

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0388b.3
(149 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
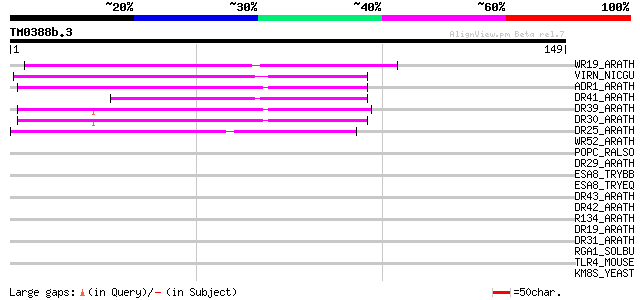
Score E
Sequences producing significant alignments: (bits) Value
WR19_ARATH (Q9SZ67) Probable WRKY transcription factor 19 (WRKY ... 69 3e-12
VIRN_NICGU (Q40392) TMV resistance protein N 61 1e-09
ADR1_ARATH (Q9FW44) Disease resistance protein ADR1 (Activated d... 46 3e-05
DR41_ARATH (Q9FKZ2) Probable disease resistance protein At5g66890 43 2e-04
DR39_ARATH (Q9LVT1) Putative disease resistance protein At5g47280 43 3e-04
DR30_ARATH (Q9LZ25) Probable disease resistance protein At5g04720 43 3e-04
DR25_ARATH (O50052) Putative disease resistance protein At4g19050 42 5e-04
WR52_ARATH (Q9FH83) Probable WRKY transcription factor 52 (WRKY ... 41 0.001
POPC_RALSO (Q9RBS2) PopC protein 40 0.001
DR29_ARATH (Q9SZA7) Probable disease resistance protein At4g33300 40 0.002
ESA8_TRYBB (P23799) Putative adenylate cyclase regulatory protei... 39 0.003
ESA8_TRYEQ (P26337) Putative adenylate cyclase regulatory protein 38 0.007
DR43_ARATH (Q9FKZ0) Probable disease resistance protein At5g66910 38 0.009
DR42_ARATH (Q9FKZ1) Probable disease resistance protein At5g66900 37 0.016
R134_ARATH (Q38834) Putative disease resistance RPP13-like prote... 36 0.035
DR19_ARATH (Q9C8T9) Putative disease resistance protein At1g63350 36 0.035
DR31_ARATH (Q9FLB4) Putative disease resistance protein At5g05400 35 0.060
RGA1_SOLBU (Q7XA42) Putative disease resistance protein RGA1 (RG... 33 0.18
TLR4_MOUSE (Q9QUK6) Toll-like receptor 4 precursor 33 0.23
KM8S_YEAST (Q03533) Probable serine/threonine-protein kinase YMR... 33 0.30
>WR19_ARATH (Q9SZ67) Probable WRKY transcription factor 19 (WRKY
DNA-binding protein 19)
Length = 1895
Score = 69.3 bits (168), Expect = 3e-12
Identities = 37/100 (37%), Positives = 57/100 (57%), Gaps = 2/100 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK++ LS S L + P + LE +D GC +LL + SI L +L FL+L+GCS L
Sbjct: 1260 LKKMRLSYSDQLTKIPRLSSATNLEHIDLEGCNSLLSLSQSISYLKKLVFLNLKGCSKLE 1319
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFRCG 104
+ + ++ +L SL +L+LSGC++L + P N G
Sbjct: 1320 N--IPSMVDLESLEVLNLSGCSKLGNFPEISPNVKELYMG 1357
Score = 30.0 bits (66), Expect = 1.9
Identities = 19/54 (35%), Positives = 30/54 (55%), Gaps = 1/54 (1%)
Query: 5 LKRLDLSNSKYLMETP-NFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSL 57
L++LDL NS++L P + + LE L+ +GC +L + S + L FL L
Sbjct: 1374 LEKLDLENSRHLKNLPTSIYKLKHLETLNLSGCISLERFPDSSRRMKCLRFLDL 1427
>VIRN_NICGU (Q40392) TMV resistance protein N
Length = 1144
Score = 60.8 bits (146), Expect = 1e-09
