

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0377b.5
(372 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
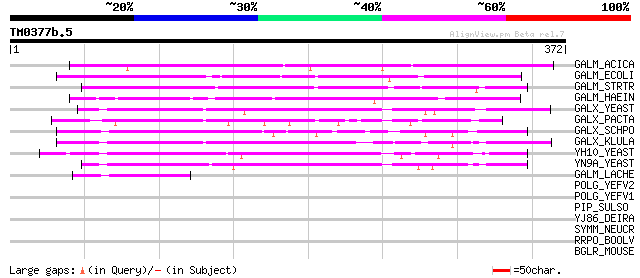
Score E
Sequences producing significant alignments: (bits) Value
GALM_ACICA (P05149) Aldose 1-epimerase precursor (EC 5.1.3.3) (M... 239 6e-63
GALM_ECOLI (P40681) Aldose 1-epimerase (EC 5.1.3.3) (Mutarotase) 179 8e-45
GALM_STRTR (P21955) Aldose 1-epimerase (EC 5.1.3.3) (Mutarotase) 167 5e-41
GALM_HAEIN (P31765) Aldose 1-epimerase (EC 5.1.3.3) (Mutarotase) 149 1e-35
GALX_YEAST (P04397) GAL10 bifunctional protein [Includes: UDP-gl... 122 2e-27
GALX_PACTA (P40801) GAL10 bifunctional protein [Includes: UDP-gl... 108 2e-23
GALX_SCHPO (Q9HDU3) GAL10 bifunctional protein [Includes: UDP-gl... 106 1e-22
GALX_KLULA (P09609) GAL10 bifunctional protein [Includes: UDP-gl... 101 3e-21
YH10_YEAST (P38893) Hypothetical 37.9 kDa protein in TWT1-FLO5 i... 94 4e-19
YN9A_YEAST (P53757) Hypothetical 37.9 kDa protein in BIO3-HXT17 ... 92 2e-18
GALM_LACHE (Q00053) Aldose 1-epimerase (EC 5.1.3.3) (Mutarotase)... 51 5e-06
POLG_YEFV2 (P19901) Genome polyprotein [Contains: Capsid protein... 32 2.0
POLG_YEFV1 (P03314) Genome polyprotein [Contains: Capsid protein... 32 2.0
PIP_SULSO (Q97UA2) Proline iminopeptidase (EC 3.4.11.5) (PIP) (P... 30 7.5
YJ86_DEIRA (Q9RSY3) Hypothetical UPF0230 protein DR1986 30 9.9
SYMM_NEUCR (Q9C2H9) Probable methionyl-tRNA synthetase, mitochon... 30 9.9
RRPO_BOOLV (Q992J0) RNA-directed RNA polymerase (EC 2.7.7.48) (R... 30 9.9
BGLR_MOUSE (P12265) Beta-glucuronidase precursor (EC 3.2.1.31) 30 9.9
>GALM_ACICA (P05149) Aldose 1-epimerase precursor (EC 5.1.3.3)
(Mutarotase)
Length = 381
Score = 239 bits (611), Expect = 6e-63
Identities = 138/335 (41%), Positives = 196/335 (58%), Gaps = 13/335 (3%)
Query: 41 LSLKVSNWGATLVSLVLPDKHGKLGDVVLGYDSLKQY----TNDSTYFGTTAGRVANRIG 96
+S+ ++G + ++ PD GK ++VLG+D LK Y T + +FG GR ANRIG
Sbjct: 48 VSVSFISFGGVITQILTPDAQGKQNNIVLGFDDLKGYEVTDTKEGIHFGGLIGRYANRIG 107
Query: 97 GAQFTLNGIHYKLIANEGNNTLHGGPRGFSDVLWKVERYVKEGDKPRITFSYHSFDGEEG 156
A+F+L+G Y L N G N+LH G GF +W+V+ V +G+ + + S +G++G
Sbjct: 108 NAKFSLDGKTYNLEKNNGPNSLHSGNPGFDKRVWQVKPLVSKGETVKASLKLTSPNGDQG 167
Query: 157 FPGDLLVTVSYILGSKNSLSIIMKAKALNKPTPVNLINHAYWNL--GNHNSGNILGEVVQ 214
