

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
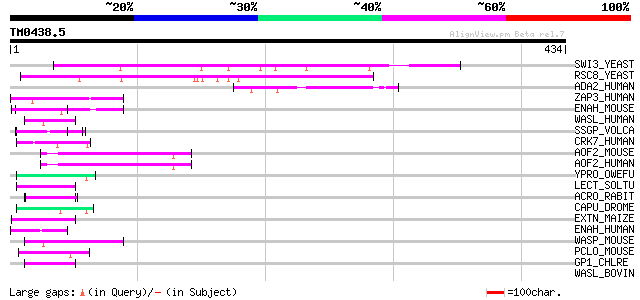
Query= TM0438.5
(434 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
SWI3_YEAST (P32591) Transcription regulatory protein SWI3 (SWI/S... 126 1e-28
RSC8_YEAST (P43609) Chromatin structure remodeling complex prote... 124 6e-28
ADA2_HUMAN (O75478) Transcriptional adapter 2-like (ADA2-like pr... 52 3e-06
ZAP3_HUMAN (P49750) Nuclear protein ZAP3 (ZAP113) 51 5e-06
ENAH_MOUSE (Q03173) Enabled protein homolog (NPC derived proline... 49 3e-05
WASL_HUMAN (O00401) Neural Wiskott-Aldrich syndrome protein (N-W... 47 1e-04
SSGP_VOLCA (P21997) Sulfated surface glycoprotein 185 precursor ... 46 2e-04
CRK7_HUMAN (Q9NYV4) Cell division cycle 2-related protein kinase... 46 2e-04
AOF2_MOUSE (Q6ZQ88) Amine oxidase flavin containing domain prote... 46 2e-04
AOF2_HUMAN (O60341) Amine oxidase flavin containing domain prote... 46 2e-04
YPRO_OWEFU (P21260) Hypothetical proline-rich protein (Fragment) 45 3e-04
LECT_SOLTU (Q9S8M0) Chitin-binding lectin 1 precursor (PL-I) 45 3e-04
ACRO_RABIT (P48038) Acrosin precursor (EC 3.4.21.10) 45 3e-04
CAPU_DROME (Q24120) Cappuccino protein 45 4e-04
EXTN_MAIZE (P14918) Extensin precursor (Proline-rich glycoprotein) 45 5e-04
ENAH_HUMAN (Q8N8S7) Enabled protein homolog 45 5e-04
WASP_MOUSE (P70315) Wiskott-Aldrich syndrome protein homolog (WASp) 44 6e-04
PCLO_MOUSE (Q9QYX7) Piccolo protein (Presynaptic cytomatrix prot... 44 8e-04
GP1_CHLRE (Q9FPQ6) Vegetative cell wall protein gp1 precursor (H... 44 8e-04
WASL_BOVIN (Q95107) Neural Wiskott-Aldrich syndrome protein (N-W... 44 0.001
>SWI3_YEAST (P32591) Transcription regulatory protein SWI3 (SWI/SNF
complex component SWI3) (Transcription factor TYE2)
Length = 825
Score = 126 bits (316), Expect = 1e-28
Identities = 94/369 (25%), Positives = 161/369 (43%), Gaps = 66/369 (17%)
Query: 35 PAKPDAPPSEPKLSATPDANVILVPSHSRWFSWDSIHQCEVRHLPEFFDSS--SKSPRVY 92
P K EP+ P A+ I++PS+S+WF+ + IH EV+ LPEFF + SK+P VY
Sbjct: 284 PKKTTITRVEPETFEIPQAHEIVIPSYSKWFNLEKIHSIEVQSLPEFFTNRIPSKTPEVY 343
Query: 93 KYYRNSIIKYFRYNPNRKITFTDVRKMLVGDVGSIRRVFEFLEAWGLINYHPSSSF---- 148
YRN ++ +R NPN + T R+ + GD ++ R+ +FL WGLINY S
Sbjct: 344 MRYRNFMVNSYRLNPNEYFSVTTARRNVSGDAAALFRLHKFLTKWGLINYQVDSKLLPKN 403
Query: 149 SKPFKWDDKDTKVDSAEPPPP----------PVRETAKRVCSGCKALCTIACFVCD---K 195
+P T+ D+ P P K++ + + T+ ++ + K
Sbjct: 404 IEPPLTSQYSTRHDAPRGLFPFESYKPSVQLPDMAKLKKMMNTSDSESTLYKYLKESKRK 463
Query: 196 YDLTLCARCYV------------RGNYRVGVSSSDFKRVEISEETKT------------- 230
YD + N V S+S + EE +T
Sbjct: 464 YDEITHPPSTTDDENGDKNDNGGKMNNEVSTSTSMTGDANLLEEGETSRPLKKVKILEQI 523
Query: 231 --EWTEKETLNLVEAISHYGDDWKRVSHQVVGRTEKECVAHFLKLPFGEQFL-----GSA 283
W++++ L++ I +G DW +V+ V ++ ++C+ FL+LP ++FL G
Sbjct: 524 DENWSKEDLQKLLKGIQEFGADWYKVAKNVGNKSPEQCILRFLQLPIEDKFLYGDGNGKG 583
Query: 284 VSDDGCELKQHADESETVASAESNKRMRLTPLADASNPIMAQAAFLSALAGPEVAQAAAQ 343
+D+G ++A P + + NP+++ AFL L P+ Q+ Q
Sbjct: 584 DNDNGLGPLKYAPH---------------LPFSKSENPVLSTIAFLVGLVNPKTVQSMTQ 628
Query: 344 AALTTLSDV 352
A+ + +
Sbjct: 629 RAIQSAESI 637
>RSC8_YEAST (P43609) Chromatin structure remodeling complex protein
RSC8 (Remodel the structure of chromatin complex subunit
8) (SWI3 homolog)
Length = 557
Score = 124 bits (310), Expect = 6e-28
Identities = 100/351 (28%), Positives = 153/351 (43%), Gaps = 75/351 (21%)
Query: 9 ATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDA--------------- 53
ATE PP++S TP P + + A A + K++ +A
Sbjct: 18 ATEEPPTTSTNTPSFPHLAQEQAKEESATLGAEVAHKKINYEQEAQKLEEKALRFLAKQT 77
Query: 54 NVILVPSHSRWFSWDSIHQCEVRHLPEFFDSSS--KSPRVYKYYRNSIIKYFRYNPNRKI 111
+ +++PS + WF IH+ E R P+FF+ SS K+P+ YK RN II +R +P +
Sbjct: 78 HPVIIPSFASWFDISKIHEIEKRSNPDFFNDSSRFKTPKAYKDTRNFIINTYRLSPYEYL 137
Query: 112 TFTDVRKMLVGDVGSIRRVFEFLEAWGLINYH------PS---SSFS------------- 149
T T VR+ + DV SI ++ FLE WGLINY PS SF+
Sbjct: 138 TITAVRRNVAMDVASIVKIHAFLEKWGLINYQIDPRTKPSLIGPSFTGHFQVVLDTPQGL 197
Query: 150 KPFKWDDKDTKV----DSAEPPPP---PVRETAKR------------------------- 177
KPF ++ + D AEP PV T K+
Sbjct: 198 KPFLPENVIKQEVEGGDGAEPQVKKEFPVNLTIKKNVYDSAQDFNALQDESRNSRQIHKV 257
Query: 178 -VCSGC-KALCTIACFVCDKYDLTLCARCYVRGNYRVGVSSSDFKRVEIS-EETKTEWTE 234
+C C + D LC+RC+ G++ SSDF R+E + K W++
Sbjct: 258 YICHTCGNESINVRYHNLRARDTNLCSRCFQEGHFGANFQSSDFIRLENNGNSVKKNWSD 317
Query: 235 KETLNLVEAISHYGDDWKRVSHQVVG-RTEKECVAHFLKLPFGEQFLGSAV 284
+E L L+E I Y D W++++ V G + ++C+ FL LP + ++ V
Sbjct: 318 QEMLLLLEGIEMYEDQWEKIADHVGGHKRVEDCIEKFLSLPIEDNYIREVV 368
>ADA2_HUMAN (O75478) Transcriptional adapter 2-like (ADA2-like
protein) (KL04P)
Length = 443
Score = 52.0 bits (123), Expect = 3e-06
Identities = 39/141 (27%), Positives = 67/141 (46%), Gaps = 23/141 (16%)
Query: 176 KRVCSGCKALCT---IACFVCDKYDLTLCARCYVRG--------NYRVGVSSSDFKRVEI 224
K C GC + I C C LC +C+ RG ++ + +SDF ++
Sbjct: 14 KPPCRGCSSYLMEPYIKCAECGPPPFFLCLQCFTRGFEYKKHQSDHTYEIMTSDFPVLDP 73
Query: 225 SEETKTEWTEKETLNLVEAISHYG-DDWKRVSHQVVGRTEKECVAHFLKLPFGEQFLGSA 283
S WT +E + L+EA+ G +W+ V++Q+ +T++EC H++K S
Sbjct: 74 S------WTAQEEMALLEAVMDCGFGNWQDVANQMCTKTKEECEKHYMKHFINNPLFAST 127
Query: 284 VSDDGCELKQHADESETVASA 304
+ LKQ A+E++T +A
Sbjct: 128 L----LNLKQ-AEEAKTADTA 143
>ZAP3_HUMAN (P49750) Nuclear protein ZAP3 (ZAP113)
Length = 1822
Score = 51.2 bits (121), Expect = 5e-06
Identities = 31/93 (33%), Positives = 44/93 (46%), Gaps = 5/93 (5%)
Query: 1 MATKSPTTATEPPPSS----SAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVI 56
M +SP + PP S S PPP PPP + P ++P PP++P S +P +
Sbjct: 141 MELESPPESPPVPPGSYMPPSQSYMPPPQPPPSYYPPTSSQPYLPPAQPSPSQSPPSQSY 200
Query: 57 LVPSHSRWFSWDSIHQCEVRHLPEFFDSSSKSP 89
L P+ S + S S Q + H + SS SP
Sbjct: 201 LAPTPS-YSSSSSSSQSYLSHSQSYLPSSQASP 232
Score = 37.4 bits (85), Expect = 0.076
Identities = 17/39 (43%), Positives = 22/39 (55%)
Query: 11 EPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSA 49
+PPP A TPPP +PPP + Q P + A P P S+
Sbjct: 424 QPPPHIQATTPPPGIPPPGVPQGIPPQLTAAPVPPASSS 462