Identities = 36/95 (37%), Positives = 49/95 (50%), Gaps = 3/95 (3%)
Query: 2 VPFLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCS 61
+P L+R+DLS SK L TP+F G LE ++ C+NL +VH S+G +++ L L C
Sbjct: 618 LPSLRRIDLSWSKRLTRTPDFTGMPNLEYVNLYQCSNLEEVHHSLGCCSKVIGLYLNDCK 677
Query: 62 SLVSLYLGTVCELNSLRILHLSGCTRLESTPNFIG 96
SL + SL L L C LE P G
Sbjct: 678 SLKRF---PCVNVESLEYLGLRSCDSLEKLPEIYG 709
>ADR1_ARATH (Q9FW44) Disease resistance protein ADR1 (Activated
disease resistance protein 1)
Length = 787
Score = 45.8 bits (107), Expect = 3e-05
Identities = 29/94 (30%), Positives = 44/94 (45%), Gaps = 1/94 (1%)
Query: 3 PFLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSS 62
P L L + + L+E + G L L T C +L++ ++ + L L L C
Sbjct: 628 PSLSDLTIDHCDDLLELKSIFGITSLNSLSITNCPRILELPKNLSNVQSLERLRLYACPE 687
Query: 63 LVSLYLGTVCELNSLRILHLSGCTRLESTPNFIG 96
L+SL + VCEL L+ + +S C L S P G
Sbjct: 688 LISLPV-EVCELPCLKYVDISQCVSLVSLPEKFG 720
>DR41_ARATH (Q9FKZ2) Probable disease resistance protein At5g66890
Length = 415
Score = 43.1 bits (100), Expect = 2e-04
Identities = 27/69 (39%), Positives = 41/69 (59%), Gaps = 1/69 (1%)
Query: 28 LERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLYLGTVCELNSLRILHLSGCTR 87
L++L T C L +V +IG L +L L L C+SL+ L T+ L++LR L +SG +
Sbjct: 281 LKKLSVTNCNKLCRVIEAIGDLRDLETLRLSSCASLLEL-PETIDRLDNLRFLDVSGGFQ 339
Query: 88 LESTPNFIG 96
L++ P IG
Sbjct: 340 LKNLPLEIG 348
>DR39_ARATH (Q9LVT1) Putative disease resistance protein At5g47280
Length = 623
Score = 42.7 bits (99), Expect = 3e-04
Identities = 31/96 (32%), Positives = 43/96 (44%), Gaps = 2/96 (2%)
Query: 3 PFLKRLDLSNSKYLMETPN-FDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCS 61
P L + + L E P+ G L + T C N+ ++ +I L L L L C
Sbjct: 463 PKLTDITIDYCDDLAELPSTICGITSLNSISITNCPNIKELPKNISKLQALQLLRLYACP 522
Query: 62 SLVSLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
L SL + +CEL L + +S C L S P IGN
Sbjct: 523 ELKSLPV-EICELPRLVYVDISHCLSLSSLPEKIGN 557
>DR30_ARATH (Q9LZ25) Probable disease resistance protein At5g04720
Length = 811
Score = 42.7 bits (99), Expect = 3e-04
Identities = 29/95 (30%), Positives = 44/95 (45%), Gaps = 2/95 (2%)
Query: 3 PFLKRLDLSNSKYLMETPN-FDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCS 61
P L L + + L+E P+ G L + T C + ++ ++ L L L L C
Sbjct: 651 PKLSDLTIDHCDDLLELPSTICGITSLNSISITNCPRIKELPKNLSKLKALQLLRLYACH 710
Query: 62 SLVSLYLGTVCELNSLRILHLSGCTRLESTPNFIG 96
L SL + +CEL L+ + +S C L S P IG
Sbjct: 711 ELNSLPV-EICELPRLKYVDISQCVSLSSLPEKIG 744
>DR25_ARATH (O50052) Putative disease resistance protein At4g19050
Length = 1181
Score = 42.0 bits (97), Expect = 5e-04
Identities = 27/93 (29%), Positives = 49/93 (52%), Gaps = 2/93 (2%)
