

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0437b.11
(528 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
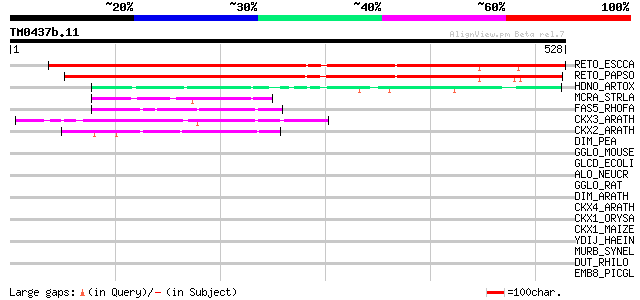
Score E
Sequences producing significant alignments: (bits) Value
RETO_ESCCA (P30986) Reticuline oxidase precursor (EC 1.21.3.3) (... 402 e-112
RETO_PAPSO (P93479) Reticuline oxidase precursor (EC 1.21.3.3) (... 397 e-110
HDNO_ARTOX (P08159) 6-hydroxy-D-nicotine oxidase (EC 1.5.3.6) (6... 87 8e-17
MCRA_STRLA (P43485) Mitomycin radical oxidase (EC 1.5.3.-) 71 8e-12
FAS5_RHOFA (P46377) Hypothetical 47.9 kDa oxidoreductase in fasc... 56 3e-07
CKX3_ARATH (Q9LTS3) Cytokinin dehydrogenase 3 precursor (EC 1.5.... 54 1e-06
CKX2_ARATH (Q9FUJ3) Cytokinin dehydrogenase 2 precursor (EC 1.5.... 50 2e-05
DIM_PEA (P93472) Cell elongation protein diminuto 43 0.002
GGLO_MOUSE (P58710) L-gulonolactone oxidase (EC 1.1.3.8) (LGO) (... 41 0.007
GLCD_ECOLI (P52075) Glycolate oxidase subunit glcD 40 0.015
ALO_NEUCR (Q7SGY1) Putative D-arabinono-1,4-lactone oxidase (EC ... 40 0.020
GGLO_RAT (P10867) L-gulonolactone oxidase (EC 1.1.3.8) (LGO) (L-... 39 0.025
DIM_ARATH (Q39085) Cell elongation protein DIMINUTO (Cell elonga... 39 0.025
CKX4_ARATH (Q9FUJ2) Cytokinin dehydrogenase 4 precursor (EC 1.5.... 39 0.043
CKX1_ORYSA (Q9LDE6) Probable cytokinin dehydrogenase precursor (... 38 0.074
CKX1_MAIZE (Q9T0N8) Cytokinin dehydrogenase 1 precursor (EC 1.5.... 37 0.13
YDIJ_HAEIN (Q57252) Protein HI1163 35 0.63
MURB_SYNEL (Q8DLV6) UDP-N-acetylenolpyruvoylglucosamine reductas... 33 1.4
DUT_RHILO (Q98C10) Deoxyuridine 5'-triphosphate nucleotidohydrol... 33 1.8
EMB8_PICGL (Q40863) Embryogenesis-associated protein EMB8 33 2.4
>RETO_ESCCA (P30986) Reticuline oxidase precursor (EC 1.21.3.3)
(Berberine-bridge-forming enzyme) (BBE)
(Tetrahydroprotoberberine synthase)
Length = 538
Score = 402 bits (1034), Expect = e-112
Identities = 213/500 (42%), Positives = 304/500 (60%), Gaps = 16/500 (3%)
Query: 38 SSIVRNSNPTEEIVLTQNSSSYESLLQSSIRNTRFLDSSVLKPNLIVTPQELSHVQAAIT 97
S + N + + S + L SI+N F +S + KP+ I+ P + I
Sbjct: 29 SCLTFNGVRNHTVFSADSDSDFNRFLHLSIQNPLFQNSLISKPSAIILPGSKEELSNTIR 88
Query: 98 CSVKQGLQVRVRSGGHDYEGLSYISNVPFLIIDLSNFRSISIDIEDESAWVQSGATLGEL 157
C K +R+RSGGH YEGLSY S+ PF++IDL N +SID+E E+AWV+SG+TLGEL
Sbjct: 89 CIRKGSWTIRLRSGGHSYEGLSYTSDTPFILIDLMNLNRVSIDLESETAWVESGSTLGEL 148
Query: 158 YYAIANQSNVHAFPAGSCPTVGIGGHFSGGGFSTIFRKHGLAADHVIDAQIIDVNGKILN 217
YYAI S+ F AG CPTVG GGH SGGGF + RK+GLAAD+V+DA +ID NG IL+
Sbjct: 149 YYAITESSSKLGFTAGWCPTVGTGGHISGGGFGMMSRKYGLAADNVVDAILIDANGAILD 208
