

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0388a.4
(1489 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
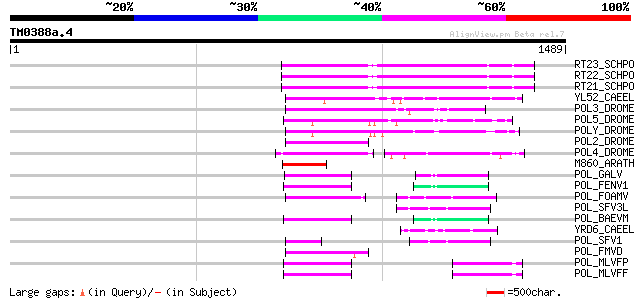
Score E
Sequences producing significant alignments: (bits) Value
RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protei... 346 2e-94
RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protei... 346 2e-94
RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protei... 346 2e-94
YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III 201 1e-50
POL3_DROME (P04323) Retrovirus-related Pol polyprotein from tran... 183 3e-45
POL5_DROME (Q8I7P9) Retrovirus-related Pol polyprotein from tran... 178 1e-43
POLY_DROME (P10401) Retrovirus-related Pol polyprotein from tran... 176 4e-43
POL2_DROME (P20825) Retrovirus-related Pol polyprotein from tran... 168 9e-41
POL4_DROME (P10394) Retrovirus-related Pol polyprotein from tran... 167 2e-40
M860_ARATH (P92523) Hypothetical mitochondrial protein AtMg00860... 114 2e-24
POL_GALV (P21414) Pol polyprotein [Contains: Protease (EC 3.4.23... 91 2e-17
POL_FENV1 (P31792) Pol polyprotein [Contains: Reverse transcript... 91 2e-17
POL_FOAMV (P14350) Pol polyprotein [Contains: Reverse transcript... 89 7e-17
POL_SFV3L (P27401) Pol polyprotein [Contains: Protease (EC 3.4.2... 87 3e-16
POL_BAEVM (P10272) Pol polyprotein [Contains: Protease (EC 3.4.2... 87 3e-16
YRD6_CAEEL (Q09575) Hypothetical protein K02A2.6 in chromosome II 87 4e-16
POL_SFV1 (P23074) Pol polyprotein [Contains: Protease (EC 3.4.23... 85 2e-15
POL_FMVD (P09523) Enzymatic polyprotein [Contains: Aspartic prot... 84 4e-15
POL_MLVFP (P26808) Pol polyprotein [Contains: Protease (EC 3.4.2... 82 1e-14
POL_MLVFF (P26809) Pol polyprotein [Contains: Protease (EC 3.4.2... 82 1e-14
>RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protein
type 3
Length = 1333
Score = 346 bits (888), Expect = 2e-94
Identities = 213/690 (30%), Positives = 350/690 (49%), Gaps = 34/690 (4%)
Query: 730 KGKELYANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLA 789
K L N +KCEF +VKF+G+ IS +G I+ ++ W+QPK ++R F+G
Sbjct: 590 KNANLIINQAKCEFHQSQVKFIGYHISEKGFTPCQENIDKVLQWKQPKNRKELRQFLGSV 649
Query: 790 GYYRRFVKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGEPYD 849
Y R+F+ ++LT PL L KK+ + WT ++ + +K+ L + +L +
Sbjct: 650 NYLRKFIPKTSQLTHPLNNLLKKDVRWKWTPTQTQAIENIKQCLVSPPVLRHFDFSKKIL 709
Query: 850 VYCDASHNGLGCVMMQEKK-----VIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHY 904
+ DAS +G V+ Q+ + Y S ++ + NY D E+ AI+ +LK WRHY
Sbjct: 710 LETDASDVAVGAVLSQKHDDDKYYPVGYYSAKMSKAQLNYSVSDKEMLAIIKSLKHWRHY 769
Query: 905 LYGSM--FTIYSDHKSL--KYLFDQKDLNMRQRRWMEFLQDYEFDLQ*HPGKTNVVADAL 960
L ++ F I +DH++L + + + N R RW FLQD+ F++ PG N +ADAL
Sbjct: 770 LESTIEPFKILTDHRNLIGRITNESEPENKRLARWQLFLQDFNFEINYRPGSANHIADAL 829
Query: 961 SRKAMHVSSLMVKELELLEAF*DLSLDVKIAPGKLSF-GMVTVSSGLLDEIKSKQETDEG 1019
SR +V E E + ++F ++++ +++ ++ D
Sbjct: 830 SR--------IVDETE--------PIPKDSEDNSINFVNQISITDDFKNQVVTEYTNDTK 873
Query: 1020 LLEWRKLVTQGKAPEFSIGSDNILRCKGRVCVPNDATMRRLILDEGHKSRLSIHPGMNKM 1079
LL + + ++ K ++ +PND + R I+ + H+ IHPG+ +
Sbjct: 874 LLNLLNNEDKRVEENIQLKDGLLINSKDQILLPNDTQLTRTIIKKYHEEGKLIHPGIELL 933
Query: 1080 YQDLKLHFWWPGMKKQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVT 1139
+ F W G++KQ+ EYV C TC K + KP G LQ + E W+S+SMDF+T
Sbjct: 934 TNIILRRFTWKGIRKQIQEYVQNCHTCQINKSRNHKPYGPLQPIPPSERPWESLSMDFIT 993
Query: 1140 ALPLTRRRFDAIWVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRD 1199
ALP + ++A++VVVDR +K A VP + + E+ A + +++ G P I++D D
Sbjct: 994 ALPESSG-YNALFVVVDRFSKMAILVPCTKSITAEQTARMFDQRVIAYFGNPKEIIADND 1052
Query: 1200 SRFTSRFWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELLP 1259
FTS+ W ++ S Y PQTDGQTERT Q++E LLR H +W + +
Sbjct: 1053 HIFTSQTWKDFAHKYNFVMKFSLPYRPQTDGQTERTNQTVEKLLRCVCSTHPNTWVDHIS 1112
Query: 1260 LIEFTYNNSFHASIGMAPYEALYGRRCQTPLCWHQDGEHLVIGPELVQQTTEEVKRIQEK 1319
L++ +YNN+ H++ M P+E ++ R L + E Q+T + + ++E
Sbjct: 1113 LVQQSYNNAIHSATQMTPFEIVH--RYSPALSPLELPSFSDKTDENSQETIQVFQTVKEH 1170
Query: 1320 MRIS*SRQKSYADNRRKEL-EFQAGDHVFLRVTPMTGVGRAIKSKKLTPKFIGPYQITER 1378
+ + + K Y D + +E+ EFQ GD V ++ T G KS KL P F GP+ + ++
Sbjct: 1171 LNTNNIKMKKYFDMKIQEIEEFQPGDLVMVK---RTKTGFLHKSNKLAPSFAGPFYVLQK 1227
Query: 1379 VGPVAYRIALPPFLSHI-HDVLHVSQLRKY 1407
GP Y + LP + H+ HVS L KY
Sbjct: 1228 SGPNNYELDLPDSIKHMFSSTFHVSHLEKY 1257
>RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protein
type 2
Length = 1333
Score = 346 bits (888), Expect = 2e-94
Identities = 213/690 (30%), Positives = 350/690 (49%), Gaps = 34/690 (4%)
Query: 730 KGKELYANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLA 789
K L N +KCEF +VKF+G+ IS +G I+ ++ W+QPK ++R F+G
Sbjct: 590 KNANLIINQAKCEFHQSQVKFIGYHISEKGFTPCQENIDKVLQWKQPKNRKELRQFLGSV 649
Query: 790 GYYRRFVKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGEPYD 849
Y R+F+ ++LT PL L KK+ + WT ++ + +K+ L + +L +
Sbjct: 650 NYLRKFIPKTSQLTHPLNNLLKKDVRWKWTPTQTQAIENIKQCLVSPPVLRHFDFSKKIL 709
Query: 850 VYCDASHNGLGCVMMQEKK-----VIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHY 904
+ DAS +G V+ Q+ + Y S ++ + NY D E+ AI+ +LK WRHY
Sbjct: 710 LETDASDVAVGAVLSQKHDDDKYYPVGYYSAKMSKAQLNYSVSDKEMLAIIKSLKHWRHY 769