Query: 1 DVPFLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGC 60
DV L +L L N + E P+ + LE D +GC L ++ S G ++ L ++L
Sbjct: 700 DVVNLNKLLLRNCSLIEELPSIEKLTHLEVFDVSGCIKLKNINGSFGEMSYLHEVNLS-- 757
Query: 61 SSLVSLYLGTVCELNSLRILHLSGCTRLESTPN 93
+ +S + EL++L+ L + C++L++ PN
Sbjct: 758 ETNLSELPDKISELSNLKELIIRKCSKLKTLPN 790
Score = 37.4 bits (85), Expect = 0.012
Identities = 26/89 (29%), Positives = 43/89 (48%), Gaps = 2/89 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK L + L PN + LE D +GCT L + S L+ L ++L + +
Sbjct: 774 LKELIIRKCSKLKTLPNLEKLTNLEIFDVSGCTELETIEGSFENLSCLHKVNLS--ETNL 831
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPN 93
+ EL++L+ L L C++L++ PN
Sbjct: 832 GELPNKISELSNLKELILRNCSKLKALPN 860
Score = 33.5 bits (75), Expect = 0.18
Identities = 19/55 (34%), Positives = 27/55 (48%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEG 59
LK L L N L PN + L D +GCTNL ++ S ++ L ++L G
Sbjct: 844 LKELILRNCSKLKALPNLEKLTHLVIFDVSGCTNLDKIEESFESMSYLCEVNLSG 898
Score = 29.6 bits (65), Expect = 2.5
Identities = 27/93 (29%), Positives = 43/93 (46%), Gaps = 3/93 (3%)
Query: 2 VPFLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQ-VHPSIGLLTELAFLSLEGC 60
+P L RL L N L P L+ LD G T+L++ + + EL L +
Sbjct: 630 MPILTRLLLRNCTRLKRLPQLRPLTNLQILDACGATDLVEMLEVCLEEKKELRILDMSK- 688
Query: 61 SSLVSLYLGTVCELNSLRILHLSGCTRLESTPN 93
+SL L T+ ++ +L L L C+ +E P+
Sbjct: 689 TSLPEL-ADTIADVVNLNKLLLRNCSLIEELPS 720
>WR52_ARATH (Q9FH83) Probable WRKY transcription factor 52 (WRKY
DNA-binding protein 52) (Disease resistance protein
RRS1) (Resistance to Ralstonia solanacearum 1 protein)
Length = 1378
Score = 40.8 bits (94), Expect = 0.001
Identities = 37/117 (31%), Positives = 51/117 (42%), Gaps = 28/117 (23%)
Query: 15 YLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLYLGTVCEL 74
+L E P + +LERL T+LL+ + S L +L L L+ CS L SL +L
Sbjct: 696 FLTEIPGLSEASKLERL-----TSLLESNSSCQDLGKLICLELKDCSCLQSLPNMANLDL 750
Query: 75 NSLRILHLSGCTRLESTPNFIGNPSHFRCGFDIVVPGNKIPYWFDSQFKGGSRIREV 131
N +L LSGC+ L S F P + + GG+ IREV
Sbjct: 751 N---VLDLSGCSSLNSIQGF--------------------PRFLKQLYLGGTAIREV 784
>POPC_RALSO (Q9RBS2) PopC protein
Length = 1024
Score = 40.4 bits (93), Expect = 0.001
Identities = 36/94 (38%), Positives = 48/94 (50%), Gaps = 4/94 (4%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQ--RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSS 62
LK L + NS L P G Q RL +L + T L + SIG L+ L L+L+ +
Sbjct: 568 LKTLTVENSP-LTSIPADIGIQCERLTQLSLSN-TQLRALPSSIGKLSNLKGLTLKNNAR 625
Query: 63 LVSLYLGTVCELNSLRILHLSGCTRLESTPNFIG 96
L L V +L S+R + LSGC RL P+ IG
Sbjct: 626 LELLSESGVRKLESVRKIDLSGCVRLTGLPSSIG 659
Score = 37.4 bits (85), Expect = 0.012