Query: 218 RNLMGEDLFWAIKGGGGSSFGVITSWKVKLVHVPPKVTIFDLPKKLD-QNVSEIFQKWQT 276
R MGED+FWAI+GGGG +G I +WK+KL+ VP KVT+F + K + + + KWQ
Sbjct: 209 RQAMGEDVFWAIRGGGGGVWGAIYAWKIKLLPVPEKVTVFRVTKNVAIDEATSLLHKWQF 268
Query: 277 IAHKLPGELFLHSVMGVSNSPKHGGKSVIVSFTGLYLGIAENLLPLMQSNFAELGLRRDD 336
+A +L E F SV+G ++ K V ++ G + G+ F ELGL +D
Sbjct: 269 VAEELE-EDFTLSVLGGADE-----KQVWLTMLGFHFGLKTVAKSTFDLLFPELGLVEED 322
Query: 337 CTEMNWIQSVLYLAGYPINASLNVLLQRNQTFGSFKAKSDYVNEPIPRAGLEGLWKMLLE 396
EM+W +S YLAG + LN + +FK K D EP+P GL + L +
Sbjct: 323 YLEMSWGESFAYLAGLETVSQLNNRFLKFDE-RAFKTKVDLTKEPLPSKAFYGLLERLSK 381
Query: 397 EDSPLLILTPYGGRMSEISESETPFPHRNKSIFGIQYLVNWDKNEETKM--HVEWMRRLY 454
E + + L +GG+MS+IS TPFPHR+ + ++Y+V W+++E+ K ++W+ ++Y
Sbjct: 382 EPNGFIALNGFGGQMSKISSDFTPFPHRSGTRLMVEYIVAWNQSEQKKKTEFLDWLEKVY 441
Query: 455 AYMKPYVSKNPRGAYLNYRDLDI-GVNRG-----NTSYEEAKCWGLKYFKSNFERLARVK 508
+MKP+VSKNPR Y+N+ DLD+ G++ G N + E ++ WG YF SN+ERL R K
Sbjct: 442 EFMKPFVSKNPRLGYVNHIDLDLGGIDWGNKTVVNNAIEISRSWGESYFLSNYERLIRAK 501
Query: 509 AQVDPSNFFRHEQSIPPLSD 528
+DP+N F H QSIPP+++
Sbjct: 502 TLIDPNNVFNHPQSIPPMAN 521
>RETO_PAPSO (P93479) Reticuline oxidase precursor (EC 1.21.3.3)
(Berberine-bridge-forming enzyme) (BBE)
(Tetrahydroprotoberberine synthase)
Length = 535
Score = 397 bits (1021), Expect = e-110
Identities = 208/483 (43%), Positives = 301/483 (62%), Gaps = 16/483 (3%)
Query: 53 TQNSSSYESLLQSSIRNTRFLDSSVLKPNLIVTPQELSHVQAAITCSVKQGLQVRVRSGG 112
T +S Y LL +S++N F +V KP+ IV P + + + C ++ +R+RSGG
Sbjct: 48 TDTNSDYFKLLHASMQNPLFAKPTVSKPSFIVMPGSKEELSSTVHCCTRESWTIRLRSGG 107
Query: 113 HDYEGLSYISNVPFLIIDLSNFRSISIDIEDESAWVQSGATLGELYYAIANQSNVHAFPA 172
H YEGLSY ++ PF+I+D+ N ISID+ E+AWV+SGATLGELYYAIA ++ F A
Sbjct: 108 HSYEGLSYTADTPFVIVDMMNLNRISIDVLSETAWVESGATLGELYYAIAQSTDTLGFTA 167
Query: 173 GSCPTVGIGGHFSGGGFSTIFRKHGLAADHVIDAQIIDVNGKILNRNLMGEDLFWAIKGG 232
G CPTVG GGH SGGGF + RK+GLAAD+V+DA +ID NG IL+R MG+D+FWAI+GG
Sbjct: 168 GWCPTVGSGGHISGGGFGMMSRKYGLAADNVVDAILIDSNGAILDREKMGDDVFWAIRGG 227
Query: 233 GGSSFGVITSWKVKLVHVPPKVTIFDLPKKLD-QNVSEIFQKWQTIAHKLPGELFLHSVM 291
GG +G I +WK+KL+ VP K+T+F + K + ++ S + KWQ +A +L E F SV+
Sbjct: 228 GGGVWGAIYAWKIKLLPVPEKLTVFRVTKNVGIEDASSLLHKWQYVADEL-DEDFTVSVL 286
Query: 292 GVSNSPKHGGKSVIVSFTGLYLGIAENLLPLMQSNFAELGLRRDDCTEMNWIQSVLYLAG 351
G N G + F GL+LG + ++ F ELGL + EM+W +S+ +L+G
Sbjct: 287 GGVN-----GNDAWLMFLGLHLGRKDAAKTIIDEKFPELGLVDKEFQEMSWGESMAFLSG 341
Query: 352 YPINASLNVLLQRNQTFGSFKAKSDYVNEPIPRAGLEGLWKMLLEEDSPLLILTPYGGRM 411
+ LN + +FK K D+ +P +ML E+ + L +GG+M
Sbjct: 342 LDTISELNNRFLKFDE-RAFKTKVDFTKVSVPLNVFRHALEMLSEQPGGFIALNGFGGKM 400
