

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
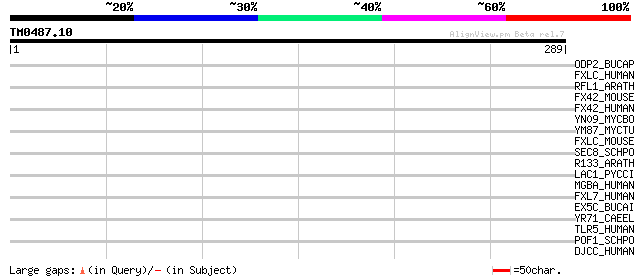
Query= TM0487.10
(289 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
ODP2_BUCAP (Q8K9T8) Dihydrolipoyllysine-residue acetyltransferas... 35 0.16
FXLC_HUMAN (Q9NXK8) F-box/LRR-repeat protein 12 (F-box and leuci... 32 1.8
RFL1_ARATH (Q8L3R3) Disease resistance protein RFL1 (RPS5-like p... 32 2.4
FX42_MOUSE (Q6PDJ6) F-box only protein 42 31 3.1
FX42_HUMAN (Q6P3S6) F-box only protein 42 31 3.1
YN09_MYCBO (P65527) Putative Na(+)/H(+) exchanger Mb2309 31 4.0
YM87_MYCTU (P65526) Putative Na(+)/H(+) exchanger Rv2287/MT2345 31 4.0
FXLC_MOUSE (Q9EPX5) F-box/LRR-repeat protein 12 (F-box and leuci... 31 4.0
SEC8_SCHPO (O74562) Exocyst complex component sec8 30 5.3
R133_ARATH (Q9STE7) Putative disease resistance RPP13-like prote... 30 5.3
LAC1_PYCCI (O59896) Laccase precursor (EC 1.10.3.2) (Benzenediol... 30 5.3
MGBA_HUMAN (Q13296) Mammaglobin A precursor (Mammaglobin 1) (Sec... 30 6.9
FXL7_HUMAN (Q9UJT9) F-box/LRR-repeat protein 7 (F-box and leucin... 30 6.9
EX5C_BUCAI (P57528) Exodeoxyribonuclease V gamma chain (EC 3.1.1... 30 6.9
YR71_CAEEL (Q09564) Hypothetical protein F43C1.1 in chromosome III 30 9.0
TLR5_HUMAN (O60602) Toll-like receptor 5 precursor (Toll/interle... 30 9.0
POF1_SCHPO (P87053) F-box/WD-repeat protein pof1 (Skp1-binding p... 30 9.0
DJCC_HUMAN (Q9UKB3) DnaJ homolog subfamily C member 12 (J domain... 30 9.0
>ODP2_BUCAP (Q8K9T8) Dihydrolipoyllysine-residue acetyltransferase
component of pyruvate dehydrogenase complex (EC
2.3.1.12) (E2) (Dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase complex)
Length = 402
Score = 35.4 bits (80), Expect = 0.16
Identities = 48/209 (22%), Positives = 88/209 (41%), Gaps = 16/209 (7%)
Query: 81 KLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICIYSLSPLNLSSIFSCKTLVVLKLE 140
K M S S + ++ + + V+ I I+ ++ NL+SI + K L +K E
Sbjct: 35 KASMEIPSPISGIVKDITIKIGEKVKTSSI----IMIFKVN--NLNSIKNEKDLNYIKTE 88
Query: 141 G-LDLNTTVSSVDLPFLKVLHLQ-VLQLSQRRCLAQLLNGCPVLEDLKAKAIYFKKYKET 198
L+ N D+ + ++H V++ R L N P + + Y
Sbjct: 89 KKLNENFLEEKKDIKKIVLVHATPVVRRLARHLNVDLKNITPSGPKNRILKEDIELYIRN 148
Query: 199 GFEYKTLSKLVRADISGTSTDLFRMEVINNVDFL---RIHEIYIRIPHDDQFTDMFHNLT 255
K+ + + + + DLF I N+ + +H+ ++ IPH QF ++ N+T
Sbjct: 149 NTSNKSSFNIEKNNTTNFHKDLFNEIPITNIQQIIGKNLHQNWVNIPHVTQFDEV--NIT 206
Query: 256 HIELGYYDYNTNWIEVVEFLNYCPKLQVL 284
+E + YNT E + N C K+ +L
Sbjct: 207 LLEEFRHKYNT---EKKQKNNMCSKITIL 232
>FXLC_HUMAN (Q9NXK8) F-box/LRR-repeat protein 12 (F-box and
leucine-rich repeat protein 12) (F-box protein FBL12)
Length = 326
Score = 32.0 bits (71), Expect = 1.8
Identities = 20/58 (34%), Positives = 25/58 (42%), Gaps = 9/58 (15%)
Query: 8 LPDEILCYILSFLPTKKAVATSILSKRWKP------LWRSVTTLDFDEANFRRTKGWH 59
LPD +L I S+LP + + S + RWK LWR V D R WH