Identities = 33/99 (33%), Positives = 49/99 (49%), Gaps = 12/99 (12%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLE---RLDFTGCTNLLQVHPSIGLLTELAFLSLEGCS 61
LK L L N+ L E + G ++LE ++D +GC L + SIG L +L L L GC+
Sbjct: 615 LKGLTLKNNARL-ELLSESGVRKLESVRKIDLSGCVRLTGLPSSIGKLPKLRTLDLSGCT 673
Query: 62 --SLVSLYLGTVCELNSLRIL---HLS---GCTRLESTP 92
S+ SL V + L ++ HL G R++ P
Sbjct: 674 GLSMASLPRSLVLPRDGLNVIFPEHLKTDVGNARIQQNP 712
>DR29_ARATH (Q9SZA7) Probable disease resistance protein At4g33300
Length = 855
Score = 40.0 bits (92), Expect = 0.002
Identities = 30/95 (31%), Positives = 42/95 (43%), Gaps = 2/95 (2%)
Query: 3 PFLKRLDLSNSKYLMETPN-FDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCS 61
P L L + + L+ P+ G L L T C L ++ ++ L L L L C
Sbjct: 695 PKLGDLTIDHCDDLVALPSSICGLTSLSCLSITNCPRLGELPKNLSKLQALEILRLYACP 754
Query: 62 SLVSLYLGTVCELNSLRILHLSGCTRLESTPNFIG 96
L +L G +CEL L+ L +S C L P IG
Sbjct: 755 ELKTLP-GEICELPGLKYLDISQCVSLSCLPEEIG 788
>ESA8_TRYBB (P23799) Putative adenylate cyclase regulatory protein
(Leucine repeat protein) (VSG expression site-associated
protein F14.9)
Length = 630
Score = 39.3 bits (90), Expect = 0.003
Identities = 32/107 (29%), Positives = 47/107 (43%), Gaps = 22/107 (20%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK L +SN K + + LE+L+ +GC + + + L+ L L + GC SLV
Sbjct: 326 LKVLSVSNCKNFKDLNGLERLVNLEKLNLSGCHGVSSLG-FVANLSNLKELDISGCESLV 384
Query: 65 S------------LYL---------GTVCELNSLRILHLSGCTRLES 90
LYL G + L+ +R L LSGC R+ S
Sbjct: 385 CFDGLQDLNNLEVLYLRDVKSFTNVGAIKNLSKMRELDLSGCERITS 431
Score = 37.0 bits (84), Expect = 0.016
Identities = 23/81 (28%), Positives = 41/81 (50%), Gaps = 2/81 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK LD+S+ + + G + LE+L +GC N+ + + + L L + GC L
Sbjct: 256 LKMLDISSCHEITDLTAIGGVRSLEKLSLSGCWNVTKGLEELCKFSNLRELDISGCLVLG 315
Query: 65 SLYLGTVCELNSLRILHLSGC 85
S + + L +L++L +S C
Sbjct: 316 SAVV--LKNLINLKVLSVSNC 334
Score = 35.0 bits (79), Expect = 0.060
Identities = 23/64 (35%), Positives = 37/64 (56%), Gaps = 2/64 (3%)
Query: 27 RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLYLGTVCELNSLRILHLSGCT 86
+L+ LD + C + + +IG + L LSL GC + V+ L +C+ ++LR L +SGC
Sbjct: 255 KLKMLDISSCHEITDL-TAIGGVRSLEKLSLSGCWN-VTKGLEELCKFSNLRELDISGCL 312
Query: 87 RLES 90
L S
Sbjct: 313 VLGS 316
Score = 32.7 bits (73), Expect = 0.30
Identities = 24/83 (28%), Positives = 37/83 (43%), Gaps = 3/83 (3%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
++ LDLS + + + + LE L GC ++ P I L L L + C +L
Sbjct: 418 MRELDLSGCERITSLSGLETLKGLEELSLEGCGEIMSFDP-IWSLYHLRVLYVSECGNLE 476
Query: 65 SLYLGTVCELNSLRILHLSGCTR 87
L G C L L ++L GC +
Sbjct: 477 DL-SGLQC-LTGLEEMYLHGCRK 497
