

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0380.7
(155 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
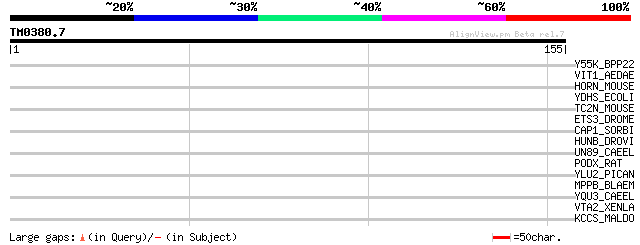
Y55K_BPP22 (P57019) Hypothetical 55.3 kDa protein in gtrB 5'regi... 33 0.25
VIT1_AEDAE (Q16927) Vitellogenin A1 precursor (VG) (PVG1) [Conta... 33 0.33
HORN_MOUSE (Q8VHD8) Hornerin 30 1.6
YDHS_ECOLI (P77148) Hypothetical protein ydhS 30 2.1
TC2N_MOUSE (Q91XT6) Membrane targeting tandem C2 domain containi... 30 2.8
ETS3_DROME (P29774) DNA-binding protein D-ETS-3 30 2.8
CAP1_SORBI (P29195) Phosphoenolpyruvate carboxylase 1 (EC 4.1.1.... 30 2.8
HUNB_DROVI (P13361) Hunchback protein 29 3.6
UN89_CAEEL (O01761) Muscle M-line assembly protein unc-89 (Uncoo... 29 4.8
PODX_RAT (Q9WTQ2) Podocalyxin precursor 29 4.8
YLU2_PICAN (P34735) Hypothetical protein in LEU2 3'region (Fragm... 28 6.2
MPPB_BLAEM (Q00302) Mitochondrial processing peptidase beta subu... 28 6.2
YQU3_CAEEL (Q09550) Hypothetical protein F26C11.3 in chromosome II 28 8.1
VTA2_XENLA (P18709) Vitellogenin A2 precursor (VTG A2) [Contains... 28 8.1
KCCS_MALDO (Q07250) Calcium/calmodulin-dependent serine/threonin... 28 8.1
>Y55K_BPP22 (P57019) Hypothetical 55.3 kDa protein in gtrB 5'region
(ORF485)
Length = 485
Score = 33.1 bits (74), Expect = 0.25
Identities = 13/35 (37%), Positives = 21/35 (59%)
Query: 86 DWTWAWILSRKPIFARDLELNDQETKLLGGSHNRG 120
DW W+ +L ++ +F+R+ L D+E KL G G
Sbjct: 433 DWMWSEVLMQRNVFSRNYRLYDKEVKLENGWKKSG 467
>VIT1_AEDAE (Q16927) Vitellogenin A1 precursor (VG) (PVG1)
[Contains: Vitellin light chain (VL); Vitellin heavy
chain (VH)]
Length = 2148
Score = 32.7 bits (73), Expect = 0.33
Identities = 22/65 (33%), Positives = 38/65 (57%), Gaps = 2/65 (3%)
Query: 8 QHRSFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRS 67
Q+R T+S++ S S S +S EH R SS++++ +++S+S+S+ SS+ S
Sbjct: 486 QYRLHRLNDTSSDSSSSDSSSSSSSESKEH--RNGTSSYSSSSSSSSSSSSSESSSYSSS 543
Query: 68 KSLSS 72
S SS
Sbjct: 544 SSSSS 548
>HORN_MOUSE (Q8VHD8) Hornerin
Length = 2496
Score = 30.4 bits (67), Expect = 1.6
Identities = 25/76 (32%), Positives = 34/76 (43%), Gaps = 8/76 (10%)
Query: 5 GHQQHRSFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSF 64
G QH+S S + Q P+S + S HG SS T + +S S+S WS
Sbjct: 1356 GRNQHQSGSR--HGSGSGQFPISGQQG---SHHG---HSSSSGTHNSGSSQSSSTQWSHG 1407
Query: 65 PRSKSLSSMGDYAGTS 80
S+ S +G Y TS
Sbjct: 1408 SGSEQSSGLGHYGSTS 1423
Score = 30.4 bits (67), Expect = 1.6
Identities = 25/76 (32%), Positives = 34/76 (43%), Gaps = 8/76 (10%)
Query: 5 GHQQHRSFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSF 64
G QH+S S + Q P+S + S HG SS T + +S S+S WS
Sbjct: 1008 GRSQHQSGSR--HGSGSGQFPISGQQG---SHHG---HSSSSGTHNSGSSQSSSTQWSHG 1059