Query: 412 SEISESETPFPHRNKSIFGIQYLVNWDKNEETKM--HVEWMRRLYAYMKPYVSKNPRGAY 469
SEIS TPFPHR + +Y++ W+++EE+K+ EW+ + Y Y++P+VSK PR Y
Sbjct: 401 SEISTDFTPFPHRKGTKLMFEYIIAWNQDEESKIGEFSEWLAKFYDYLEPFVSKEPRVGY 460
Query: 470 LNYRDLDIG----VNRGNT--SYEEAKCWGLKYFKSNFERLARVKAQVDPSNFFRHEQSI 523
+N+ DLDIG N+ +T + E A+ WG +YF SN+ERL + K +DP+N F H QSI
Sbjct: 461 VNHIDLDIGGIDWRNKSSTTNAVEIARNWGERYFSSNYERLVKAKTLIDPNNVFNHPQSI 520
Query: 524 PPL 526
PP+
Sbjct: 521 PPM 523
>HDNO_ARTOX (P08159) 6-hydroxy-D-nicotine oxidase (EC 1.5.3.6)
(6-HDNO)
Length = 458
Score = 87.4 bits (215), Expect = 8e-17
Identities = 110/464 (23%), Positives = 186/464 (39%), Gaps = 62/464 (13%)
Query: 79 KPNLIVTPQELSHVQAAITCSVKQGLQVRVRSGGHDYEGLSYISNVPFLIIDLSNFRSIS 138
+P+LI V ++ + GL++ VRSGGH+ G Y +N +++DL SI
Sbjct: 37 RPSLIARCLSAGDVAKSVRYACDNGLEISVRSGGHNPNG--YATNDGGIVLDLRLMNSIH 94
Query: 139 IDIEDESAWVQSGATLGELYYAIANQSNVHAFPAGSCPTVGIGGHFSGGGFSTIFRKHGL 198
ID A + G G+L A A G P VG G GG + K+GL
Sbjct: 95 IDTAGSRARIGGGVISGDLVKEAAKFGL--AAVTGMHPKVGFCGLALNGGVGFLTPKYGL 152
Query: 199 AADHVIDAQIIDVNGKILN-RNLMGEDLFWAIKGGGGSSFGVITSWKVKLVHVPPKVTIF 257
A+D+++ A ++ G ++ + +LFWA++ G G +FGV+T +V+L +
Sbjct: 153 ASDNILGATLVTATGDVIYCSDDERPELFWAVR-GAGPNFGVVTEVEVQL---------Y 202
Query: 258 DLPKKLDQNVSEIFQKWQTIAHKLPGELFLHSVMGVSNSPKHGGKSVIVSFTGLYLGIAE 317
+LP+K+ F W +L G L S++ N + + +++G+ E
Sbjct: 203 ELPRKMLAG----FITWAPSVSELAG--LLTSLLDALN------EMADHIYPSVFVGVDE 250
Query: 318 NLLPLMQSNFAELG---LRRDDCTEMNWIQSVLYLAGYPINASLNV-----LLQRNQTFG 369
N P + LG + D + + G ++ S+ V ++ N G
Sbjct: 251 NRAPSVTVCVGHLGGLDIAERDIARLRGL-------GRTVSDSIAVRSYDEVVALNAEVG 303
Query: 370 SFKAKSDYVNEPIPRAGLEGLWKMLLEEDSPLLILTPYGGRMSEISESETPF------PH 423
SF+ + A + + + + P G ++ PF P
Sbjct: 304 SFEDGMSNLWIDREIAMPNARFAEAIAGNLDKFVSEPASGGSVKLEIEGMPFGNPKRTPA 363
Query: 424 RNKSIFGIQYLVNWD-KNEETKMHVEWMRRL-YAYMKPYVSKNPRGAYLNYRDLDIGVNR 481
R++ G+ L W ++ + E R L A ++ V+ + G
Sbjct: 364 RHRDAMGVLALAEWSGAAPGSEKYPELARELDAALLRAGVTTSGFGLL------------ 411
Query: 482 GNTSYEEAKCWGLKYFKSNFERLARVKAQVDPSNFFRHEQSIPP 525
N S A+ Y + RLA VK + DP N FRH +I P
Sbjct: 412 NNNSEVTAEMVAEVYKPEVYCRLAAVKREYDPENRFRHNYNIDP 455
>MCRA_STRLA (P43485) Mitomycin radical oxidase (EC 1.5.3.-)
Length = 447
Score = 70.9 bits (172), Expect = 8e-12
Identities = 47/175 (26%), Positives = 84/175 (47%), Gaps = 12/175 (6%)
Query: 79 KPNLIVTPQELSHVQAAITCSVKQGLQVRVRSGGHDYEGLSYISNVPFLIIDLSNFRSIS 138
+P +V + V AA+ + +Q V V + GH +S ++++ +S
Sbjct: 29 RPAYVVEAADEQEVAAAVRLAAEQKRPVGVMATGHGPS----VSADDAVLVNTRRMEGVS 84
Query: 139 IDIEDESAWVQSGATLGELYYAIANQSNVHAFPA--GSCPTVGIGGHFSGGGFSTIFRKH 196
+D +AW+++GA + + + H GS P VG G+ GGG + R+
