

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0388a.2
(422 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
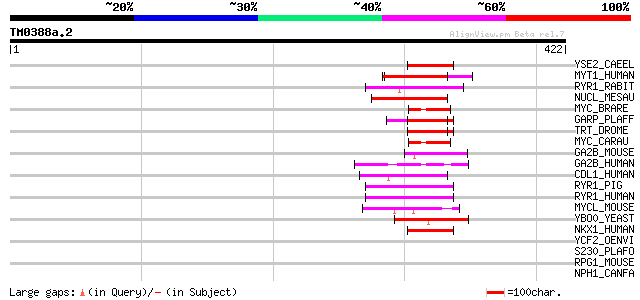
Score E
Sequences producing significant alignments: (bits) Value
YSE2_CAEEL (Q09936) Hypothetical protein C53C9.2 in chromosome X 48 5e-05
MYT1_HUMAN (Q01538) Myelin transcription factor 1 (MYT1) (MYTI) ... 48 5e-05
RYR1_RABIT (P11716) Ryanodine receptor 1 (Skeletal muscle-type r... 46 2e-04
NUCL_MESAU (P08199) Nucleolin (Protein C23) 46 2e-04
MYC_BRARE (P52160) Myc protein (c-myc) (zc-Myc) 46 2e-04
GARP_PLAFF (P13816) Glutamic acid-rich protein precursor 46 2e-04
TRT_DROME (P19351) Troponin T, skeletal muscle (Upheld protein) ... 45 3e-04
MYC_CARAU (P49709) Myc protein (c-myc) 45 5e-04
GA2B_MOUSE (O08795) Glucosidase II beta subunit precursor (Prote... 45 5e-04
GA2B_HUMAN (P14314) Glucosidase II beta subunit precursor (Prote... 45 5e-04
CDL1_HUMAN (P21127) PITSLRE serine/threonine-protein kinase CDC2... 45 5e-04
RYR1_PIG (P16960) Ryanodine receptor 1 (Skeletal muscle-type rya... 44 6e-04
RYR1_HUMAN (P21817) Ryanodine receptor 1 (Skeletal muscle-type r... 44 6e-04
MYCL_MOUSE (P10166) L-myc proto-oncogene protein 44 6e-04
YBO0_YEAST (P38222) Hypothetical 62.6 kDa protein in CDS1-RPL4A ... 44 8e-04
NKX1_HUMAN (O60721) Sodium/potassium/calcium exchanger 1 precurs... 44 8e-04
YCF2_OENVI (P31569) Protein ycf2 (Fragment) 44 0.001
S230_PLAFO (Q08372) Transmission-blocking target antigen S230 pr... 44 0.001
RPG1_MOUSE (Q9EPQ2) X-linked retinitis pigmentosa GTPase regulat... 44 0.001
NPH1_CANFA (Q9TU19) Nephrocystin 1 (Fragment) 44 0.001
>YSE2_CAEEL (Q09936) Hypothetical protein C53C9.2 in chromosome X
Length = 397
Score = 47.8 bits (112), Expect = 5e-05
Identities = 22/35 (62%), Positives = 28/35 (79%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE EEPA +++EEEEEEEEEEE E+ E
Sbjct: 352 EEEEEEEKIEEPAAKEEEEEEEEEEEEEEEELEEE 386
Score = 42.4 bits (98), Expect = 0.002
Identities = 20/31 (64%), Positives = 24/31 (76%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+E+EEEE+ EE E+ EEEEEEEEEEE E
Sbjct: 367 EEEEEEEEEEEEEEEELEEEEEEEEEEEEDE 397
Score = 41.6 bits (96), Expect = 0.004
Identities = 21/36 (58%), Positives = 26/36 (71%)
Query: 296 EWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEEN 331
E A +E+EEEE+ EE E +EEEEEEEEEEE+
Sbjct: 361 EEPAAKEEEEEEEEEEEEEEEELEEEEEEEEEEEED 396
Score = 40.8 bits (94), Expect = 0.007
Identities = 20/44 (45%), Positives = 30/44 (67%)
Query: 287 KERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
+E + +E A+ +E+EEEE+ EE ++EEEEEEEEEE+
Sbjct: 353 EEEEEEKIEEPAAKEEEEEEEEEEEEEEEELEEEEEEEEEEEED 396
Score = 40.4 bits (93), Expect = 0.009
Identities = 22/51 (43%), Positives = 31/51 (60%)
Query: 280 PVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
PV P +E + ++ +E+EEEE+ EE E++ EEEEEEEEEE
Sbjct: 343 PVEPEPEEEEEEEEEEKIEEPAAKEEEEEEEEEEEEEEEELEEEEEEEEEE 393
Score = 38.5 bits (88), Expect = 0.033
Identities = 18/30 (60%), Positives = 23/30 (76%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
++EEEE+ EE E++E EEEEEEEEE E
Sbjct: 366 KEEEEEEEEEEEEEEEELEEEEEEEEEEEE 395
Score = 38.1 bits (87), Expect = 0.043
Identities = 19/35 (54%), Positives = 23/35 (65%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEV 338
E+EEEE+ EE E +EEEEEEEEE E E+
Sbjct: 349 EEEEEEEEEEKIEEPAAKEEEEEEEEEEEEEEEEL 383
Score = 37.0 bits (84), Expect = 0.095
Identities = 18/36 (50%), Positives = 23/36 (63%)
Query: 302 PQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
P+ +EEEE+ EE E+ +EEEEEEEE E E
Sbjct: 346 PEPEEEEEEEEEEKIEEPAAKEEEEEEEEEEEEEEE 381
Score = 31.6 bits (70), Expect = 4.0
Identities = 14/33 (42%), Positives = 20/33 (60%)
Query: 298 TARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
++R + D + EP + EEEEEEEEEE+
Sbjct: 327 SSRQSEIDSQSVKASEPVEPEPEEEEEEEEEEK 359
Score = 31.6 bits (70), Expect = 4.0
Identities = 15/32 (46%), Positives = 21/32 (64%)
Query: 295 KEWTARMPQEDEEEEDPEEPAGEDKEEEEEEE 326
KE +E+EEEE+ E E++EEEEE+E
Sbjct: 366 KEEEEEEEEEEEEEEEELEEEEEEEEEEEEDE 397
Score = 31.2 bits (69), Expect = 5.2