Query: 65 PRSKSLSSMGDYAGTS 80
S+ S +G Y TS
Sbjct: 1060 SGSEQSSGLGHYGSTS 1075
Score = 30.4 bits (67), Expect = 1.6
Identities = 25/76 (32%), Positives = 34/76 (43%), Gaps = 8/76 (10%)
Query: 5 GHQQHRSFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSF 64
G QH+S S + Q P+S + S HG SS T + +S S+S WS
Sbjct: 1704 GRSQHQSGSR--HGSGSGQFPISGQQG---SHHG---HSSSSGTHNSGSSQSSSTQWSHG 1755
Query: 65 PRSKSLSSMGDYAGTS 80
S+ S +G Y TS
Sbjct: 1756 SGSEQSSGLGHYGSTS 1771
Score = 30.4 bits (67), Expect = 1.6
Identities = 25/76 (32%), Positives = 34/76 (43%), Gaps = 8/76 (10%)
Query: 5 GHQQHRSFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSF 64
G QH+S S + Q P+S + S HG SS T + +S S+S WS
Sbjct: 659 GRSQHQSGSR--HGSGSGQFPISGQQG---SHHG---HSSSSGTHNSGSSQSSSTQWSHG 710
Query: 65 PRSKSLSSMGDYAGTS 80
S+ S +G Y TS
Sbjct: 711 SGSEQSSGLGHYGSTS 726
Score = 30.4 bits (67), Expect = 1.6
Identities = 25/76 (32%), Positives = 34/76 (43%), Gaps = 8/76 (10%)
Query: 5 GHQQHRSFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSF 64
G QH+S S + Q P+S + S HG SS T + +S S+S WS
Sbjct: 2394 GRSQHQSGSR--HGSGSGQFPISGQQG---SHHG---HSSSSGTHNSGSSQSSSTQWSHG 2445
Query: 65 PRSKSLSSMGDYAGTS 80
S+ S +G Y TS
Sbjct: 2446 SGSEQSSGLGHYGSTS 2461
Score = 30.4 bits (67), Expect = 1.6
Identities = 25/76 (32%), Positives = 34/76 (43%), Gaps = 8/76 (10%)
Query: 5 GHQQHRSFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSF 64
G QH+S S + Q P+S + S HG SS T + +S S+S WS
Sbjct: 2052 GRSQHQSGSR--HGSGSGQFPISGQQG---SHHG---HSSSSGTHNSGSSQSSSTQWSHG 2103
Query: 65 PRSKSLSSMGDYAGTS 80
S+ S +G Y TS
Sbjct: 2104 SGSEQSSGLGHYGSTS 2119
Score = 28.5 bits (62), Expect = 6.2
Identities = 24/76 (31%), Positives = 33/76 (42%), Gaps = 8/76 (10%)
Query: 5 GHQQHRSFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSF 64
G QH+S S + Q P+S + S HG SS T + +S S+S WS
Sbjct: 2223 GRSQHQSGSR--HGSGSGQFPISGQQG---SHHG---HSSSSGTHNSGSSQSSSTQWSHG 2274
Query: 65 PRSKSLSSMGDYAGTS 80
S+ S +G Y S
Sbjct: 2275 SGSEQSSGLGQYGSPS 2290
Score = 28.1 bits (61), Expect = 8.1
Identities = 17/55 (30%), Positives = 26/55 (46%)
Query: 30 RSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKSLSSMGDYAGTSIKNW 84
+S+ S+ HG SS + T S+ +S + S SS G Y TS +N+
Sbjct: 328 QSITSANHGSHSNQSSCSGTRECGSSESSMKKTHVSGSGHSSSTGKYTSTSGQNY 382
>YDHS_ECOLI (P77148) Hypothetical protein ydhS
Length = 534
Score = 30.0 bits (66), Expect = 2.1
Identities = 17/57 (29%), Positives = 24/57 (41%), Gaps = 1/57 (1%)
Query: 90 AWILSRKPIFARD-LELNDQETKLLGGSHNRGTWRHVFYKLRSEIRRFIPPTNSHHH 145
AW RK D E N QE + + WR+V +L ++ +P N H H
Sbjct: 316 AWFAERKQRDPFDWAEKNLQEVERNKREKHTVPWRYVILRLHEAVQEIVPHLNEHDH 372
>TC2N_MOUSE (Q91XT6) Membrane targeting tandem C2 domain containing