>ESA8_TRYEQ (P26337) Putative adenylate cyclase regulatory protein
Length = 630
Score = 38.1 bits (87), Expect = 0.007
Identities = 27/96 (28%), Positives = 43/96 (44%), Gaps = 3/96 (3%)
Query: 1 DVPFLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGC 60
D+ L+ L L + K ++ LD +GC + + + L L LSLEGC
Sbjct: 391 DLNNLEVLYLRDVKSFTNVGAIKNLSKMRELDLSGCERITSLS-GLETLKGLEELSLEGC 449
Query: 61 SSLVSLYLGTVCELNSLRILHLSGCTRLESTPNFIG 96
++S + L+ LR+L++S C LE G
Sbjct: 450 GEIMSF--DPIWSLHHLRVLYVSECGNLEDLSGLEG 483
Score = 38.1 bits (87), Expect = 0.007
Identities = 31/107 (28%), Positives = 47/107 (42%), Gaps = 22/107 (20%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK L +SN K + + L++L+ +GC + + + L+ L L + GC SLV
Sbjct: 326 LKVLSVSNCKNFKDLNGLERLVNLDKLNLSGCHGVSSLG-FVANLSNLKELDISGCESLV 384
Query: 65 S------------LYL---------GTVCELNSLRILHLSGCTRLES 90
LYL G + L+ +R L LSGC R+ S
Sbjct: 385 CFDGLQDLNNLEVLYLRDVKSFTNVGAIKNLSKMRELDLSGCERITS 431
Score = 32.7 bits (73), Expect = 0.30
Identities = 22/64 (34%), Positives = 37/64 (57%), Gaps = 2/64 (3%)
Query: 27 RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLYLGTVCELNSLRILHLSGCT 86
+L+ L ++ C + + +IG + L LSL GC + V+ L +C+ ++LR L +SGC
Sbjct: 255 KLKVLRYSSCHEITDL-TAIGGMRSLEKLSLSGCWN-VTKGLEELCKFSNLRELDISGCL 312
Query: 87 RLES 90
L S
Sbjct: 313 VLGS 316
Score = 32.3 bits (72), Expect = 0.39
Identities = 22/81 (27%), Positives = 39/81 (47%), Gaps = 2/81 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK L S+ + + G + LE+L +GC N+ + + + L L + GC L
Sbjct: 256 LKVLRYSSCHEITDLTAIGGMRSLEKLSLSGCWNVTKGLEELCKFSNLRELDISGCLVLG 315
Query: 65 SLYLGTVCELNSLRILHLSGC 85
S + + L +L++L +S C
Sbjct: 316 SAVV--LKNLINLKVLSVSNC 334
Score = 32.3 bits (72), Expect = 0.39
Identities = 27/107 (25%), Positives = 43/107 (39%), Gaps = 24/107 (22%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHP-------------SIGLLTE 51
++ LDLS + + + + LE L GC ++ P G L +
Sbjct: 418 MRELDLSGCERITSLSGLETLKGLEELSLEGCGEIMSFDPIWSLHHLRVLYVSECGNLED 477
Query: 52 LAFLSLEGCSSLVSLYL---------GTVCELNSLRILHLSGCTRLE 89
L+ LEG + L LYL G + L ++ ++ LS C LE
Sbjct: 478 LS--GLEGITGLEELYLHGCRKCTNFGPIWNLRNVCVVELSCCENLE 522
>DR43_ARATH (Q9FKZ0) Probable disease resistance protein At5g66910
Length = 815
Score = 37.7 bits (86), Expect = 0.009
Identities = 25/69 (36%), Positives = 34/69 (49%), Gaps = 1/69 (1%)
Query: 28 LERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLYLGTVCELNSLRILHLSGCTR 87
L+ L T C L Q+ +IG L+ L L + C +L L T L++LR L +S C
Sbjct: 681 LKTLSITNCNKLSQLPEAIGNLSRLEVLRMCSCMNLSELPEATE-RLSNLRSLDISHCLG 739
Query: 88 LESTPNFIG 96
L P IG
Sbjct: 740 LRKLPQEIG 748
Score = 35.4 bits (80), Expect = 0.046