Sbjct: 85 VDAARATAWIEAGAR----WRKVLEHTAPHGLAPLNGSSPNVGAVGYLVGGGAGLLGRRF 140
Query: 197 GLAADHVIDAQIIDVNGKILNRNL-MGEDLFWAIKGGGGSSFGVITSWKVKLVHV 250
G AADHV +++ +G++ + DLFWA++ GG +FG++ +V L V
Sbjct: 141 GYAADHVRRLRLVTADGRLRDVTAGTDPDLFWAVR-GGKDNFGLVVGMEVDLFPV 194
Score = 34.7 bits (78), Expect = 0.63
Identities = 16/30 (53%), Positives = 19/30 (63%)
Query: 496 YFKSNFERLARVKAQVDPSNFFRHEQSIPP 525
Y ++F RL VKAQ DP N FR +IPP
Sbjct: 413 YTPADFARLRAVKAQYDPDNMFRVNFNIPP 442
>FAS5_RHOFA (P46377) Hypothetical 47.9 kDa oxidoreductase in
fasciation locus (ORF5)
Length = 438
Score = 55.8 bits (133), Expect = 3e-07
Identities = 44/182 (24%), Positives = 80/182 (43%), Gaps = 6/182 (3%)
Query: 79 KPNLIVTPQELSHVQAAITCSVKQGLQVRVRSGGHDYEGLSYISNVPFLIIDLSNFRSIS 138
KP ++V P+ ++ VQ A+ + + L + VR GH G +++D+ F ++
Sbjct: 26 KPPVVVVPRTVADVQEALRYTAARNLSLAVRGSGHSTYGQCQADGG--VVLDMKRFNTVH 83
Query: 139 IDIEDESAWVQSGATLGELYYAIANQSNVHAFPAGSCPTVGIGGHFSGGGFSTIFRKHGL 198
D+ A + +G ++ A ++ T +GG S GGF GL
Sbjct: 84 -DVRSGQATIDAGVRWSDVVAATLSRQQTPPVLTDYLGTT-VGGTLSVGGFGGSSHGFGL 141
Query: 199 AADHVIDAQIIDVNGKILNRN-LMGEDLFWAIKGGGGSSFGVITSWKVKLVHVPPKVTIF 257
D+V ++ +G + + +LF A++GG G FGVI + ++L V +
Sbjct: 142 QTDNVDSLAVVTGSGDFRECSAVSNSELFDAVRGGLG-QFGVIVNATIRLTAAHESVRQY 200
Query: 258 DL 259
L
Sbjct: 201 KL 202
>CKX3_ARATH (Q9LTS3) Cytokinin dehydrogenase 3 precursor (EC
1.5.99.12) (Cytokinin oxidase 3) (CKO 3)
Length = 523
Score = 53.5 bits (127), Expect = 1e-06
Identities = 69/307 (22%), Positives = 131/307 (42%), Gaps = 30/307 (9%)
Query: 6 TKLALLSITIFISIFLETSISFDIE-KPFLHCFSSIVRNSNPTEEIVLTQNSSSYESLLQ 64
+++ L++ITI I I L T I+ + +P+ +I+ ++ + LT +SSS ES
Sbjct: 8 SQVRLIAITIVIIITLSTPITTNTSPQPW-----NILSHNEFAGK--LTSSSSSVESAA- 59
Query: 65 SSIRNTRFLDSSVLKPNLIVTPQELSHVQAAITCSVKQGLQVRVRSGGHDYEGLSYISNV 124
T F + + P+ ++ P + + I S L + + GH + S
Sbjct: 60 -----TDFGHVTKIFPSAVLIPSSVEDITDLIKLSFDSQLSFPLAARGHGHSHRGQASAK 114
Query: 125 PFLIIDLSNFRSISIDIEDESAWVQSGATLGELYYAIANQSNVHAFPAGSCPT------- 177
+++++ + + I+ + L+ + N++ G P
Sbjct: 115 DGVVVNMRSMVNRDRGIKVSRTCLYVDVDAAWLWIEVLNKT----LELGLTPVSWTDYLY 170
Query: 178 VGIGGHFSGGGFSTIFRKHGLAADHVIDAQIIDVNGKILNRNL-MGEDLFWAIKGGGGSS 236
+ +GG S GG S ++G +V++ +I G+I + M DLF+A+ GG G
Sbjct: 171 LTVGGTLSNGGISGQTFRYGPQITNVLEMDVITGKGEIATCSKDMNSDLFFAVLGGLG-Q 229
Query: 237 FGVITSWKVKLVHVPPKVTIFDLPKKLDQNVSEIFQKWQTIAHKLPGELFLHSVMGVSNS 296
FG+IT ++KL P + + L + SE + + + K G FL + V +
Sbjct: 230 FGIITRARIKLEVAPKRAKWL---RFLYIDFSEFTRDQERVISKTDGVDFLEGSIMVDHG 286
Query: 297 PKHGGKS 303
P +S
Sbjct: 287 PPDNWRS 293
>CKX2_ARATH (Q9FUJ3) Cytokinin dehydrogenase 2 precursor (EC