protein 1 (Tandem C2 protein in nucleus) (Tac2-N)
Length = 489
Score = 29.6 bits (65), Expect = 2.8
Identities = 22/80 (27%), Positives = 43/80 (53%), Gaps = 5/80 (6%)
Query: 7 QQHRSFFTIPTTSNNKQLPVSHSRSVDS-SEHGLRKRLSSFNTTPAAASASASAVWSSFP 65
Q+H SF ++P++S++++ +RS+D+ + G + L N ++SA +W +
Sbjct: 186 QRHDSFSSVPSSSSSRKNSQGSNRSLDTITLSGDERDLGRLN-VKLFYNSSAEQIWITVL 244
Query: 66 RSKSL---SSMGDYAGTSIK 82
+ + + SS GD SIK
Sbjct: 245 QCRDISWPSSYGDTPTISIK 264
>ETS3_DROME (P29774) DNA-binding protein D-ETS-3
Length = 490
Score = 29.6 bits (65), Expect = 2.8
Identities = 26/88 (29%), Positives = 38/88 (42%), Gaps = 6/88 (6%)
Query: 2 SSIGHQQHR-----SFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASAS 56
SS H +H+ S TTS++ P S S + D + + SS NT+ AS
Sbjct: 192 SSSSHVEHKVRADKSTLDCATTSSHAAAPSSSSSASDHQQGRISGSKSS-NTSGTGGGAS 250
Query: 57 ASAVWSSFPRSKSLSSMGDYAGTSIKNW 84
AS S S SS G ++ T + +
Sbjct: 251 ASGGGGSALYSSYKSSWGSHSSTQSQGY 278
>CAP1_SORBI (P29195) Phosphoenolpyruvate carboxylase 1 (EC 4.1.1.31)
(PEPCase 1) (CP21)
Length = 960
Score = 29.6 bits (65), Expect = 2.8
Identities = 13/45 (28%), Positives = 23/45 (50%)
Query: 73 MGDYAGTSIKNWWDWTWAWILSRKPIFARDLELNDQETKLLGGSH 117
+G YA S + DW + + ++P+F DL ++ +LG H
Sbjct: 465 IGSYAEWSEEKRQDWLLSELRGKRPLFGSDLPQTEETADVLGTFH 509
>HUNB_DROVI (P13361) Hunchback protein
Length = 816
Score = 29.3 bits (64), Expect = 3.6
Identities = 18/65 (27%), Positives = 31/65 (47%), Gaps = 4/65 (6%)
Query: 6 HQQHRSFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFP 65
++QH ++++ +N KQ P+SH HG + + N+ A+S S + S P
Sbjct: 13 YEQHNAWYSSMFAANIKQEPLSHHH----HHHGQQHHDNHSNSNSGASSPRQSPLPSPIP 68
Query: 66 RSKSL 70
S L
Sbjct: 69 PSTQL 73
>UN89_CAEEL (O01761) Muscle M-line assembly protein unc-89
(Uncoordinated protein 89)
Length = 6632
Score = 28.9 bits (63), Expect = 4.8
Identities = 17/86 (19%), Positives = 39/86 (44%), Gaps = 4/86 (4%)
Query: 3 SIGHQQHRSFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWS 62
+I +++++S IP ++ ++ V S V SSE + ++ +TTP S
Sbjct: 2028 TIKNEENKSSLIIPNAQDSGKITVEASNEVGSSESSAQLTVNPPSTTPIVVDGPKSVTIK 2087
Query: 63 SFPRSKSLSSMGDYAGTSIKNWWDWT 88
++ +++ + ++K WT
Sbjct: 2088 ETETAEFKATISGFPAPTVK----WT 2109
>PODX_RAT (Q9WTQ2) Podocalyxin precursor
Length = 485
Score = 28.9 bits (63), Expect = 4.8
Identities = 18/59 (30%), Positives = 25/59 (41%)
Query: 14 TIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKSLSS 72
T+PTTSN+ Q S +S D L L N + + S S P + S +S
Sbjct: 102 TVPTTSNSGQTVSSGGKSSDKITTALPTSLGPVNASSQPTDLNTSTKLPSTPTTNSTAS 160
>YLU2_PICAN (P34735) Hypothetical protein in LEU2 3'region
(Fragment)
Length = 373
Score = 28.5 bits (62), Expect = 6.2
Identities = 15/46 (32%), Positives = 29/46 (62%)