Query: 905 LYGSM--FTIYSDHKSL--KYLFDQKDLNMRQRRWMEFLQDYEFDLQ*HPGKTNVVADAL 960
L ++ F I +DH++L + + + N R RW FLQD+ F++ PG N +ADAL
Sbjct: 770 LESTIEPFKILTDHRNLIGRITNESEPENKRLARWQLFLQDFNFEINYRPGSANHIADAL 829
Query: 961 SRKAMHVSSLMVKELELLEAF*DLSLDVKIAPGKLSF-GMVTVSSGLLDEIKSKQETDEG 1019
SR +V E E + ++F ++++ +++ ++ D
Sbjct: 830 SR--------IVDETE--------PIPKDSEDNSINFVNQISITDDFKNQVVTEYTNDTK 873
Query: 1020 LLEWRKLVTQGKAPEFSIGSDNILRCKGRVCVPNDATMRRLILDEGHKSRLSIHPGMNKM 1079
LL + + ++ K ++ +PND + R I+ + H+ IHPG+ +
Sbjct: 874 LLNLLNNEDKRVEENIQLKDGLLINSKDQILLPNDTQLTRTIIKKYHEEGKLIHPGIELL 933
Query: 1080 YQDLKLHFWWPGMKKQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVT 1139
+ F W G++KQ+ EYV C TC K + KP G LQ + E W+S+SMDF+T
Sbjct: 934 TNIILRRFTWKGIRKQIQEYVQNCHTCQINKSRNHKPYGPLQPIPPSERPWESLSMDFIT 993
Query: 1140 ALPLTRRRFDAIWVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRD 1199
ALP + ++A++VVVDR +K A VP + + E+ A + +++ G P I++D D
Sbjct: 994 ALPESSG-YNALFVVVDRFSKMAILVPCTKSITAEQTARMFDQRVIAYFGNPKEIIADND 1052
Query: 1200 SRFTSRFWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELLP 1259
FTS+ W ++ S Y PQTDGQTERT Q++E LLR H +W + +
Sbjct: 1053 HIFTSQTWKDFAHKYNFVMKFSLPYRPQTDGQTERTNQTVEKLLRCVCSTHPNTWVDHIS 1112
Query: 1260 LIEFTYNNSFHASIGMAPYEALYGRRCQTPLCWHQDGEHLVIGPELVQQTTEEVKRIQEK 1319
L++ +YNN+ H++ M P+E ++ R L + E Q+T + + ++E
Sbjct: 1113 LVQQSYNNAIHSATQMTPFEIVH--RYSPALSPLELPSFSDKTDENSQETIQVFQTVKEH 1170
Query: 1320 MRIS*SRQKSYADNRRKEL-EFQAGDHVFLRVTPMTGVGRAIKSKKLTPKFIGPYQITER 1378
+ + + K Y D + +E+ EFQ GD V ++ T G KS KL P F GP+ + ++
Sbjct: 1171 LNTNNIKMKKYFDMKIQEIEEFQPGDLVMVK---RTKTGFLHKSNKLAPSFAGPFYVLQK 1227
Query: 1379 VGPVAYRIALPPFLSHI-HDVLHVSQLRKY 1407
GP Y + LP + H+ HVS L KY
Sbjct: 1228 SGPNNYELDLPDSIKHMFSSTFHVSHLEKY 1257
>RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protein
type 1
Length = 1333
Score = 346 bits (888), Expect = 2e-94
Identities = 213/690 (30%), Positives = 350/690 (49%), Gaps = 34/690 (4%)
Query: 730 KGKELYANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLA 789
K L N +KCEF +VKF+G+ IS +G I+ ++ W+QPK ++R F+G
Sbjct: 590 KNANLIINQAKCEFHQSQVKFIGYHISEKGFTPCQENIDKVLQWKQPKNRKELRQFLGSV 649
Query: 790 GYYRRFVKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGEPYD 849
Y R+F+ ++LT PL L KK+ + WT ++ + +K+ L + +L +
Sbjct: 650 NYLRKFIPKTSQLTHPLNNLLKKDVRWKWTPTQTQAIENIKQCLVSPPVLRHFDFSKKIL 709
Query: 850 VYCDASHNGLGCVMMQEKK-----VIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHY 904
+ DAS +G V+ Q+ + Y S ++ + NY D E+ AI+ +LK WRHY
Sbjct: 710 LETDASDVAVGAVLSQKHDDDKYYPVGYYSAKMSKAQLNYSVSDKEMLAIIKSLKHWRHY 769
Query: 905 LYGSM--FTIYSDHKSL--KYLFDQKDLNMRQRRWMEFLQDYEFDLQ*HPGKTNVVADAL 960
L ++ F I +DH++L + + + N R RW FLQD+ F++ PG N +ADAL
Sbjct: 770 LESTIEPFKILTDHRNLIGRITNESEPENKRLARWQLFLQDFNFEINYRPGSANHIADAL 829
Query: 961 SRKAMHVSSLMVKELELLEAF*DLSLDVKIAPGKLSF-GMVTVSSGLLDEIKSKQETDEG 1019
SR +V E E + ++F ++++ +++ ++ D
Sbjct: 830 SR--------IVDETE--------PIPKDSEDNSINFVNQISITDDFKNQVVTEYTNDTK 873
Query: 1020 LLEWRKLVTQGKAPEFSIGSDNILRCKGRVCVPNDATMRRLILDEGHKSRLSIHPGMNKM 1079
LL + + ++ K ++ +PND + R I+ + H+ IHPG+ +
Sbjct: 874 LLNLLNNEDKRVEENIQLKDGLLINSKDQILLPNDTQLTRTIIKKYHEEGKLIHPGIELL 933
Query: 1080 YQDLKLHFWWPGMKKQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVT 1139
+ F W G++KQ+ EYV C TC K + KP G LQ + E W+S+SMDF+T
Sbjct: 934 TNIILRRFTWKGIRKQIQEYVQNCHTCQINKSRNHKPYGPLQPIPPSERPWESLSMDFIT 993
Query: 1140 ALPLTRRRFDAIWVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRD 1199
ALP + ++A++VVVDR +K A VP + + E+ A + +++ G P I++D D
Sbjct: 994 ALPESSG-YNALFVVVDRFSKMAILVPCTKSITAEQTARMFDQRVIAYFGNPKEIIADND 1052
Query: 1200 SRFTSRFWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELLP 1259
FTS+ W ++ S Y PQTDGQTERT Q++E LLR H +W + +
Sbjct: 1053 HIFTSQTWKDFAHKYNFVMKFSLPYRPQTDGQTERTNQTVEKLLRCVCSTHPNTWVDHIS 1112
Query: 1260 LIEFTYNNSFHASIGMAPYEALYGRRCQTPLCWHQDGEHLVIGPELVQQTTEEVKRIQEK 1319
L++ +YNN+ H++ M P+E ++ R L + E Q+T + + ++E
Sbjct: 1113 LVQQSYNNAIHSATQMTPFEIVH--RYSPALSPLELPSFSDKTDENSQETIQVFQTVKEH 1170
Query: 1320 MRIS*SRQKSYADNRRKEL-EFQAGDHVFLRVTPMTGVGRAIKSKKLTPKFIGPYQITER 1378
+ + + K Y D + +E+ EFQ GD V ++ T G KS KL P F GP+ + ++
Sbjct: 1171 LNTNNIKMKKYFDMKIQEIEEFQPGDLVMVK---RTKTGFLHKSNKLAPSFAGPFYVLQK 1227
Query: 1379 VGPVAYRIALPPFLSHI-HDVLHVSQLRKY 1407
GP Y + LP + H+ HVS L KY
Sbjct: 1228 SGPNNYELDLPDSIKHMFSSTFHVSHLEKY 1257
>YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III
Length = 2186
Score = 201 bits (512), Expect = 1e-50
Identities = 180/684 (26%), Positives = 311/684 (45%), Gaps = 74/684 (10%)
Query: 739 SKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLAGYYRRFVKD 798
SKC +EV++LGH ++ GV K + + + +P +++SF+GL GYYR+F+ +
Sbjct: 1129 SKCHIAKKEVEYLGHKVTLDGVETQEVKTDKMKQFSRPTNVKELQSFLGLVGYYRKFILN 1188
Query: 799 YAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALP------KEGEPYDVYC 852
+A++ S LT L + W ++ E +FQE+KK + +LA P K P+ +Y
Sbjct: 1189 FAQIASSLTSLISAKVAWIWEKEQEIAFQELKKLVCQTPVLAQPDVEAALKGDRPFMIYT 1248
Query: 853 DASHNGLGCVMMQE-----KKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLYG 907
DAS G+G V+ QE + IA+AS+ L E Y DLE A+++AL+ ++ +YG
Sbjct: 1249 DASRKGIGAVLAQEGPDGQQHPIAFASKALSPAETRYHITDLEALAMMFALRRFKTIIYG 1308