FPG L V V Y L +N I KAK ++PT VNL NH+Y+NL +N +L VVQ
Sbjct: 168 FPGKLDVEVIYSLSDQNEFKIEYKAKT-DQPTVVNLTNHSYFNLSGAGNNPYGVLDHVVQ 226
Query: 215 IFGSRITVLDSHLIPTGEIASVKGTPYDFLKPQV----VGARINQLPKTNGYDINYVLDG 270
+ RI V D + +PTGEIASV GTP+DF P+ + A QL GYD +V++
Sbjct: 227 LNAGRILVTDQNSLPTGEIASVAGTPFDFRMPKAIVKDIRANNQQLAYGYGYDQTWVIN- 285
Query: 271 GKSKGLIKLAAIVMDKKSGRVMKLFTNAPGLQFYTANK-VKNEKGKGGFVYQPRAGLCLE 329
KS+G + LAAIV+D KS R M++ T P +Q YTA+ + N G G +Y+ L LE
Sbjct: 286 QKSQGKLNLAAIVVDPKSKRTMQVLTTEPSVQMYTADHLLGNIVGANGVLYRQADALALE 345
Query: 330 SQVFPDSVNHPNFPSSIVTPEKPYKHVMLLKFSTK 364
+Q FPDS N P FPS+ + P + Y V + KF +
Sbjct: 346 TQHFPDSPNQPTFPSTRLNPNQTYNSVTVFKFGVQ 380
>GALM_ECOLI (P40681) Aldose 1-epimerase (EC 5.1.3.3) (Mutarotase)
Length = 346
Score = 179 bits (455), Expect = 8e-45
Identities = 110/317 (34%), Positives = 169/317 (52%), Gaps = 14/317 (4%)
Query: 32 RLFELKKG-DLSLKVSNWGATLVSLVLPDKHGKLGDVVLGYDSLKQYTNDSTYFGTTAGR 90
RL L+ + + + +WGATL+S +P G + + +LG S + Y + + + G + GR
Sbjct: 16 RLLTLRNNAGMVVTLMDWGATLLSARIPLSDGSVREALLGCASPECYQDQAAFLGASIGR 75
Query: 91 VANRIGGAQFTLNGIHYKLIANEGNNTLHGGPRGFSDVLWKVERYVKEGDKPRITFSYHS 150
ANRI +++T +G L ++G N LHGGP GF W++ V + D+ ++ F+ S
Sbjct: 76 YANRIANSRYTFDGETVTLSPSQGVNQLHGGPEGFDKRRWQI---VNQNDR-QVLFALSS 131
Query: 151 FDGEEGFPGDLLVTVSYILGSKNSLSIIMKAKALNKPTPVNLINHAYWNLGNHNSGNILG 210
DG++GFPG+L TV Y L N +SI +A ++KP PVN+ NH Y+NL S ++
Sbjct: 132 DDGDQGFPGNLGATVQYRLTDDNRISITYRA-TVDKPCPVNMTNHVYFNLDGEQS-DVRN 189
Query: 211 EVVQIFGSRITVLDSHLIPTGEIASVKGTPYDFLKPQVVGARI---NQLPKTNGYDINYV 267
+QI +D IP + SV GT +DF +++ + + K GYD ++
Sbjct: 190 HKLQILADEYLPVDEGGIPHDGLKSVAGTSFDFRSAKIIASEFLADDDQRKVKGYDHAFL 249
Query: 268 LDG-GKSKGLIKLAAIVMDKKSGRVMKLFTNAPGLQFYTANKVKNEKGKGGFVYQPRAGL 326
L G K K+AA V +K++T AP LQFY+ N + +G Y GL
Sbjct: 250 LQAKGDGK---KVAAHVWSADEKLQLKVYTTAPALQFYSGNFLGGTPSRGTEPYADWQGL 306
Query: 327 CLESQVFPDSVNHPNFP 343
LES+ PDS NHP +P
Sbjct: 307 ALESEFLPDSPNHPEWP 323
>GALM_STRTR (P21955) Aldose 1-epimerase (EC 5.1.3.3) (Mutarotase)
Length = 348
Score = 167 bits (422), Expect = 5e-41
Identities = 105/305 (34%), Positives = 164/305 (53%), Gaps = 16/305 (5%)
Query: 49 GATLVSLVLPDKHGKLGDVVLGYDSLKQYTNDSTYFGTTAGRVANRIGGAQFTLNGIHYK 108
GATL ++P + G L ++VLG+ + Y ++ + GRVA RIG A +T N + Y
