

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0391.11
(490 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
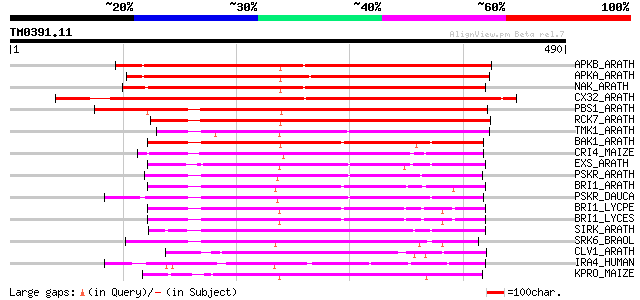
Score E
Sequences producing significant alignments: (bits) Value
APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor ... 427 e-119
APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor ... 424 e-118
NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK ... 422 e-117
CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx3... 361 2e-99
PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC 2.7... 335 2e-91
RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase RLC... 320 7e-87
TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precur... 212 2e-54
BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated rec... 209 1e-53
CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4 pr... 206 8e-53
EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase E... 204 4e-52
PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (... 202 2e-51
BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC ... 201 3e-51
PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.... 195 2e-49
BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.... 194 6e-49
BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precurso... 194 6e-49
SIRK_ARATH (O64483) Senescence-induced receptor-like serine/thre... 187 4e-47
SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase rec... 184 4e-46
CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (... 176 1e-43
IRA4_HUMAN (Q9NWZ3) Interleukin-1 receptor-associated kinase-4 (... 174 4e-43
KPRO_MAIZE (P17801) Putative receptor protein kinase ZmPK1 precu... 174 5e-43
>APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor (EC
2.7.1.-)
Length = 412
Score = 427 bits (1097), Expect = e-119
Identities = 211/335 (62%), Positives = 267/335 (78%), Gaps = 5/335 (1%)
Query: 94 ESSTTTSNEESIASTPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFK 153
+S + S+ SI + P+ E L+ + +L+ FTF LK ATRNFRP+S+LGEGGFG VFK
Sbjct: 27 DSLGSKSSSVSIRTNPRTEGEILQ-SPNLKSFTFAELKAATRNFRPDSVLGEGGFGSVFK 85
Query: 154 GWIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIED 213
GWI+E KPGTG+ +AVK LN +G QGH+EWLAE+NYLG HPNLVKLIG+C+ED
Sbjct: 86 GWIDEQTLTASKPGTGVVIAVKKLNQDGWQGHQEWLAEVNYLGQFSHPNLVKLIGYCLED 145
Query: 214 DQRLLVYEFMPRGSLENHLFRRPL---PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRD 270
+ RLLVYEFMPRGSLENHLFRR PL W++R+K+ALGAAKGLAFLH +++ +IYRD
Sbjct: 146 EHRLLVYEFMPRGSLENHLFRRGSYFQPLSWTLRLKVALGAAKGLAFLH-NAETSVIYRD 204
Query: 271 FKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDV 330
FKTSNILLD+EYNAKLSDFGLAKDGP G+K+HVSTR+MGTYGYAAPEY+ TGHL++KSDV
Sbjct: 205 FKTSNILLDSEYNAKLSDFGLAKDGPTGDKSHVSTRIMGTYGYAAPEYLATGHLTTKSDV 264
Query: 331 YSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQ 390
YS+GVVLLE+L+GRR++DK RP GE LVEWARP+L ++R F++ID RL+ +S++ A
Sbjct: 265 YSYGVVLLEVLSGRRAVDKNRPPGEQKLVEWARPLLANKRKLFRVIDNRLQDQYSMEEAC 324
Query: 391 KAAQLAAQCLSRDPKARPLMSEVVHTLKPLPNLKD 425
K A LA +CL+ + K RP M+EVV L+ + L +
Sbjct: 325 KVATLALRCLTFEIKLRPNMNEVVSHLEHIQTLNE 359
>APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor (EC
2.7.1.-)
Length = 410
Score = 424 bits (1089), Expect = e-118
Identities = 209/323 (64%), Positives = 261/323 (80%), Gaps = 5/323 (1%)
Query: 104 SIASTPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAP 163
S+ +P+ E L+ + +L+ F+F LK ATRNFRP+S+LGEGGFGCVFKGWI+E
Sbjct: 36 SVRPSPRTEGEILQ-SPNLKSFSFAELKSATRNFRPDSVLGEGGFGCVFKGWIDEKSLTA 94
Query: 164 VKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFM 223
+PGTGL +AVK LN +G QGH+EWLAE+NYLG H +LVKLIG+C+ED+ RLLVYEFM