Query: 908 SMFTIYSDHKSLKYLFDQKDLNMRQRRWMEFLQDYEFDLQ*HPGKTNVVADALSRKAMHV 967
+ T+++DHK L L L R RW + +++ + GK N VADALSR
Sbjct: 1309 TAITVFTDHKPLISLLKGSPLADRLWRWSIEILEFDVKIVYLAGKANAVADALSRGGCPP 1368
Query: 968 SSLMVKELELLEAF*DLSLDVKIAPGKLSFGMVTVSSGLLDEIKSKQETDEGLLEWRKLV 1027
+ L ++ + L + + + + SS L+ +K + DEG W++++
Sbjct: 1369 NELEEEQTKELTSI--------VNAIQTELPDILDSSCWLERLKGE---DEG---WKEVI 1414
Query: 1028 -------TQGKAPEFSIGSDNILRC---------------KGRVCVPNDATMRRLILDEG 1065
T+G I S+ L + R VP +R +L E
Sbjct: 1415 AALEGGKTKGTFKIVGIESEISLEYYKIVGGVLKNTEIEEQSRSVVPE--KIRTPLLKEL 1472
Query: 1066 HKSRLSIHPGMNKMYQDLKLHFWWPGMKKQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDV 1125
H+ L+ H G+ KM++ + F+WP M+ V V TC C A +H K L +
Sbjct: 1473 HEGMLAGHFGIKKMWRMVHRKFYWPQMRVCVENCVRTCAKCLCAN-DHSKLTSSLTPYRM 1531
Query: 1126 PEWKWDSISMDFV-TALPLTRRRFDAIWVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKI 1184
+ + ++ D + L + R+ I ++D TK VPI + K E + + + +
Sbjct: 1532 -TFPLEIVACDLMDVGLSVQGNRY--ILTIIDLFTKYGTAVPIP-DKKAETVLKAFVERW 1587
Query: 1185 VRLHG-VPSSIVSDRDSRFTSRFWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLL 1243
G +P +++D+ F + + L + + YN + +G ER +++ ++
Sbjct: 1588 AIGEGRIPLKLLTDQGKEFVNGLFAQFTHMLKIEHITTKGYNSRANGAVERFNKTIMHIM 1647
Query: 1244 RACVLDHKGSWDELLPLIEFTYNNSFHASIGMAPYEALYGRRCQTPLCWHQDGEHLV--- 1300
+ WD+ + + YNN H + G P ++GR PL GE V
Sbjct: 1648 KKKTAVPM-EWDDQVVYAVYAYNNCVHENTGETPMFLMHGRDVMGPL--EMSGEDAVGIN 1704
Query: 1301 --IGPELVQQTTEEVKRIQEKMRIS*SRQ----KSYADNR---RKELEFQAGDHVFLRVT 1351
E T+E+ ++Q+ + R+ KS D + +K Q G V L +
Sbjct: 1705 YADMDEYKHLLTQELLKVQKIAKEHAMREQESYKSLFDQKYASKKHRFPQPGSRVLLEI- 1763
Query: 1352 PMTGVGRAIKSKKLTPKFIGPYQI 1375
P +G + KL K+ GPY++
Sbjct: 1764 PSEKLG--AQCPKLVNKWSGPYRV 1785
>POL3_DROME (P04323) Retrovirus-related Pol polyprotein from
transposon 17.6 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1058
Score = 183 bits (464), Expect = 3e-45
Identities = 146/574 (25%), Positives = 260/574 (44%), Gaps = 50/574 (8%)
Query: 740 KCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLAGYYRRFVKDY 799
KCEF +E FLGHV++ G+ +P KIE+I + P +I++F+GL GYYR+F+ ++
Sbjct: 400 KCEFLKQETTFLGHVLTPDGIKPNPEKIEAIQKYPIPTKPKEIKAFLGLTGYYRKFIPNF 459
Query: 800 AKLTSPLTQLTKKNQPFAWTE-KCEESFQEMKKRLTTALILALPKEGEPYDVYCDASHNG 858
A + P+T+ KKN T + + +F+++K ++ IL +P + + + DAS
Sbjct: 460 ADIAKPMTKCLKKNMKIDTTNPEYDSAFKKLKYLISEDPILKVPDFTKKFTLTTDASDVA 519
Query: 859 LGCVMMQEKKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLYGSMFTIYSDHKS 918
LG V+ Q+ ++Y SR L HE NY T + EL AIV+A K +RHYL G F I SDH+
Sbjct: 520 LGAVLSQDGHPLSYISRTLNEHEINYSTIEKELLAIVWATKTFRHYLLGRHFEISSDHQP 579
Query: 919 LKYLFDQKDLNMRQRRWMEFLQDYEFDLQ*HPGKTNVVADALSRKAMHVSSLMVKELELL 978
L +L+ KD N + RW L +++FD++ GK N VADALSR + + L +
Sbjct: 580 LSWLYRMKDPNSKLTRWRVKLSEFDFDIKYIKGKENCVADALSRIKLEETYLSEQTQHSA 639
Query: 979 EAF*DLSLDVKIAPGKLSFGMVTVSSGLLDEIKSKQETDEGLLEWRKLVTQGKAPEFSIG 1038
E + + P V S G D +K + ++T+ KA ++ I
Sbjct: 640 EEDNSDLIFITERPLNTFNRQVIFSKGPPDIKVTKYFKKHITQIFYDIMTREKAEQYLI- 698
Query: 1039 SDNILRCKGRVCVPNDATMRRLILDEGHKSRLS--------------------------- 1071
D+ K + + +DA ++ HK ++
Sbjct: 699 -DHFCGKKSALYIESDADFE--VIQAAHKLAINTKYTKILRSTILLKNITTYAEFKELIL 755
Query: 1072 ------IHPGMNKMYQDLKLHFWWPGMKKQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDV 1125
+HPG+ K + +++P + + ++ C C AK EH+ ++
Sbjct: 756 TAHEKLLHPGIQKTTKLFGETYYFPNSQLLIQNIINECSICNLAKTEHRNTDMPTKTTPK 815
Query: 1126 PEWKWDSISMDFVTALPLTRRRFDAIWVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIV 1185
PE + +D ++ + + + +D +K A I +E + + +I
Sbjct: 816 PEHCREKFMIDIYSS---EGKHYVS---CIDIYSKFATLEEIKTKDWIE--CKNALMRIF 867
Query: 1186 RLHGVPSSIVSDRDSRFTSRFWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRA 1245
G P + +DRD F+S ++ +L+L++ D ER +++ + +R
Sbjct: 868 NQLGKPKLLKADRDGAFSSLALKRWLESEEVELQLNTTKTGVAD--IERLHKTINEKIRI 925
Query: 1246 CVL--DHKGSWDELLPLIEFTYNNSFHASIGMAP 1277
D + ++ ++ + + H + G P
Sbjct: 926 IKTSDDEETKLSKMETVLNIYNHKTKHDTTGQTP 959
>POL5_DROME (Q8I7P9) Retrovirus-related Pol polyprotein from
transposon opus [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1003
Score = 178 bits (451), Expect = 1e-43
Identities = 169/683 (24%), Positives = 290/683 (41%), Gaps = 98/683 (14%)
Query: 734 LYANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLAGYYR 793
L N K F +V+FLG+++++ G+ DP K+ +I P +++ F+G+ YYR
Sbjct: 310 LQVNLEKSHFLDTQVEFLGYIVTADGIKADPKKVRAISEMPPPTSVKELKRFLGMTSYYR 369
Query: 794 RFVKDYAKLTSPLTQLTK-----------KNQPFAWTEKCEESFQEMKKRLTTALILALP 842
+F++DYAK+ PLT LT+ P E +SF ++K L ++ ILA P
Sbjct: 370 KFIQDYAKVAKPLTNLTRGLYANIKSSQSSKVPITLDETALQSFNDLKSILCSSEILAFP 429
Query: 843 KEGEPYDVYCDASHNGLGCVMMQE----KKVIAYASRQLKIHEKNYPTHDLELAAIVYAL 898
+P+ + DAS+ +G V+ Q+ + IAY SR L E+NY T + E+ AI+++L
Sbjct: 430 CFTKPFHLTTDASNWAIGAVLSQDDQGRDRPIAYISRSLNKTEENYATIEKEMLAIIWSL 489
Query: 899 KIWRHYLYGS-MFTIYSDHKSLKYLFDQKDLNMRQRRWMEFLQDYEFDLQ*HPGKTNVVA 957
R YLYG+ +Y+DH+ L + ++ N + +RW +++Y +L PGK+NVVA
Sbjct: 490 DNLRAYLYGAGTIKVYTDHQPLTFALGNRNFNAKLKRWKARIEEYNCELIYKPGKSNVVA 549
Query: 958 DALSR---------------------------KAMHVSSLMVKELE----------LLEA 980
DALSR A+H SS ++ +E + +
Sbjct: 550 DALSRIPPQLNQLSTDLDANPEDDMQSLATAHSALHDSSRLIPHVESPINVFKNQLIFDT 609