1.5.99.12) (Cytokinin oxidase 2) (CKO 2)
Length = 501
Score = 49.7 bits (117), Expect = 2e-05
Identities = 46/224 (20%), Positives = 95/224 (41%), Gaps = 20/224 (8%)
Query: 50 IVLTQNSSSYESLLQSSIRNTRFLDSSVLK-------------PNLIVTPQELSHVQAAI 96
+++T++S+ + L S+ T D S++ P ++ P + + +
Sbjct: 14 LMITKSSNGIKIDLPKSLNLTLSTDPSIISAASHDFGNITTVTPGGVICPSSTADISRLL 73
Query: 97 TCSV--KQGLQVRVRSGGHDYEGLSYISNVPFLIIDLSNFRSISIDIEDESAWVQSGATL 154
+ K QV R GH G + +S +I++++ + + + + A V +G
Sbjct: 74 QYAANGKSTFQVAARGQGHSLNGQASVSGG--VIVNMTCITDVVVSKDKKYADVAAGTLW 131
Query: 155 GELYYAIANQSNVHAFPAGSCPTVGIGGHFSGGGFSTIFRKHGLAADHVIDAQIIDVNGK 214
++ A + V + +GG S GG ++G +V++ +I G+
Sbjct: 132 VDVLKKTA-EKGVSPVSWTDYLHITVGGTLSNGGIGGQVFRNGPLVSNVLELDVITGKGE 190
Query: 215 ILN-RNLMGEDLFWAIKGGGGSSFGVITSWKVKLVHVPPKVTIF 257
+L + +LF+ + GG G FG+IT ++ L H P + F
Sbjct: 191 MLTCSRQLNPELFYGVLGGLGQ-FGIITRARIVLDHAPKRAKWF 233
>DIM_PEA (P93472) Cell elongation protein diminuto
Length = 567
Score = 43.1 bits (100), Expect = 0.002
Identities = 36/125 (28%), Positives = 62/125 (48%), Gaps = 5/125 (4%)
Query: 129 IDLSNFRSI-SIDIEDESAWVQSGATLGELYYAIANQSNVHAFPAGSCPTVGIGGHFSGG 187
+DLS FR+I ID E A V+ +G++ + N+ + +GG +G
Sbjct: 109 VDLSPFRNILDIDKERMIARVEPLVNMGQIT-RVTVPMNLALAVVAELDDLTVGGLINGY 167
Query: 188 GFSTIFRKHGLAADHVIDAQIIDVNGKILNRNLMGE--DLFWAIKGGGGSSFGVITSWKV 245
G K+GL +D V+ +II +G ++ E DLF+AI G + G++ + +V
Sbjct: 168 GIEGSSHKYGLFSDTVVAFEIILADGSLVKATKDNEYSDLFYAIPWSQG-TLGLLVAAEV 226
Query: 246 KLVHV 250
KL+ +
Sbjct: 227 KLIPI 231
>GGLO_MOUSE (P58710) L-gulonolactone oxidase (EC 1.1.3.8) (LGO)
(L-gulono-gamma-lactone oxidase) (GLO)
Length = 439
Score = 41.2 bits (95), Expect = 0.007
Identities = 31/137 (22%), Positives = 62/137 (44%), Gaps = 4/137 (2%)
Query: 80 PNLIVTPQELSHVQAAITCSVKQGLQVRVRSGGHDYEGLSYISNVPFLIIDLSNFRSISI 139
P + P + V+ + + +Q +V+V GGH ++ F+I R + +
Sbjct: 20 PEMYYQPTSVGEVREVLALARQQNKKVKVVGGGHSPSDIACTDG--FMIHMGKMNRVLQV 77
Query: 140 DIEDESAWVQSGATLGELYYAIANQSNVHAFPAGSCPTVGIGGHFSGGGFSTIFRKHGLA 199
D E + V++G L +L+ + ++ + G+ V +GG G +T KHG+
Sbjct: 78 DKEKKQVTVEAGILLTDLHPQL-DKHGLALSNLGAVSDVTVGGVIGSGTHNTGI-KHGIL 135
Query: 200 ADHVIDAQIIDVNGKIL 216
A V+ ++ +G +L
Sbjct: 136 ATQVVALTLMKADGTVL 152
>GLCD_ECOLI (P52075) Glycolate oxidase subunit glcD
Length = 499
Score = 40.0 bits (92), Expect = 0.015
Identities = 48/202 (23%), Positives = 84/202 (40%), Gaps = 12/202 (5%)
Query: 79 KPNLIVTPQELSHVQAAITCSVKQGLQVRVRSGGHDYEGLSYISNVPFLIIDLSNFRSI- 137
+P L+V P+++ V A + + + V R G G + L++ ++ F+ I
Sbjct: 55 RPLLVVLPKQMEQVTAILAVCHRLRVPVVTRGAGTGLSGGALPLEKGVLLV-MARFKEIL 113