Sbjct: 7 LPDSVLLEIFSYLPVRDRIRISRVCHRWKRLVDDRWLWRHV---DLTLYTMRPKVMWH 61
>RFL1_ARATH (Q8L3R3) Disease resistance protein RFL1 (RPS5-like
protein 1) (pNd13/pNd14)
Length = 885
Score = 31.6 bits (70), Expect = 2.4
Identities = 33/123 (26%), Positives = 51/123 (40%), Gaps = 18/123 (14%)
Query: 96 NVNVWVNAAVQRGGFEHVDICIYSLSPLNLSSIFSCKTLVVLKLEGLDLNTTVSSVDLPF 155
N+ VW G +H++ I S +S+ + L KLE L+L L
Sbjct: 770 NLRVW--------GCKHLEDII---SKEKAASVLEKEILPFQKLECLNL------YQLSE 812
Query: 156 LKVLHLQVLQLSQRRCLAQLLNGCPVLEDLKAKAIYFKKYKETGFEYKTLSKLVRADISG 215
LK ++ L + RCL +LN CP L L + K +E +YK + R +
Sbjct: 813 LKSIYWNALPFQRLRCL-DILNNCPKLRKLPLDSKSVVKVEEFVIKYKEKKWIERVEWED 871
Query: 216 TST 218
+T
Sbjct: 872 EAT 874
>FX42_MOUSE (Q6PDJ6) F-box only protein 42
Length = 717
Score = 31.2 bits (69), Expect = 3.1
Identities = 16/39 (41%), Positives = 26/39 (66%), Gaps = 1/39 (2%)
Query: 5 ISSLPDEILCYILSFL-PTKKAVATSILSKRWKPLWRSV 42
+S LP+E+L YILSFL P ++ +++ K+W L + V
Sbjct: 47 MSELPEEVLEYILSFLSPYQEHKTAALVCKQWYRLIKGV 85
>FX42_HUMAN (Q6P3S6) F-box only protein 42
Length = 717
Score = 31.2 bits (69), Expect = 3.1
Identities = 16/39 (41%), Positives = 26/39 (66%), Gaps = 1/39 (2%)
Query: 5 ISSLPDEILCYILSFL-PTKKAVATSILSKRWKPLWRSV 42
+S LP+E+L YILSFL P ++ +++ K+W L + V
Sbjct: 47 MSELPEEVLEYILSFLSPYQEHKTAALVCKQWYRLIKGV 85
>YN09_MYCBO (P65527) Putative Na(+)/H(+) exchanger Mb2309
Length = 542
Score = 30.8 bits (68), Expect = 4.0
Identities = 27/91 (29%), Positives = 42/91 (45%), Gaps = 3/91 (3%)
Query: 128 IFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLSQRRCLAQLLNGCPVLEDLKA 187
+ S LV + ++G L T V +P V H LQL++ R L+ P + D
Sbjct: 395 VVSVVILVTVLVQGTSLPTVVRWARMPE-DVAHANELQLARTRSAQAALDALPTVADELG 453
Query: 188 KAIYFKKYKETGFEYKTLSKLVRADISGTST 218
A K+ E EY+ + LV AD + ++T
Sbjct: 454 VAPDLVKHLEK--EYEERAVLVMADGADSAT 482
>YM87_MYCTU (P65526) Putative Na(+)/H(+) exchanger Rv2287/MT2345
Length = 542
Score = 30.8 bits (68), Expect = 4.0
Identities = 27/91 (29%), Positives = 42/91 (45%), Gaps = 3/91 (3%)
Query: 128 IFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLSQRRCLAQLLNGCPVLEDLKA 187
+ S LV + ++G L T V +P V H LQL++ R L+ P + D
Sbjct: 395 VVSVVILVTVLVQGTSLPTVVRWARMPE-DVAHANELQLARTRSAQAALDALPTVADELG 453
Query: 188 KAIYFKKYKETGFEYKTLSKLVRADISGTST 218
A K+ E EY+ + LV AD + ++T
Sbjct: 454 VAPDLVKHLEK--EYEERAVLVMADGADSAT 482
>FXLC_MOUSE (Q9EPX5) F-box/LRR-repeat protein 12 (F-box and
leucine-rich repeat protein 12) (F-box protein FBL12)
Length = 326
Score = 30.8 bits (68), Expect = 4.0
Identities = 20/58 (34%), Positives = 25/58 (42%), Gaps = 9/58 (15%)
Query: 8 LPDEILCYILSFLPTKKAVATSILSKRWKP------LWRSVTTLDFDEANFRRTKGWH 59
LPD +L I S+LP + + S + RWK LWR V D R WH
Sbjct: 7 LPDLVLLEIFSYLPVRDRIRISRVCHRWKRLVDDRWLWRHV---DLTLYTMRPKVMWH 61
>SEC8_SCHPO (O74562) Exocyst complex component sec8
