

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0378b.13
(282 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
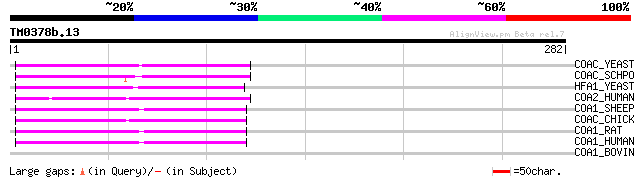
Score E
Sequences producing significant alignments: (bits) Value
COAC_YEAST (Q00955) Acetyl-CoA carboxylase (EC 6.4.1.2) (ACC) [I... 70 6e-12
COAC_SCHPO (P78820) Acetyl-CoA carboxylase (EC 6.4.1.2) (ACC) [I... 63 7e-10
HFA1_YEAST (P32874) HFA1 protein 56 1e-07
COA2_HUMAN (O00763) Acetyl-CoA carboxylase 2 (EC 6.4.1.2) (ACC-b... 48 2e-05
COA1_SHEEP (Q28559) Acetyl-CoA carboxylase 1 (EC 6.4.1.2) (ACC-a... 46 9e-05
COAC_CHICK (P11029) Acetyl-CoA carboxylase (EC 6.4.1.2) (ACC) [I... 46 1e-04
COA1_RAT (P11497) Acetyl-CoA carboxylase 1 (EC 6.4.1.2) (ACC-alp... 45 2e-04
COA1_HUMAN (Q13085) Acetyl-CoA carboxylase 1 (EC 6.4.1.2) (ACC-a... 44 3e-04
COA1_BOVIN (Q9TTS3) Acetyl-CoA carboxylase 1 (EC 6.4.1.2) (ACC-a... 43 0.001
>COAC_YEAST (Q00955) Acetyl-CoA carboxylase (EC 6.4.1.2) (ACC)
[Includes: Biotin carboxylase (EC 6.3.4.14)]
Length = 2233
Score = 70.1 bits (170), Expect = 6e-12
Identities = 45/119 (37%), Positives = 67/119 (55%), Gaps = 1/119 (0%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+ + G +H IY +EE A T L +D LL+ ++DP +L L +LV
Sbjct: 664 LIAIGGKSHTIYWKEEVAATRLSVDSMTTLLEVENDPTQLRTPSPGKLVKFLVENGEHII 723
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF 122
+ PYAE+EV+KM MPL+S + I+Q G + A +++A + LDDPS V+ A PF
Sbjct: 724 KGQ-PYAEIEVMKMQMPLVSQENGIVQLLKQPGSTIVAGDIMAIMTLDDPSKVKHALPF 781
>COAC_SCHPO (P78820) Acetyl-CoA carboxylase (EC 6.4.1.2) (ACC)
[Includes: Biotin carboxylase (EC 6.3.4.14)]
Length = 2280
Score = 63.2 bits (152), Expect = 7e-10
Identities = 41/121 (33%), Positives = 67/121 (54%), Gaps = 5/121 (4%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVS--KPLS 61
L+ L+G+++ +Y +E GT + ID + +L+ ++DP +L L +LV + +
Sbjct: 674 LVLLNGHSYTVYYRDEVTGTRISIDNLSCMLEQENDPTQLRTPSPGKLVRFLVETGEHIK 733
Query: 62 FAE*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEP 121
E YAEVEV+KM MPL++ ++Q G + A +++ L LDDPS V A P
Sbjct: 734 AGE---AYAEVEVMKMIMPLVATEDGVVQLIKQPGASLDAGDILGILTLDDPSRVTHALP 790
Query: 122 F 122
F
Sbjct: 791 F 791
>HFA1_YEAST (P32874) HFA1 protein
Length = 2273
Score = 55.8 bits (133), Expect = 1e-07
Identities = 37/117 (31%), Positives = 64/117 (54%), Gaps = 3/117 (2%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLV-SKPLSF 62
L+ +DG H +Y +++ GT L ID L+ + +P ++ L +LV S F
Sbjct: 733 LISVDGKCHTVYWKDDIRGTRLSIDSNTIFLEAELNPTQVISPTPGKLVKYLVRSGDHVF 792
Query: 63 AE*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKA 119
A YAE+E++KM MPL++ + +I+ G ++A ++IA+L LD PS ++
Sbjct: 793 A--GQQYAEIEIMKMQMPLVAKSDGVIELLRQPGSIIEAGDVIAKLTLDSPSKANES 847
>COA2_HUMAN (O00763) Acetyl-CoA carboxylase 2 (EC 6.4.1.2)
(ACC-beta) [Includes: Biotin carboxylase (EC 6.3.4.14)]
Length = 2483
Score = 48.1 bits (113), Expect = 2e-05
Identities = 38/119 (31%), Positives = 59/119 (48%), Gaps = 2/119 (1%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
LL +GN++ Y++EE + I + + + ++DP L L V
Sbjct: 857 LLSYNGNSYTTYMKEEV-DSYRTIGNKTCVFEKENDPTVLRSPSAGKLTQITVEDG-GHV 914
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF 122
E YAE+EV+KM M L +++ G ++A ++ARL+LDDPS V AEPF
Sbjct: 915 EAGRRYAEMEVMKMIMTLNVQERGRVKYIKRPGAVLEAGCVVARLELDDPSKVHPAEPF 973
>COA1_SHEEP (Q28559) Acetyl-CoA carboxylase 1 (EC 6.4.1.2)
(ACC-alpha) [Includes: Biotin carboxylase (EC 6.3.4.14)]
Length = 2346
Score = 46.2 bits (108), Expect = 9e-05