Query: 981 F*DLSLDVKIAPGKLSFGMVTVSSGLLDEIKSKQETDEGLLEWRKLVTQG-KAPEF---- 1035
L PG + L D S Q R ++ G K PE
Sbjct: 610 TRSKYLCEHPFPGYTRHLIPLKDGSLADLTNSLQSC------LRPVIINGVKIPEAHLQR 663
Query: 1036 --SIGSDNILRCKGRVC--VPNDATMRRLILDEGHKSRLSIHPGMNKMYQDLKLHFWWPG 1091
SI N L K R+ + D + I + K H G ++ L +++P
Sbjct: 664 FQSICLANFLLYKIRITQRLVADVSGAEEICEIIEKEHRRAHRGPTEIRLQLLEKYYFPR 723
Query: 1092 MKKQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTRRRFDAI 1151
M + S+C C K E LQ +P + + + +D L +R + +
Sbjct: 724 MSSTIRLQTSSCQCCKLYKYERHPNKPNLQPTPIPNYPCEILHIDI---FALEKRLYLS- 779
Query: 1152 WVVVDRLTKTAHFVPISLNYKVEKLAEIHIAK--IVRLH--GVPSSIVSDRDSRFTSRFW 1207
+D+ +K A + ++ A +H+ + + LH P +VSD +
Sbjct: 780 --CIDKFSKFAKL------FHLQSKASVHLRETLVEALHYFTAPKVLVSDNERGLLCPTV 831
Query: 1208 GALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWD--ELLPLIEFTY 1265
++L L + + +GQ ER + ++ R C+ D ++ EL+ + Y
Sbjct: 832 LNYLRSLDIDLYYAPTQKSEVNGQVERFHSTFLEIYR-CLKDELPTFKPVELVHIAVDRY 890
Query: 1266 NNSFHASIGMAPYEALYGRRCQTPLCWHQDGEHLVIGPELVQQTTEEVKRIQEKMRIS*S 1325
N S H+ P + + R + D +QT E++K + E +I +
Sbjct: 891 NTSVHSVTNRKPADVFFDRSSRVNYQGLTD---------FRRQTLEDIKGLIEYKQIRGN 941
Query: 1326 RQKSYADNRRKELEFQAGDHVFL 1348
++ NR + + GD VF+
Sbjct: 942 MARN--KNRDEPKSYGPGDEVFV 962
>POLY_DROME (P10401) Retrovirus-related Pol polyprotein from
transposon gypsy [Contains: Reverse transcriptase (EC
2.7.7.49); Endonuclease]
Length = 1035
Score = 176 bits (446), Expect = 4e-43
Identities = 168/676 (24%), Positives = 288/676 (41%), Gaps = 91/676 (13%)
Query: 740 KCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLAGYYRRFVKDY 799
K F+ E V++LG ++S G DP K+++I + +P +RSF+GLA YYR F+KD+
Sbjct: 375 KTRFFKESVEYLGFIVSKDGTKSDPEKVKAIQEYPEPDCVYKVRSFLGLASYYRVFIKDF 434
Query: 800 AKLTSPLTQLTK-----------KNQPFAWTEKCEESFQEMKKRLTTA-LILALPKEGEP 847
A + P+T + K K P + E +FQ ++ L + +IL P +P
Sbjct: 435 AAIARPITDILKGENGSVSKHMSKKIPVEFNETQRNAFQRLRNILASEDVILKYPDFKKP 494
Query: 848 YDVYCDASHNGLGCVMMQEKKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLYG 907
+D+ DAS +G+G V+ QE + I SR LK E+NY T++ EL AIV+AL +++LYG
Sbjct: 495 FDLTTDASASGIGAVLSQEGRPITMISRTLKQPEQNYATNERELLAIVWALGKLQNFLYG 554
Query: 908 SM-FTIYSDHKSLKYLFDQKDLNMRQRRWMEFLQDYEFDLQ*HPGKTNVVADALSRKAMH 966
S I++DH+ L + ++ N + +RW ++ + + PGK N VADALSR+ ++
Sbjct: 555 SREINIFTDHQPLTFAVADRNTNAKIKRWKSYIDQHNAKVFYKPGKENFVADALSRQNLN 614
Query: 967 V--------SSLMVKELEL----------LEAF*D-LSLDVKIAPGKLSFGM-------- 999
++ + EL L L F + + L+ P K + +
Sbjct: 615 ALQNEPQSDAATIHSELSLTYTVETTDKPLNCFRNQIILEAARFPLKRNLVLFRSKSRHL 674
Query: 1000 --VTVSSGLLDEIKSKQETDEGLLEWRKLVTQGKAPEFSIG---SDNILRCKGRVCVPND 1054
T S LL +K D L T I + CK V D
Sbjct: 675 ISFTDKSWLLKTLKEVVNPDVVNAIHCDLPTLASFQHDLIAHFPATQFRHCKNVVLDITD 734
Query: 1055 ATMR-RLILDEGHKSRLSIHPGMNKMYQDLKLHFWWPGMKKQVAEYVSTCLTC*NAKVEH 1113
+ ++ E +++ + + ++ +D +++P M E V+ C C AK +
Sbjct: 735 KNEQIEIVTAEHNRAHRAAQENIKQVLRD----YYFPKMGSLAKEVVANCRVCTQAKYDR 790
Query: 1114 QKPAGKLQSLDVPEWKWDSISMDFVTALPLTRRRFDAIWVVVDRLTKTAHFVPISLNYKV 1173
+L +P + + + +D + T R+ +D+ +K A P+ V
Sbjct: 791 HPKKQELGETPIPSYTGEMVHIDIFS----TDRKL--FLTCIDKFSKYAIVQPVVSRTIV 844
Query: 1174 EKLAEIHIAKIVRLHGVPSSIVSDRDSRFTSR-FWGALQQALGTKLRLSSAYNPQTDGQT 1232
+ A + +I+ L ++ D + F S L+ + G + + + ++GQ
Sbjct: 845 DITAP--LLQIINLFPNIKTVYCDNEPAFNSETVTSMLKNSFGIDIVNAPPLHSSSNGQV 902
Query: 1233 ERTIQSLEDLLRACVLDHK-GSWDELLPLIEFTYNNSFHASIGMAPYEALYGRRCQTPLC 1291
ER +L ++ R LD K EL+ YN + H+ P E
Sbjct: 903 ERFHSTLAEIARCLKLDKKTNDTVELILRATIEYNKTVHSVTRERPIE------------ 950
Query: 1292 WHQDGEHLVIGPELVQQTTEEVKRIQEKMRIS*SRQKSYADNRRKELEFQAGDHVFLRVT 1351
V+ P ++ E R+ + + S R NR F+ G+ VF++
Sbjct: 951 --------VVHPGAHERCLEIKARLVKAQQDSIGRNNPSRQNR----VFEVGERVFVKNN 998
Query: 1352 PMTGVGRAIKSKKLTP 1367
G KLTP
Sbjct: 999 KRLG-------NKLTP 1007
>POL2_DROME (P20825) Retrovirus-related Pol polyprotein from
transposon 297 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1059
Score = 168 bits (426), Expect = 9e-41
Identities = 91/225 (40%), Positives = 135/225 (59%), Gaps = 3/225 (1%)
Query: 740 KCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLAGYYRRFVKDY 799
KCEF +E FLGH+++ G+ +P K+++I+S+ P +IR+F+GL GYYR+F+ +Y
Sbjct: 399 KCEFLKKEANFLGHIVTPDGIKPNPIKVKAIVSYPIPTKDKEIRAFLGLTGYYRKFIPNY 458
Query: 800 AKLTSPLTQLTKKNQPFAWTEKCE--ESFQEMKKRLTTALILALPKEGEPYDVYCDASHN 857
A + P+T KK T+K E E+F+++K + IL LP + + + DAS+
Sbjct: 459 ADIAKPMTSCLKKRTKID-TQKLEYIEAFEKLKALIIRDPILQLPDFEKKFVLTTDASNL 517
Query: 858 GLGCVMMQEKKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLYGSMFTIYSDHK 917
LG V+ Q I++ SR L HE NY + EL AIV+A K +RHYL G F I SDH+
Sbjct: 518 ALGAVLSQNGHPISFISRTLNDHELNYSAIEKELLAIVWATKTFRHYLLGRQFLIASDHQ 577
Query: 918 SLKYLFDQKDLNMRQRRWMEFLQDYEFDLQ*HPGKTNVVADALSR 962
L++L + K+ + RW L +Y+F + GK N VADALSR
Sbjct: 578 PLRWLHNLKEPGAKLERWRVRLSEYQFKIDYIKGKENSVADALSR 622
Score = 42.4 bits (98), Expect = 0.010
Identities = 51/227 (22%), Positives = 96/227 (41%), Gaps = 19/227 (8%)
Query: 1055 ATMRRLILDEGHKSRLSIHPGMNKMYQDLKLHFWWPGMKKQVAEYVSTCLTC*NAKVEHQ 1114
A + +IL K +HPG+ KM + K + ++P + + ++ C C AK EH+
Sbjct: 748 AEFKEIILQSHEKL---LHPGIQKMTKLFKENHFFPNSQLLIQNIINECNICNLAKTEHR 804