Identities = 26/88 (29%), Positives = 42/88 (47%), Gaps = 3/88 (3%)
Query: 1 DVPFLKRLDLSNSKYLMETPNFDGS-QRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEG 59
+V LK L ++N L + P G+ RLE L C NL ++ + L+ L L +
Sbjct: 677 EVVSLKTLSITNCNKLSQLPEAIGNLSRLEVLRMCSCMNLSELPEATERLSNLRSLDISH 736
Query: 60 CSSLVSL--YLGTVCELNSLRILHLSGC 85
C L L +G + +L ++ + SGC
Sbjct: 737 CLGLRKLPQEIGKLQKLENISMRKCSGC 764
>DR42_ARATH (Q9FKZ1) Probable disease resistance protein At5g66900
Length = 809
Score = 37.0 bits (84), Expect = 0.016
Identities = 27/70 (38%), Positives = 35/70 (49%), Gaps = 3/70 (4%)
Query: 28 LERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLYLGTVCE-LNSLRILHLSGCT 86
L+ L T C L Q+ +IG L+ L L L CSS+ L E L++LR L +S C
Sbjct: 675 LKTLSITNCNKLSQLPEAIGNLSRLEVLRL--CSSMNLSELPEATEGLSNLRFLDISHCL 732
Query: 87 RLESTPNFIG 96
L P IG
Sbjct: 733 GLRKLPQEIG 742
Score = 28.1 bits (61), Expect = 7.4
Identities = 28/104 (26%), Positives = 49/104 (46%), Gaps = 8/104 (7%)
Query: 2 VPFLKRLDLSN-SKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGL-----LTELAFL 55
+P LKR+ L S L++ P S L++L C+ + + + L++L +
Sbjct: 596 LPNLKRIRLEKVSITLLDIPQLQLSS-LKKLSLVMCSFGEVFYDTEDIVVSNALSKLQEI 654
Query: 56 SLEGCSSLVSLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPS 99
++ C L L + E+ SL+ L ++ C +L P IGN S
Sbjct: 655 DIDYCYDLDELPYW-ISEIVSLKTLSITNCNKLSQLPEAIGNLS 697
>R134_ARATH (Q38834) Putative disease resistance RPP13-like protein
4
Length = 852
Score = 35.8 bits (81), Expect = 0.035
Identities = 21/57 (36%), Positives = 31/57 (53%), Gaps = 1/57 (1%)
Query: 8 LDLSNSKYLMETP-NFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
L LSN+ L++ P + + L+ LD + C NL Q+ P I L +L L + C SL
Sbjct: 591 LSLSNTHPLIQFPRSMEDLHNLQILDASYCQNLKQLQPCIVLFKKLLVLDMTNCGSL 647
>DR19_ARATH (Q9C8T9) Putative disease resistance protein At1g63350
Length = 898
Score = 35.8 bits (81), Expect = 0.035
Identities = 33/85 (38%), Positives = 45/85 (52%), Gaps = 6/85 (7%)
Query: 2 VPFLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGL--LTELAFLSLEG 59
+P L LDLS + YL E PN G L L + ++ H GL L +L L LE
Sbjct: 560 MPKLAVLDLSGNYYLSELPN--GISELVSLQYLNLSSTGIRHLPKGLQELKKLIHLYLER 617
Query: 60 CSSLVSLYLGTVCELNSLRILHLSG 84
S L S+ +G C L++L++L LSG
Sbjct: 618 TSQLGSM-VGISC-LHNLKVLKLSG 640
>DR31_ARATH (Q9FLB4) Putative disease resistance protein At5g05400
Length = 874
Score = 35.0 bits (79), Expect = 0.060
Identities = 26/81 (32%), Positives = 41/81 (50%), Gaps = 3/81 (3%)
Query: 2 VPFLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCS 61
VP L LDLS + L+E P+F L L+ + CT + + + L L +L+LE
Sbjct: 550 VPILMVLDLSLNPNLIELPSFSPLYSLRFLNLS-CTGITSLPDGLYALRNLLYLNLEHTY 608
Query: 62 SLVSLYLGTVCELNSLRILHL 82
L +Y + +L +L +L L