Identities = 14/44 (31%), Positives = 25/44 (56%)
Query: 287 KERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
K++ + + + + + +P EP E++EEEEEEE+ EE
Sbjct: 319 KKKFEERESSRQSEIDSQSVKASEPVEPEPEEEEEEEEEEKIEE 362
Score = 31.2 bits (69), Expect = 5.2
Identities = 18/39 (46%), Positives = 20/39 (51%)
Query: 299 ARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
A P E E EE+ EE E EE +EEEEE E E
Sbjct: 340 ASEPVEPEPEEEEEEEEEEKIEEPAAKEEEEEEEEEEEE 378
Score = 30.8 bits (68), Expect = 6.8
Identities = 19/62 (30%), Positives = 25/62 (39%)
Query: 276 EKRLPVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVV 335
E+ P S + K++ R E + A E E E EEEEEEE E +
Sbjct: 301 EETKPPGSASSVDPFGHYKKKFEERESSRQSEIDSQSVKASEPVEPEPEEEEEEEEEEKI 360
Query: 336 NE 337
E
Sbjct: 361 EE 362
>MYT1_HUMAN (Q01538) Myelin transcription factor 1 (MYT1) (MYTI)
(Proteolipid protein binding protein) (PLPB1)
Length = 1121
Score = 47.8 bits (112), Expect = 5e-05
Identities = 27/67 (40%), Positives = 37/67 (54%)
Query: 286 SKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRAD 345
SK++ ++E +E+EEEE+ EE E++EEEEEEEEEEE E E A
Sbjct: 249 SKQKGILSHEEEDEEEEEEEEEEEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEAA 308
Query: 346 PSPAWME 352
P + E
Sbjct: 309 PDVIFQE 315
Score = 44.3 bits (103), Expect = 6e-04
Identities = 23/50 (46%), Positives = 32/50 (64%)
Query: 284 LLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+LS E + +E +++EEEE+ EE E++EEEEEEEEEEE E
Sbjct: 254 ILSHEEEDEEEEEEEEEEEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 303
Score = 42.0 bits (97), Expect = 0.003
Identities = 25/64 (39%), Positives = 34/64 (53%), Gaps = 3/64 (4%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSPAWMEAAFGRMFLRQ 362
+EDEEEE+ EE E++E+EEEEEEEEE E E + E A + ++
Sbjct: 259 EEDEEEEEEEE---EEEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEAAPDVIFQE 315
Query: 363 DRLH 366
D H
Sbjct: 316 DTSH 319
Score = 40.0 bits (92), Expect = 0.011
Identities = 20/46 (43%), Positives = 28/46 (60%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSP 348
+E+EEEE+ EE E++EEEEEEEEE + E + A +P
Sbjct: 281 EEEEEEEEEEEEEEEEEEEEEEEEEEAAPDVIFQEDTSHTSAQKAP 326
>RYR1_RABIT (P11716) Ryanodine receptor 1 (Skeletal muscle-type
ryanodine receptor) (RyR1) (RYR-1) (Skeletal muscle
calcium release channel)
Length = 5037
Score = 46.2 bits (108), Expect = 2e-04
Identities = 28/79 (35%), Positives = 40/79 (50%), Gaps = 4/79 (5%)
Query: 271 PVHVIEKRLPVAPLLSKERIAQLYK----EWTARMPQEDEEEEDPEEPAGEDKEEEEEEE 326
PV + L V + E + Q+ K E +E+EEEE+ EE ED+EE+EE+E
Sbjct: 1840 PVLKLVSTLLVMGIFGDEDVKQILKMIEPEVFTEEEEEEEEEEEEEEEEEEDEEEKEEDE 1899
Query: 327 EEEENVEVVNEVVTPPRAD 345
EEEE + E P +
Sbjct: 1900 EEEEKEDAEKEEEEAPEGE 1918
Score = 34.7 bits (78), Expect = 0.47
Identities = 18/32 (56%), Positives = 24/32 (74%), Gaps = 2/32 (6%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEE--EEEEENVE 333
E+E+EED EE ED E+EEEE E E+E++E
Sbjct: 1892 EEEKEEDEEEEEKEDAEKEEEEAPEGEKEDLE 1923
Score = 30.4 bits (67), Expect = 8.9
Identities = 19/47 (40%), Positives = 24/47 (50%), Gaps = 2/47 (4%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEV--VTPPRADPSP 348
+ EEEE EP E + E E+ +EE E V E P +A PSP
Sbjct: 4480 DGEEEELVPEPEPEPEPEPEKADEENGEKEEVPEAPPEPPKKAPPSP 4526
>NUCL_MESAU (P08199) Nucleolin (Protein C23)
Length = 713
Score = 46.2 bits (108), Expect = 2e-04
Identities = 21/58 (36%), Positives = 36/58 (61%)
Query: 276 EKRLPVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
E+ + + P K+ A++ + +ED+++E+ +E ED+EEEE+EEEEEE E
Sbjct: 212 EEAMEITPAKGKKAPAKVVPVKAKNVAEEDDDDEEEDEDEEEDEEEEEDEEEEEEEEE 269
Score = 40.0 bits (92), Expect = 0.011
Identities = 22/46 (47%), Positives = 30/46 (64%), Gaps = 6/46 (13%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSPA 349
EDE+EE+ EE ED EEEE++ EEEE +E +TP + +PA
Sbjct: 188 EDEDEEEDEEEEEED-EEEEDDSEEEEAME-----ITPAKGKKAPA 227
Score = 33.9 bits (76), Expect = 0.81
Identities = 13/25 (52%), Positives = 21/25 (84%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEE 328
EDE+++D E+ + ED+E+EEE+E E
Sbjct: 145 EDEDDDDDEDDSDEDEEDEEEDEFE 169