Score = 35.0 bits (79), Expect = 0.38
Identities = 22/64 (34%), Positives = 32/64 (49%), Gaps = 8/64 (12%)
Query: 17 SAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSHSRWFSWDSIHQCEVR 76
S+ PPPP+PPP P A P+A P S+T A PS S + S+ H +++
Sbjct: 11 SSHYPPPPVPPP----PPVALPEASPGPGYSSSTTPA----APSSSGFMSFREQHLAQLQ 62
Query: 77 HLPE 80
L +
Sbjct: 63 QLQQ 66
Score = 34.7 bits (78), Expect = 0.49
Identities = 19/59 (32%), Positives = 25/59 (42%), Gaps = 1/59 (1%)
Query: 15 SSSAGTPPPP-LPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSHSRWFSWDSIHQ 72
S +A P P LPPPP+H P P P +S P + H R S + + Q
Sbjct: 1402 SETAAIPSAPVLPPPPVHSSIPPPGPVPMGMPPMSKPPPVQQTVDYGHGRDISTNKVEQ 1460
Score = 34.3 bits (77), Expect = 0.64
Identities = 16/38 (42%), Positives = 19/38 (49%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSA 49
PP SS PP+P PP P P P P +P+ SA
Sbjct: 1336 PPARSSVPVTRPPVPIPPPPPPPPLPPPPPVIKPQTSA 1373
Score = 32.7 bits (73), Expect = 1.9
Identities = 13/26 (50%), Positives = 14/26 (53%)
Query: 4 KSPTTATEPPPSSSAGTPPPPLPPPP 29
+S T PP PPPPLPPPP
Sbjct: 1339 RSSVPVTRPPVPIPPPPPPPPLPPPP 1364
Score = 32.7 bits (73), Expect = 1.9
Identities = 18/36 (50%), Positives = 19/36 (52%), Gaps = 9/36 (25%)
Query: 21 PPPPLPPPPL--------HQPQPAKPDAPPSEPKLS 48
PPPPLPPPP+ QP P P PP P LS
Sbjct: 82 PPPPLPPPPVMPGGGYGDWQP-PPPPMPPPPGPALS 116
Score = 32.3 bits (72), Expect = 2.4
Identities = 17/39 (43%), Positives = 19/39 (48%), Gaps = 3/39 (7%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSE 44
P T+ P SS PPLPPP P P P+ P SE
Sbjct: 337 PATSQVPESPSSE---EPPLPPPNEEVPPPLPPEEPQSE 372
>ENAH_MOUSE (Q03173) Enabled protein homolog (NPC derived
proline-rich protein 1) (NDPP-1)
Length = 802
Score = 48.5 bits (114), Expect = 3e-05
Identities = 32/100 (32%), Positives = 41/100 (41%), Gaps = 19/100 (19%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPD---------------APPSEPKLSA 49
S T PPP+S PPPP PPPP P P P +PP L++
Sbjct: 426 SVTYPVSPPPTSGPAAPPPPPPPPPPPPPPPLPPPPLPPLASLSHCGSQASPPPGTPLAS 485
Query: 50 TPDANVILVPSHSRWFSWDSIHQCEVRHLPEFFDSSSKSP 89
TP + ++PS S + E PE DSS+ P
Sbjct: 486 TPSSKPSVLPSPSA----GAPASAETPLNPELGDSSASEP 521
Score = 48.1 bits (113), Expect = 4e-05
Identities = 21/44 (47%), Positives = 22/44 (49%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
A P PPP S G PPPP PPPPL P P PP+ P
Sbjct: 560 ALPPPPGPPPPPPLPSTGPPPPPPPPPPLPNQAPPPPPPPPAPP 603
Score = 38.9 bits (89), Expect = 0.026
Identities = 18/41 (43%), Positives = 19/41 (45%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPP 42
A+ P PPP T PPP PPPP P A P PP
Sbjct: 558 ASALPPPPGPPPPPPLPSTGPPPPPPPPPPLPNQAPPPPPP 598
Score = 37.7 bits (86), Expect = 0.058
Identities = 18/46 (39%), Positives = 21/46 (45%), Gaps = 4/46 (8%)
Query: 6 PTTATEPP----PSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKL 47
P PP P+ ++ PPPP PPPP P P PP P L
Sbjct: 543 PAPPPPPPLPSGPAYASALPPPPGPPPPPPLPSTGPPPPPPPPPPL 588
Score = 37.0 bits (84), Expect = 0.099
Identities = 21/48 (43%), Positives = 22/48 (45%), Gaps = 7/48 (14%)
Query: 8 TATEPPPSSSAGTPPPPL----PPPPLHQPQPAKPDAPPSEPKLSATP 51
+A PPP PPPPL PPPP P P APP P A P
Sbjct: 559 SALPPPPGPP---PPPPLPSTGPPPPPPPPPPLPNQAPPPPPPPPAPP 603
Score = 33.9 bits (76), Expect = 0.84
Identities = 17/46 (36%), Positives = 20/46 (42%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P + P +S+ PP P PPPPL P P PP A P
Sbjct: 549 PPLPSGPAYASALPPPPGPPPPPPLPSTGPPPPPPPPPPLPNQAPP 594
Score = 32.3 bits (72), Expect = 2.4
Identities = 18/51 (35%), Positives = 18/51 (35%), Gaps = 8/51 (15%)
Query: 9 ATEPPPSSSAGTPPPPLPPPPLHQ--------PQPAKPDAPPSEPKLSATP 51
A P P PP P PPPPL P P P PP P P
Sbjct: 530 AESPTPQGLVLGPPAPPPPPPLPSGPAYASALPPPPGPPPPPPLPSTGPPP 580
Score = 31.6 bits (70), Expect = 4.2
Identities = 17/49 (34%), Positives = 17/49 (34%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDAN 54
P PPP S LPPPP P P P P P P N
Sbjct: 542 PPAPPPPPPLPSGPAYASALPPPPGPPPPPPLPSTGPPPPPPPPPPLPN 590
>WASL_HUMAN (O00401) Neural Wiskott-Aldrich syndrome protein
(N-WASP)
Length = 505
Score = 46.6 bits (109), Expect = 1e-04
Identities = 21/46 (45%), Positives = 23/46 (49%), Gaps = 6/46 (13%)
Query: 12 PPPSSSAGTPPPPL------PPPPLHQPQPAKPDAPPSEPKLSATP 51
PPP S+G PPPP PPPP P A P PPS P + P
Sbjct: 291 PPPPHSSGPPPPPARGRGAPPPPPSRAPTAAPPPPPPSRPSVEVPP 336
Score = 39.3 bits (90), Expect = 0.020
Identities = 21/51 (41%), Positives = 25/51 (48%), Gaps = 6/51 (11%)
Query: 3 TKSPTTATEPPPSS--SAGTPPPP----LPPPPLHQPQPAKPDAPPSEPKL 47
+++PT A PPP S S PPPP PPPP P A PP P +
Sbjct: 315 SRAPTAAPPPPPPSRPSVEVPPPPPNRMYPPPPPALPSSAPSGPPPPPPSV 365
Score = 38.1 bits (87), Expect = 0.044
Identities = 22/47 (46%), Positives = 24/47 (50%), Gaps = 8/47 (17%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQ---PQPAK----PDAPPSEPKLSATP 51
PPP S G PPPP PPPP P PA+ P PPS +A P
Sbjct: 278 PPPPPSRGGPPPP-PPPPHSSGPPPPPARGRGAPPPPPSRAPTAAPP 323
Score = 35.0 bits (79), Expect = 0.38
Identities = 19/55 (34%), Positives = 24/55 (43%), Gaps = 5/55 (9%)
Query: 1 MATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANV 55
+ + +P+ PPPS P P PPPP P P PP P L + D V
Sbjct: 350 LPSSAPSGPPPPPPSVLGVGPVAPPPPPP-----PPPPPGPPPPPGLPSDGDHQV 399
Score = 35.0 bits (79), Expect = 0.38
Identities = 16/40 (40%), Positives = 19/40 (47%), Gaps = 8/40 (20%)
Query: 12 PPPSSSAGTPPPPL--------PPPPLHQPQPAKPDAPPS 43
PPP+ G PPPP PPPP +P P PP+
Sbjct: 301 PPPARGRGAPPPPPSRAPTAAPPPPPPSRPSVEVPPPPPN 340
Score = 35.0 bits (79), Expect = 0.38
Identities = 20/54 (37%), Positives = 22/54 (40%), Gaps = 14/54 (25%)
Query: 6 PTTATEPPP-----SSSAGTPPPP---------LPPPPLHQPQPAKPDAPPSEP 45
P PPP S+ +G PPPP PPPP P P P PP P
Sbjct: 339 PNRMYPPPPPALPSSAPSGPPPPPPSVLGVGPVAPPPPPPPPPPPGPPPPPGLP 392
Score = 34.3 bits (77), Expect = 0.64
Identities = 21/56 (37%), Positives = 25/56 (44%), Gaps = 7/56 (12%)
Query: 12 PPPS--SSAGTPPPPLPPP-----PLHQPQPAKPDAPPSEPKLSATPDANVILVPS 60
PPP+ SSA + PPP PP P+ P P P PP P P VP+
Sbjct: 346 PPPALPSSAPSGPPPPPPSVLGVGPVAPPPPPPPPPPPGPPPPPGLPSDGDHQVPT 401