Length = 1088
Score = 30.4 bits (67), Expect = 5.3
Identities = 22/101 (21%), Positives = 47/101 (45%), Gaps = 9/101 (8%)
Query: 28 TSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSRFVQSVHRVIHTPDFL--------QPI 79
T +K ++ S+ L FD R + +++ S+++ ++ PD++ I
Sbjct: 869 TDFYKSAYKEVFDSLQRLQFDALLLIRMEV-RLQYIHSINQSVNLPDYVVEYRGRPDASI 927
Query: 80 QKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICIYSL 120
L + V+ + ++ + +N W V +G E VD +YS+
Sbjct: 928 MALNSTIVTTNLKLETCLNEWERRFVFQGLSELVDSSLYSI 968
>R133_ARATH (Q9STE7) Putative disease resistance RPP13-like protein
3
Length = 847
Score = 30.4 bits (67), Expect = 5.3
Identities = 31/113 (27%), Positives = 44/113 (38%), Gaps = 10/113 (8%)
Query: 122 PLNLSSIFSCKTLVVLKLE------GLDLNTTVSSVDLPFLKVLHLQVLQLSQRRCLAQL 175
PLN S K L VLKLE + TT+ VD+ F + L ++ +
Sbjct: 701 PLNFVSFSKPKNLRVLKLEMRNFKLSSESRTTIGLVDVNFPSLESLTLVGTTLEENSMPA 760
Query: 176 LNGCPVLEDLKAKAIYFKKYKETGFEYKTLSKLVRADIS----GTSTDLFRME 224
L P LEDL K + K + +L ++S G D R+E
Sbjct: 761 LQKLPRLEDLVLKDCNYSGVKIMSISAQGFGRLKNLEMSMERRGHGLDELRIE 813
>LAC1_PYCCI (O59896) Laccase precursor (EC 1.10.3.2)
(Benzenediol:oxygen oxidoreductase) (Urishiol oxidase)
(Ligninolytic phenoloxidase)
Length = 518
Score = 30.4 bits (67), Expect = 5.3
Identities = 20/67 (29%), Positives = 30/67 (43%), Gaps = 16/67 (23%)
Query: 213 ISGTSTDLFRMEVINNVDFLRIHEIYIRIPHDDQFTDMFHNLTHIELGYYDYNTNWIEVV 272
I+G D F++ VI+N+ H T H H G++ + TNW + V
Sbjct: 57 IAGQKGDRFQLNVIDNLT-----------NHTMLKTTSIH--WH---GFFQHGTNWADGV 100
Query: 273 EFLNYCP 279
F+N CP
Sbjct: 101 SFVNQCP 107
>MGBA_HUMAN (Q13296) Mammaglobin A precursor (Mammaglobin 1)
(Secretoglobin family 2A member 2)
Length = 93
Score = 30.0 bits (66), Expect = 6.9
Identities = 25/98 (25%), Positives = 50/98 (50%), Gaps = 7/98 (7%)
Query: 156 LKVLHLQVLQLSQRRCLAQLLNGCPVLEDLKAKAIYFKKYKETGFEYK-TLSKLVRADIS 214
+K+L + +L + C A +GCP+LE++ +K I + K EYK L + + + +
Sbjct: 1 MKLLMVLMLAALSQHCYAG--SGCPLLENVISKTINPQVSKT---EYKELLQEFIDDNAT 55
Query: 215 GTSTDLFRMEVINNVD-FLRIHEIYIRIPHDDQFTDMF 251
+ D + +N D L E+++++ +D D+F
Sbjct: 56 TNAIDELKECFLNQTDETLSNVEVFMQLIYDSSLCDLF 93
>FXL7_HUMAN (Q9UJT9) F-box/LRR-repeat protein 7 (F-box and
leucine-rich repeat protein 7) (F-box protein FBL6/FBL7)
Length = 491
Score = 30.0 bits (66), Expect = 6.9
Identities = 18/56 (32%), Positives = 27/56 (48%), Gaps = 10/56 (17%)
Query: 5 ISSLPDEILCYILSFLPTKKAVATSILSKRW------KPLWRSV----TTLDFDEA 50
I LPD + I SFLPT + + + +RW LWR++ T++ D A
Sbjct: 114 IDRLPDHSMVQIFSFLPTNQLCRCARVCRRWYNLAWDPRLWRTIRLTGETINVDRA 169
>EX5C_BUCAI (P57528) Exodeoxyribonuclease V gamma chain (EC
3.1.11.5)
Length = 1070
Score = 30.0 bits (66), Expect = 6.9
Identities = 15/55 (27%), Positives = 26/55 (47%), Gaps = 3/55 (5%)
Query: 238 YIRIPHDDQFTDMFHNLTHIELGYYDYNTNWIEVVE---FLNYCPKLQVLVINQV 289
Y R + D +D F N T IE WIE+++ NY K+ + ++ ++