Score = 33.1 bits (74), Expect = 1.4
Identities = 13/32 (40%), Positives = 22/32 (68%)
Query: 299 ARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
A+ DE+E+D ++ D++EE+EEE+E E
Sbjct: 138 AKKEDSDEDEDDDDDEDDSDEDEEDEEEDEFE 169
Score = 31.6 bits (70), Expect = 4.0
Identities = 17/31 (54%), Positives = 21/31 (66%), Gaps = 9/31 (29%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+++EEEED EEEEEEEEEEE V+
Sbjct: 253 EDEEEEED---------EEEEEEEEEEEPVK 274
Score = 30.8 bits (68), Expect = 6.8
Identities = 12/33 (36%), Positives = 22/33 (66%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVV 335
++ +E+ED ++ + E+EE+EEE+E VV
Sbjct: 141 EDSDEDEDDDDDEDDSDEDEEDEEEDEFEPPVV 173
>MYC_BRARE (P52160) Myc protein (c-myc) (zc-Myc)
Length = 408
Score = 46.2 bits (108), Expect = 2e-04
Identities = 23/32 (71%), Positives = 27/32 (83%), Gaps = 3/32 (9%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVV 335
EDEEEED EE E++EEEEEEEEEEE ++VV
Sbjct: 208 EDEEEEDEEE---EEEEEEEEEEEEEEEIDVV 236
Score = 34.7 bits (78), Expect = 0.47
Identities = 16/24 (66%), Positives = 18/24 (74%)
Query: 310 DPEEPAGEDKEEEEEEEEEEENVE 333
D E+ ED+EEEEEEEEEEE E
Sbjct: 206 DSEDEEEEDEEEEEEEEEEEEEEE 229
>GARP_PLAFF (P13816) Glutamic acid-rich protein precursor
Length = 678
Score = 46.2 bits (108), Expect = 2e-04
Identities = 22/35 (62%), Positives = 27/35 (76%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+EDEEEE+ EE E++EEEEEEEEEEE E +E
Sbjct: 573 EEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEDE 607
Score = 44.7 bits (104), Expect = 5e-04
Identities = 24/51 (47%), Positives = 30/51 (58%)
Query: 287 KERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
KE ++ +E E+E EED EE E++EEEEEEEEEEE E E
Sbjct: 552 KEESKEVQEESKEVQEDEEEVEEDEEEEEEEEEEEEEEEEEEEEEEEEEEE 602
Score = 44.3 bits (103), Expect = 6e-04
Identities = 19/31 (61%), Positives = 25/31 (80%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+ED++EED +E ED+E+EEEEEEEEE E
Sbjct: 635 EEDDDEEDDDEDEDEDEEDEEEEEEEEEESE 665
Score = 43.9 bits (102), Expect = 8e-04
Identities = 20/35 (57%), Positives = 28/35 (79%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE+ + +E
Sbjct: 578 EEEEEEEEEEEEEEEEEEEEEEEEEEEEDEDEEDE 612
Score = 43.5 bits (101), Expect = 0.001
Identities = 21/34 (61%), Positives = 26/34 (75%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
E+EEEE+ EE E++EEEEEEEEEEE E +E
Sbjct: 576 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEDEDE 609
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/35 (51%), Positives = 27/35 (76%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
++D+EE+D EE ED++E+EE+EEEEE E +E
Sbjct: 631 EDDDEEDDDEEDDDEDEDEDEEDEEEEEEEEEESE 665
Score = 41.6 bits (96), Expect = 0.004
Identities = 18/35 (51%), Positives = 27/35 (76%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEE+E+EE+ + E
Sbjct: 583 EEEEEEEEEEEEEEEEEEEEEEEDEDEEDEDDAEE 617
Score = 40.4 bits (93), Expect = 0.009
Identities = 18/45 (40%), Positives = 31/45 (68%)
Query: 287 KERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEEN 331
+E + + +E +E+EEEE+ EE E++EEEE+E+EE+E+
Sbjct: 569 EEEVEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEDEDEEDED 613
Score = 38.9 bits (89), Expect = 0.025
Identities = 15/31 (48%), Positives = 25/31 (80%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+E+EEEE+ EE ED+E+E++ EE+E++ E
Sbjct: 593 EEEEEEEEEEEEEDEDEEDEDDAEEDEDDAE 623
Score = 38.9 bits (89), Expect = 0.025
Identities = 16/31 (51%), Positives = 23/31 (73%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEV 334
+DEE++D +E E+ EEEEEEEEEE ++
Sbjct: 638 DDEEDDDEDEDEDEEDEEEEEEEEEESEKKI 668
Score = 38.1 bits (87), Expect = 0.043
Identities = 15/31 (48%), Positives = 24/31 (77%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+ED++EED +E ++ E+E+EE+EEEE E
Sbjct: 630 EEDDDEEDDDEEDDDEDEDEDEEDEEEEEEE 660
Score = 36.2 bits (82), Expect = 0.16
Identities = 15/35 (42%), Positives = 25/35 (70%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E+ E+EE+E++ EE+ + E
Sbjct: 590 EEEEEEEEEEEEEEEEDEDEEDEDDAEEDEDDAEE 624
Score = 36.2 bits (82), Expect = 0.16