Query: 1115 KPAGKLQSLDVPEWKWDSISMDFVTALPLTRRRFDAIWVVVDRLTKTAHFVPISLNYKVE 1174
L+ PE + +D ++ + + + +D +K A I +E
Sbjct: 805 NTKMPLKITPNPEHCREKFVVDIYSS---EGKHYIS---CIDIYSKFATLEQIKTKDWIE 858
Query: 1175 KLAEIHIAKIVRLHGVPSSIVSDRDSRFTSRFWGALQQALGTKLRLSSAYNPQTDGQTER 1234
+ +I G P + +DRD F+S + +L+L++A N D ER
Sbjct: 859 --CRNALMRIFNQLGKPKLLKADRDGAFSSLALKRWLEEEEVELQLNTAKNGVAD--VER 914
Query: 1235 TIQSLEDLLRACVLDHKGSWDELLPLIE---FTYNNSF-HASIGMAP 1277
+++ + +R +++ + L IE +TYN H + G P
Sbjct: 915 LHKTINEKIR--IINSSDDEEVKLSKIETILYTYNQKIKHDTTGQRP 959
>POL4_DROME (P10394) Retrovirus-related Pol polyprotein from
transposon 412 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1237
Score = 167 bits (424), Expect = 2e-40
Identities = 97/269 (36%), Positives = 146/269 (54%), Gaps = 15/269 (5%)
Query: 712 RRVKKNTKSICVKC*EYCKGKELYANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIM 771
+ + KN + KC EY L + KC F+M EV FLGH + +G+ D K + I
Sbjct: 484 KHMLKNLTEVFGKCREY----NLKLHPEKCSFFMHEVTFLGHKCTDKGILPDDKKYDVIQ 539
Query: 772 SWEQPKIASDIRSFVGLAGYYRRFVKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKK 831
++ P A R FV YYRRF+K++A + +T+L KKN PF WT++C+++F +K
Sbjct: 540 NYPVPHDADSARRFVAFCNYYRRFIKNFADYSRHITRLCKKNVPFEWTDECQKAFIHLKS 599
Query: 832 RLTTALILALPKEGEPYDVYCDASHNGLGCVMMQ----EKKVIAYASRQLKIHEKNYPTH 887
+L +L P + + + DAS G V+ Q + +AYASR E N T
Sbjct: 600 QLINPTLLQYPDFSKEFCITTDASKQACGAVLTQNHNGHQLPVAYASRAFTKGESNKSTT 659
Query: 888 DLELAAIVYALKIWRHYLYGSMFTIYSDHKSLKYLFDQKDLNMRQRRWMEFLQDYEFDLQ 947
+ ELAAI +A+ +R Y+YG FT+ +DH+ L YLF + + + R L++Y F ++
Sbjct: 660 EQELAAIHWAIIHFRPYIYGKHFTVKTDHRPLTYLFSMVNPSSKLTRIRLELEEYNFTVE 719
Query: 948 *HPGKTNVVADALSRKAMHVSSLMVKELE 976
GK N VADALSR + +KEL+
Sbjct: 720 YLKGKDNHVADALSR-------ITIKELK 741
Score = 99.4 bits (246), Expect = 7e-20
Identities = 95/398 (23%), Positives = 168/398 (41%), Gaps = 40/398 (10%)
Query: 1007 LDEIKSKQETDEGLLE--------WRKLVTQGKAPEFSIGSDNILRCKGRVCVPNDATM- 1057
LD+ + E G+ + W+K+ +F + IL+ +V + N T
Sbjct: 829 LDQFLQRLELQAGIYDISQIKMAPWKKIFEHVSIDKFKNMGNKILK-NLKVALLNPVTQI 887
Query: 1058 -----RRLILDEGHKSRLSI-HPGMNKMYQDLKLHFWWPGMKKQVAEYVSTCLTC*NAKV 1111
+ IL H + H G+ K +K H++W M K + EYV C C AK
Sbjct: 888 NNEKEKEAILSTLHDDPIQGGHTGITKTLAKVKRHYYWKNMSKYIKEYVRKCQKCQKAKT 947
Query: 1112 -EHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTRRRFDAIWVVVDRLTKTAHFVPISLN 1170
+H K + + PE +D + +D + LP + + ++ LTK +PI+ N
Sbjct: 948 TKHTKTP--MTITETPEHAFDRVVVDTIGPLPKSENGNEYAVTLICDLTKYLVAIPIA-N 1004
Query: 1171 YKVEKLAEIHIAKIVRLHGVPSSIVSDRDSRFTSRFWGALQQALGTKLRLSSAYNPQTDG 1230
+ +A+ + +G + ++D + + + L + L K S+A++ QT G
Sbjct: 1005 KSAKTVAKAIFESFILKYGPMKTFITDMGTEYKNSIITDLCKYLKIKNITSTAHHHQTVG 1064
Query: 1231 QTERTIQSLEDLLRACVLDHKGSWDELLPLIEFTYNNSFHASIGMAPYEALYGRRCQTPL 1290
ER+ ++L + +R+ + K WD L + +N + PYE ++GR P
Sbjct: 1065 VVERSHRTLNEYIRSYISTDKTDWDVWLQYFVYCFNTTQSMVHNYCPYELVFGRTSNLPK 1124
Query: 1291 CWHQDGEHLVIGPELVQQTTEEVK--------RIQEKMRIS*SRQKSYADNRRKELEFQA 1342
H + H + + +E K R ++ + + K D + K++E +
Sbjct: 1125 --HFNKLHSIEPIYNIDDYAKESKYRLEVAYARARKLLEAHKEKNKENYDLKVKDIELEV 1182
Query: 1343 GDHVFLRVTPMTGVGRAIKSKKLTPKFIGPYQITERVG 1380
GD V LR VG KL K+ GPY+I E +G
Sbjct: 1183 GDKVLLR----NEVGH-----KLDFKYTGPYKI-ESIG 1210
>M860_ARATH (P92523) Hypothetical mitochondrial protein AtMg00860
(ORF158)
Length = 158
Score = 114 bits (286), Expect = 2e-24
Identities = 53/118 (44%), Positives = 79/118 (66%), Gaps = 3/118 (2%)
Query: 733 ELYANGSKCEFWMEEVKFLGH--VISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLAG 790
+ YAN KC F ++ +LGH +IS +GV+ DP+K+E+++ W +PK +++R F+GL G
Sbjct: 15 QFYANRKKCAFGQPQIAYLGHRHIISGEGVSADPAKLEAMVGWPEPKNTTELRGFLGLTG 74
Query: 791 YYRRFVKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGEPY 848
YYRRFVK+Y K+ PLT+L KKN WTE +F+ +K +TT +LALP P+
Sbjct: 75 YYRRFVKNYGKIVRPLTELLKKNS-LKWTEMAALAFKALKGAVTTLPVLALPDLKLPF 131
>POL_GALV (P21414) Pol polyprotein [Contains: Protease (EC
3.4.23.-); Reverse transcriptase/ribonuclease H (EC
2.7.7.49) (EC 3.1.26.4) (RT); Integrase (IN)]
Length = 1165
Score = 90.9 bits (224), Expect = 2e-17
Identities = 53/184 (28%), Positives = 93/184 (49%), Gaps = 4/184 (2%)
Query: 737 NGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLAGYYRRFV 796
+ K + EV +LG+++ + P++ ++M P +R F+G AG+ R ++
Sbjct: 372 SAKKAQLCQREVTYLGYLLKEGKRWLTPARKATVMKIPVPTTPRQVREFLGTAGFCRLWI 431
Query: 797 KDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGEPYDVYCDASH 856
+A L +PL LTK++ PF WTE+ +++F +KK L +A LALP +P+ +Y D
Sbjct: 432 PGFASLAAPLYPLTKESIPFIWTEEHQQAFDHIKKALLSAPALALPDLTKPFTLYIDERA 491
Query: 857 NGLGCVMMQE----KKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLYGSMFTI 912
V+ Q ++ +AY S++L +PT +AA+ LK G T+
Sbjct: 492 GVARGVLTQTLGPWRRPVAYLSKKLDPVASGWPTCLKAVAAVALLLKDADKLTLGQNVTV 551
Query: 913 YSDH 916
+ H
Sbjct: 552 IASH 555
Score = 63.9 bits (154), Expect = 3e-09
Identities = 55/196 (28%), Positives = 82/196 (41%), Gaps = 8/196 (4%)
Query: 1090 PGMKKQVAEYVSTCLTC*NAK-VEHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTRRRF 1148
P ++ V E S C C V + GK Q D P W+ +DF P R
Sbjct: 839 PNLQSAVREVTSQCQACAMTNAVTTYRETGKRQRGDRPGVYWE---VDFTEIKP-GRYGN 894
Query: 1149 DAIWVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRDSRFTSRFWG 1208