Identities = 36/118 (30%), Positives = 60/118 (50%), Gaps = 3/118 (2%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLS-F 62
LL DG+++ Y++EE + I + + + ++DP L L ++V F
Sbjct: 715 LLSYDGSSYTTYMKEEVDRYRITIGNKTCVFEKENDPSVLRSPSAGKLIQYIVEDGGHVF 774
Query: 63 AE*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAE 120
A YAE+EV+KM M L + S I + G + +IA++ LD+PS V++AE
Sbjct: 775 AGQC--YAEIEVMKMVMTLTAAESGCIHYVKRPGAALDPGCVIAKMQLDNPSKVQQAE 830
>COAC_CHICK (P11029) Acetyl-CoA carboxylase (EC 6.4.1.2) (ACC)
[Includes: Biotin carboxylase (EC 6.3.4.14)]
Length = 2324
Score = 45.8 bits (107), Expect = 1e-04
Identities = 35/117 (29%), Positives = 58/117 (48%), Gaps = 1/117 (0%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
LL DG+++ Y++EE + I + + + ++DP L L ++V
Sbjct: 715 LLSYDGSSYTTYMKEEVDRYRITIGNKTCVFEKENDPSILRSPSAGKLIQYVVEDG-GHV 773
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAE 120
+AE+EV+KM M L + S I + G + +IA+L LDDPS V++AE
Sbjct: 774 FAGQCFAEIEVMKMVMTLTAGESGCIHYVKRPGAVLDPGCVIAKLQLDDPSRVQQAE 830
>COA1_RAT (P11497) Acetyl-CoA carboxylase 1 (EC 6.4.1.2) (ACC-alpha)
[Includes: Biotin carboxylase (EC 6.3.4.14)]
Length = 2345
Score = 45.1 bits (105), Expect = 2e-04
Identities = 35/118 (29%), Positives = 60/118 (50%), Gaps = 3/118 (2%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLS-F 62
LL DG+++ Y++EE + I + + + ++DP + L ++V F
Sbjct: 714 LLSYDGSSYTTYMKEEVDRYRITIGNKTCVFEKENDPSVMRSPSAGKLIQYIVEDGGHVF 773
Query: 63 AE*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAE 120
A YAE+EV+KM M L + S I + G + +IA++ LD+PS V++AE
Sbjct: 774 AGQC--YAEIEVMKMVMTLTAVESGCIHYVKRPGAALDPGCVIAKMQLDNPSKVQQAE 829
>COA1_HUMAN (Q13085) Acetyl-CoA carboxylase 1 (EC 6.4.1.2)
(ACC-alpha) [Includes: Biotin carboxylase (EC 6.3.4.14)]
Length = 2346
Score = 44.3 bits (103), Expect = 3e-04
Identities = 34/118 (28%), Positives = 60/118 (50%), Gaps = 3/118 (2%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLS-F 62
LL DG+++ Y++EE + I + + + ++DP + L ++V F
Sbjct: 715 LLSYDGSSYTTYMKEEVDRYRITIGNKTCVFEKENDPSVMRSPSAGKLIQYIVEDGGHVF 774
Query: 63 AE*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAE 120
A YAE+EV+KM M L + S I + G + ++A++ LD+PS V++AE
Sbjct: 775 AGQC--YAEIEVMKMVMTLTAVESGCIHYVKRPGAALDPGCVLAKMQLDNPSKVQQAE 830
>COA1_BOVIN (Q9TTS3) Acetyl-CoA carboxylase 1 (EC 6.4.1.2)
(ACC-alpha) [Includes: Biotin carboxylase (EC 6.3.4.14)]
Length = 2346
Score = 42.7 bits (99), Expect = 0.001
Identities = 35/118 (29%), Positives = 59/118 (49%), Gaps = 3/118 (2%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLS-F 62
LL D +++ Y++EE + I + + + ++DP L L ++V F
Sbjct: 715 LLSYDVSSYTTYMKEEVDRYRITIGNKTCVFEKENDPSVLRSPSAGKLIQYIVEDGGHVF 774
Query: 63 AE*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAE 120
A YAE+EV+KM M L + S I + G + +IA++ LD+PS V++AE
Sbjct: 775 AGQC--YAEIEVMKMVMTLTAAESGCIHYVKRPGAALDPGCVIAKMQLDNPSKVQQAE 830
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.359 0.161 0.584
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,927,287
Number of Sequences: 164201
Number of extensions: 1001416
Number of successful extensions: 3564
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 3544
Number of HSP's gapped (non-prelim): 15
length of query: 282
length of database: 59,974,054
effective HSP length: 109
effective length of query: 173
effective length of database: 42,076,145
effective search space: 7279173085
effective search space used: 7279173085
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0378b.13