Score = 33.5 bits (75), Expect = 1.1
Identities = 19/50 (38%), Positives = 24/50 (48%), Gaps = 13/50 (26%)
Query: 9 ATEPPPSSSAGTPPPP---------LPPPPLHQ----PQPAKPDAPPSEP 45
A PPPS + PPP +PPPP ++ P PA P + PS P
Sbjct: 309 APPPPPSRAPTAAPPPPPPSRPSVEVPPPPPNRMYPPPPPALPSSAPSGP 358
>SSGP_VOLCA (P21997) Sulfated surface glycoprotein 185 precursor
(SSG 185)
Length = 485
Score = 46.2 bits (108), Expect = 2e-04
Identities = 22/55 (40%), Positives = 24/55 (43%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVP 59
SP+ PPP PPPP PPPP P P P PP P P + VP
Sbjct: 257 SPSPPPPPPPPPPPPPPPPPSPPPPPPPPPPPPPPPPPPSPSPPRKPPSPSPPVP 311
Score = 45.8 bits (107), Expect = 2e-04
Identities = 21/41 (51%), Positives = 22/41 (53%), Gaps = 2/41 (4%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
SP + PPPS S PPPP PPPP P P P PP P
Sbjct: 247 SPRPPSPPPPSPSPPPPPPPPPPPP--PPPPPSPPPPPPPP 285
Score = 45.1 bits (105), Expect = 4e-04
Identities = 22/53 (41%), Positives = 25/53 (46%), Gaps = 1/53 (1%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPS-EPKLSATPDANVIL 57
P PPPS PPPP PPPP P P+ P PPS P + P +L
Sbjct: 268 PPPPPPPPPSPPPPPPPPPPPPPPPPPPSPSPPRKPPSPSPPVPPPPSPPSVL 320
Score = 43.1 bits (100), Expect = 0.001
Identities = 19/46 (41%), Positives = 21/46 (45%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P + PPP PPPP PPP P P P PP P S +P
Sbjct: 254 PPPSPSPPPPPPPPPPPPPPPPPSPPPPPPPPPPPPPPPPPPSPSP 299
Score = 42.0 bits (97), Expect = 0.003
Identities = 18/46 (39%), Positives = 21/46 (45%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P + + PPP PPPP PP P P P P PP P + P
Sbjct: 255 PPSPSPPPPPPPPPPPPPPPPPSPPPPPPPPPPPPPPPPPPSPSPP 300
Score = 42.0 bits (97), Expect = 0.003
Identities = 21/55 (38%), Positives = 23/55 (41%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPS 60
P PPP P PP PPPP P P P PS P+ +P V PS
Sbjct: 261 PPPPPPPPPPPPPPPPSPPPPPPPPPPPPPPPPPPSPSPPRKPPSPSPPVPPPPS 315
Score = 40.8 bits (94), Expect = 0.007
Identities = 18/46 (39%), Positives = 21/46 (45%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
PT ++ PP + PP P PP P P P P PP P S P
Sbjct: 235 PTASSRPPSPPPSPRPPSPPPPSPSPPPPPPPPPPPPPPPPPSPPP 280
Score = 38.9 bits (89), Expect = 0.026
Identities = 18/40 (45%), Positives = 19/40 (47%), Gaps = 2/40 (5%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
PPPS +PPPP P PP P P P PP P P
Sbjct: 244 PPPSPRPPSPPPPSPSPP--PPPPPPPPPPPPPPPSPPPP 281
Score = 35.8 bits (81), Expect = 0.22
Identities = 19/52 (36%), Positives = 22/52 (41%), Gaps = 5/52 (9%)
Query: 5 SPTTATEPPPSSSAGTPPPP-----LPPPPLHQPQPAKPDAPPSEPKLSATP 51
SP + P +SS PPP PPPP P P P PP P +P
Sbjct: 227 SPLPPSPQPTASSRPPSPPPSPRPPSPPPPSPSPPPPPPPPPPPPPPPPPSP 278
Score = 32.0 bits (71), Expect = 3.2
Identities = 13/41 (31%), Positives = 16/41 (38%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
+P + PP + PP PPP P P P P P
Sbjct: 223 APNNSPLPPSPQPTASSRPPSPPPSPRPPSPPPPSPSPPPP 263
Score = 32.0 bits (71), Expect = 3.2
Identities = 15/46 (32%), Positives = 17/46 (36%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P PPP +P PP PP P P P P P + P
Sbjct: 282 PPPPPPPPPPPPPPSPSPPRKPPSPSPPVPPPPSPPSVLPAATGFP 327
>CRK7_HUMAN (Q9NYV4) Cell division cycle 2-related protein kinase 7
(EC 2.7.1.37) (CDC2-related protein kinase 7) (CrkRS)
Length = 1490
Score = 45.8 bits (107), Expect = 2e-04
Identities = 26/63 (41%), Positives = 35/63 (55%), Gaps = 6/63 (9%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPA---KPDAPPSEPKLSATPDANVILVP--S 60
PT A+ PPP + TPPP PP P P PA +P PPS+P S P ++ +P +
Sbjct: 530 PTIASPPPPLPTT-TPPPQTPPLPPLPPIPALPQQPPLPPSQPAFSQVPASSTSTLPPST 588
Query: 61 HSR 63
HS+
Sbjct: 589 HSK 591
Score = 31.6 bits (70), Expect = 4.2
Identities = 14/38 (36%), Positives = 16/38 (41%)
Query: 10 TEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKL 47
T P A PP + PP P+P P PP P L
Sbjct: 1244 TPTMPQEEAAACPPHILPPEKRPPEPPGPPPPPPPPPL 1281
>AOF2_MOUSE (Q6ZQ88) Amine oxidase flavin containing domain protein
2 (AOF2 protein) (BRAF-HDAC complex protein BHC110)
Length = 853
Score = 45.8 bits (107), Expect = 2e-04
Identities = 31/122 (25%), Positives = 51/122 (41%), Gaps = 12/122 (9%)
Query: 25 LPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSHSRWFSWDSIHQCEVRHLPEFFDS 84
LPPPP P APP E S + + + + D + E P+
Sbjct: 152 LPPPP--------PQAPPEEENESEPEEPSGVEGAAFQSRLPHDRMTSQEAACFPDIISG 203
Query: 85 SSKSPRVYKYYRNSIIKYFRYNPNRKITFTDVRKMLVGDVGS----IRRVFEFLEAWGLI 140
++ +V+ + RN ++ + NP ++TF + L S + RV +LE GLI
Sbjct: 204 PQQTQKVFLFIRNRTLQLWLDNPKIQLTFEATLQQLEAPYNSDTVLVHRVHSYLERHGLI 263
Query: 141 NY 142
N+
Sbjct: 264 NF 265
>AOF2_HUMAN (O60341) Amine oxidase flavin containing domain protein
2 (AOF2 protein) (BRAF-HDAC complex protein BHC110)
Length = 852
Score = 45.8 bits (107), Expect = 2e-04
Identities = 31/122 (25%), Positives = 51/122 (41%), Gaps = 12/122 (9%)
Query: 25 LPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSHSRWFSWDSIHQCEVRHLPEFFDS 84
LPPPP P APP E S + + + + D + E P+
Sbjct: 151 LPPPP--------PQAPPEEENESEPEEPSGVEGAAFQSRLPHDRMTSQEAACFPDIISG 202
Query: 85 SSKSPRVYKYYRNSIIKYFRYNPNRKITFTDVRKMLVGDVGS----IRRVFEFLEAWGLI 140
++ +V+ + RN ++ + NP ++TF + L S + RV +LE GLI
Sbjct: 203 PQQTQKVFLFIRNRTLQLWLDNPKIQLTFEATLQQLEAPYNSDTVLVHRVHSYLERHGLI 262
Query: 141 NY 142
N+
Sbjct: 263 NF 264
>YPRO_OWEFU (P21260) Hypothetical proline-rich protein (Fragment)
Length = 141
Score = 45.4 bits (106), Expect = 3e-04
Identities = 25/67 (37%), Positives = 27/67 (39%), Gaps = 5/67 (7%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILV-----PS 60
P PPP PPPP PPPP P P P PP P A N+ L S
Sbjct: 18 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPRRARIHHNIPLFLRFFKKS 77
Query: 61 HSRWFSW 67
+S F W
Sbjct: 78 YSSHFHW 84
Score = 43.5 bits (101), Expect = 0.001
Identities = 19/43 (44%), Positives = 19/43 (44%)
Query: 3 TKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
T P PPP PPPP PPPP P P P PP P
Sbjct: 8 TPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 50