Query: 27 SHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKSLSS 72
SHS + D++ SS +++ +++S S+S+ SS P+S ++SS
Sbjct: 115 SHSSTDDATSTSSTSTTSSSSSSLSSSSTSSSSKQSSSPQSSTMSS 160
>MPPB_BLAEM (Q00302) Mitochondrial processing peptidase beta
subunit, mitochondrial precursor (EC 3.4.24.64)
(Beta-MPP) (BeMPP1)
Length = 465
Score = 28.5 bits (62), Expect = 6.2
Identities = 13/50 (26%), Positives = 26/50 (52%), Gaps = 5/50 (10%)
Query: 35 SEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKSLSSMGDYAGTSIKNW 84
++H L +SFNTT S + +W + +S + ++ D A +++ W
Sbjct: 317 AKHNLANSFTSFNTT-----YSDTGLWGIYIQSNNRDNLDDLAHFTVREW 361
>YQU3_CAEEL (Q09550) Hypothetical protein F26C11.3 in chromosome II
Length = 1240
Score = 28.1 bits (61), Expect = 8.1
Identities = 29/121 (23%), Positives = 46/121 (37%), Gaps = 2/121 (1%)
Query: 2 SSIGHQQHRSFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVW 61
SS G Q + T P+T+ + S + S + SS T P +AS S S+
Sbjct: 235 SSTGSTQTTTPSTEPSTTITTPMEQSSTVSSVQKTRTSEDKPSSSTTVPTSASTSESSTS 294
Query: 62 SSFPRSKSLSSMGDYAGTSIKNWWDWTWAWILSRKPIFARDLELNDQETKLLGGSHNRGT 121
S + S S+ + S + + + S L N+Q T G H+ T
Sbjct: 295 SPMAETSSSSTTSQSSPASTSTVPE--SSTVGSTPTTGLTTLSTNEQSTSTSSGGHSTST 352
Query: 122 W 122
+
Sbjct: 353 F 353
>VTA2_XENLA (P18709) Vitellogenin A2 precursor (VTG A2) [Contains:
Lipovitellin I; Lipovitellin II; Phosvitin]
Length = 1807
Score = 28.1 bits (61), Expect = 8.1
Identities = 17/53 (32%), Positives = 31/53 (58%)
Query: 19 SNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKSLS 71
S K+ H+R +D+ RK+ SS +++ +++S+S+S+ SS S S S
Sbjct: 1101 SKQKKKNKIHNRRLDAEVVEARKQQSSLSSSSSSSSSSSSSSSSSSSSSSSSS 1153
>KCCS_MALDO (Q07250) Calcium/calmodulin-dependent
serine/threonine-protein kinase (EC 2.7.1.123)
Length = 415
Score = 28.1 bits (61), Expect = 8.1
Identities = 19/54 (35%), Positives = 28/54 (51%), Gaps = 3/54 (5%)
Query: 26 VSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSS--FPRSKSLSS-MGDY 76
V S D + + RL SFN +A+ ++VW+S F R+K L S +G Y
Sbjct: 309 VGLSAREDQMDAEIVSRLQSFNARRKLRAAAIASVWTSSIFLRTKKLKSLLGSY 362
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.127 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,605,753
Number of Sequences: 164201
Number of extensions: 676734
Number of successful extensions: 1923
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 1906
Number of HSP's gapped (non-prelim): 25
length of query: 155
length of database: 59,974,054
effective HSP length: 101
effective length of query: 54
effective length of database: 43,389,753
effective search space: 2343046662
effective search space used: 2343046662
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0380.7