Sbjct: 609 MLKRIY--EIHDLPNLEVLKL 627
>RGA1_SOLBU (Q7XA42) Putative disease resistance protein RGA1
(RGA3-blb)
Length = 1025
Score = 33.5 bits (75), Expect = 0.18
Identities = 32/92 (34%), Positives = 41/92 (43%), Gaps = 2/92 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
L+ L+L NS + L LD +G + + + L L L L C SL
Sbjct: 573 LRVLNLRNSNLNQLPSSIGDLVHLRYLDLSGNFRIRNLPKRLCKLQNLQTLDLHYCDSLS 632
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIG 96
L T +L SLR L L GC+ L STP IG
Sbjct: 633 CLPKQT-SKLGSLRNLLLDGCS-LTSTPPRIG 662
Score = 33.1 bits (74), Expect = 0.23
Identities = 28/90 (31%), Positives = 39/90 (43%), Gaps = 2/90 (2%)
Query: 5 LKRLDLSNSKYLMETPN--FDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSS 62
L LD+S++ P F L+ L + NL ++ S+ L L L E C +
Sbjct: 891 LTSLDISDNVEATSLPEEMFKSLANLKYLKISFFRNLKELPTSLASLNALKSLKFEFCDA 950
Query: 63 LVSLYLGTVCELNSLRILHLSGCTRLESTP 92
L SL V L SL L +S C L+ P
Sbjct: 951 LESLPEEGVKGLTSLTELSVSNCMMLKCLP 980
>TLR4_MOUSE (Q9QUK6) Toll-like receptor 4 precursor
Length = 835
Score = 33.1 bits (74), Expect = 0.23
Identities = 30/95 (31%), Positives = 46/95 (47%), Gaps = 16/95 (16%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTEL-AFLSLEGCSSL 63
L+ LDLS + ++ + NF G + L+ LDF H ++ +TE AFLSLE L
Sbjct: 399 LRHLDLSFNGAIIMSANFMGLEELQHLDFQ--------HSTLKRVTEFSAFLSLEKLLYL 450
Query: 64 VSLYLGTVCE-------LNSLRILHLSGCTRLEST 91
Y T + L SL L ++G + ++T
Sbjct: 451 DISYTNTKIDFDGIFLGLTSLNTLKMAGNSFKDNT 485
>KM8S_YEAST (Q03533) Probable serine/threonine-protein kinase
YMR291W (EC 2.7.1.37)
Length = 586
Score = 32.7 bits (73), Expect = 0.30
Identities = 23/78 (29%), Positives = 36/78 (45%), Gaps = 1/78 (1%)
Query: 3 PFLKR-LDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCS 61
PF+K SNS +TPNF + R+ T + L+ T LA+L+++G S
Sbjct: 349 PFIKDYFATSNSLNTKDTPNFSFHPTIRRVSSTASMHTLRSPSKSRKTTTLAYLNMDGGS 408
Query: 62 SLVSLYLGTVCELNSLRI 79
S S + +L L +
Sbjct: 409 SETSTAFSSKMDLPDLYV 426
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.325 0.143 0.458
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,601,190
Number of Sequences: 164201
Number of extensions: 735024
Number of successful extensions: 2426
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 39
Number of HSP's successfully gapped in prelim test: 33
Number of HSP's that attempted gapping in prelim test: 2320
Number of HSP's gapped (non-prelim): 121
length of query: 149
length of database: 59,974,054
effective HSP length: 100
effective length of query: 49
effective length of database: 43,553,954
effective search space: 2134143746
effective search space used: 2134143746
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0388b.3