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/40 (45%), Positives = 18/40 (45%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 13 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 52
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/40 (45%), Positives = 18/40 (45%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 17 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 56
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/40 (45%), Positives = 18/40 (45%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 14 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 53
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/40 (45%), Positives = 18/40 (45%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 12 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 51
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/40 (45%), Positives = 18/40 (45%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 15 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 54
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/40 (45%), Positives = 18/40 (45%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 16 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 55
Score = 42.4 bits (98), Expect = 0.002
Identities = 19/41 (46%), Positives = 19/41 (46%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
S T PPP PPPP PPPP P P P PP P
Sbjct: 6 SLTPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 46
>LECT_SOLTU (Q9S8M0) Chitin-binding lectin 1 precursor (PL-I)
Length = 323
Score = 45.4 bits (106), Expect = 3e-04
Identities = 20/46 (43%), Positives = 24/46 (51%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P + PPPS + PP P PPPP P P+ P PS P A+P
Sbjct: 156 PPPPSPPPPSPPSPPPPSPPPPPPPSPPPPSPPPPSPSPPPPPASP 201
Score = 42.7 bits (99), Expect = 0.002
Identities = 19/40 (47%), Positives = 20/40 (49%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
PPP S P PP PPPP P P PPS P S +P
Sbjct: 155 PPPPPSPPPPSPPSPPPPSPPPPPPPSPPPPSPPPPSPSP 194
Score = 41.2 bits (95), Expect = 0.005
Identities = 19/43 (44%), Positives = 21/43 (48%), Gaps = 2/43 (4%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKL 47
SP + PPP + PPPP PPPP P P PP P L
Sbjct: 168 SPPPPSPPPPPPPS--PPPPSPPPPSPSPPPPPASPPPPPPAL 208
Score = 39.3 bits (90), Expect = 0.020
Identities = 17/40 (42%), Positives = 19/40 (47%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P + PPP S PPP PPP P P+ P P S P
Sbjct: 163 PPSPPSPPPPSPPPPPPPSPPPPSPPPPSPSPPPPPASPP 202
Score = 37.4 bits (85), Expect = 0.076
Identities = 17/42 (40%), Positives = 21/42 (49%), Gaps = 1/42 (2%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPA-KPDAPPSEP 45
SP + P P + PPPP PPP P P+ P PP+ P
Sbjct: 160 SPPPPSPPSPPPPSPPPPPPPSPPPPSPPPPSPSPPPPPASP 201
Score = 37.4 bits (85), Expect = 0.076
Identities = 15/34 (44%), Positives = 16/34 (46%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
PPP + PP P PPP P P P PP P
Sbjct: 154 PPPPPPSPPPPSPPSPPPPSPPPPPPPSPPPPSP 187
Score = 36.6 bits (83), Expect = 0.13
Identities = 18/52 (34%), Positives = 22/52 (41%), Gaps = 2/52 (3%)
Query: 6 PTTATEPPPSSSAGTPP--PPLPPPPLHQPQPAKPDAPPSEPKLSATPDANV 55
P+ + PPPS PP PP PPP P P +PP P P +
Sbjct: 164 PSPPSPPPPSPPPPPPPSPPPPSPPPPSPSPPPPPASPPPPPPALPYPQCGI 215
Score = 35.4 bits (80), Expect = 0.29
Identities = 14/30 (46%), Positives = 15/30 (49%)
Query: 16 SSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
S P PP PPPP P P+ P PP P
Sbjct: 145 SQCKLPSPPPPPPPPSPPPPSPPSPPPPSP 174
Score = 34.3 bits (77), Expect = 0.64
Identities = 17/32 (53%), Positives = 18/32 (56%), Gaps = 2/32 (6%)
Query: 5 SPTTATEPPPSSSAGTPP--PPLPPPPLHQPQ 34
SP + PPPS S PP PP PPP L PQ
Sbjct: 181 SPPPPSPPPPSPSPPPPPASPPPPPPALPYPQ 212
>ACRO_RABIT (P48038) Acrosin precursor (EC 3.4.21.10)
Length = 431
Score = 45.4 bits (106), Expect = 3e-04
Identities = 19/41 (46%), Positives = 22/41 (53%)
Query: 13 PPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDA 53
PP++ A PPPP PPPP P P P PP P + P A
Sbjct: 346 PPAAQAPPPPPPPPPPPPPPPPPPPPPPPPPPPASTKPPQA 386
Score = 43.9 bits (102), Expect = 8e-04
Identities = 18/40 (45%), Positives = 20/40 (50%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P P ++ PPPP PPPP P P P PP P S P
Sbjct: 344 PQPPAAQAPPPPPPPPPPPPPPPPPPPPPPPPPPPASTKP 383
Score = 43.1 bits (100), Expect = 0.001
Identities = 19/45 (42%), Positives = 22/45 (48%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSAT 50
P PP +A PPPP PPPP P P P PP P ++T
Sbjct: 337 PHPHPRPPQPPAAQAPPPPPPPPPPPPPPPPPPPPPPPPPPPAST 381
Score = 38.5 bits (88), Expect = 0.034
Identities = 18/41 (43%), Positives = 19/41 (45%), Gaps = 4/41 (9%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPK 46
P A PPP PPPP PPP P P P PP+ K
Sbjct: 346 PPAAQAPPPPPPPPPPPPPPPPP----PPPPPPPPPPASTK 382
Score = 37.4 bits (85), Expect = 0.076
Identities = 16/34 (47%), Positives = 16/34 (47%)
Query: 9 ATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPP 42
A PPP PPPP PPPP P P PP
Sbjct: 351 APPPPPPPPPPPPPPPPPPPPPPPPPPPASTKPP 384
Score = 36.2 bits (82), Expect = 0.17
Identities = 17/40 (42%), Positives = 19/40 (47%), Gaps = 3/40 (7%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAP 41
A ++P PPP PPPP PPPP P PA P
Sbjct: 348 AAQAPPPPPPPPPPPPPPPPPPPPPPPP---PPPASTKPP 384
Score = 33.1 bits (74), Expect = 1.4
Identities = 19/56 (33%), Positives = 21/56 (36%), Gaps = 3/56 (5%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPL---PPPPLHQPQPAKPDAPPSEPKLSATPDAN 54
A+ P P P P PP PPPP P P P PP P P A+
Sbjct: 325 ASGPPHPHPHPHPHPHPRPPQPPAAQAPPPPPPPPPPPPPPPPPPPPPPPPPPPAS 380
Score = 30.4 bits (67), Expect = 9.3
Identities = 13/31 (41%), Positives = 14/31 (44%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPA 36
P PPP PPPP PPP +P A
Sbjct: 356 PPPPPPPPPPPPPPPPPPPPPPPASTKPPQA 386
>CAPU_DROME (Q24120) Cappuccino protein
Length = 1059
Score = 45.1 bits (105), Expect = 4e-04