Identities = 14/28 (50%), Positives = 23/28 (82%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
+E+EEEE+ EE ED++EE+E++ EE+
Sbjct: 591 EEEEEEEEEEEEEEEDEDEEDEDDAEED 618
Score = 35.8 bits (81), Expect = 0.21
Identities = 14/29 (48%), Positives = 23/29 (79%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEEN 331
+E+EEEED +E +D EE+E++ EE+E+
Sbjct: 599 EEEEEEEDEDEEDEDDAEEDEDDAEEDED 627
Score = 35.4 bits (80), Expect = 0.28
Identities = 15/39 (38%), Positives = 25/39 (63%)
Query: 299 ARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
A ++D EE+D EE E+ ++E+E+E+EE+ E E
Sbjct: 622 AEEDEDDAEEDDDEEDDDEEDDDEDEDEDEEDEEEEEEE 660
Score = 34.3 bits (77), Expect = 0.62
Identities = 14/30 (46%), Positives = 22/30 (72%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
EDEE+ED E +D EE+E++ EE+++ E
Sbjct: 607 EDEEDEDDAEEDEDDAEEDEDDAEEDDDEE 636
Score = 33.5 bits (75), Expect = 1.1
Identities = 24/74 (32%), Positives = 40/74 (53%), Gaps = 10/74 (13%)
Query: 272 VHVIEKRLPVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKE--------EEE 323
V+V+ +R ++K A+L K+ + +E++++E+ +E E KE EE+
Sbjct: 518 VNVVPRRDNHKKKMAKIEEAELQKQ--KHVDKEEDKKEESKEVQEESKEVQEDEEEVEED 575
Query: 324 EEEEEEENVEVVNE 337
EEEEEEE E E
Sbjct: 576 EEEEEEEEEEEEEE 589
Score = 33.5 bits (75), Expect = 1.1
Identities = 13/28 (46%), Positives = 21/28 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
+E+EEEE+ E+ ED EE+E++ EE+
Sbjct: 598 EEEEEEEEDEDEEDEDDAEEDEDDAEED 625
Score = 33.5 bits (75), Expect = 1.1
Identities = 12/31 (38%), Positives = 24/31 (76%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+E+EEE++ EE + +E+E++ EE+E++ E
Sbjct: 600 EEEEEEDEDEEDEDDAEEDEDDAEEDEDDAE 630
Score = 32.3 bits (72), Expect = 2.4
Identities = 11/28 (39%), Positives = 22/28 (78%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEEN 331
ED+ EED ++ +D EE+++EE+++E+
Sbjct: 619 EDDAEEDEDDAEEDDDEEDDDEEDDDED 646
Score = 31.6 bits (70), Expect = 4.0
Identities = 12/35 (34%), Positives = 24/35 (68%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ +E ++ + EE+E++ EE+ + E
Sbjct: 597 EEEEEEEEEDEDEEDEDDAEEDEDDAEEDEDDAEE 631
Score = 31.2 bits (69), Expect = 5.2
Identities = 15/29 (51%), Positives = 22/29 (75%), Gaps = 3/29 (10%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENV 332
EDE+EED EE E++EEEE E++ + N+
Sbjct: 647 EDEDEEDEEE---EEEEEEESEKKIKRNL 672
Score = 30.8 bits (68), Expect = 6.8
Identities = 14/32 (43%), Positives = 23/32 (71%), Gaps = 1/32 (3%)
Query: 303 QEDEEEEDPEEPA-GEDKEEEEEEEEEEENVE 333
+EDE+EED ++ ED EE+E++ EE++ E
Sbjct: 604 EEDEDEEDEDDAEEDEDDAEEDEDDAEEDDDE 635
Score = 30.8 bits (68), Expect = 6.8
Identities = 13/34 (38%), Positives = 26/34 (76%), Gaps = 3/34 (8%)
Query: 303 QEDEEEEDPEE---PAGEDKEEEEEEEEEEENVE 333
+++E+E+D EE A ED+++ EE+++EE++ E
Sbjct: 607 EDEEDEDDAEEDEDDAEEDEDDAEEDDDEEDDDE 640
Score = 30.4 bits (67), Expect = 8.9
Identities = 9/29 (31%), Positives = 23/29 (79%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEEN 331
+EDE++ + +E E+ ++EE+++EE+++
Sbjct: 616 EEDEDDAEEDEDDAEEDDDEEDDDEEDDD 644
>TRT_DROME (P19351) Troponin T, skeletal muscle (Upheld protein)
(Intended thorax protein)
Length = 396
Score = 45.4 bits (106), Expect = 3e-04
Identities = 21/31 (67%), Positives = 25/31 (79%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+EDEEEE+ EE E++EEEEEEEEEEE E
Sbjct: 364 EEDEEEEEEEEEEEEEEEEEEEEEEEEEEEE 394
Score = 44.7 bits (104), Expect = 5e-04
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+EDEE+E+ EE E++EEEEEEEEEEE E E
Sbjct: 358 EEDEEDEEDEEEEEEEEEEEEEEEEEEEEEEEEEE 392
Score = 44.7 bits (104), Expect = 5e-04
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+EDEE+E+ EE E++EEEEEEEEEEE E E
Sbjct: 361 EEDEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 395
Score = 43.5 bits (101), Expect = 0.001
Identities = 20/35 (57%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+++E+EED EE E++EEEEEEEEEEE E E
Sbjct: 359 EDEEDEEDEEEEEEEEEEEEEEEEEEEEEEEEEEE 393
Score = 43.1 bits (100), Expect = 0.001
Identities = 20/30 (66%), Positives = 24/30 (79%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