Sbjct: 95 SRPGTGLVIAVKKLNQDGWQGHQEWLAEVNYLGQFSHRHLVKLIGYCLEDEHRLLVYEFM 154
Query: 224 PRGSLENHLFRRPL---PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDA 280
PRGSLENHLFRR L PL W +R+K+ALGAAKGLAFLH R +IYRDFKTSNILLD+
Sbjct: 155 PRGSLENHLFRRGLYFQPLSWKLRLKVALGAAKGLAFLHSSETR-VIYRDFKTSNILLDS 213
Query: 281 EYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEM 340
EYNAKLSDFGLAKDGP G+K+HVSTRVMGT+GYAAPEY+ TGHL++KSDVYSFGVVLLE+
Sbjct: 214 EYNAKLSDFGLAKDGPIGDKSHVSTRVMGTHGYAAPEYLATGHLTTKSDVYSFGVVLLEL 273
Query: 341 LTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCL 400
L+GRR++DK RP+GE NLVEWA+P L ++R F++ID RL+ +S++ A K A L+ +CL
Sbjct: 274 LSGRRAVDKNRPSGERNLVEWAKPYLVNKRKIFRVIDNRLQDQYSMEEACKVATLSLRCL 333
Query: 401 SRDPKARPLMSEVVHTLKPLPNL 423
+ + K RP MSEVV L+ + +L
Sbjct: 334 TTEIKLRPNMSEVVSHLEHIQSL 356
>NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK (EC
2.7.1.37)
Length = 389
Score = 422 bits (1084), Expect = e-117
Identities = 207/324 (63%), Positives = 260/324 (79%), Gaps = 5/324 (1%)
Query: 100 SNEESIASTPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEEN 159
S+ S + P+ E L+ A+ L+ F+ + LK ATRNFRP+S++GEGGFGCVFKGWI+E+
Sbjct: 32 SSTASFSYMPRTEGEILQNAN-LKNFSLSELKSATRNFRPDSVVGEGGFGCVFKGWIDES 90
Query: 160 GTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLV 219
AP KPGTG+ +AVK LN G QGH+EWLAE+NYLG L HPNLVKLIG+C+E++ RLLV
Sbjct: 91 SLAPSKPGTGIVIAVKRLNQEGFQGHREWLAEINYLGQLDHPNLVKLIGYCLEEEHRLLV 150
Query: 220 YEFMPRGSLENHLFRRPL---PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNI 276
YEFM RGSLENHLFRR PL W+ R+++ALGAA+GLAFLH ++Q +IYRDFK SNI
Sbjct: 151 YEFMTRGSLENHLFRRGTFYQPLSWNTRVRMALGAARGLAFLH-NAQPQVIYRDFKASNI 209
Query: 277 LLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVV 336
LLD+ YNAKLSDFGLA+DGP G+ +HVSTRVMGT GYAAPEY+ TGHLS KSDVYSFGVV
Sbjct: 210 LLDSNYNAKLSDFGLARDGPMGDNSHVSTRVMGTQGYAAPEYLATGHLSVKSDVYSFGVV 269
Query: 337 LLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLA 396
LLE+L+GRR+IDK +P GEHNLV+WARP L ++R +++DPRL+G +S+ A K A LA
Sbjct: 270 LLELLSGRRAIDKNQPVGEHNLVDWARPYLTNKRRLLRVMDPRLQGQYSLTRALKIAVLA 329
Query: 397 AQCLSRDPKARPLMSEVVHTLKPL 420
C+S D K+RP M+E+V T++ L
Sbjct: 330 LDCISIDAKSRPTMNEIVKTMEEL 353
>CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx32,
chloroplast precursor (EC 2.7.1.37)
Length = 419
Score = 361 bits (927), Expect = 2e-99
Identities = 194/408 (47%), Positives = 264/408 (64%), Gaps = 20/408 (4%)
Query: 41 SFFGSCIPSRPKVDSSISGTSTHNGNSIKSVITQSEENKSGFEKNTEETEAPPESSTTTS 100
SFF S PS+ + S + + NG S T A SS
Sbjct: 6 SFFSSSSPSKTGLHSHATTNNHSNGTEFSST-----------------TGATTNSSVGQQ 48
Query: 101 NEESIASTPKLF-SEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEEN 159
++ S ST + S +L + +L+ + F LK AT+NF+P+S+LG+GGFG V++GW++
Sbjct: 49 SQFSDISTGIISDSGKLLESPNLKVYNFLDLKTATKNFKPDSMLGQGGFGKVYRGWVDAT 108
Query: 160 GTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLV 219
AP + G+G+ VA+K LN QG EW +E+N+LG L H NLVKL+G+C ED + LLV
Sbjct: 109 TLAPSRVGSGMIVAIKRLNSESVQGFAEWRSEVNFLGMLSHRNLVKLLGYCREDKELLLV 168
Query: 220 YEFMPRGSLENHLFRRPLPLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLD 279
YEFMP+GSLE+HLFRR P PW +R+KI +GAA+GLAFLH QR +IYRDFK SNILLD
Sbjct: 169 YEFMPKGSLESHLFRRNDPFPWDLRIKIVIGAARGLAFLHS-LQREVIYRDFKASNILLD 227
Query: 280 AEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLE 339
+ Y+AKLSDFGLAK GP EK+HV+TR+MGTYGYAAPEY+ TGHL KSDV++FGVVLLE
Sbjct: 228 SNYDAKLSDFGLAKLGPADEKSHVTTRIMGTYGYAAPEYMATGHLYVKSDVFAFGVVLLE 287
Query: 340 MLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQC 399
++TG + + KRP G+ +LV+W RP L ++ QI+D ++G ++ K A + A++ C
Sbjct: 288 IMTGLTAHNTKRPRGQESLVDWLRPELSNKHRVKQIMDKGIKGQYTTKVATEMARITLSC 347
Query: 400 LSRDPKARPLMSEVVHTLKPLPNLKDMAISSYHFQTARVDRTMSMPNH 447