Identities = 25/74 (33%), Positives = 30/74 (39%), Gaps = 14/74 (18%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKP---------DAPPSEPKLSATPDANVI 56
P PPP ++ G PPPP PPPP P P PP P +SA+P I
Sbjct: 505 PPPPPPPPPLANYGAPPPPPPPPPGSGSAPPPPPPAPIEGGGGIPPPPPPMSASPSKTTI 564
Query: 57 LV-----PSHSRWF 65
P+ WF
Sbjct: 565 SPAPLPDPAEGNWF 578
Score = 40.4 bits (93), Expect = 0.009
Identities = 20/44 (45%), Positives = 22/44 (49%), Gaps = 3/44 (6%)
Query: 8 TATEPPPSSSAGTPPPPLPPPPLH---QPQPAKPDAPPSEPKLS 48
+A E A PPPP PPPPLH P P P PP P L+
Sbjct: 472 SAKEDGEKPHAVAPPPPPPPPPLHAFVAPPPPPPPPPPPPPPLA 515
Score = 39.3 bits (90), Expect = 0.020
Identities = 18/41 (43%), Positives = 20/41 (47%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
+P PPP + PPPP PPPP P A APP P
Sbjct: 484 APPPPPPPPPLHAFVAPPPPPPPPPPPPPPLANYGAPPPPP 524
Score = 35.4 bits (80), Expect = 0.29
Identities = 19/53 (35%), Positives = 21/53 (38%), Gaps = 3/53 (5%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQP---QPAKPDAPPSEPKLSATP 51
A P PP + PPPP PPPP P A P PP P + P
Sbjct: 482 AVAPPPPPPPPPLHAFVAPPPPPPPPPPPPPPLANYGAPPPPPPPPPGSGSAP 534
Score = 33.1 bits (74), Expect = 1.4
Identities = 18/46 (39%), Positives = 18/46 (39%), Gaps = 6/46 (13%)
Query: 6 PTTATEPPPSSSAGTPPPPLP------PPPLHQPQPAKPDAPPSEP 45
P A PP PPPP P PPP P P APP P
Sbjct: 493 PLHAFVAPPPPPPPPPPPPPPLANYGAPPPPPPPPPGSGSAPPPPP 538
>EXTN_MAIZE (P14918) Extensin precursor (Proline-rich glycoprotein)
Length = 267
Score = 44.7 bits (104), Expect = 5e-04
Identities = 19/50 (38%), Positives = 25/50 (50%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
ATK PT PP + + PP P P PP + P P P P+ P + +P
Sbjct: 119 ATKPPTPKPTPPTYTPSPKPPTPKPTPPTYTPSPKPPTPKPTPPTYTPSP 168
Score = 43.5 bits (101), Expect = 0.001
Identities = 19/45 (42%), Positives = 22/45 (48%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPK 46
ATK PT PP + + PP P P PP + P P P P PK
Sbjct: 82 ATKPPTPKPTPPTYTPSPKPPTPKPTPPTYTPSPKPPATKPPTPK 126
Score = 42.0 bits (97), Expect = 0.003
Identities = 18/45 (40%), Positives = 22/45 (48%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPK 46
A+K PT PP + + PP P P PP + P P P P PK
Sbjct: 45 ASKPPTPKPTPPTYTPSPKPPTPKPTPPTYTPSPKPPATKPPTPK 89
Score = 40.8 bits (94), Expect = 0.007
Identities = 19/50 (38%), Positives = 23/50 (46%), Gaps = 1/50 (2%)
Query: 3 TKSPTTAT-EPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
T PT T P P A PP P P PP + P P P P+ P + +P
Sbjct: 29 TPKPTPPTYTPSPKPPASKPPTPKPTPPTYTPSPKPPTPKPTPPTYTPSP 78
Score = 40.0 bits (92), Expect = 0.012
Identities = 17/43 (39%), Positives = 20/43 (45%)
Query: 4 KSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPK 46
K PT PP + + PP P P PP + P P P P PK
Sbjct: 188 KPPTPKPTPPTYTPSPKPPTPKPTPPTYTPSPKPPATKPPTPK 230
Score = 40.0 bits (92), Expect = 0.012
Identities = 19/50 (38%), Positives = 23/50 (46%), Gaps = 1/50 (2%)
Query: 3 TKSPTTAT-EPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
T PT T P P A PP P P PP + P P P P+ P + +P
Sbjct: 103 TPKPTPPTYTPSPKPPATKPPTPKPTPPTYTPSPKPPTPKPTPPTYTPSP 152
Score = 40.0 bits (92), Expect = 0.012
Identities = 19/50 (38%), Positives = 23/50 (46%), Gaps = 1/50 (2%)
Query: 3 TKSPTTAT-EPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
T PT T P P A PP P P PP + P P P P+ P + +P
Sbjct: 66 TPKPTPPTYTPSPKPPATKPPTPKPTPPTYTPSPKPPTPKPTPPTYTPSP 115
Score = 39.7 bits (91), Expect = 0.015
Identities = 16/46 (34%), Positives = 22/46 (47%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
PT PP + + PP P P PP + P P P P+ P + +P
Sbjct: 174 PTPKPTPPTYTPSPKPPTPKPTPPTYTPSPKPPTPKPTPPTYTPSP 219
Score = 38.9 bits (89), Expect = 0.026
Identities = 19/51 (37%), Positives = 23/51 (44%), Gaps = 3/51 (5%)
Query: 4 KSPTTATEPP---PSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
K PT PP PS T P P P PP + P P P P+ P + +P
Sbjct: 153 KPPTPKPTPPTYTPSPKPPTHPTPKPTPPTYTPSPKPPTPKPTPPTYTPSP 203
Score = 38.5 bits (88), Expect = 0.034
Identities = 16/42 (38%), Positives = 20/42 (47%)
Query: 4 KSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
K PT PP + + PP P P PP + P P P P +P
Sbjct: 137 KPPTPKPTPPTYTPSPKPPTPKPTPPTYTPSPKPPTHPTPKP 178
Score = 36.2 bits (82), Expect = 0.17
Identities = 20/50 (40%), Positives = 22/50 (44%), Gaps = 3/50 (6%)
Query: 3 TKSPTTAT-EPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
T PT T P P A PP P P PP + P P P P P + TP
Sbjct: 207 TPKPTPPTYTPSPKPPATKPPTPKPTPPTYTPTPKPPATKP--PTYTPTP 254
Score = 35.8 bits (81), Expect = 0.22
Identities = 14/35 (40%), Positives = 18/35 (51%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPK 46
PP + + PP P P PP + P P P + P PK
Sbjct: 18 PPTYTPSPKPPTPKPTPPTYTPSPKPPASKPPTPK 52
Score = 33.9 bits (76), Expect = 0.84
Identities = 17/44 (38%), Positives = 20/44 (44%), Gaps = 2/44 (4%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
ATK PT +P P + TP PP PP + P P P P
Sbjct: 223 ATKPPTP--KPTPPTYTPTPKPPATKPPTYTPTPPVSHTPSPPP 264
Score = 32.3 bits (72), Expect = 2.4
Identities = 19/53 (35%), Positives = 21/53 (38%), Gaps = 7/53 (13%)
Query: 6 PTTATEPPPSSSAGTPPP--PLPPPPLHQPQPAKPDAP-----PSEPKLSATP 51
PT P P + TPP P P PP +P KP P P P TP
Sbjct: 19 PTYTPSPKPPTPKPTPPTYTPSPKPPASKPPTPKPTPPTYTPSPKPPTPKPTP 71
>ENAH_HUMAN (Q8N8S7) Enabled protein homolog
Length = 591
Score = 44.7 bits (104), Expect = 5e-04
Identities = 21/45 (46%), Positives = 23/45 (50%), Gaps = 1/45 (2%)
Query: 1 MATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
+A P PPP S G PPPP PPPPL P P PP+ P
Sbjct: 328 VALPPPPGPPPPPPLPSTGPPPPP-PPPPLPNQVPPPPPPPPAPP 371
Score = 38.9 bits (89), Expect = 0.026
Identities = 18/51 (35%), Positives = 21/51 (40%)
Query: 1 MATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
+A P P +S PPPP PPPP P P PP P + P
Sbjct: 311 LAPPPPPPLPPGPAQASVALPPPPGPPPPPPLPSTGPPPPPPPPPLPNQVP 361
Score = 38.9 bits (89), Expect = 0.026
Identities = 19/45 (42%), Positives = 20/45 (44%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSAT 50
P PPP T PPP PPPP Q P PP P L A+
Sbjct: 331 PPPPGPPPPPPLPSTGPPPPPPPPPLPNQVPPPPPPPPAPPLPAS 375