E+EEEE+ EE E++EEEEEEEEEEE E
Sbjct: 367 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 396
Score = 43.1 bits (100), Expect = 0.001
Identities = 21/44 (47%), Positives = 29/44 (65%)
Query: 287 KERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
+E + + E +E+EEEE+ EE E++EEEEEEEEEEE
Sbjct: 352 EEEVVEEEDEEDEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 395
Score = 42.7 bits (99), Expect = 0.002
Identities = 19/28 (67%), Positives = 24/28 (84%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
+E+EEEE+ EE E++EEEEEEEEEEE
Sbjct: 369 EEEEEEEEEEEEEEEEEEEEEEEEEEEE 396
Score = 42.7 bits (99), Expect = 0.002
Identities = 20/34 (58%), Positives = 25/34 (72%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
E+E+EED E+ E++EEEEEEEEEEE E E
Sbjct: 357 EEEDEEDEEDEEEEEEEEEEEEEEEEEEEEEEEE 390
Score = 41.6 bits (96), Expect = 0.004
Identities = 21/50 (42%), Positives = 32/50 (64%)
Query: 288 ERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
E I + +E + +E++EE++ +E E++EEEEEEEEEEE E E
Sbjct: 342 EDIVEDDEEVEEEVVEEEDEEDEEDEEEEEEEEEEEEEEEEEEEEEEEEE 391
>MYC_CARAU (P49709) Myc protein (c-myc)
Length = 399
Score = 44.7 bits (104), Expect = 5e-04
Identities = 22/32 (68%), Positives = 27/32 (83%), Gaps = 3/32 (9%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVV 335
EDEEEE+ EE E++EEEEEEEEEEE ++VV
Sbjct: 207 EDEEEEEEEE---EEEEEEEEEEEEEEEIDVV 235
Score = 36.6 bits (83), Expect = 0.12
Identities = 21/50 (42%), Positives = 29/50 (58%), Gaps = 4/50 (8%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEE----ENVEVVNEVVTPPRADPSP 348
+++EEEE+ EE E++EEEEEEEE + E + NE PSP
Sbjct: 207 EDEEEEEEEEEEEEEEEEEEEEEEEIDVVTVEKRQKRNEADVSDSRYPSP 256
Score = 33.1 bits (74), Expect = 1.4
Identities = 15/24 (62%), Positives = 18/24 (74%)
Query: 310 DPEEPAGEDKEEEEEEEEEEENVE 333
D E+ E++EEEEEEEEEEE E
Sbjct: 205 DSEDEEEEEEEEEEEEEEEEEEEE 228
Score = 31.2 bits (69), Expect = 5.2
Identities = 14/23 (60%), Positives = 16/23 (68%)
Query: 315 AGEDKEEEEEEEEEEENVEVVNE 337
+G D E+EEEEEEEEE E E
Sbjct: 202 SGSDSEDEEEEEEEEEEEEEEEE 224
>GA2B_MOUSE (O08795) Glucosidase II beta subunit precursor (Protein
kinase C substrate, 60.1 kDa protein, heavy chain)
(PKCSH) (80K-H protein)
Length = 521
Score = 44.7 bits (104), Expect = 5e-04
Identities = 24/50 (48%), Positives = 29/50 (58%), Gaps = 2/50 (4%)
Query: 301 MPQEDE--EEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSP 348
+P+E E EE+ P P E++EEEEEE EEEE E E PP P P
Sbjct: 291 VPEETEPKEEKPPVLPPTEEEEEEEEEPEEEEEEEEEEEEAPPPLQPPQP 340
>GA2B_HUMAN (P14314) Glucosidase II beta subunit precursor (Protein
kinase C substrate, 60.1 kDa protein, heavy chain)
(PKCSH) (80K-H protein)
Length = 527
Score = 44.7 bits (104), Expect = 5e-04
Identities = 32/87 (36%), Positives = 44/87 (49%), Gaps = 14/87 (16%)
Query: 263 AVRDRARRPVHVIEKRLPVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEE 322
A+RD+ R + P AP L++ KE +P EEE+ EE E++EE
Sbjct: 275 AIRDKYRSEALPTDLPAPSAPDLTEP------KEEQPPVPSSPTEEEEEEE---EEEEEA 325
Query: 323 EEEEEEEENVEVVNEVVTPPRADPSPA 349
EEEEEEE+ +E PP + P PA
Sbjct: 326 EEEEEEED-----SEEAPPPLSPPQPA 347
>CDL1_HUMAN (P21127) PITSLRE serine/threonine-protein kinase CDC2L1
(EC 2.7.1.37) (Galactosyltransferase associated protein
kinase p58/GTA) (Cell division cycle 2-like 1) (CLK-1)
(CDK11) (p58 CLK-1)
Length = 795
Score = 44.7 bits (104), Expect = 5e-04
Identities = 28/77 (36%), Positives = 39/77 (50%), Gaps = 10/77 (12%)
Query: 267 RARRPVHVIEKRLPVAPLLS----------KERIAQLYKEWTARMPQEDEEEEDPEEPAG 316
RA++P + E+++ LLS K A+ + +E+EEEE+ EE G
Sbjct: 258 RAQKPAQLKEEKMEERDLLSDLQDISDSERKTSSAESSSAESGSGSEEEEEEEEEEEEEG 317
Query: 317 EDKEEEEEEEEEEENVE 333
EE EEEEEEEE E
Sbjct: 318 STSEESEEEEEEEEEEE 334
Score = 34.7 bits (78), Expect = 0.47
Identities = 15/35 (42%), Positives = 23/35 (64%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE G + EE E+ EE + E ++E
Sbjct: 325 EEEEEEEEEEEETGSNSEEASEQSAEEVSEEEMSE 359
>RYR1_PIG (P16960) Ryanodine receptor 1 (Skeletal muscle-type
ryanodine receptor) (RyR1) (RYR-1) (Skeletal muscle
calcium release channel)
Length = 5035
