

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
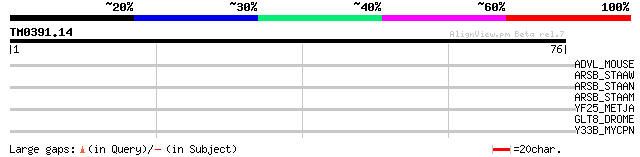
Query= TM0391.14
(76 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
ADVL_MOUSE (O88398) Advillin (p92) (Actin-binding protein DOC6) 28 3.3
ARSB_STAAW (Q8NW09) Arsenical pump membrane protein (Arsenic eff... 28 4.3
ARSB_STAAN (P63620) Arsenical pump membrane protein (Arsenic eff... 28 4.3
ARSB_STAAM (P63619) Arsenical pump membrane protein (Arsenic eff... 28 4.3
YF25_METJA (Q58920) Hypothetical protein MJ1525 27 7.3
GLT8_DROME (Q9VUT6) Polypeptide N-acetylgalactosaminyltransferas... 27 7.3
Y33B_MYCPN (P75302) Hypothetical protein MG335.2 homolog (P01_or... 27 9.5
>ADVL_MOUSE (O88398) Advillin (p92) (Actin-binding protein DOC6)
Length = 819
Score = 28.5 bits (62), Expect = 3.3
Identities = 12/22 (54%), Positives = 13/22 (58%)
Query: 23 LELKDVEYQWHAFIKNGNTVYV 44
LEL VEYQWH F G+ V
Sbjct: 405 LELVPVEYQWHGFFYGGDCYLV 426
>ARSB_STAAW (Q8NW09) Arsenical pump membrane protein (Arsenic efflux
pump protein)
Length = 429
Score = 28.1 bits (61), Expect = 4.3
Identities = 12/35 (34%), Positives = 20/35 (56%)
Query: 22 VLELKDVEYQWHAFIKNGNTVYVGLLWIPAVLIYL 56
VL KDV+ W + K G + + +L+I + +YL
Sbjct: 390 VLTQKDVKISWGTYFKTGIIITIPVLFITLIGLYL 424
>ARSB_STAAN (P63620) Arsenical pump membrane protein (Arsenic efflux
pump protein)
Length = 429
Score = 28.1 bits (61), Expect = 4.3
Identities = 12/35 (34%), Positives = 20/35 (56%)
Query: 22 VLELKDVEYQWHAFIKNGNTVYVGLLWIPAVLIYL 56
VL KDV+ W + K G + + +L+I + +YL
Sbjct: 390 VLTQKDVKISWGTYFKTGIIITIPVLFITLIGLYL 424
>ARSB_STAAM (P63619) Arsenical pump membrane protein (Arsenic efflux
pump protein)
Length = 429
Score = 28.1 bits (61), Expect = 4.3
Identities = 12/35 (34%), Positives = 20/35 (56%)
Query: 22 VLELKDVEYQWHAFIKNGNTVYVGLLWIPAVLIYL 56
VL KDV+ W + K G + + +L+I + +YL
Sbjct: 390 VLTQKDVKISWGTYFKTGIIITIPVLFITLIGLYL 424
>YF25_METJA (Q58920) Hypothetical protein MJ1525
Length = 933
Score = 27.3 bits (59), Expect = 7.3
Identities = 13/38 (34%), Positives = 21/38 (55%), Gaps = 1/38 (2%)
Query: 28 VEYQWHAFIKNGNTVYVGLLWIPAVLIYLMDIQIWYAI 65
+ Y WH+ I T+ W+PAVL L+ I I++ +
Sbjct: 131 IYYIWHS-IDLTVTIMNAAFWVPAVLGMLLGIPIYFVV 167
>GLT8_DROME (Q9VUT6) Polypeptide N-acetylgalactosaminyltransferase 8
(EC 2.4.1.41) (Protein-UDP
acetylgalactosaminyltransferase 8)
(UDP-GalNAc:polypeptide
N-acetylgalactosaminyltransferase 8) (pp-GaNTase 8)
Length = 590
Score = 27.3 bits (59), Expect = 7.3
Identities = 13/40 (32%), Positives = 19/40 (47%)
Query: 36 IKNGNTVYVGLLWIPAVLIYLMDIQIWYAIYSSLKMKLWI 75
I NG +G W + Y ++IW A L +KLW+
Sbjct: 296 IMNGGLFAIGREWFSELGGYDKGLKIWGAEQFELSLKLWL 335
>Y33B_MYCPN (P75302) Hypothetical protein MG335.2 homolog
(P01_orf341)
Length = 341
Score = 26.9 bits (58), Expect = 9.5
Identities = 14/47 (29%), Positives = 23/47 (48%), Gaps = 3/47 (6%)
Query: 29 EYQWHAFIKNGNTVYVGLLWIPAVLIYLMDIQIWYAIYSSLKMKLWI 75
+Y W+ I NT + L + V ++ DI +W+ I + LWI
Sbjct: 168 QYIWNIVI---NTAFFKALDLQFVNRFIEDIAVWFPIMFKAQKVLWI 211
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.333 0.146 0.478
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,639,953
Number of Sequences: 164201
Number of extensions: 280078
Number of successful extensions: 1118
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 1113
Number of HSP's gapped (non-prelim): 7
length of query: 76
length of database: 59,974,054
effective HSP length: 52
effective length of query: 24
effective length of database: 51,435,602
effective search space: 1234454448
effective search space used: 1234454448
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0391.14