+ DPK RP M EVV L+ + L + S Q A + + S P+H
Sbjct: 348 IEPDPKNRPHMKEVVEVLEHIQGLNVVPNRSSTKQ-AVANSSRSSPHH 394
>PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC
2.7.1.37) (AvrPphB susceptible protein 1)
Length = 456
Score = 335 bits (859), Expect = 2e-91
Identities = 179/356 (50%), Positives = 232/356 (64%), Gaps = 18/356 (5%)
Query: 76 EENKSGFEKNTEETEAPPESSTTTSNEESIASTPKLFSEELKVAS------SLRKFTFNG 129
+E+ G +K ++ T + S + E+ + T EL + + F F
Sbjct: 19 DESNHGQKKQSQPTVSNNISGLPSGGEKLSSKTNGGSKRELLLPRDGLGQIAAHTFAFRE 78
Query: 130 LKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWL 189
L AT NF P++ LGEGGFG V+KG ++ TG VAVK L+ NG QG++E+L
Sbjct: 79 LAAATMNFHPDTFLGEGGFGRVYKGRLDS---------TGQVVAVKQLDRNGLQGNREFL 129
Query: 190 AELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRPLP---LPWSIRMK 246
E+ L L HPNLV LIG+C + DQRLLVYEFMP GSLE+HL P L W++RMK
Sbjct: 130 VEVLMLSLLHHPNLVNLIGYCADGDQRLLVYEFMPLGSLEDHLHDLPPDKEALDWNMRMK 189
Query: 247 IALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTR 306
IA GAAKGL FLH+ + P+IYRDFK+SNILLD ++ KLSDFGLAK GP G+K+HVSTR
Sbjct: 190 IAAGAAKGLEFLHDKANPPVIYRDFKSSNILLDEGFHPKLSDFGLAKLGPTGDKSHVSTR 249
Query: 307 VMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVL 366
VMGTYGY APEY MTG L+ KSDVYSFGVV LE++TGR++ID + P+GE NLV WARP+
Sbjct: 250 VMGTYGYCAPEYAMTGQLTVKSDVYSFGVVFLELITGRKAIDSEMPHGEQNLVAWARPLF 309
Query: 367 GHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLPN 422
RR F ++ DPRL+G F + +A +A+ C+ RPL+++VV L L N
Sbjct: 310 NDRRKFIKLADPRLKGRFPTRALYQALAVASMCIQEQAATRPLIADVVTALSYLAN 365
>RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase
RLCKVII (EC 2.7.1.37)
Length = 423
Score = 320 bits (819), Expect = 7e-87
Identities = 167/303 (55%), Positives = 209/303 (68%), Gaps = 12/303 (3%)
Query: 125 FTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQG 184
FTF L AT NFR + LGEGGFG VFKG IE+ VA+K L+ NG QG
Sbjct: 91 FTFQELAEATGNFRSDCFLGEGGFGKVFKGTIEK---------LDQVVAIKQLDRNGVQG 141
Query: 185 HKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRPL---PLPW 241
+E++ E+ L HPNLVKLIGFC E DQRLLVYE+MP+GSLE+HL P PL W
Sbjct: 142 IREFVVEVLTLSLADHPNLVKLIGFCAEGDQRLLVYEYMPQGSLEDHLHVLPSGKKPLDW 201
Query: 242 SIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKT 301
+ RMKIA GAA+GL +LH+ P+IYRD K SNILL +Y KLSDFGLAK GP G+KT
Sbjct: 202 NTRMKIAAGAARGLEYLHDRMTPPVIYRDLKCSNILLGEDYQPKLSDFGLAKVGPSGDKT 261
Query: 302 HVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEW 361
HVSTRVMGTYGY AP+Y MTG L+ KSD+YSFGVVLLE++TGR++ID + + NLV W
Sbjct: 262 HVSTRVMGTYGYCAPDYAMTGQLTFKSDIYSFGVVLLELITGRKAIDNTKTRKDQNLVGW 321
Query: 362 ARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLP 421
ARP+ RR F +++DP L+G + V+G +A ++A C+ P RP++S+VV L L
Sbjct: 322 ARPLFKDRRNFPKMVDPLLQGQYPVRGLYQALAISAMCVQEQPTMRPVVSDVVLALNFLA 381
Query: 422 NLK 424
+ K
Sbjct: 382 SSK 384
>TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precursor
(EC 2.7.1.-)
Length = 942
Score = 212 bits (539), Expect = 2e-54
Identities = 116/302 (38%), Positives = 180/302 (59%), Gaps = 19/302 (6%)
Query: 130 LKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHN--GHQGHKE 187
L+ T NF +++LG GGFG V+KG + + G +AVK + + +G E
Sbjct: 581 LRSVTNNFSSDNILGSGGFGVVYKGELHD----------GTKIAVKRMENGVIAGKGFAE 630
Query: 188 WLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRP----LPLPWSI 243
+ +E+ L + H +LV L+G+C++ +++LLVYE+MP+G+L HLF PL W
Sbjct: 631 FKSEIAVLTKVRHRHLVTLLGYCLDGNEKLLVYEYMPQGTLSRHLFEWSEEGLKPLLWKQ 690
Query: 244 RMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHV 303
R+ +AL A+G+ +LH + + I+RD K SNILL + AK++DFGL + PEG K +
Sbjct: 691 RLTLALDVARGVEYLHGLAHQSFIHRDLKPSNILLGDDMRAKVADFGLVRLAPEG-KGSI 749