Score = 44.3 bits (103), Expect = 6e-04
Identities = 26/67 (38%), Positives = 37/67 (54%)
Query: 271 PVHVIEKRLPVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
PV + L V + E + Q+ K + E+EEEE+ EE E+ EEE+EE+EEEE
Sbjct: 1841 PVLKLVSTLLVMGIFGDEDVKQILKMIEPEVFTEEEEEEEEEEEEEEEDEEEKEEDEEEE 1900
Query: 331 NVEVVNE 337
E +E
Sbjct: 1901 AREKEDE 1907
Score = 39.3 bits (90), Expect = 0.019
Identities = 18/42 (42%), Positives = 27/42 (63%)
Query: 292 QLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+++ E +E+EEEE+ EE ED+EEE E+E+EE E
Sbjct: 1870 EVFTEEEEEEEEEEEEEEEDEEEKEEDEEEEAREKEDEEKEE 1911
Score = 35.4 bits (80), Expect = 0.28
Identities = 24/66 (36%), Positives = 32/66 (48%), Gaps = 2/66 (3%)
Query: 304 EDEEEEDPEEPAGE--DKEEEEEEEEEEENVEVVNEVVTPPRADPSPAWMEAAFGRMFLR 361
E+E+EED EE A E D+E+EEEE E E E + E + + S F
Sbjct: 1890 EEEKEEDEEEEAREKEDEEKEEEETAEGEKEEYLEEGLLQMKLPESVKLQMCNLLEYFCD 1949
Query: 362 QDRLHR 367
Q+ HR
Sbjct: 1950 QELQHR 1955
Score = 32.3 bits (72), Expect = 2.4
Identities = 14/28 (50%), Positives = 20/28 (71%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
+E++EEE+ E E+KEEEE E E+E
Sbjct: 1893 KEEDEEEEAREKEDEEKEEEETAEGEKE 1920
>RYR1_HUMAN (P21817) Ryanodine receptor 1 (Skeletal muscle-type
ryanodine receptor) (RyR1) (RYR-1) (Skeletal muscle
calcium release channel)
Length = 5038
Score = 44.3 bits (103), Expect = 6e-04
Identities = 26/67 (38%), Positives = 36/67 (52%)
Query: 271 PVHVIEKRLPVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
PV + L V + E + Q+ K + E+EEEED EE E+ EEE+EE+EEE
Sbjct: 1840 PVLKLVSTLLVMGIFGDEDVKQILKMIEPEVFTEEEEEEDEEEEGEEEDEEEKEEDEEET 1899
Query: 331 NVEVVNE 337
E +E
Sbjct: 1900 AQEKEDE 1906
Score = 35.4 bits (80), Expect = 0.28
Identities = 16/27 (59%), Positives = 21/27 (77%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEE 330
E+E+EED EE A E ++EE+EEEE E
Sbjct: 1889 EEEKEEDEEETAQEKEDEEKEEEEAAE 1915
Score = 35.0 bits (79), Expect = 0.36
Identities = 13/26 (50%), Positives = 22/26 (84%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEE 328
+++EE+E+ EE ++KE+EE+EEEE
Sbjct: 1887 EDEEEKEEDEEETAQEKEDEEKEEEE 1912
Score = 32.3 bits (72), Expect = 2.4
Identities = 16/32 (50%), Positives = 23/32 (71%), Gaps = 1/32 (3%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEE-EEEENVE 333
+E++EEE +E E+KEEEE E E+EE +E
Sbjct: 1892 KEEDEEETAQEKEDEEKEEEEAAEGEKEEGLE 1923
>MYCL_MOUSE (P10166) L-myc proto-oncogene protein
Length = 368
Score = 44.3 bits (103), Expect = 6e-04
Identities = 31/86 (36%), Positives = 43/86 (49%), Gaps = 19/86 (22%)
Query: 269 RRPVHVIEKRLPVAPLLSKERIA--QLYKEWTARMPQED----------EEEEDPEEPAG 316
R+PV + + P+ P + I+ Q + AR P E +EE PE A
Sbjct: 184 RKPVTITVRADPLDPCMKHFHISIHQQQHNYAARFPPESCSQEGDPEPGPQEEAPEIEAP 243
Query: 317 EDKEEEEEEEEEEENVEVVNEVVTPP 342
++KEEEEEEEEEE E+V+PP
Sbjct: 244 KEKEEEEEEEEEE-------EIVSPP 262
>YBO0_YEAST (P38222) Hypothetical 62.6 kDa protein in CDS1-RPL4A
intergenic region
Length = 552
Score = 43.9 bits (102), Expect = 8e-04
Identities = 23/63 (36%), Positives = 39/63 (61%), Gaps = 6/63 (9%)
Query: 293 LYKEWTARMPQEDEEEEDPEEPAGE-----DKEEEEEEEEEEENVEV-VNEVVTPPRADP 346
L+ W A+M QEDEE+ED + + + EEEEEEEE ++++E ++++ +P
Sbjct: 388 LFSAWYAQMRQEDEEDEDGQAKSDNLSDDIESEEEEEEEEGDDSLESWLSQLYIDSSGEP 447
Query: 347 SPA 349
SP+
Sbjct: 448 SPS 450
>NKX1_HUMAN (O60721) Sodium/potassium/calcium exchanger 1 precursor
(Na(+)/K(+)/Ca(2+)-exchange protein 1) (Retinal rod
Na-Ca+K exchanger)
Length = 1099
Score = 43.9 bits (102), Expect = 8e-04
Identities = 20/35 (57%), Positives = 27/35 (77%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ +E E++E+EEEEEEEEE E NE
Sbjct: 860 EEEEEEEEEQEEEEEEEEQEEEEEEEEEEEEKGNE 894
Score = 40.8 bits (94), Expect = 0.007
Identities = 18/31 (58%), Positives = 25/31 (80%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+E+E+EE+ EE E++EEEEEEEEE+ N E
Sbjct: 865 EEEEQEEEEEEEEQEEEEEEEEEEEEKGNEE 895
Score = 40.0 bits (92), Expect = 0.011
Identities = 19/33 (57%), Positives = 23/33 (69%)
Query: 305 DEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
D EEE+ EE E++EEEEE+EEEEE E E