Query: 138 SIDIEDESAWVQSGATLGELYYAIANQSNVHAFPAGSCPTVGIGGHFSGGGFSTIFRKHG 197
I+ A VQ G + A+A + +A S IGG+ + K+G
Sbjct: 114 DINPVGRRARVQPGVRNLAISQAVAPHNLYYAPDPSSQIACSIGGNVAENAGGVHCLKYG 173
Query: 198 LAADHVIDAQIIDVNGKILN-----RNLMGEDLFWAIKGGGGSSFGVITSWKVKLVHVPP 252
L +++ ++ ++G+ L + G DL A+ G GV T VKL+ PP
Sbjct: 174 LTVHNLLKIEVQTLDGEALTLGSDALDSPGFDLL-ALFTGSEGMLGVTTEVTVKLLPKPP 232
Query: 253 KVTI----FDLPKKLDQNVSEI 270
+ FD +K V +I
Sbjct: 233 VARVLLASFDSVEKAGLAVGDI 254
>ALO_NEUCR (Q7SGY1) Putative D-arabinono-1,4-lactone oxidase (EC
1.1.3.37) (ALO) (L-galactono-gamma-lactone oxidase)
Length = 556
Score = 39.7 bits (91), Expect = 0.020
Identities = 52/199 (26%), Positives = 89/199 (44%), Gaps = 15/199 (7%)
Query: 80 PNLIVTPQELSHVQAAITCSVKQGLQVRVRSGGHDYEGLSYISNVPFLIIDLSNFRS-IS 138
P L + P+ +S +Q + + +V GH ++ S+ +++L NF IS
Sbjct: 51 PELYIQPESVSEIQKVVRLARHARRRVTTTGCGHSPSDITCTSS---WLVNLDNFNKVIS 107
Query: 139 IDIEDESAWVQSGATLGELYYAIANQSNVHAFPA-GSCPTVGIGGHFSGGGFSTIFRKHG 197
+D VQ+G L +L + + A P+ GS I G S G + KHG
Sbjct: 108 VDHLTGLVVVQAGIRLYQLSDELDRRGL--ALPSLGSINEQSIAGAISTGTHGSGI-KHG 164
Query: 198 LAADHVIDAQIIDVNGKILNRNLMGE-DLFWAIKGGGGSSFGVITSWKVKLVHVPPKVTI 256
L + + + +I NG+ L+ + + DLF A G + G+IT K V P ++
Sbjct: 165 LVGESITELKITLANGETLSCSPEDKPDLFRAALISLG-ALGIITEVTFKAV---PAFSL 220
Query: 257 FDLPKKLDQNVSEIFQKWQ 275
+ +D + IF+KW+
Sbjct: 221 -AWSQAIDSD-KRIFEKWE 237
>GGLO_RAT (P10867) L-gulonolactone oxidase (EC 1.1.3.8) (LGO)
(L-gulono-gamma-lactone oxidase) (GLO)
Length = 439
Score = 39.3 bits (90), Expect = 0.025
Identities = 30/137 (21%), Positives = 62/137 (44%), Gaps = 4/137 (2%)
Query: 80 PNLIVTPQELSHVQAAITCSVKQGLQVRVRSGGHDYEGLSYISNVPFLIIDLSNFRSISI 139
P + P + V+ + + +Q +V+V GGH ++ F+I R + +
Sbjct: 20 PEVYYQPTSVEEVREVLALAREQKKKVKVVGGGHSPSDIACTDG--FMIHMGKMNRVLQV 77
Query: 140 DIEDESAWVQSGATLGELYYAIANQSNVHAFPAGSCPTVGIGGHFSGGGFSTIFRKHGLA 199
D E + V++G L +L+ + ++ + G+ V + G G +T KHG+
Sbjct: 78 DKEKKQITVEAGILLADLHPQL-DEHGLAMSNLGAVSDVTVAGVIGSGTHNTGI-KHGIL 135
Query: 200 ADHVIDAQIIDVNGKIL 216
A V+ ++ +G++L
Sbjct: 136 ATQVVALTLMTADGEVL 152
>DIM_ARATH (Q39085) Cell elongation protein DIMINUTO (Cell
elongation protein Dwarf1)
Length = 561
Score = 39.3 bits (90), Expect = 0.025
Identities = 35/139 (25%), Positives = 66/139 (47%), Gaps = 5/139 (3%)
Query: 129 IDLSNFRSI-SIDIEDESAWVQSGATLGELYYAIANQSNVHAFPAGSCPTVGIGGHFSGG 187
+DL FR+I I+ E +A V+ +G++ A N+ + +GG +G
Sbjct: 110 VDLGEFRNILEINKEKMTARVEPLVNMGQISRATVPM-NLSLAVVAELDDLTVGGLINGY 168
Query: 188 GFSTIFRKHGLAADHVIDAQIIDVNGKILNRNLMGE--DLFWAIKGGGGSSFGVITSWKV 245
G +GL AD V +I+ G+++ E DL++AI G + G++ + ++
