), Expect = 3e-39
Identities = 107/273 (39%), Positives = 155/273 (56%), Gaps = 22/273 (8%)
Query: 3 NSFSISLLTTLLPFILLITTVKSD-SFSFNFPRFESDTKTILSDGDANTTG-GVLQLTKK 60
NSF + LL+ F+LL+ V S S SF+FP+F + ++ DA T GVLQLT
Sbjct: 10 NSFLL-LLSISFFFLLLLNKVNSTGSLSFSFPKFAPNQPYLIFQRDALVTSTGVLQLTNV 68
Query: 61 DLYGIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGL 120
+ G+PS+ S G+ + + D+ TG VA F T FSF++ DG F++A +
Sbjct: 69 -VNGVPSRKSLGRALYAAPFQIWDSTTGNVASFVTSFSFIIQAPNPATTADGLAFFLAPV 127
Query: 121 DFDFPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTN-VWDPSYPSNSPHVGIDINS 179
D P D GG LG+F KN + +NQ+VAVEFD+F+N WDP+ H+GI++NS
Sbjct: 128 DTQ-PLD--LGGMLGIF--KNGYFNKSNQIVAVEFDTFSNRHWDPT----GRHMGINVNS 178
Query: 180 IRSVATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVL 239
I SV T PW GE+ ISY +++K L+ +VYP+ ++ + VD+ VL
Sbjct: 179 IVSVKTVPWN---WANGEVANVFISYEASTKSLTASLVYPS--LETSFIIHAIVDVKDVL 233
Query: 240 SDWVLVGFTAATG---SVSETHDILSWAFNSFL 269
+WV GF+A TG +T+D+LSW+F S L
Sbjct: 234 PEWVRFGFSATTGIDTGYVQTNDVLSWSFESNL 266
>CONA_CANGL (P14894) Concanavalin A precursor (Con A)
Length = 290
Score = 159 bits (403), Expect = 5e-39
Identities = 104/285 (36%), Positives = 151/285 (52%), Gaps = 31/285 (10%)
Query: 3 NSFSISLLTTLLPFILLITTVKS-----DSFSFNFPRFESDTKTILSDGDANT-TGGVLQ 56
+S + + T + F++++ V S ++ F F +F D K ++ GDA T T G L+
Sbjct: 7 SSLFLPIFTFITMFLMVVNKVSSSTHETNALHFMFNQFSKDQKDLILQGDATTGTDGNLE 66
Query: 57 LTKKDLYGIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFY 116
LT+ G P +S G+ F+ VH+ ++ + VA F F+F++ S P DG F+
Sbjct: 67 LTRVSSNGSPQGSSVGRALFYAPVHIWES-SAVVASFDATFTFLIKSPDSHP-ADGIAFF 124
Query: 117 IAGLDFDFPEDPKEGGFLGLFDPKNAFNTS----------ANQVVAVEFDSF--TNVWDP 164
I+ +D P G LGLF N S A+ +VAVE D++ T++ DP
Sbjct: 125 ISNIDSSIPSG-STGRLLGLFPDANVIRNSTTIDFNAAYNADTIVAVELDTYPNTDIGDP 183
Query: 165 SYPSNSPHVGIDINSIRSVATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVK 224
+YP H+GIDI S+RS TA W + G++G A I Y S K LS V YPN
Sbjct: 184 NYP----HIGIDIKSVRSKKTAKWNMQ---NGKVGTAHIIYNSVGKRLSAVVSYPNGD-- 234
Query: 225 IETFVAYEVDLGAVLSDWVLVGFTAATGSVSETHDILSWAFNSFL 269
V+Y+VDL VL +WV VG +A+TG ET+ ILSW+F S L
Sbjct: 235 -SATVSYDVDLDNVLPEWVRVGLSASTGLYKETNTILSWSFTSKL 278
>LCS1_ROBPS (Q41162) Seed agglutinin I precursor (RPSAI) (LECRPAS1)
Length = 285
Score = 155 bits (393), Expect = 7e-38
Identities = 105/273 (38%), Positives = 153/273 (55%), Gaps = 22/273 (8%)