+ V +D + P + +I + +I+ G+P + SD F ++
Sbjct: 895 KYLLVFIDTFSGWVEAFPTKTETALIVCKKI-LEEILPRFGIPKVLGSDNGPAFVAQVSQ 953
Query: 1209 ALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKG-SWDELLPLIEFTYNN 1267
L LG +L AY PQ+ GQ ER +++++ L L+ G W LLPL N
Sbjct: 954 GLATQLGINWKLHCAYRPQSSGQVERMNRTIKETLTKLALETGGKDWVTLLPLALLRARN 1013
Query: 1268 SFHASIGMAPYEALYG 1283
+ G+ PYE LYG
Sbjct: 1014 T-PGRFGLTPYEILYG 1028
>POL_FENV1 (P31792) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)] (Fragment)
Length = 1046
Score = 90.9 bits (224), Expect = 2e-17
Identities = 51/185 (27%), Positives = 95/185 (50%), Gaps = 4/185 (2%)
Query: 736 ANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLAGYYRRF 795
A+ K + +V +LG+++S + P +IE++ P+ ++R F+G AG+ R +
Sbjct: 229 ASAKKAQICQTKVTYLGYILSEGKRWLTPGRIETVAHIPPPQNPREVREFLGTAGFCRLW 288
Query: 796 VKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGEPYDVYCDAS 855
+ +A+L +PL LTK++ PF W EK + +F+ +K+ L +A L LP +P+ ++ D
Sbjct: 289 IPGFAELAAPLYALTKESAPFTWQEKHQSAFEALKEALLSAPALGLPDTSKPFTLFIDEK 348
Query: 856 HNGLGCVMMQE----KKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLYGSMFT 911
V+ Q+ K+ +AY S++L +P +AA +K G T
Sbjct: 349 QGIAKGVLTQKLGPWKRPVAYLSKKLDPVAAGWPPCLRIMAATAMLVKDSAKLTLGQPLT 408
Query: 912 IYSDH 916
+ + H
Sbjct: 409 VITPH 413
Score = 54.7 bits (130), Expect = 2e-06
Identities = 53/202 (26%), Positives = 81/202 (39%), Gaps = 8/202 (3%)
Query: 1084 KLHFWWPGMKKQVAEYVSTCLTC*NAKVEHQK-PAGKLQSLDVPEWKWDSISMDFVTALP 1142
K F P + + S C C + P GK + P W+ +DF P
Sbjct: 718 KTDFLIPKAGTLIEQVTSACKVCQQVNAGATRVPEGKRTRGNRPGVYWE---IDFTEVKP 774
Query: 1143 LTRRRFDAIWVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRDSRF 1202
+ + V VD + P + +A+ + +I G+P I SD F
Sbjct: 775 -HYAGYKYLLVFVDTFSGWVEAYP-TRQETAHMVAKKILEEIFPRFGLPKVIGSDNGPAF 832
Query: 1203 TSRFWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLD-HKGSWDELLPLI 1261
S+ L + LG +L AY PQ+ GQ ER +++++ L L+ W LL L
Sbjct: 833 VSQVSQGLARTLGINWKLHCAYRPQSSGQVERMNRTIKETLTKLTLETGLKDWRRLLSLA 892
Query: 1262 EFTYNNSFHASIGMAPYEALYG 1283
N+ + G+ PYE LYG
Sbjct: 893 LLRARNTPN-RFGLTPYEILYG 913
>POL_FOAMV (P14350) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)]
Length = 886
Score = 89.4 bits (220), Expect = 7e-17
Identities = 62/268 (23%), Positives = 119/268 (44%), Gaps = 10/268 (3%)
Query: 1038 GSDNILRCKGRVCVPNDATMRRLILDEGHKSRLSIHPGMNKMYQDLKLHFWWPGMKKQVA 1097
G + R +G +P + ++++L + + H G + +WWP M+K V
Sbjct: 589 GKVKVSRPEGVKIIPPQSDRQKIVLQAHNLA----HTGREATLLKIANLYWWPNMRKDVV 644
Query: 1098 EYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTRRRFDAIWVVVDR 1157
+ + C C + K +G + D P+ +D +D++ LP ++ + + VVVD
Sbjct: 645 KQLGRCQQCLITNASN-KASGPILRPDRPQKPFDKFFIDYIGPLPPSQG-YLYVLVVVDG 702
Query: 1158 LTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRDSRFTSRFWGALQQALGTK 1217
+T P + +++ + +P I SD+ + FTS + + G
Sbjct: 703 MTGFTWLYPTKAPSTSATVKSLNVLTSI---AIPKVIHSDQGAAFTSSTFAEWAKERGIH 759
Query: 1218 LRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELLPLIEFTYNNSFHASIGMAP 1277
L S+ Y+PQ+ + ER ++ LL ++ W +LLP+++ NN++ + P
Sbjct: 760 LEFSTPYHPQSGSKVERKNSDIKRLLTKLLVGRPTKWYDLLPVVQLALNNTYSPVLKYTP 819
Query: 1278 YEALYGRRCQTPLCWHQDGEHLVIGPEL 1305
++ L+G TP +QD L EL
Sbjct: 820 HQLLFGIDSNTPFA-NQDTLDLTREEEL 846
Score = 51.2 bits (121), Expect = 2e-05
Identities = 55/223 (24%), Positives = 94/223 (41%), Gaps = 11/223 (4%)
Query: 740 KCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLAGYYRRFVKDY 799
K E + V+FLG I+ +G + + +++ PK ++S +GL + R F+ ++
Sbjct: 151 KSEIGQKTVEFLGFNITKEGRGLTDTFKTKLLNITPPKDLKQLQSILGLLNFARNFIPNF 210
Query: 800 AKLTSPLTQL--TKKNQPFAWTEKCEESFQEMKKRLTTALIL--ALPKEGEPYDVYCDAS 855
A+L PL L + K + W+E+ + + + L TA L LP++ V S
Sbjct: 211 AELVQPLYNLIASAKGKYIEWSEENTKQLNMVIEALNTASNLEERLPEQRLVIKVNTSPS 270
Query: 856 HNGLGCVMMQEKKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLYGSMFTIYSD 915
+ KK I Y + E + + L + AL G +YS
Sbjct: 271 AGYVRYYNETGKKPIMYLNYVFSKAELKFSMLEKLLTTMHKALIKAMDLAMGQEILVYSP 330
Query: 916 HKSLKY-----LFDQKDLNMRQRRWMEFLQDYEFDLQ*HPGKT 953
S+ L ++K L +R WM +L+D +Q H KT
Sbjct: 331 IVSMTKIQKTPLPERKALPIRWITWMTYLEDPR--IQFHYDKT 371
>POL_SFV3L (P27401) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1157
Score = 87.4 bits (215), Expect = 3e-16
Identities = 62/254 (24%), Positives = 111/254 (43%), Gaps = 13/254 (5%)
Query: 1038 GSDNILRCKGRVCVPNDATMRRLILDEGHKSRLSIHPGMNKMYQDLKLHFWWPGMKKQVA 1097
G + R G+ +P + ++IL + + H G + + + +WWP ++K V
Sbjct: 799 GQVMVTRPNGKRIIPPKSDRPQIILQAHNIA----HTGRDSTFLKVSSKYWWPNLRKDVV 854
Query: 1098 EYVSTCLTC*--NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTRRRFDAIWVVV 1155
+ + C C NA P + + P +D +D++ LP + + VVV
Sbjct: 855 KVIRQCKQCLVTNAATLAAPPILRPER---PVKPFDKFFIDYIGPLPPSNGYLHVL-VVV 910
Query: 1156 DRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRDSRFTSRFWGALQQALG 1215
D +T P A + ++ VP I SD+ + FTS + + G
Sbjct: 911 DSMTGFVWLYPTKAP---STSATVKALNMLTSIAVPKVIHSDQGAAFTSATFADWAKNKG 967
Query: 1216 TKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELLPLIEFTYNNSFHASIGM 1275
+L S+ Y+PQ+ G+ ER ++ LL ++ W +LLP+++ NNS+ S
Sbjct: 968 IQLEFSTPYHPQSSGKVERKNSDIKRLLTKLLVGRPAKWYDLLPVVQLALNNSYSPSSKY 1027
Query: 1276 APYEALYGRRCQTP 1289