Query: 304 STRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEW-A 362
TR+ GT+GY APEY +TG +++K DVYSFGV+L+E++TGR+S+D+ +P +LV W
Sbjct: 750 ETRIAGTFGYLAPEYAVTGRVTTKVDVYSFGVILMELITGRKSLDESQPEESIHLVSWFK 809
Query: 363 RPVLGHRRMFFQIIDPRLE-GHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLP 421
R + F + ID ++ ++ A+LA C +R+P RP M V+ L L
Sbjct: 810 RMYINKEASFKKAIDTTIDLDEETLASVHTVAELAGHCCAREPYQRPDMGHAVNILSSLV 869
Query: 422 NL 423
L
Sbjct: 870 EL 871
>BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated
receptor kinase 1 precursor (EC 2.7.1.37)
(BRI1-associated receptor kinase 1) (Somatic
embryogenesis receptor-like kinase 3)
Length = 615
Score = 209 bits (532), Expect = 1e-53
Identities = 121/303 (39%), Positives = 186/303 (60%), Gaps = 18/303 (5%)
Query: 122 LRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNG 181
L++F+ L+VA+ NF +++LG GGFG V+KG + + G VAVK L
Sbjct: 274 LKRFSLRELQVASDNFSNKNILGRGGFGKVYKGRLAD----------GTLVAVKRLKEER 323
Query: 182 HQGHK-EWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRPL--- 237
QG + ++ E+ + +H NL++L GFC+ +RLLVY +M GS+ + L RP
Sbjct: 324 TQGGELQFQTEVEMISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERPESQP 383
Query: 238 PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPE 297
PL W R +IALG+A+GLA+LH+ II+RD K +NILLD E+ A + DFGLAK +
Sbjct: 384 PLDWPKRQRIALGSARGLAYLHDHCDPKIIHRDVKAANILLDEEFEAVVGDFGLAK-LMD 442
Query: 298 GEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHN 357
+ THV+T V GT G+ APEY+ TG S K+DV+ +GV+LLE++TG+R+ D R + +
Sbjct: 443 YKDTHVTTAVRGTIGHIAPEYLSTGKSSEKTDVFGYGVMLLELITGQRAFDLARLANDDD 502
Query: 358 --LVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVH 415
L++W + +L +++ ++D L+G++ + ++ Q+A C P RP MSEVV
Sbjct: 503 VMLLDWVKGLLKEKKL-EALVDVDLQGNYKDEEVEQLIQVALLCTQSSPMERPKMSEVVR 561
Query: 416 TLK 418
L+
Sbjct: 562 MLE 564
>CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4
precursor (EC 2.7.1.-)
Length = 901
Score = 206 bits (525), Expect = 8e-53
Identities = 123/309 (39%), Positives = 186/309 (59%), Gaps = 16/309 (5%)
Query: 114 EELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVA 173
E+LK+ + ++F++ L+ AT F +S +G+G F CVFKG + + GT + V
Sbjct: 483 EDLKIRRA-QEFSYEELEQATGGFSEDSQVGKGSFSCVFKGILRD--------GTVVAVK 533
Query: 174 VKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLF 233
I + + KE+ EL+ L L H +L+ L+G+C + +RLLVYEFM GSL HL
Sbjct: 534 RAIKASDVKKSSKEFHNELDLLSRLNHAHLLNLLGYCEDGSERLLVYEFMAHGSLYQHLH 593
Query: 234 RRPLPLP----WSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDF 289
+ L W+ R+ IA+ AA+G+ +LH + P+I+RD K+SNIL+D ++NA+++DF
Sbjct: 594 GKDPNLKKRLNWARRVTIAVQAARGIEYLHGYACPPVIHRDIKSSNILIDEDHNARVADF 653
Query: 290 GLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDK 349
GL+ GP T +S GT GY PEY +L++KSDVYSFGVVLLE+L+GR++ID
Sbjct: 654 GLSILGPADSGTPLSELPAGTLGYLDPEYYRLHYLTTKSDVYSFGVVLLEILSGRKAIDM 713
Query: 350 KRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPL 409
+ G N+VEWA P++ F I+DP L ++ +K A +A +C+ K RP
Sbjct: 714 QFEEG--NIVEWAVPLI-KAGDIFAILDPVLSPPSDLEALKKIASVACKCVRMRGKDRPS 770
Query: 410 MSEVVHTLK 418
M +V L+
Sbjct: 771 MDKVTTALE 779
>EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase EXS
precursor (EC 2.7.1.37) (Extra sporogenous cells protein)
(EXCESS MICROSPOROCYTES1 protein)
Length = 1192
Score = 204 bits (519), Expect = 4e-52
Identities = 126/304 (41%), Positives = 179/304 (58%), Gaps = 18/304 (5%)
Query: 122 LRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNG 181
L K + AT +F ++++G+GGFG V+K + PG TVAVK L+
Sbjct: 902 LLKVRLGDIVEATDHFSKKNIIGDGGFGTVYKACL---------PGEK-TVAVKKLSEAK 951
Query: 182 HQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRP---LP 238
QG++E++AE+ LG + HPNLV L+G+C +++LLVYE+M GSL++ L +