Score = 35.8 bits (81), Expect = 0.22
Identities = 20/48 (41%), Positives = 21/48 (43%), Gaps = 6/48 (12%)
Query: 7 TTATEPPPSSSAGTPPPPLP---PPPLHQPQPAKPDAPPSEPKLSATP 51
+ A PPP PPPPLP PPP P P PP P A P
Sbjct: 327 SVALPPPPGPP---PPPPLPSTGPPPPPPPPPLPNQVPPPPPPPPAPP 371
>WASP_MOUSE (P70315) Wiskott-Aldrich syndrome protein homolog (WASp)
Length = 520
Score = 44.3 bits (103), Expect = 6e-04
Identities = 29/89 (32%), Positives = 40/89 (44%), Gaps = 11/89 (12%)
Query: 12 PPPSSS-AGTPPPPL---------PPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSH 61
PPP++ +G PPPPL PPPP P P P + P+ P L TP + P
Sbjct: 388 PPPATGRSGPPPPPLPGAGGPPAPPPPPPPPPPPPCPGSGPAPPPLPPTPVSGGSPAPGG 447
Query: 62 SRWFSWDSIHQ-CEVRHLPEFFDSSSKSP 89
R D I Q ++ P ++S + P
Sbjct: 448 GRGALLDQIRQGIQLNKTPGALENSVQQP 476
Score = 35.0 bits (79), Expect = 0.38
Identities = 15/35 (42%), Positives = 17/35 (47%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPA 36
A P PPP +G PPPLPP P+ PA
Sbjct: 410 APPPPPPPPPPPPCPGSGPAPPPLPPTPVSGGSPA 444
Score = 31.2 bits (69), Expect = 5.4
Identities = 14/33 (42%), Positives = 15/33 (45%), Gaps = 3/33 (9%)
Query: 13 PPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
PP G PPPP P P P P + PP P
Sbjct: 360 PPVPMGGAPPPPTPRGP---PPPGRGGPPPPPP 389
>PCLO_MOUSE (Q9QYX7) Piccolo protein (Presynaptic cytomatrix protein)
(Aczonin) (Brain-derived HLMN protein)
Length = 5038
Score = 43.9 bits (102), Expect = 8e-04
Identities = 23/58 (39%), Positives = 29/58 (49%), Gaps = 3/58 (5%)
Query: 8 TATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPK---LSATPDANVILVPSHS 62
+A PPP PPPP PPPP PA PP+ PK +A P A +V +H+
Sbjct: 2302 SAQPPPPPPPPPPPPPPPPPPPPPPLPPATSPKPPTYPKRKLAAAAPVAPTAIVTAHA 2359
Score = 31.6 bits (70), Expect = 4.2
Identities = 16/48 (33%), Positives = 22/48 (45%), Gaps = 2/48 (4%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKP--DAPPSEPKLSAT 50
+P TA +PP + + P PP P Q P D P +PK + T
Sbjct: 628 TPATAPQPPVAEALPKPAPPKKPSAALPEQAKAPVADVEPKQPKTTET 675
Score = 30.8 bits (68), Expect = 7.1
Identities = 17/47 (36%), Positives = 20/47 (42%), Gaps = 8/47 (17%)
Query: 17 SAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSHSR 63
SA PPPP PPPP P PP P P A P++ +
Sbjct: 2302 SAQPPPPPPPPPP--------PPPPPPPPPPPPLPPATSPKPPTYPK 2340
>GP1_CHLRE (Q9FPQ6) Vegetative cell wall protein gp1 precursor
(Hydroxyproline-rich glycoprotein 1)
Length = 555
Score = 43.9 bits (102), Expect = 8e-04
Identities = 18/40 (45%), Positives = 20/40 (50%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
PPP S PPPP PP P + P P P +PP P P
Sbjct: 264 PPPPPSPPPPPPPRPPFPANTPMPPSPPSPPPSPAPPTPP 303
Score = 42.7 bits (99), Expect = 0.002
Identities = 22/56 (39%), Positives = 24/56 (42%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPS 60
SP P P A TP PP PP P P P P PPS S P + + PS
Sbjct: 269 SPPPPPPPRPPFPANTPMPPSPPSPPPSPAPPTPPTPPSPSPPSPVPPSPAPVPPS 324
Score = 42.0 bits (97), Expect = 0.003
Identities = 20/46 (43%), Positives = 24/46 (51%), Gaps = 1/46 (2%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P+ A PPS + +P PP PP P+ P PA P P PK A P
Sbjct: 221 PSPAPPSPPSPAPPSPSPPAPPSPV-PPSPAPPSPAPPSPKPPAPP 265
Score = 42.0 bits (97), Expect = 0.003
Identities = 19/56 (33%), Positives = 26/56 (45%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPS 60
+P + PPS + PP P PP P P P+ +P P S +P + I PS
Sbjct: 319 APVPPSPAPPSPAPSPPPSPAPPTPSPSPSPSPSPSPSPSPSPSPSPSPSPIPSPS 374
Score = 41.6 bits (96), Expect = 0.004
Identities = 21/50 (42%), Positives = 23/50 (46%), Gaps = 3/50 (6%)
Query: 5 SPTTATEPPPSSSAGTPPPP---LPPPPLHQPQPAKPDAPPSEPKLSATP 51
SP + PPS PPPP PPPP P PA PPS P +P
Sbjct: 248 SPAPPSPAPPSPKPPAPPPPPSPPPPPPPRPPFPANTPMPPSPPSPPPSP 297
Score = 41.2 bits (95), Expect = 0.005
Identities = 20/52 (38%), Positives = 25/52 (47%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDA 53
A SP + PPS + +P PP P PP P P +PPS S +P A
Sbjct: 82 APPSPAPPSPAPPSPAPPSPAPPSPAPPSPAPPSPAPPSPPSPAPPSPSPPA 133
Score = 40.8 bits (94), Expect = 0.007
Identities = 21/57 (36%), Positives = 25/57 (43%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSHS 62
PT T P PS + PP P P PP P P PPS + +P + PS S
Sbjct: 300 PTPPTPPSPSPPSPVPPSPAPVPPSPAPPSPAPSPPPSPAPPTPSPSPSPSPSPSPS 356
Score = 40.8 bits (94), Expect = 0.007
Identities = 20/51 (39%), Positives = 24/51 (46%), Gaps = 3/51 (5%)
Query: 2 ATKSPTTATEP---PPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSA 49
A SP+ P PPS + +P PP P PP P P+ P PP P A
Sbjct: 232 APPSPSPPAPPSPVPPSPAPPSPAPPSPKPPAPPPPPSPPPPPPPRPPFPA 282
Score = 40.8 bits (94), Expect = 0.007
Identities = 22/58 (37%), Positives = 28/58 (47%), Gaps = 1/58 (1%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLH-QPQPAKPDAPPSEPKLSATPDANVILVPSHS 62
P+ A PP+ + +PP P+PP P P PA P PS P A P + PS S
Sbjct: 295 PSPAPPTPPTPPSPSPPSPVPPSPAPVPPSPAPPSPAPSPPPSPAPPTPSPSPSPSPS 352
Score = 40.0 bits (92), Expect = 0.012
Identities = 22/59 (37%), Positives = 26/59 (43%), Gaps = 1/59 (1%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPS 60
A SP + PPS + +P PP P PP P PA P P P + P L PS
Sbjct: 92 APPSPAPPSPAPPSPAPPSPAPPSPAPP-SPPSPAPPSPSPPAPPSPSPPSPAPPLPPS 149
Score = 39.7 bits (91), Expect = 0.015
Identities = 22/59 (37%), Positives = 26/59 (43%), Gaps = 1/59 (1%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPL-HQPQPAKPDAPPSEPKLSATPDANVILVPSHS 62
SP + PPS + +P PP P PP P PA P P P A P + PS S
Sbjct: 80 SPAPPSPAPPSPAPPSPAPPSPAPPSPAPPSPAPPSPAPPSPPSPAPPSPSPPAPPSPS 138
Score = 39.7 bits (91), Expect = 0.015
Identities = 22/61 (36%), Positives = 27/61 (44%), Gaps = 3/61 (4%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSH 61
A SP + PPS + PP P PP P P P+ P P P A P + + PS
Sbjct: 107 APPSPAPPSPAPPSPPSPAPPSPSPPAP---PSPSPPSPAPPLPPSPAPPSPSPPVPPSP 163
Query: 62 S 62
S
Sbjct: 164 S 164
Score = 39.3 bits (90), Expect = 0.020
Identities = 21/50 (42%), Positives = 22/50 (44%), Gaps = 4/50 (8%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
A SP PPP S PPPP PP P P +PPS P A P