Sbjct: 169 GIEGSSHIYGLFADTVEAYEIVLAGGELVRATRDNEYSDLYYAIPWSQG-TLGLLVAAEI 227
Query: 246 KLVHVPPKVTIFDLPKKLD 264
+L+ V + + +P K D
Sbjct: 228 RLIKVKEYMRLTYIPVKGD 246
>CKX4_ARATH (Q9FUJ2) Cytokinin dehydrogenase 4 precursor (EC
1.5.99.12) (Cytokinin oxidase 4) (CKO 4)
Length = 524
Score = 38.5 bits (88), Expect = 0.043
Identities = 22/78 (28%), Positives = 40/78 (51%), Gaps = 2/78 (2%)
Query: 178 VGIGGHFSGGGFSTIFRKHGLAADHVIDAQIIDVNGKILNRN-LMGEDLFWAIKGGGGSS 236
+ +GG S G +HG +V + +I G+++ + + +LF+ + GG G
Sbjct: 177 LSVGGTLSNAGIGGQTFRHGPQISNVHELDVITGKGEMMTCSPKLNPELFYGVLGGLGQ- 235
Query: 237 FGVITSWKVKLVHVPPKV 254
FG+IT ++ L H P +V
Sbjct: 236 FGIITRARIALDHAPTRV 253
>CKX1_ORYSA (Q9LDE6) Probable cytokinin dehydrogenase precursor (EC
1.5.99.12) (Cytokinin oxidase) (CKO)
Length = 532
Score = 37.7 bits (86), Expect = 0.074
Identities = 46/193 (23%), Positives = 83/193 (42%), Gaps = 19/193 (9%)
Query: 70 TRFLDSSVLKPNLIVTPQELSHVQAAITCSVKQGLQVRVRSGGHDYEGLSYISN-VPFLI 128
T L ++VL P +P +++ + A + + V R GH G + + V +
Sbjct: 65 TAALPAAVLFPG---SPGDVAELLRAAYAAPGRPFTVSFRGRGHSTMGQALAAGGVVVHM 121
Query: 129 IDLSNFRSISIDIEDESAWVQSGATLGELYYAIANQSNVHAFPAGSCPT-------VGIG 181
+ + I++ + A+V +G GE + ++ A G P + +G
Sbjct: 122 QSMGGGGAPRINVSADGAYVDAG---GEQLWVDVLRA---ALARGVAPRSWTDYLHLTVG 175
Query: 182 GHFSGGGFSTIFRKHGLAADHVIDAQIIDVNGKILN-RNLMGEDLFWAIKGGGGSSFGVI 240
G S G S +HG +V++ +I +G+ + + DLF A+ GG G FGVI
Sbjct: 176 GTLSNAGVSGQTYRHGPQISNVLELDVITGHGETVTCSKAVNSDLFDAVLGGLG-QFGVI 234
Query: 241 TSWKVKLVHVPPK 253
T +V + P +
Sbjct: 235 TRARVAVEPAPAR 247
>CKX1_MAIZE (Q9T0N8) Cytokinin dehydrogenase 1 precursor (EC
1.5.99.12) (Cytokinin oxidase 1) (CKO 1)
Length = 534
Score = 37.0 bits (84), Expect = 0.13
Identities = 23/75 (30%), Positives = 39/75 (51%), Gaps = 2/75 (2%)
Query: 180 IGGHFSGGGFSTIFRKHGLAADHVIDAQIIDVNGKILN-RNLMGEDLFWAIKGGGGSSFG 238
+GG S G S +HG +V++ +I +G+++ + DLF A+ GG G FG
Sbjct: 175 VGGTLSNAGISGQAFRHGPQISNVLEMDVITGHGEMVTCSKQLNADLFDAVLGGLG-QFG 233
Query: 239 VITSWKVKLVHVPPK 253
VIT ++ + P +
Sbjct: 234 VITRARIAVEPAPAR 248
>YDIJ_HAEIN (Q57252) Protein HI1163
Length = 1027
Score = 34.7 bits (78), Expect = 0.63
Identities = 34/176 (19%), Positives = 71/176 (40%), Gaps = 19/176 (10%)
Query: 44 SNPTEEIVLTQNSSSYESLLQSSIRNTRFLDSSVLKPNLIVTPQELSHVQAAITCSVKQG 103
+N + + L ++S Y+ L Q+ +L P + ++ + Q
Sbjct: 33 TNYADRLSLATDNSVYQQLPQA-----------ILFPKTVA---DIVRITKLANLPEYQS 78
Query: 104 LQVRVRSGGHDYEGLSYISNVPFLIIDLSNFRSISIDIEDESAWV--QSGATLGELYYAI 161
+ R GG G S +N+ I+DLS + +++ + WV Q+G +L +
Sbjct: 79 ISFTPRGGGTGTNGQSINNNI---IVDLSRHMTAILELNVKERWVRVQAGVVKDQLNQFL 135