Sbjct: 952 TQGNREFMAEMETLGKVKHPNLVSLLGYCSFSEEKLLVYEYMVNGSLDHWLRNQTGMLEV 1011
Query: 239 LPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEG 298
L WS R+KIA+GAA+GLAFLH II+RD K SNILLD ++ K++DFGLA+
Sbjct: 1012 LDWSKRLKIAVGAARGLAFLHHGFIPHIIHRDIKASNILLDGDFEPKVADFGLARLISAC 1071
Query: 299 EKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSI--DKKRPNGEH 356
E +HVST + GT+GY PEY + ++K DVYSFGV+LLE++TG+ D K G
Sbjct: 1072 E-SHVSTVIAGTFGYIPPEYGQSARATTKGDVYSFGVILLELVTGKEPTGPDFKESEG-G 1129
Query: 357 NLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHT 416
NLV WA + + +IDP L + Q+A CL+ P RP M +V+
Sbjct: 1130 NLVGWAIQKINQGKA-VDVIDPLLVSVALKNSQLRLLQIAMLCLAETPAKRPNMLDVLKA 1188
Query: 417 LKPL 420
LK +
Sbjct: 1189 LKEI 1192
>PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (EC
2.7.1.37) (Phytosulfokine LRR receptor kinase)
Length = 1008
Score = 202 bits (514), Expect = 2e-51
Identities = 119/301 (39%), Positives = 173/301 (56%), Gaps = 15/301 (4%)
Query: 120 SSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNH 179
S+ ++ +++ L +T +F +++G GGFG V+K + + G VA+K L+
Sbjct: 717 SNDKELSYDDLLDSTNSFDQANIIGCGGFGMVYKATLPD----------GKKVAIKKLSG 766
Query: 180 NGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRR---P 236
+ Q +E+ AE+ L HPNLV L GFC + RLL+Y +M GSL+ L R P
Sbjct: 767 DCGQIEREFEAEVETLSRAQHPNLVLLRGFCFYKNDRLLIYSYMENGSLDYWLHERNDGP 826
Query: 237 LPLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGP 296
L W R++IA GAAKGL +LHE I++RD K+SNILLD +N+ L+DFGLA+
Sbjct: 827 ALLKWKTRLRIAQGAAKGLLYLHEGCDPHILHRDIKSSNILLDENFNSHLADFGLARLMS 886
Query: 297 EGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEH 356
E THVST ++GT GY PEY + K DVYSFGVVLLE+LT +R +D +P G
Sbjct: 887 PYE-THVSTDLVGTLGYIPPEYGQASVATYKGDVYSFGVVLLELLTDKRPVDMCKPKGCR 945
Query: 357 NLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHT 416
+L+ W + H ++ DP + + K + ++A CLS +PK RP ++V
Sbjct: 946 DLISWV-VKMKHESRASEVFDPLIYSKENDKEMFRVLEIACLCLSENPKQRPTTQQLVSW 1004
Query: 417 L 417
L
Sbjct: 1005 L 1005
>BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC
2.7.1.37) (AtBRI1) (Brassinosteroid LRR receptor kinase)
Length = 1196
Score = 201 bits (512), Expect = 3e-51
Identities = 126/306 (41%), Positives = 176/306 (57%), Gaps = 22/306 (7%)
Query: 122 LRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNG 181
LRK TF L AT F +SL+G GGFG V+K +++ G VA+K L H
Sbjct: 868 LRKLTFADLLQATNGFHNDSLIGSGGFGDVYKAILKD----------GSAVAIKKLIHVS 917
Query: 182 HQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLF---RRPLP 238
QG +E++AE+ +G + H NLV L+G+C D+RLLVYEFM GSLE+ L + +
Sbjct: 918 GQGDREFMAEMETIGKIKHRNLVPLLGYCKVGDERLLVYEFMKYGSLEDVLHDPKKAGVK 977
Query: 239 LPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEG 298
L WS R KIA+G+A+GLAFLH + II+RD K+SN+LLD A++SDFG+A+
Sbjct: 978 LNWSTRRKIAIGSARGLAFLHHNCSPHIIHRDMKSSNVLLDENLEARVSDFGMAR-LMSA 1036
Query: 299 EKTHVSTRVM-GTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHN 357
TH+S + GT GY PEY + S+K DVYS+GVVLLE+LTG+R D G++N
Sbjct: 1037 MDTHLSVSTLAGTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKRPTDSP-DFGDNN 1095
Query: 358 LVEWARPVLGHRRM-FFQIIDPRLEGHFSVKGAQ--KAAQLAAQCLSRDPKARPLMSEVV 414
LV W + H ++ + DP L + + ++A CL RP M +V+
Sbjct: 1096 LVGWVKQ---HAKLRISDVFDPELMKEDPALEIELLQHLKVAVACLDDRAWRRPTMVQVM 1152
Query: 415 HTLKPL 420
K +
Sbjct: 1153 AMFKEI 1158
>PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.37)
(Phytosulfokine LRR receptor kinase)
Length = 1021
Score = 195 bits (496), Expect = 2e-49
Identities = 119/338 (35%), Positives = 183/338 (53%), Gaps = 19/338 (5%)
Query: 84 KNTEETEAPPESSTTTSNEESIASTPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLL 143
+ T E PE E + + LF + S + + + + +T +F +++