P++ L+G TP
Sbjct: 1028 TPHQLLFGIDSNTP 1041
Score = 44.7 bits (104), Expect = 0.002
Identities = 24/99 (24%), Positives = 48/99 (48%), Gaps = 2/99 (2%)
Query: 740 KCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLAGYYRRFVKDY 799
K E EV+FLG I+ +G + + + +++ P+ ++S +GL + R F+ ++
Sbjct: 362 KSEIAQHEVEFLGFNITKEGRGLTETFKQKLLNITPPRDLKQLQSILGLLNFARNFIPNF 421
Query: 800 AKLTSPLTQL--TKKNQPFAWTEKCEESFQEMKKRLTTA 836
++L PL + T + WT + Q + L +A
Sbjct: 422 SELVKPLYNIIATANGKYITWTTDNSQQLQNIISMLNSA 460
>POL_BAEVM (P10272) Pol polyprotein [Contains: Protease (EC
3.4.23.-); Reverse transcriptase/ribonuclease H (EC
2.7.7.49) (EC 3.1.26.4) (RT); Integrase (IN)]
Length = 1189
Score = 87.4 bits (215), Expect = 3e-16
Identities = 50/185 (27%), Positives = 94/185 (50%), Gaps = 4/185 (2%)
Query: 736 ANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLAGYYRRF 795
A+ K + +V +LG+++S + P +IE++ P+ ++R F+G AG+ R +
Sbjct: 372 ASAKKAQICQTKVTYLGYILSEGKRWLTPGRIETVARIPPPRNPREVREFLGTAGFCRLW 431
Query: 796 VKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGEPYDVYCDAS 855
+ +A+L +PL LTK++ PF W + + +F+ +KK L +A L LP +P+ ++ D
Sbjct: 432 IPGFAELAAPLYALTKESTPFTWQTEHQLAFEALKKALLSAPALGLPDTSKPFTLFLDER 491
Query: 856 HNGLGCVMMQE----KKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLYGSMFT 911
V+ Q+ K+ +AY S++L +P +AA +K G T
Sbjct: 492 QGIAKGVLTQKLGPWKRPVAYLSKKLDPVAAGWPPCLRIMAATAMLVKDSAKLTLGQPLT 551
Query: 912 IYSDH 916
+ + H
Sbjct: 552 VITPH 556
Score = 56.2 bits (134), Expect = 6e-07
Identities = 54/202 (26%), Positives = 82/202 (39%), Gaps = 8/202 (3%)
Query: 1084 KLHFWWPGMKKQVAEYVSTCLTC*NAKVEHQK-PAGKLQSLDVPEWKWDSISMDFVTALP 1142
K F P + + S C C + PAGK + P W+ +DF P
Sbjct: 861 KTDFLIPRASTLIEQVTSACKVCQQVNAGATRVPAGKRTRGNRPGVYWE---IDFTEVKP 917
Query: 1143 LTRRRFDAIWVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRDSRF 1202
+ + V VD + P + +A+ + +I G+P I SD F
Sbjct: 918 -HYAGYKYLLVFVDTFSGWVEAFP-TRQETAHIVAKKILEEIFPRFGLPKVIGSDNGPAF 975
Query: 1203 TSRFWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLD-HKGSWDELLPLI 1261
S+ L + LG +L AY PQ+ GQ ER +++++ L L+ W LL L
Sbjct: 976 VSQVSQGLARILGINWKLHCAYRPQSSGQVERMNRTIKETLTKLTLETGLKDWRRLLSLA 1035
Query: 1262 EFTYNNSFHASIGMAPYEALYG 1283
N+ + G+ PYE LYG
Sbjct: 1036 LLRARNTPN-RFGLTPYEILYG 1056
>YRD6_CAEEL (Q09575) Hypothetical protein K02A2.6 in chromosome II
Length = 1268
Score = 86.7 bits (213), Expect = 4e-16
Identities = 71/266 (26%), Positives = 126/266 (46%), Gaps = 27/266 (10%)
Query: 1048 RVCVPNDATMRRLILDEGHKSRLSIHPGMNKMYQDLKLHFWWPGMKKQVAEYVSTCLTC* 1107
RV VP ++++++L + H+ HPG+ +M Q + +W G+ + V C C
Sbjct: 775 RVIVPK--SLQKIVLKQLHEG----HPGIVQMKQKARSFVFWRGLDSDIENMVRHCNNCQ 828
Query: 1108 -NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTRRRFDAIWVVVDRLTKTAHFVP 1166
N+K+ P L VPE W I +DF L + VVVD TK A
Sbjct: 829 ENSKMPRVVP---LNPWPVPEAPWKRIHIDFAGPL-----NGCYLLVVVDAKTKYAE--- 877
Query: 1167 ISLNYKVEKLAEIHIAK-IVRLHGVPSSIVSDRDSRFTSRFWGALQQALGTKLRLSSAYN 1225
+ L + + I + + I +HG P +I+SD ++ TS + + Q+ G + + S+ Y
Sbjct: 878 VKLTRSISAVTTIDLLEEIFSIHGYPETIISDNGTQLTSHLFAQMCQSHGIEHKTSAVYY 937
Query: 1226 PQTDGQTERTIQSLEDLLRACVLDHKGSW---DELLPLIEFTYNNSFHASI-GMAPYEAL 1281
P+++G ER + D L+ + KG ++L +Y N+ H+++ G P E
Sbjct: 938 PRSNGAAERFV----DTLKRGIAKIKGEGSVNQQILNKFLISYRNTPHSALNGSTPAECH 993
Query: 1282 YGRRCQTPLCWHQDGEHLVIGPELVQ 1307
+GR+ +T + + ++ P+L Q
Sbjct: 994 FGRKIRTTMSLLMPTDRVLKVPKLTQ 1019
Score = 38.5 bits (88), Expect = 0.14
Identities = 21/63 (33%), Positives = 32/63 (50%), Gaps = 1/63 (1%)
Query: 737 NGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLAGYYRRFV 796
+ KC F ++V FLG V G D K E+I S + P + SF+G A + R +
Sbjct: 626 SAEKCAFAQKQVTFLGFV-DEHGRRPDSKKTEAIRSMKAPTDQKQLASFLGAADWLSRMM 684
Query: 797 KDY 799
+D+
Sbjct: 685 QDH 687
>POL_SFV1 (P23074) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1161
Score = 84.7 bits (208), Expect = 2e-15
Identities = 52/217 (23%), Positives = 96/217 (43%), Gaps = 5/217 (2%)
Query: 1073 HPGMNKMYQDLKLHFWWPGMKKQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDS 1132
H G + + + +WWP ++K V + + C C + L+ + P +D
Sbjct: 828 HTGRDATFLKVSSKYWWPNLRKDVVKSIRQCKQCLVTNATNLTSPPILRPVK-PLKPFDK 886
Query: 1133 ISMDFVTALPLTRRRFDAIWVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPS 1192
+D++ LP + + VVVD +T P A + ++ +P
Sbjct: 887 FYIDYIGPLPPSNGYLHVL-VVVDSMTGFVWLYPTKAP---STSATVKALNMLTSIAIPK 942
Query: 1193 SIVSDRDSRFTSRFWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKG 1252
+ SD+ + FTS + + G +L S+ Y+PQ+ G+ ER ++ LL ++
Sbjct: 943 VLHSDQGAAFTSSTFADWAKEKGIQLEFSTPYHPQSSGKVERKNSDIKRLLTKLLIGRPA 1002
Query: 1253 SWDELLPLIEFTYNNSFHASIGMAPYEALYGRRCQTP 1289
W +LLP+++ NNS+ S P++ L+G TP
Sbjct: 1003 KWYDLLPVVQLALNNSYSPSSKYTPHQLLFGVDSNTP 1039
Score = 47.0 bits (110), Expect = 4e-04
Identities = 29/99 (29%), Positives = 49/99 (49%), Gaps = 2/99 (2%)
Query: 740 KCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLAGYYRRFVKDY 799
K E EV+FLG I+ +G + + + +++ PK ++S +GL + R F+ +Y
Sbjct: 360 KSEIAQREVEFLGFNITKEGRGLTDTFKQKLLNITPPKDLKQLQSILGLLNFARNFIPNY 419
Query: 800 AKLTSPL-TQLTKKNQPF-AWTEKCEESFQEMKKRLTTA 836
++L PL T + N F +WTE Q + L A
Sbjct: 420 SELVKPLYTIVANANGKFISWTEDNSNQLQHIISVLNQA 458