Sbjct: 858 DSEEEEEEEEEQEEEEEEEEQEEEEEEEEEEEE 890
Score = 40.0 bits (92), Expect = 0.011
Identities = 18/30 (60%), Positives = 23/30 (76%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+ EEEE+ EE E++EEEE+EEEEEE E
Sbjct: 858 DSEEEEEEEEEQEEEEEEEEQEEEEEEEEE 887
Score = 31.2 bits (69), Expect = 5.2
Identities = 16/34 (47%), Positives = 20/34 (58%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+++ ED G D EEEEEEEEE+E E E
Sbjct: 844 DEKGVEDGGGSDGGDSEEEEEEEEEQEEEEEEEE 877
>YCF2_OENVI (P31569) Protein ycf2 (Fragment)
Length = 630
Score = 43.5 bits (101), Expect = 0.001
Identities = 25/58 (43%), Positives = 35/58 (60%), Gaps = 4/58 (6%)
Query: 284 LLSKERIAQLYKEWTARMPQE----DEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
LL ++ +L +E + +E DEEEE+P+E E EEEEEEEEEEE + + E
Sbjct: 440 LLEAQQEDELLEEEDEELDEEEDELDEEEEEPKEEEDELHEEEEEEEEEEEEEDELQE 497
>S230_PLAFO (Q08372) Transmission-blocking target antigen S230
precursor
Length = 3135
Score = 43.5 bits (101), Expect = 0.001
Identities = 21/34 (61%), Positives = 26/34 (75%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+DEEEE+ EE E++EEEEEEEEEEE + V E
Sbjct: 278 DDEEEEEEEEEEEEEEEEEEEEEEEEEYDDYVYE 311
Score = 39.7 bits (91), Expect = 0.015
Identities = 19/34 (55%), Positives = 25/34 (72%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
++EEEE+ EE E++EEEEEEEEEE + V E
Sbjct: 279 DEEEEEEEEEEEEEEEEEEEEEEEEEYDDYVYEE 312
Score = 39.3 bits (90), Expect = 0.019
Identities = 21/41 (51%), Positives = 27/41 (65%), Gaps = 3/41 (7%)
Query: 305 DEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRAD 345
D+EEE+ EE E++EEEEEEEEEEE E ++ V D
Sbjct: 278 DDEEEEEEE---EEEEEEEEEEEEEEEEEEYDDYVYEESGD 315
Score = 38.5 bits (88), Expect = 0.033
Identities = 17/30 (56%), Positives = 24/30 (79%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENV 332
+E+EEEE+ EE E++EEEEEEEE ++ V
Sbjct: 280 EEEEEEEEEEEEEEEEEEEEEEEEEYDDYV 309
Score = 35.0 bits (79), Expect = 0.36
Identities = 18/33 (54%), Positives = 20/33 (60%)
Query: 305 DEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
DEE+ P + D EEEEEEEEEEE E E
Sbjct: 265 DEEDMSPRDNFVIDDEEEEEEEEEEEEEEEEEE 297
Score = 33.5 bits (75), Expect = 1.1
Identities = 27/81 (33%), Positives = 37/81 (45%), Gaps = 14/81 (17%)
Query: 303 QEDEEEEDPEEPAGEDKEEE-------------EEEEEEEENVEVVNEVVTPPRADPSPA 349
+E+EEEE+ EE E++EEE EE+ +EE EV E D
Sbjct: 285 EEEEEEEEEEEEEEEEEEEEYDDYVYEESGDETEEQLQEEHQEEVGAESSEESFNDEDED 344
Query: 350 WMEAAFGRMFLRQDRLHRDLD 370
+EA G M +R D + D D
Sbjct: 345 SVEARDGDM-IRVDEYYEDQD 364
>RPG1_MOUSE (Q9EPQ2) X-linked retinitis pigmentosa GTPase
regulator-interacting protein 1 (RPGR-interacting protein
1)
Length = 1331
Score = 43.5 bits (101), Expect = 0.001
Identities = 25/63 (39%), Positives = 36/63 (56%), Gaps = 6/63 (9%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSPAWMEAAFGRMFLRQ 362
+E+E+EE+ EE ED++EEEEEEEEEE E +E A + W+ F Q
Sbjct: 953 EEEEKEEEKEEEEEEDEKEEEEEEEEEEEEEEEDENKDVLEASFTEEWVP------FFSQ 1006
Query: 363 DRL 365
D++
Sbjct: 1007 DQI 1009
Score = 42.7 bits (99), Expect = 0.002
Identities = 21/39 (53%), Positives = 27/39 (68%)
Query: 295 KEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+E R +E++EEE EE ++KEEEEEEEEEEE E
Sbjct: 946 EEEEEREEEEEKEEEKEEEEEEDEKEEEEEEEEEEEEEE 984
Score = 42.7 bits (99), Expect = 0.002
Identities = 19/28 (67%), Positives = 23/28 (81%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
+E+EEEE+ EE E+KEEEEEEE EEE
Sbjct: 927 EEEEEEEEEEEEVKEEKEEEEEEEREEE 954
Score = 40.8 bits (94), Expect = 0.007
Identities = 20/35 (57%), Positives = 24/35 (68%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E +EEEEEE EEEE E E
Sbjct: 928 EEEEEEEEEEEVKEEKEEEEEEEREEEEEKEEEKE 962
Score = 38.9 bits (89), Expect = 0.025
Identities = 19/35 (54%), Positives = 24/35 (68%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE EE E++EE EEEEE+EE E E
Sbjct: 932 EEEEEEEVKEEKEEEEEEEREEEEEKEEEKEEEEE 966
Score = 38.5 bits (88), Expect = 0.033
Identities = 19/39 (48%), Positives = 26/39 (65%)
Query: 299 ARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
A P+ +E+EE+ E +++E EEEEEEEEE EV E