Sbjct: 694 RTTSRGEVDPEKKADADEIELGSRSVVLFHNK----DSNNELSLDDILKSTSSFNQANII 749
Query: 144 GEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNL 203
G GGFG V+K + + G VA+K L+ + Q +E+ AE+ L HPNL
Sbjct: 750 GCGGFGLVYKATLPD----------GTKVAIKRLSGDTGQMDREFQAEVETLSRAQHPNL 799
Query: 204 VKLIGFCIEDDQRLLVYEFMPRGSLENHLFRR---PLPLPWSIRMKIALGAAKGLAFLHE 260
V L+G+C + +LL+Y +M GSL+ L + P L W R++IA GAA+GLA+LH+
Sbjct: 800 VHLLGYCNYKNDKLLIYSYMDNGSLDYWLHEKVDGPPSLDWKTRLRIARGAAEGLAYLHQ 859
Query: 261 DSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVM 320
+ I++RD K+SNILL + A L+DFGLA+ + THV+T ++GT GY PEY
Sbjct: 860 SCEPHILHRDIKSSNILLSDTFVAHLADFGLARLILPYD-THVTTDLVGTLGYIPPEYGQ 918
Query: 321 TGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRL 380
+ K DVYSFGVVLLE+LTGRR +D +P G +L+ W + +R +I DP +
Sbjct: 919 ASVATYKGDVYSFGVVLLELLTGRRPMDVCKPRGSRDLISWVLQMKTEKRE-SEIFDPFI 977
Query: 381 EGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLK 418
+ ++A +CL +PK RP ++V L+
Sbjct: 978 YDKDHAEEMLLVLEIACRCLGENPKTRPTTQQLVSWLE 1015
>BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.37)
(Brassinosteroid LRR receptor kinase)
Length = 1207
Score = 194 bits (492), Expect = 6e-49
Identities = 124/306 (40%), Positives = 174/306 (56%), Gaps = 22/306 (7%)
Query: 122 LRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNG 181
LRK TF L AT F +SL+G GGFG V+K +++ G VA+K L H
Sbjct: 873 LRKLTFADLLEATNGFHNDSLVGSGGFGDVYKAQLKD----------GSVVAIKKLIHVS 922
Query: 182 HQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRP---LP 238
QG +E+ AE+ +G + H NLV L+G+C ++RLLVYE+M GSLE+ L R +
Sbjct: 923 GQGDREFTAEMETIGKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKTGIK 982
Query: 239 LPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEG 298
L W R KIA+GAA+GLAFLH + II+RD K+SN+LLD A++SDFG+A+
Sbjct: 983 LNWPARRKIAIGAARGLAFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMAR-LMSA 1041
Query: 299 EKTHVSTRVM-GTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHN 357
TH+S + GT GY PEY + S+K DVYS+GVVLLE+LTG++ D G++N
Sbjct: 1042 MDTHLSVSTLAGTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKQPTD-SADFGDNN 1100
Query: 358 LVEWARPVLGHRRMFFQIIDPRL---EGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVV 414
LV W + L + + D L + ++ Q ++A CL RP M +V+
Sbjct: 1101 LVGWVK--LHAKGKITDVFDRELLKEDASIEIELLQH-LKVACACLDDRHWKRPTMIQVM 1157
Query: 415 HTLKPL 420
K +
Sbjct: 1158 AMFKEI 1163
>BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precursor (EC
2.7.1.37) (tBRI1) (Altered brassinolide sensitivity 1)
(Systemin receptor SR160)
Length = 1207
Score = 194 bits (492), Expect = 6e-49
Identities = 124/306 (40%), Positives = 174/306 (56%), Gaps = 22/306 (7%)
Query: 122 LRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNG 181
LRK TF L AT F +SL+G GGFG V+K +++ G VA+K L H
Sbjct: 873 LRKLTFADLLEATNGFHNDSLVGSGGFGDVYKAQLKD----------GSVVAIKKLIHVS 922
Query: 182 HQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRP---LP 238
QG +E+ AE+ +G + H NLV L+G+C ++RLLVYE+M GSLE+ L R +
Sbjct: 923 GQGDREFTAEMETIGKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKIGIK 982
Query: 239 LPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEG 298
L W R KIA+GAA+GLAFLH + II+RD K+SN+LLD A++SDFG+A+
Sbjct: 983 LNWPARRKIAIGAARGLAFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMAR-LMSA 1041
Query: 299 EKTHVSTRVM-GTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHN 357
TH+S + GT GY PEY + S+K DVYS+GVVLLE+LTG++ D G++N
Sbjct: 1042 MDTHLSVSTLAGTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKQPTD-SADFGDNN 1100
Query: 358 LVEWARPVLGHRRMFFQIIDPRL---EGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVV 414
LV W + L + + D L + ++ Q ++A CL RP M +V+