Sbjct: 255 APPSPKPPAPPPPPS----PPPPPPPRPPFPANTPMPPSPPSPPPSPAPP 300
Score = 39.3 bits (90), Expect = 0.020
Identities = 20/60 (33%), Positives = 28/60 (46%), Gaps = 3/60 (5%)
Query: 6 PTTATEPPPSSSAGTP---PPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSHS 62
P + PPS + +P PPP P PP P P+ +P P S +P + +PS S
Sbjct: 315 PPSPAPVPPSPAPPSPAPSPPPSPAPPTPSPSPSPSPSPSPSPSPSPSPSPSPSPIPSPS 374
Score = 38.9 bits (89), Expect = 0.026
Identities = 20/49 (40%), Positives = 22/49 (44%), Gaps = 3/49 (6%)
Query: 6 PTTATEPPPSSSAGTPP---PPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P+ + PPS S PP PP P PP P KP APP P P
Sbjct: 226 PSPPSPAPPSPSPPAPPSPVPPSPAPPSPAPPSPKPPAPPPPPSPPPPP 274
Score = 38.9 bits (89), Expect = 0.026
Identities = 19/50 (38%), Positives = 23/50 (46%), Gaps = 1/50 (2%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
A SP + PPS + +PP P PP P P P P P P L +P
Sbjct: 102 APPSPAPPSPAPPSPAPPSPPSPAPPSP-SPPAPPSPSPPSPAPPLPPSP 150
Score = 38.9 bits (89), Expect = 0.026
Identities = 23/56 (41%), Positives = 29/56 (51%), Gaps = 3/56 (5%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPL--HQPQPAKPDAPPSEPKLSATPDANVILVP 59
P+ A PPS + +P PP+PP P P PA P +PPS S +P A VP
Sbjct: 192 PSPAPPVPPSPAPPSPAPPVPPSPAPPSPPSPA-PPSPPSPAPPSPSPPAPPSPVP 246
Score = 38.9 bits (89), Expect = 0.026
Identities = 21/56 (37%), Positives = 25/56 (44%), Gaps = 3/56 (5%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPS 60
SP+ P P+ + TPP P PP P P PA P P P A P + PS
Sbjct: 162 SPSPPVPPSPAPPSPTPPSPSPPVP---PSPAPPSPAPPVPPSPAPPSPAPPVPPS 214
Score = 38.5 bits (88), Expect = 0.034
Identities = 18/47 (38%), Positives = 22/47 (46%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
SP + PPS + +P PP P PP P P +PPS S P
Sbjct: 42 SPAPPSPAPPSPAPPSPAPPSPAPPSPGPPSPAPPSPPSPAPPSPAP 88
Score = 38.5 bits (88), Expect = 0.034
Identities = 22/51 (43%), Positives = 25/51 (48%), Gaps = 2/51 (3%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPD-APPSEPKLSATP 51
A SP + PPS + +PP P PP P P PA P APPS S P
Sbjct: 59 APPSPAPPSPGPPSPAPPSPPSPAPPSPA-PPSPAPPSPAPPSPAPPSPAP 108
Score = 38.1 bits (87), Expect = 0.044
Identities = 22/51 (43%), Positives = 25/51 (48%), Gaps = 2/51 (3%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPD-APPSEPKLSATP 51
A SP + PPS + +P PP P PP P PA P APPS S P
Sbjct: 49 APPSPAPPSPAPPSPAPPSPGPPSPAPP-SPPSPAPPSPAPPSPAPPSPAP 98
Score = 38.1 bits (87), Expect = 0.044
Identities = 20/50 (40%), Positives = 22/50 (44%), Gaps = 3/50 (6%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
A SPT + PP + PP P PP P P PA P P P A P
Sbjct: 172 APPSPTPPSPSPPVPPSPAPPSPAPPVP---PSPAPPSPAPPVPPSPAPP 218
Score = 38.1 bits (87), Expect = 0.044
Identities = 22/51 (43%), Positives = 24/51 (46%), Gaps = 2/51 (3%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPD-APPSEPKLSATP 51
A SP + PPS +P PP PP P P PA P APPS S P
Sbjct: 54 APPSPAPPSPAPPSPGPPSPAPPSPPSPA-PPSPAPPSPAPPSPAPPSPAP 103
Score = 37.7 bits (86), Expect = 0.058
Identities = 17/42 (40%), Positives = 21/42 (49%), Gaps = 1/42 (2%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPL-HQPQPAKPDAPPSEP 45
SP + PP+ + PP P PP P P+P P PPS P
Sbjct: 230 SPAPPSPSPPAPPSPVPPSPAPPSPAPPSPKPPAPPPPPSPP 271
Score = 37.7 bits (86), Expect = 0.058
Identities = 22/51 (43%), Positives = 24/51 (46%), Gaps = 2/51 (3%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPD-APPSEPKLSATP 51
A SP + PPS + PP P PP P P PA P APPS S P
Sbjct: 64 APPSPGPPSPAPPSPPSPAPPSPAPPSPA-PPSPAPPSPAPPSPAPPSPAP 113
Score = 37.4 bits (85), Expect = 0.076
Identities = 19/50 (38%), Positives = 22/50 (44%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
A SP + PPS + +P PP P PP P APPS S P
Sbjct: 44 APPSPAPPSPAPPSPAPPSPAPPSPGPPSPAPPSPPSPAPPSPAPPSPAP 93
Score = 37.0 bits (84), Expect = 0.099
Identities = 16/42 (38%), Positives = 18/42 (42%)
Query: 4 KSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
+ P A P P S PP P PP P P P+ P P P
Sbjct: 277 RPPFPANTPMPPSPPSPPPSPAPPTPPTPPSPSPPSPVPPSP 318
Score = 37.0 bits (84), Expect = 0.099
Identities = 18/46 (39%), Positives = 21/46 (45%), Gaps = 3/46 (6%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P+ A PPS + +PP P PP P P PA P P P P
Sbjct: 205 PSPAPPVPPSPAPPSPPSPAPPSP---PSPAPPSPSPPAPPSPVPP 247
Score = 36.6 bits (83), Expect = 0.13
Identities = 19/44 (43%), Positives = 24/44 (54%), Gaps = 5/44 (11%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKP----DAPPSEP 45
P+ + PPS + +P PP+PP P P PA P APPS P
Sbjct: 179 PSPSPPVPPSPAPPSPAPPVPPSPA-PPSPAPPVPPSPAPPSPP 221
Score = 36.6 bits (83), Expect = 0.13
Identities = 23/61 (37%), Positives = 28/61 (45%), Gaps = 8/61 (13%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQ-------PQPAKPDAPPSEPKLSATPDANVILV 58
P+ + PPS S +P PPLPP P P P+ P PPS S TP + V
Sbjct: 127 PSPSPPAPPSPSPPSPAPPLPPSPAPPSPSPPVPPSPSPP-VPPSPAPPSPTPPSPSPPV 185
Query: 59 P 59
P
Sbjct: 186 P 186
Score = 36.6 bits (83), Expect = 0.13
Identities = 20/48 (41%), Positives = 23/48 (47%), Gaps = 2/48 (4%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPD-APPSEPKLSATP 51
SP + P P+ + PP P PP P P PA P APPS S P
Sbjct: 72 SPAPPSPPSPAPPSPAPPSPAPPSPA-PPSPAPPSPAPPSPAPPSPAP 118
Score = 36.2 bits (82), Expect = 0.17
Identities = 18/56 (32%), Positives = 22/56 (39%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPS 60
SP + PP + PP P PP P P+ P P P A P + PS
Sbjct: 188 SPAPPSPAPPVPPSPAPPSPAPPVPPSPAPPSPPSPAPPSPPSPAPPSPSPPAPPS 243
Score = 36.2 bits (82), Expect = 0.17
Identities = 23/64 (35%), Positives = 28/64 (42%), Gaps = 4/64 (6%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPP---PPLHQPQPAKPDAPPSEPKLSATPDANVILV 58
A +P T P P S P P+PP PP P P APP+ P S +P +
Sbjct: 298 APPTPPTPPSPSPPSPVPPSPAPVPPSPAPPSPAPSPPPSPAPPT-PSPSPSPSPSPSPS 356
Query: 59 PSHS 62
PS S
Sbjct: 357 PSPS 360
Score = 36.2 bits (82), Expect = 0.17
Identities = 18/48 (37%), Positives = 22/48 (45%), Gaps = 2/48 (4%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQ--PQPAKPDAPPSEPKLSATP 51
P+ A PPS + +P PP+PP P P PA P P P P