>POL_FMVD (P09523) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 666
Score = 83.6 bits (205), Expect = 4e-15
Identities = 63/235 (26%), Positives = 110/235 (46%), Gaps = 14/235 (5%)
Query: 740 KCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSW-EQPKIASDIRSFVGLAGYYRRFVKD 798
K + E++ FLG I +E+I + ++ + ++ F+G+ Y ++
Sbjct: 430 KANLFKEKINFLGLEIDKGTHCPQNHILENIHKFPDRLEDKKHLQRFLGVLTYAETYIPK 489
Query: 799 YAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGEPYDVYCDASHNG 858
A++ PL KK+ + WT+ + +++KK L + L LPK + + DAS +
Sbjct: 490 LAEIRKPLQVKLKKDVTWNWTQSDSDYVKKIKKNLGSFPKLYLPKPEDHLIIETDASDSF 549
Query: 859 LGCVMMQE-----KKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLYGSMFTIY 913
G V+ + + Y+S K EKNY ++D EL A+ + + YL FT+
Sbjct: 550 WGGVLKARALDGVELICRYSSGSFKQAEKNYHSNDKELLAVKQVITKFSAYLTPVRFTVR 609
Query: 914 SDHKSLKYLF------DQKDLNMRQRRWMEFLQDYEFDLQ*HPGKTNVVADALSR 962
+D+K+ Y D K R RW + Y+FD++ G NV+AD L+R
Sbjct: 610 TDNKNFTYFLRINLKGDSK--QGRLVRWQNWFSKYQFDVEHLEGVKNVLADCLTR 662
>POL_MLVFP (P26808) Pol polyprotein [Contains: Protease (EC
3.4.23.-); Reverse transcriptase/ribonuclease H (EC
2.7.7.49) (EC 3.1.26.4) (RT); Integrase (IN)]
Length = 1204
Score = 81.6 bits (200), Expect = 1e-14
Identities = 49/185 (26%), Positives = 92/185 (49%), Gaps = 4/185 (2%)
Query: 736 ANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLAGYYRRF 795
A+ K + ++VK+LG+++ + ++ E++M PK +R F+G AG+ R +
Sbjct: 379 ASAKKAQICQKQVKYLGYLLKEGQRWLTEARKETVMGQPTPKTPRQLREFLGTAGFCRLW 438
Query: 796 VKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGEPYDVYCDAS 855
+ +A++ +PL LTK F W ++++QE+K+ L TA L LP +P++++ D
Sbjct: 439 IPGFAEMAAPLYPLTKTGTLFKWGPDQQKAYQEIKQALLTAPALGLPDLTKPFELFVDEK 498
Query: 856 HNGLGCVMMQE----KKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLYGSMFT 911
V+ Q+ ++ +AY S++L +P +AAI K G
Sbjct: 499 QGYAKGVLTQKLGPWRRPVAYLSKKLDPVAAGWPPCLRMVAAIAVLTKDAGKLTMGQPLV 558
Query: 912 IYSDH 916
I + H
Sbjct: 559 ILAPH 563
Score = 52.8 bits (125), Expect = 7e-06
Identities = 52/191 (27%), Positives = 82/191 (42%), Gaps = 17/191 (8%)
Query: 1189 GVPSSIVSDRDSRFTSRFWGALQQALGTKLRLSSAYNPQTDGQTERTIQSL-EDLLRACV 1247
G+P + +D F S+ + LG +L AY PQ+ GQ ER +++ E L + +
Sbjct: 972 GMPQVLGTDNGPAFVSKVSQTVADLLGVDWKLHCAYRPQSSGQVERMNRTIKETLTKLTL 1031
Query: 1248 LDHKGSWDELLPLIEFTYNNSFHASIGMAPYEALYGRRCQTPLCWHQDGEHLVI--GPEL 1305
W LLPL + N+ G+ PYE LYG PL D + + P L
Sbjct: 1032 ATGSRDWVLLLPLALYRARNT-PGPHGLTPYEILYG--APPPLVNFPDPDMAKVTHNPSL 1088
Query: 1306 VQQTTEEVKRIQ-EKMRIS*SRQKSYADNRRKELEFQAGDHVFLRVTPMTGVGRAIKSKK 1364
Q + + +Q E R + + D F+ GD V++ R ++K
Sbjct: 1089 -QAHLQALYLVQHEVWRPLAAAYQEQLDRPVVPHPFRVGDTVWV---------RRHQTKN 1138
Query: 1365 LTPKFIGPYQI 1375
L P++ GPY +
Sbjct: 1139 LEPRWKGPYTV 1149
>POL_MLVFF (P26809) Pol polyprotein [Contains: Protease (EC
3.4.23.-); Reverse transcriptase/ribonuclease H (EC
2.7.7.49) (EC 3.1.26.4) (RT); Integrase (IN)]
Length = 1204
Score = 81.6 bits (200), Expect = 1e-14
Identities = 49/185 (26%), Positives = 92/185 (49%), Gaps = 4/185 (2%)
Query: 736 ANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLAGYYRRF 795
A+ K + ++VK+LG+++ + ++ E++M PK +R F+G AG+ R +
Sbjct: 379 ASAKKAQICQKQVKYLGYLLKEGQRWLTEARKETVMGQPTPKTPRQLREFLGTAGFCRLW 438
Query: 796 VKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGEPYDVYCDAS 855
+ +A++ +PL LTK F W ++++QE+K+ L TA L LP +P++++ D
Sbjct: 439 IPGFAEMAAPLYPLTKTGTLFEWGPDQQKAYQEIKQALLTAPALGLPDLTKPFELFVDEK 498
Query: 856 HNGLGCVMMQE----KKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLYGSMFT 911
V+ Q+ ++ +AY S++L +P +AAI K G
Sbjct: 499 QGYAKGVLTQKLGPWRRPVAYLSKKLDPVAAGWPPCLRMVAAIAVLTKDAGKLTMGQPLV 558
Query: 912 IYSDH 916
I + H
Sbjct: 559 ILAPH 563
Score = 52.8 bits (125), Expect = 7e-06
Identities = 52/191 (27%), Positives = 82/191 (42%), Gaps = 17/191 (8%)
Query: 1189 GVPSSIVSDRDSRFTSRFWGALQQALGTKLRLSSAYNPQTDGQTERTIQSL-EDLLRACV 1247
G+P + +D F S+ + LG +L AY PQ+ GQ ER +++ E L + +
Sbjct: 972 GMPQVLGTDNGPAFVSKVSQTVADLLGVDWKLHCAYRPQSSGQVERMNRTIKETLTKLTL 1031
Query: 1248 LDHKGSWDELLPLIEFTYNNSFHASIGMAPYEALYGRRCQTPLCWHQDGEHLVI--GPEL 1305
W LLPL + N+ G+ PYE LYG PL D + + P L
Sbjct: 1032 ATGSRDWVLLLPLALYRARNT-PGPHGLTPYEILYG--APPPLVNFPDPDMAKVTHNPSL 1088
Query: 1306 VQQTTEEVKRIQ-EKMRIS*SRQKSYADNRRKELEFQAGDHVFLRVTPMTGVGRAIKSKK 1364
Q + + +Q E R + + D F+ GD V++ R ++K
Sbjct: 1089 -QAHLQALYLVQHEVWRPLAAAYQEQLDRPVVPHPFRVGDTVWV---------RRHQTKN 1138
Query: 1365 LTPKFIGPYQI 1375
L P++ GPY +
Sbjct: 1139 LEPRWKGPYTV 1149
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.342 0.148 0.511
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 158,411,066
Number of Sequences: 164201
Number of extensions: 6124449
Number of successful extensions: 17759
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 40
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 17604
Number of HSP's gapped (non-prelim): 91
length of query: 1489
length of database: 59,974,054
effective HSP length: 123
effective length of query: 1366
effective length of database: 39,777,331
effective search space: 54335834146
effective search space used: 54335834146
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 72 (32.3 bits)
Lotus: description of TM0388a.4