Query: 3 NSFSISLLTTLLPFILLITTVKSD-SFSFNFPRFESDTKTILSDGDANTTG-GVLQLTKK 60
NSF + LL+ F+LL+ V S S SF+FP+F + ++ DA T GVLQLT
Sbjct: 10 NSFPL-LLSISFFFLLLLNKVNSTGSLSFSFPKFAPNQPYLIFQRDALVTSTGVLQLTNV 68
Query: 61 DLYGIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGL 120
+ G+P + S G+ + + DN TG VA F T FSF++ DG F++A +
Sbjct: 69 -VNGVPPRRSIGRALYAAPFQIWDNTTGNVASFVTSFSFIIQAPNPATTADGLAFFLAPV 127
Query: 121 DFDFPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNV-WDPSYPSNSPHVGIDINS 179
D P D GG LG+F K+ +NQ+VAVEFD+F+N+ +DP H+GI++NS
Sbjct: 128 DTQ-PGD--LGGMLGIF--KDGSYNKSNQIVAVEFDTFSNIHFDP----KGRHMGINVNS 178
Query: 180 IRSVATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVL 239
I SV T PW GE+ ISY +++K L+ +VYP+ ++ + VD+ VL
Sbjct: 179 IVSVKTVPWN---WTNGEVANVFISYEASTKSLNASLVYPS--LETSFIIHAIVDVKDVL 233
Query: 240 SDWVLVGFTAATG---SVSETHDILSWAFNSFL 269
+WV GF+A TG +T+D+LSW+F S L
Sbjct: 234 PEWVRFGFSATTGIDTGYVQTNDVLSWSFESNL 266
>LEC_VICFA (P02871) Favin (Lectin)
Length = 233
Score = 154 bits (389), Expect = 2e-37
Identities = 95/243 (39%), Positives = 138/243 (56%), Gaps = 22/243 (9%)
Query: 29 SFNFPRFESDTKTILSDGDANTTGGVLQLTKKDLYGIPSKNSFGQTTFFGAVHLSDNRTG 88
SF+ P+F D ++ G TT L LTK KN+ G+ + +H+ D+ TG
Sbjct: 6 SFSIPKFRPDQPNLIFQGGGYTTKEKLTLTKA------VKNTVGRALYSLPIHIWDSETG 59
Query: 89 KVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFDFPEDPKEGGFLGLFDPKNAFNTSAN 148
VADFTT F FV++ DGFTF+IA +D + GG+LG+F+ K+ T+
Sbjct: 60 NVADFTTTFIFVIDAPNGYNVADGFTFFIAPVD---TKPQTGGGYLGVFNGKDYDKTA-- 114
Query: 149 QVVAVEFDSFTN-VWDPSYPSNSPHVGIDINSIRSVATAPWPIDLVPQGEIGKALISYRS 207
Q VAVEFD+F N WDPS + H+GID+N+I+S++T W + GE IS+ +
Sbjct: 115 QTVAVEFDTFYNAAWDPS--NGKRHIGIDVNTIKSISTKSWNLQ---NGEEAHVAISFNA 169
Query: 208 ASKVLSVFVVYPNSPVKIETFVAYE-VDLGAVLSDWVLVGFTAATGSVSETHDILSWAFN 266
+ VLSV ++YPN + + E V L V+ +WV +GF+A TG+ TH++LSW F
Sbjct: 170 TTNVLSVTLLYPN----LTGYTLSEVVPLKDVVPEWVRIGFSATTGAEYATHEVLSWTFL 225
Query: 267 SFL 269
S L
Sbjct: 226 SEL 228
>LCB1_ROBPS (Q41159) Bark agglutinin I, polypeptide A precursor
(RPbAI) (LECRPA1)
Length = 285
Score = 154 bits (388), Expect = 3e-37
Identities = 102/267 (38%), Positives = 150/267 (55%), Gaps = 21/267 (7%)
Query: 9 LLTTLLPFILLITTVKSD-SFSFNFPRFESDTKTILSDGDANTTG-GVLQLTKKDLYGIP 66
LL+ F+LL+ V S S SF+FP+F + ++ DA T GVLQLT + G+P
Sbjct: 15 LLSISFFFLLLLNKVNSTGSLSFSFPKFAPNQPYLIFQRDALVTSTGVLQLTNV-VNGVP 73