Sbjct: 140 PSPAPPLPPSPAPPSPSPPVPPSPSPPVPPSPAPPSPTPPSPSPPVPP 187
Score = 36.2 bits (82), Expect = 0.17
Identities = 20/49 (40%), Positives = 23/49 (46%), Gaps = 4/49 (8%)
Query: 6 PTTATEPPPSSSAGTPPPPLPP---PPLHQPQPAKPDAPPSEPKLSATP 51
P+ + PPS S PP P PP PPL P PA P P P + P
Sbjct: 119 PSPPSPAPPSPSPPAPPSPSPPSPAPPL-PPSPAPPSPSPPVPPSPSPP 166
Score = 35.8 bits (81), Expect = 0.22
Identities = 18/46 (39%), Positives = 21/46 (45%), Gaps = 1/46 (2%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P+ + PPS S PP P PP P P P+ P P P A P
Sbjct: 153 PSPSPPVPPSPSPPVPPSPAPPSPT-PPSPSPPVPPSPAPPSPAPP 197
Score = 35.8 bits (81), Expect = 0.22
Identities = 21/51 (41%), Positives = 24/51 (46%), Gaps = 3/51 (5%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQ--PQPAKPD-APPSEPKLSATPDA 53
P+ A PPS + +PP P PP P P P P APPS S P A
Sbjct: 213 PSPAPPSPPSPAPPSPPSPAPPSPSPPAPPSPVPPSPAPPSPAPPSPKPPA 263
Score = 35.0 bits (79), Expect = 0.38
Identities = 15/46 (32%), Positives = 21/46 (45%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P + P P+ + PP P PPP P P +P P + P + P
Sbjct: 246 PPSPAPPSPAPPSPKPPAPPPPPSPPPPPPPRPPFPANTPMPPSPP 291
Score = 35.0 bits (79), Expect = 0.38
Identities = 18/48 (37%), Positives = 21/48 (43%), Gaps = 1/48 (2%)
Query: 5 SPTTATEPPPSSSAGTPP-PPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
SP + PP + +PP PP P PP P P PPS S P
Sbjct: 149 SPAPPSPSPPVPPSPSPPVPPSPAPPSPTPPSPSPPVPPSPAPPSPAP 196
Score = 35.0 bits (79), Expect = 0.38
Identities = 21/58 (36%), Positives = 24/58 (41%), Gaps = 4/58 (6%)
Query: 6 PTTATEPPPSSSAGTPP---PPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPS 60
P + PPS S PP PP+PP P P P P P P A P + PS
Sbjct: 145 PLPPSPAPPSPSPPVPPSPSPPVPPSPA-PPSPTPPSPSPPVPPSPAPPSPAPPVPPS 201
Score = 34.7 bits (78), Expect = 0.49
Identities = 21/56 (37%), Positives = 25/56 (44%), Gaps = 8/56 (14%)
Query: 6 PTTATEPPPSSSAGTPPP-------PLPPP-PLHQPQPAKPDAPPSEPKLSATPDA 53
P+ A PPPS + TP P P P P P P P+ P PK S +P A
Sbjct: 328 PSPAPSPPPSPAPPTPSPSPSPSPSPSPSPSPSPSPSPSPSPIPSPSPKPSPSPVA 383
Score = 34.7 bits (78), Expect = 0.49
Identities = 21/52 (40%), Positives = 24/52 (45%), Gaps = 2/52 (3%)
Query: 3 TKSPTTATEPPPSSSAGTPPPPLP-PPPLHQPQPAKPDAPPSEPKLSATPDA 53
T SP+ + P PS S P P P P P+ P P KP P KL DA
Sbjct: 342 TPSPSPSPSPSPSPSPSPSPSPSPSPSPIPSPSP-KPSPSPVAVKLVWADDA 392
Score = 34.7 bits (78), Expect = 0.49
Identities = 16/46 (34%), Positives = 19/46 (40%), Gaps = 1/46 (2%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P + P P+ + PP P PP P P P P P P A P
Sbjct: 40 PPSPAPPSPAPPSPAPPSPAPPSPA-PPSPGPPSPAPPSPPSPAPP 84
Score = 33.5 bits (75), Expect = 1.1
Identities = 17/59 (28%), Positives = 23/59 (38%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPS 60
A SP + P P+ + +PP P P P P P P P P + + PS
Sbjct: 112 APPSPAPPSPPSPAPPSPSPPAPPSPSPPSPAPPLPPSPAPPSPSPPVPPSPSPPVPPS 170
Score = 32.7 bits (73), Expect = 1.9
Identities = 18/58 (31%), Positives = 21/58 (36%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSHS 62
SP+ P P+ + PP P P P P P P P A P PS S
Sbjct: 180 SPSPPVPPSPAPPSPAPPVPPSPAPPSPAPPVPPSPAPPSPPSPAPPSPPSPAPPSPS 237
Score = 31.2 bits (69), Expect = 5.4
Identities = 15/50 (30%), Positives = 21/50 (42%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
A SP + P P+ +P P P P P P+ +P P S +P
Sbjct: 326 APPSPAPSPPPSPAPPTPSPSPSPSPSPSPSPSPSPSPSPSPSPIPSPSP 375
>WASL_BOVIN (Q95107) Neural Wiskott-Aldrich syndrome protein
(N-WASP)
Length = 505
Score = 43.5 bits (101), Expect = 0.001
Identities = 23/55 (41%), Positives = 26/55 (46%), Gaps = 9/55 (16%)
Query: 6 PTTATE---PPPSSSAGTPPPPL------PPPPLHQPQPAKPDAPPSEPKLSATP 51
P+ PPP S+G PPPP PPPP P A P PPS P + A P
Sbjct: 282 PSRGGPPPPPPPPHSSGPPPPPARGRGAPPPPPSRAPTAAPPPPPPSRPGVGAPP 336
Score = 40.4 bits (93), Expect = 0.009
Identities = 19/44 (43%), Positives = 25/44 (56%), Gaps = 1/44 (2%)
Query: 3 TKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQ-PAKPDAPPSEP 45
+++PT A PPP S G PP PP ++ P PA P + PS P
Sbjct: 315 SRAPTAAPPPPPPSRPGVGAPPPPPNRMYPPPLPALPSSAPSGP 358
Score = 38.1 bits (87), Expect = 0.044
Identities = 22/48 (45%), Positives = 24/48 (49%), Gaps = 10/48 (20%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQ----PQPAK----PDAPPSEPKLSATP 51
PPP S G PPPP PPP H P PA+ P PPS +A P
Sbjct: 278 PPPPPSRGGPPPP--PPPPHSSGPPPPPARGRGAPPPPPSRAPTAAPP 323
Score = 37.4 bits (85), Expect = 0.076
Identities = 16/33 (48%), Positives = 19/33 (57%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSE 44
PPP S +G+ PP PPPP P P P PS+
Sbjct: 362 PPPLSVSGSVAPPPPPPPPPPPGPPPPPGLPSD 394
Score = 37.4 bits (85), Expect = 0.076
Identities = 20/53 (37%), Positives = 24/53 (44%), Gaps = 11/53 (20%)
Query: 12 PPPSSSAGTPPPPL--------PPPPLHQPQPAKPDAPPSE---PKLSATPDA 53
PPP+ G PPPP PPPP +P P PP+ P L A P +
Sbjct: 301 PPPARGRGAPPPPPSRAPTAAPPPPPPSRPGVGAPPPPPNRMYPPPLPALPSS 353
Score = 35.8 bits (81), Expect = 0.22
Identities = 24/66 (36%), Positives = 26/66 (39%), Gaps = 12/66 (18%)
Query: 2 ATKSPTTATEPPP----SSSAGTPPPPLPPP--------PLHQPQPAKPDAPPSEPKLSA 49
A P PPP SSA + PPP PPP P P P P PP P L +
Sbjct: 334 APPPPPNRMYPPPLPALPSSAPSGPPPPPPPLSVSGSVAPPPPPPPPPPPGPPPPPGLPS 393
Query: 50 TPDANV 55
D V
Sbjct: 394 DGDHQV 399
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.315 0.131 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 54,159,697
Number of Sequences: 164201
Number of extensions: 2474203
Number of successful extensions: 36419
Number of sequences better than 10.0: 797
Number of HSP's better than 10.0 without gapping: 448
Number of HSP's successfully gapped in prelim test: 387
Number of HSP's that attempted gapping in prelim test: 21336
Number of HSP's gapped (non-prelim): 7218
length of query: 434
length of database: 59,974,054
effective HSP length: 113
effective length of query: 321
effective length of database: 41,419,341
effective search space: 13295608461
effective search space used: 13295608461
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0438.5