Query: 162 ANQSNVHAFPAGSCPTVGIGGHFSGGGFSTIFRKHGLAADHVIDAQIIDVNGKILN 217
A + +GG + ++G ++HV+ + + +NG+IL+
Sbjct: 136 KPHGLFFAPELSTSNRATLGGMINTDASGQGSLQYGKTSNHVLALRAVLINGEILD 191
>MURB_SYNEL (Q8DLV6) UDP-N-acetylenolpyruvoylglucosamine reductase
(EC 1.1.1.158) (UDP-N-acetylmuramate dehydrogenase)
Length = 301
Score = 33.5 bits (75), Expect = 1.4
Identities = 32/133 (24%), Positives = 55/133 (41%), Gaps = 7/133 (5%)
Query: 83 IVTPQELSHVQAAITCSVKQGLQVRVRSGGHDYEGLSYISNVPFLIIDLSNFRSISIDIE 142
+V P+ + +QAA + ++GL + V G + L VP L+I + R ++ D+E
Sbjct: 34 LVEPRTIPELQAAYIWAQEEGLPITVLGAGSNL--LVSDRGVPGLVISTKHLRYLTTDLE 91
Query: 143 DESAWVQSGATLGELYYAIANQSNVHAFPAGSCPTVGIGGHFSGGGFSTIFRKHGLAADH 202
V +G L +L + A G VGI G G A
Sbjct: 92 TGQLTVGAGYPLPKLAHHTAKLG-----WRGLEWVVGIPGTVGGAVVMNAGAHGSCTAQR 146
Query: 203 VIDAQIIDVNGKI 215
++ A I++ +G +
Sbjct: 147 LVSAVILEPDGTL 159
>DUT_RHILO (Q98C10) Deoxyuridine 5'-triphosphate nucleotidohydrolase
(EC 3.6.1.23) (dUTPase) (dUTP pyrophosphatase)
Length = 161
Score = 33.1 bits (74), Expect = 1.8
Identities = 27/109 (24%), Positives = 49/109 (44%), Gaps = 3/109 (2%)
Query: 82 LIVTPQELSHVQAAITCSVKQGL--QVRVRSGGHDYEGLSYISNVPFLIIDL-SNFRSIS 138
L++ P + S V + + +G+ QVR RSG GL+ +++ + D + +
Sbjct: 51 LLILPGKRSLVPTGLILEIPEGMEGQVRPRSGLAFKHGLTVLNSPGTVDSDYRGEVKVLL 110
Query: 139 IDIEDESAWVQSGATLGELYYAIANQSNVHAFPAGSCPTVGIGGHFSGG 187
I++ DE V G + ++ +A+ Q+ V G GG S G
Sbjct: 111 INLGDEDFAVTRGMRIAQIVFAVVTQAAVEERSLAGGTARGSGGFGSTG 159
>EMB8_PICGL (Q40863) Embryogenesis-associated protein EMB8
Length = 457
Score = 32.7 bits (73), Expect = 2.4
Identities = 20/67 (29%), Positives = 35/67 (51%), Gaps = 3/67 (4%)
Query: 379 NEPI-PRAGLEGLWKMLLEEDSPLLILTPYGGRMSEISESETPFPHRNKSIFGIQYLVNW 437
N+PI P G+ W+ + E + LL++TP GG + ++ + PF ++YL
Sbjct: 360 NDPIAPSRGIP--WEDIKENPNCLLVVTPNGGHLGWVAGDDAPFGAPWTDPLVMEYLEVL 417
Query: 438 DKNEETK 444
+KN+ K
Sbjct: 418 EKNQIEK 424
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.137 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 64,101,234
Number of Sequences: 164201
Number of extensions: 2822320
Number of successful extensions: 6227
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 19
Number of HSP's that attempted gapping in prelim test: 6140
Number of HSP's gapped (non-prelim): 83
length of query: 528
length of database: 59,974,054
effective HSP length: 115
effective length of query: 413
effective length of database: 41,090,939
effective search space: 16970557807
effective search space used: 16970557807
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0437b.11