Query: 67 SKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFDFPE 126
S S G+ + + D+ TG VA F T FSF++ DG F++A +D P
Sbjct: 74 SGKSLGRALYAAPFQIWDSTTGNVASFVTSFSFIIQAPNPTTTADGLAFFLAPVDTQ-PL 132
Query: 127 DPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNV-WDPSYPSNSPHVGIDINSIRSVAT 185
D GG LG+F K+ + +NQ+VAVEFD+F+N+ +DP H+GI++NSI S+ T
Sbjct: 133 D--VGGMLGIF--KDGYFNKSNQIVAVEFDTFSNIHFDP----KGRHMGINVNSIVSIKT 184
Query: 186 APWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDWVLV 245
PW GE+ ISY +++K L+ +VYP+ ++ V VD+ VL +WV
Sbjct: 185 VPWN---WTNGEVANVFISYEASTKSLTASLVYPS--LETSFIVHAIVDVKDVLPEWVRF 239
Query: 246 GFTAATG---SVSETHDILSWAFNSFL 269
GF+A TG +T+D+LSW+F S L
Sbjct: 240 GFSATTGIDKGYVQTNDVLSWSFESNL 266
>LEC2_ULEEU (P22973) Anti-H(O) lectin II (UEA-II)
Length = 249
Score = 153 bits (386), Expect = 5e-37
Identities = 99/253 (39%), Positives = 141/253 (55%), Gaps = 21/253 (8%)
Query: 25 SDSFSFNFPRFESDTKTILSDGDAN-TTGGVLQLTKKDLYGIPSKNSFGQTTFFGAVHLS 83
SD SFNF +F + K I+ GDA+ +T GVL++TK P+ S G+ + + +
Sbjct: 3 SDDLSFNFDKFVPNQKNIIFQGDASVSTKGVLEVTKVSK---PTTRSIGRALYAAPIQIW 59
Query: 84 DNRTGKVADFTTEFSFVVNPKGSEPHG--DGFTFYIAGLDFDFPEDPKEGGFLGLFDPKN 141
D+ TGKVA F T FSFVV + E DG F++A + P G F GLF N
Sbjct: 60 DSITGKVASFATSFSFVVKDEPDEKIDGVDGLAFFLAPANSQIPSGSSAGMF-GLFCSSN 118
Query: 142 AFNTSANQVVAVEFDSFT----NVWDPSYPSNSPHVGIDINSIRSVATAPWPIDLVPQGE 197
+ S+NQ++AVEFDS+ N WDP + H+GID+NSI+S+ T D GE
Sbjct: 119 D-SKSSNQIIAVEFDSYFGKTYNPWDPDFK----HIGIDVNSIKSIKTVK---DDWRNGE 170
Query: 198 IGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDWVLVGFTAATGSVSE- 256
+ +I+YR+ +K L+V + YP+ A VDL A+L +WV VGF+ G+ ++
Sbjct: 171 VADVVITYRAPTKSLTVSLSYPSDGTS-NIVTASSVDLKAILPEWVSVGFSGGVGNAAKF 229
Query: 257 THDILSWAFNSFL 269
HD+LSW F S L
Sbjct: 230 DHDVLSWYFTSNL 242
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,644,445
Number of Sequences: 164201
Number of extensions: 1458925
Number of successful extensions: 3436
Number of sequences better than 10.0: 79
Number of HSP's better than 10.0 without gapping: 68
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 3084
Number of HSP's gapped (non-prelim): 93
length of query: 269
length of database: 59,974,054
effective HSP length: 108
effective length of query: 161
effective length of database: 42,240,346
effective search space: 6800695706
effective search space used: 6800695706
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0541.11