Sbjct: 1101 LVGWVK--LHAKGKITDVFDRELLKEDASIEIELLQH-LKVACACLDDRHWKRPTMIQVM 1157
Query: 415 HTLKPL 420
K +
Sbjct: 1158 AMFKEI 1163
>SIRK_ARATH (O64483) Senescence-induced receptor-like
serine/threonine kinase precursor (FLG22-induced
receptor-like kinase 1)
Length = 876
Score = 187 bits (476), Expect = 4e-47
Identities = 113/299 (37%), Positives = 172/299 (56%), Gaps = 16/299 (5%)
Query: 123 RKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGH 182
R F ++ + T NF E ++G+GGFG V+ G I G VAVK+L+
Sbjct: 562 RYFKYSEVVNITNNF--ERVIGKGGFGKVYHGVIN-----------GEQVAVKVLSEESA 608
Query: 183 QGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLF-RRPLPLPW 241
QG+KE+ AE++ L + H NL L+G+C E + +L+YE+M +L ++L +R L W
Sbjct: 609 QGYKEFRAEVDLLMRVHHTNLTSLVGYCNEINHMVLIYEYMANENLGDYLAGKRSFILSW 668
Query: 242 SIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKT 301
R+KI+L AA+GL +LH + PI++RD K +NILL+ + AK++DFGL++
Sbjct: 669 EERLKISLDAAQGLEYLHNGCKPPIVHRDVKPTNILLNEKLQAKMADFGLSRSFSVEGSG 728
Query: 302 HVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEW 361
+ST V G+ GY PEY T ++ KSDVYS GVVLLE++TG+ +I + H + +
Sbjct: 729 QISTVVAGSIGYLDPEYYSTRQMNEKSDVYSLGVVLLEVITGQPAIASSKTEKVH-ISDH 787
Query: 362 ARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPL 420
R +L + + I+D RL + V A K +++A C RP MS+VV LK +
Sbjct: 788 VRSILANGDI-RGIVDQRLRERYDVGSAWKMSEIALACTEHTSAQRPTMSQVVMELKQI 845
>SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase
receptor precursor (EC 2.7.1.37) (S-receptor kinase)
(SRK)
Length = 849
Score = 184 bits (467), Expect = 4e-46
Identities = 118/324 (36%), Positives = 179/324 (54%), Gaps = 25/324 (7%)
Query: 103 ESIASTPKLFSEELKVAS-SLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGT 161
E + S+ + FS E K L + AT NF + LG+GGFG V+KG + +
Sbjct: 493 EMVLSSKREFSGEYKFEELELPLIEMETVVKATENFSSCNKLGQGGFGIVYKGRLLD--- 549
Query: 162 APVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYE 221
G +AVK L+ QG E++ E+ + L H NLV+++G CIE D+++L+YE
Sbjct: 550 -------GKEIAVKRLSKTSVQGTDEFMNEVTLIARLQHINLVQVLGCCIEGDEKMLIYE 602
Query: 222 FMPRGSLENHLF--RRPLPLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLD 279
++ SL+++LF R L W+ R I G A+GL +LH+DS+ II+RD K SNILLD
Sbjct: 603 YLENLSLDSYLFGKTRRSKLNWNERFDITNGVARGLLYLHQDSRFRIIHRDLKVSNILLD 662
Query: 280 AEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLE 339
K+SDFG+A+ E + +V+GTYGY +PEY M G S KSDV+SFGV++LE
Sbjct: 663 KNMIPKISDFGMARIFERDETEANTMKVVGTYGYMSPEYAMYGIFSEKSDVFSFGVIVLE 722
Query: 340 MLTGRRSIDKKRPNGEHNLVE--WARPVLGHRRMFFQIIDPRL-------EGHFSVKGAQ 390
+++G+++ + E++L+ W+R G +I+DP + F +
Sbjct: 723 IVSGKKNRGFYNLDYENDLLSYVWSRWKEGRA---LEIVDPVIVDSLSSQPSIFQPQEVL 779
Query: 391 KAAQLAAQCLSRDPKARPLMSEVV 414
K Q+ C+ + RP MS VV
Sbjct: 780 KCIQIGLLCVQELAEHRPAMSSVV 803
>CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (EC
2.7.1.-)
Length = 980
Score = 176 bits (446), Expect = 1e-43
Identities = 108/295 (36%), Positives = 160/295 (53%), Gaps = 27/295 (9%)
Query: 138 RPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGD 197
+ E+++G+GG G V++G + N +K G G H + AE+ LG
Sbjct: 693 KEENIIGKGGAGIVYRGSMPNNVDVAIKRLVG--------RGTGRSDHG-FTAEIQTLGR 743
Query: 198 LLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLF-RRPLPLPWSIRMKIALGAAKGLA 256
+ H ++V+L+G+ D LL+YE+MP GSL L + L W R ++A+ AAKGL
Sbjct: 744 IRHRHIVRLLGYVANKDTNLLLYEYMPNGSLGELLHGSKGGHLQWETRHRVAVEAAKGLC 803
Query: 257 FLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAP 316
+LH D I++RD K++NILLD+++ A ++DFGLAK +G + + + G+YGY AP
Sbjct: 804 YLHHDCSPLILHRDVKSNNILLDSDFEAHVADFGLAKFLVDGAASECMSSIAGSYGYIAP 863
Query: 317 EYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEH----NLVEWARPV------L 366
EY T + KSDVYSFGVVLLE++ G K+P GE ++V W R