Sbjct: 904 AEKPEGEEKEEEGGEEEVKEEEVEEEEEEEEEEEEVKEE 942
Score = 38.5 bits (88), Expect = 0.033
Identities = 20/35 (57%), Positives = 23/35 (65%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E EEEE+ EE E KEE+EEEEEEE E E
Sbjct: 924 EEVEEEEEEEEEEEEVKEEKEEEEEEEREEEEEKE 958
Score = 38.5 bits (88), Expect = 0.033
Identities = 20/35 (57%), Positives = 23/35 (65%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E EEE EE E+ EEEEEEEEEEE V+ E
Sbjct: 910 EEKEEEGGEEEVKEEEVEEEEEEEEEEEEVKEEKE 944
Score = 38.1 bits (87), Expect = 0.043
Identities = 18/34 (52%), Positives = 24/34 (69%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
E+EEEE+ EE ++++EEEEEEE EE E E
Sbjct: 927 EEEEEEEEEEEEVKEEKEEEEEEEREEEEEKEEE 960
Score = 37.7 bits (86), Expect = 0.056
Identities = 19/53 (35%), Positives = 32/53 (59%)
Query: 285 LSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+ +E + + +E ++E+EE+ EE E++E+EEE+EEEEE E E
Sbjct: 921 VKEEEVEEEEEEEEEEEEVKEEKEEEEEEEREEEEEKEEEKEEEEEEDEKEEE 973
Score = 35.4 bits (80), Expect = 0.28
Identities = 18/35 (51%), Positives = 22/35 (62%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEV 338
E E E+ EE GE++ +EEE EEEEE E EV
Sbjct: 905 EKPEGEEKEEEGGEEEVKEEEVEEEEEEEEEEEEV 939
Score = 34.7 bits (78), Expect = 0.47
Identities = 20/36 (55%), Positives = 24/36 (66%), Gaps = 2/36 (5%)
Query: 304 EDEEEEDPEEPAGEDK--EEEEEEEEEEENVEVVNE 337
E++EEE EE E++ EEEEEEEEEEE E E
Sbjct: 910 EEKEEEGGEEEVKEEEVEEEEEEEEEEEEVKEEKEE 945
Score = 34.7 bits (78), Expect = 0.47
Identities = 18/35 (51%), Positives = 22/35 (62%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E EEE EE E++EEEEEEEE +E E E
Sbjct: 914 EEGGEEEVKEEEVEEEEEEEEEEEEVKEEKEEEEE 948
Score = 33.9 bits (76), Expect = 0.81
Identities = 16/35 (45%), Positives = 24/35 (67%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+ +EE+ EE E++EEEE +EE+EE E E
Sbjct: 918 EEEVKEEEVEEEEEEEEEEEEVKEEKEEEEEEERE 952
Score = 31.2 bits (69), Expect = 5.2
Identities = 20/52 (38%), Positives = 27/52 (51%), Gaps = 8/52 (15%)
Query: 296 EWTAR-MPQEDEEEEDPEEPAGEDKEEE-------EEEEEEEENVEVVNEVV 339
+W + + E + + E+P GE+KEEE EEE EEEE E E V
Sbjct: 888 DWKSHYLAPEGFQMSEAEKPEGEEKEEEGGEEEVKEEEVEEEEEEEEEEEEV 939
>NPH1_CANFA (Q9TU19) Nephrocystin 1 (Fragment)
Length = 565
Score = 43.5 bits (101), Expect = 0.001
Identities = 18/29 (62%), Positives = 25/29 (86%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEEN 331
+E+EEEE+ EE GE++E EEEEEE++EN
Sbjct: 6 EEEEEEEESEEGGGEEEESEEEEEEKQEN 34
Score = 38.5 bits (88), Expect = 0.033
Identities = 19/31 (61%), Positives = 21/31 (67%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+EDEEEE+ EE EEEE EEEEEE E
Sbjct: 3 EEDEEEEEEEESEEGGGEEEESEEEEEEKQE 33
Score = 37.0 bits (84), Expect = 0.095
Identities = 16/30 (53%), Positives = 22/30 (73%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
E+EEEE+ E G ++EE EEEEEE++ E
Sbjct: 6 EEEEEEEESEEGGGEEEESEEEEEEKQENE 35
Score = 35.4 bits (80), Expect = 0.28
Identities = 17/34 (50%), Positives = 22/34 (64%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
E+E+EE+ EE E+ EEEE EEEE + NE
Sbjct: 2 EEEDEEEEEEEESEEGGGEEEESEEEEEEKQENE 35
Score = 35.4 bits (80), Expect = 0.28
Identities = 16/28 (57%), Positives = 21/28 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
+E++EEE+ EE + E EEEE EEEEE
Sbjct: 2 EEEDEEEEEEEESEEGGGEEEESEEEEE 29
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.136 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 55,964,671
Number of Sequences: 164201
Number of extensions: 2672964
Number of successful extensions: 41482
Number of sequences better than 10.0: 769
Number of HSP's better than 10.0 without gapping: 603
Number of HSP's successfully gapped in prelim test: 185
Number of HSP's that attempted gapping in prelim test: 22584
Number of HSP's gapped (non-prelim): 7163
length of query: 422
length of database: 59,974,054
effective HSP length: 113
effective length of query: 309
effective length of database: 41,419,341
effective search space: 12798576369
effective search space used: 12798576369
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0388a.2