Sbjct: 541 YWRSLYSDLVSDFFQNNTKIEKSIQIIQKKWIEIIDDSLSSNYLKKISINILKKI 595
>YR71_CAEEL (Q09564) Hypothetical protein F43C1.1 in chromosome III
Length = 1036
Score = 29.6 bits (65), Expect = 9.0
Identities = 28/102 (27%), Positives = 47/102 (45%), Gaps = 11/102 (10%)
Query: 76 LQPIQKLRMSFVSRHSEVPS-----NVNVWVNAA-----VQRGGFEHVDICIYSLSPLNL 125
LQ +Q L +SF ++ S++P + +W A + R G + + L+
Sbjct: 265 LQNLQSLDISF-NQFSQIPPCLFHLTLEMWRLAGNNIEKIDRVGEMQIQKIDLRRNVLDT 323
Query: 126 SSIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLS 167
S + + L L + +TV +L FLKV+H + LQLS
Sbjct: 324 SFRLDIENITHLDLRDNSMISTVHLTNLRFLKVIHCERLQLS 365
>TLR5_HUMAN (O60602) Toll-like receptor 5 precursor
(Toll/interleukin-1 receptor-like protein 3)
Length = 858
Score = 29.6 bits (65), Expect = 9.0
Identities = 47/191 (24%), Positives = 88/191 (45%), Gaps = 21/191 (10%)
Query: 103 AAVQRGGFEHVDICIYSLSPLNLSSIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQ 162
A + R H+D+ + LN S +F +TL LK+ L N D F + +LQ
Sbjct: 283 AGLARSSVRHLDLSHGFVFSLN-SRVF--ETLKDLKVLNLAYNKINKIADEAFYGLDNLQ 339
Query: 163 VLQLSQRRCLAQLLN----GCPVLEDLKAKAIYFKKYKETGFEYKTLSKLVRADISGTS- 217
VL LS L +L + G P + + + + ++ F++ L KL D+ +
Sbjct: 340 VLNLSYN-LLGELCSSNFYGLPKVAYIDLQKNHIAIIQDQTFKF--LEKLQTLDLRDNAL 396
Query: 218 TDLFRMEVINNVDFLRIHEIYIRIPHDDQFTDMFHNLTHIELGYYDYNTNWIEVVEFLNY 277
T + + I ++ FL +++ + +P ++ NL H+ + ++++ FL
Sbjct: 397 TTIHFIPSIPDI-FLSGNKL-VTLPK----INLTANLIHLS----ENRLENLDILYFLLR 446
Query: 278 CPKLQVLVINQ 288
P LQ+L++NQ
Sbjct: 447 VPHLQILILNQ 457
>POF1_SCHPO (P87053) F-box/WD-repeat protein pof1 (Skp1-binding
protein 1)
Length = 605
Score = 29.6 bits (65), Expect = 9.0
Identities = 16/36 (44%), Positives = 20/36 (55%)
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPL 38
D +S LP EI ILSFL + + +SK WK L
Sbjct: 108 DFLSLLPVEISFRILSFLDARSLCQAAQVSKHWKEL 143
>DJCC_HUMAN (Q9UKB3) DnaJ homolog subfamily C member 12 (J domain
containing protein 1)
Length = 198
Score = 29.6 bits (65), Expect = 9.0
Identities = 13/30 (43%), Positives = 17/30 (56%)
Query: 33 KRWKPLWRSVTTLDFDEANFRRTKGWHSRF 62
K KPL +SV+ + D + F GWH RF
Sbjct: 150 KEPKPLEKSVSPQNSDSSGFADVNGWHLRF 179
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.325 0.139 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,476,580
Number of Sequences: 164201
Number of extensions: 1281182
Number of successful extensions: 3877
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 14
Number of HSP's that attempted gapping in prelim test: 3870
Number of HSP's gapped (non-prelim): 20
length of query: 289
length of database: 59,974,054
effective HSP length: 109
effective length of query: 180
effective length of database: 42,076,145
effective search space: 7573706100
effective search space used: 7573706100
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0487.10