Sbjct: 864 EYAYTLKVDEKSDVYSFGVVLLELIAG------KKPVGEFGEGVDIVRWVRNTEEEITQP 917
Query: 367 GHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLP 421
+ I+DPRL G + + ++A C+ + ARP M EVVH L P
Sbjct: 918 SDAAIVVAIVDPRLTG-YPLTSVIHVFKIAMMCVEEEAAARPTMREVVHMLTNPP 971
>IRA4_HUMAN (Q9NWZ3) Interleukin-1 receptor-associated kinase-4 (EC
2.7.1.37) (IRAK-4) (NY-REN-64 antigen)
Length = 460
Score = 174 bits (442), Expect = 4e-43
Identities = 117/345 (33%), Positives = 179/345 (50%), Gaps = 34/345 (9%)
Query: 84 KNTEETEAPPESSTTTSNEESIASTPKLFSEELKVASSLRKFTFNGLKVATRNF--RPES 141
+N E++ PP+SS+ + ++ T F+F LK T NF RP S
Sbjct: 139 QNLEQSYMPPDSSSPENKSLEVSDT------------RFHSFSFYELKNVTNNFDERPIS 186
Query: 142 L----LGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGD 197
+ +GEGGFG V+KG++ A K + + + L Q E+ +
Sbjct: 187 VGGNKMGEGGFGVVYKGYVNNTTVAVKKLAAMVDITTEELKQQFDQ-------EIKVMAK 239
Query: 198 LLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHL--FRRPLPLPWSIRMKIALGAAKGL 255
H NLV+L+GF + D LVY +MP GSL + L PL W +R KIA GAA G+
Sbjct: 240 CQHENLVELLGFSSDGDDLCLVYVYMPNGSLLDRLSCLDGTPPLSWHMRCKIAQGAANGI 299
Query: 256 AFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAA 315
FLHE+ I+RD K++NILLD + AK+SDFGLA+ + +T +++R++GT Y A
Sbjct: 300 NFLHENHH---IHRDIKSANILLDEAFTAKISDFGLARASEKFAQTVMTSRIVGTTAYMA 356
Query: 316 PEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQI 375
PE + G ++ KSD+YSFGVVLLE++TG ++D+ R L++ + +
Sbjct: 357 PE-ALRGEITPKSDIYSFGVVLLEIITGLPAVDEHRE--PQLLLDIKEEIEDEEKTIEDY 413
Query: 376 IDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPL 420
ID ++ S + +A+QCL RP + +V L+ +
Sbjct: 414 IDKKMNDADST-SVEAMYSVASQCLHEKKNKRPDIKKVQQLLQEM 457
>KPRO_MAIZE (P17801) Putative receptor protein kinase ZmPK1
precursor (EC 2.7.1.37)
Length = 817
Score = 174 bits (441), Expect = 5e-43
Identities = 119/309 (38%), Positives = 167/309 (53%), Gaps = 22/309 (7%)
Query: 118 VASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKIL 177
+ S+ R++++ L ATR F+ E LG G G V+KG +E++ VAVK L
Sbjct: 517 MTSNFRRYSYRELVKATRKFKVE--LGRGESGTVYKGVLEDDRH----------VAVKKL 564
Query: 178 NHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRP- 236
N QG + + AEL+ +G + H NLV++ GFC E RLLV E++ GSL N LF
Sbjct: 565 E-NVRQGKEVFQAELSVIGRINHMNLVRIWGFCSEGSHRLLVSEYVENGSLANILFSEGG 623
Query: 237 -LPLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDG 295
+ L W R IALG AKGLA+LH + +I+ D K NILLD + K++DFGL K
Sbjct: 624 NILLDWEGRFNIALGVAKGLAYLHHECLEWVIHCDVKPENILLDQAFEPKITDFGLVKLL 683
Query: 296 PEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGE 355
G T + V GT GY APE+V + +++K DVYS+GVVLLE+LTG R + E
Sbjct: 684 NRGGSTQNVSHVRGTLGYIAPEWVSSLPITAKVDVYSYGVVLLELLTGTRVSELVGGTDE 743
Query: 356 -HNLVEWARPVL-----GHRRMFFQ-IIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARP 408
H+++ +L G + + +D +L + A+ +LA CL D RP
Sbjct: 744 VHSMLRKLVRMLSAKLEGEEQSWIDGYLDSKLNRPVNYVQARTLIKLAVSCLEEDRSKRP 803
Query: 409 LMSEVVHTL 417
M V TL
Sbjct: 804 TMEHAVQTL 812
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 60,927,256
Number of Sequences: 164201
Number of extensions: 2671552
Number of successful extensions: 11436
Number of sequences better than 10.0: 1599
Number of HSP's better than 10.0 without gapping: 1058
Number of HSP's successfully gapped in prelim test: 541
Number of HSP's that attempted gapping in prelim test: 7743
Number of HSP's gapped (non-prelim): 2027
length of query: 490
length of database: 59,974,054
effective HSP length: 114
effective length of query: 376
effective length of database: 41,255,140
effective search space: 15511932640
effective search space used: 15511932640
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0391.11