Sbjct: 35 GATLQEFLVPMETGALKNIVLGFSDFEDYYKNNLCACQSIGRVAGRIGKASYTHNMVLYS 94
Query: 109 LIANEGNNTLHGGPRGFSDVLWKVERYVKEGDKPRITFSYHSFDGEEGFPGDLLVTVSYI 168
L NEG+N LHGGP+G W + + D F + +GFPGD+ V++SY
Sbjct: 95 LPKNEGDNCLHGGPKGMQVQNWNYVTNLND-DYVETKFIRRLYSSVDGFPGDVTVSISYR 153
Query: 169 LGSKNSLSIIMKAKALNKPTPVNLINHAYWNLGNHNSGNILGEVVQIFGSRITVLDSHLI 228
L + N L+I+ +A + + T N NH Y+NL + ++ +QI+ LDS LI
Sbjct: 154 LNNNNRLTILFEAFDVTESTVFNPTNHVYFNLSDKQ--DLSSHELQIYSDYRLELDSELI 211
Query: 229 PTGEIASVKGTPYDFLKPQVVGARINQLPKTNGYDINYVLDGGKSKGLIKLAAIVMDKKS 288
PTG+ +V T YDF K + RI NG+D +V+ GG +K AI+ DK+S
Sbjct: 212 PTGQKINVDETNYDFRKTTDLLPRIE---ANNGFDDAFVVGGGTCDH-VKEVAILHDKES 267
Query: 289 GRVMKLFTNAPGLQFYTANKVKN------EKGKGGFVYQPRAGLCLESQVFPDSVNHPNF 342
G +++F+N GL +T + +++ +KGK + + R + +E+Q PD+VNH F
Sbjct: 268 GDGIEIFSNRNGLVIFTMDDIEDNIYFARDKGK---MAKRREAIAMEAQTLPDAVNHKGF 324
Query: 343 PSSIV 347
I+
Sbjct: 325 GDIIL 329
>GALM_HAEIN (P31765) Aldose 1-epimerase (EC 5.1.3.3) (Mutarotase)
Length = 340
Score = 149 bits (376), Expect = 1e-35
Identities = 100/305 (32%), Positives = 157/305 (50%), Gaps = 15/305 (4%)
Query: 41 LSLKVSNWGATLVSLVLPDKHGKLGDVVLGYDSLKQYTNDSTYFGTTAGRVANRIGGAQF 100
+ ++ +WGAT +S +P + L +V+LG + Y +Y G + GR ANRI AQF
Sbjct: 26 MRVQFMDWGATWLSCKVP-VNDTLREVLLGC-KVDNYPTHQSYLGASVGRYANRIANAQF 83
Query: 101 TLNGIHYKLIANEGNNTLHGGPRGFSDVLWKVERYVKEGDKPRITFSYHSFDGEEGFPGD 160
LNG KL +N+G + LHGG GF W ++ E + + FS HS DG++GFPG+
Sbjct: 84 ELNGELIKLSSNQGKHQLHGG-EGFDKRRWNIQ----ECGENFVCFSLHSVDGDQGFPGN 138
Query: 161 LLVTVSYILGSKNSLSIIMKAKALNKPTPVNLINHAYWNLGNHNSGNILGE-VVQIFGSR 219
+ V+V+Y L NS+ I A +K T +NL NH Y+NL N G+ + E +++
Sbjct: 139 VDVSVTYTLTGDNSVK-IEYAGMCDKDTALNLTNHTYFNLENAEQGSDVREHTLRLNADF 197
Query: 220 ITVLDSHLIPTGEIASVKGTPYDF--LKPQVVGARINQLPKTNGYDINYVLDGGKSKGLI 277
+D+ IP + V T +DF KP T GYD +++++ K +
Sbjct: 198 YLPVDNEGIPNSPLKHVVNTSFDFRIAKPIKQDFLQGDQQATKGYDHSFIVNKAWQKPCV 257
Query: 278 KLAAIVMDKKSGRVMKLFTNAPGLQFYTANKVKNEKGKGGFVYQPRAGLCLESQVFPDSV 337
L + D +++ T+ LQ YT N + + G +Y +G+ LE+Q PD+
Sbjct: 258 LLTSPTGDLS----LEVRTSQAALQVYTGNYLAGTPTRNGELYADFSGIALETQCLPDTP 313
Query: 338 NHPNF 342
NHP +
Sbjct: 314 NHPEW 318
>GALX_YEAST (P04397) GAL10 bifunctional protein [Includes:
UDP-glucose 4-epimerase (EC 5.1.3.2) (Galactowaldenase);
Aldose 1-epimerase (EC 5.1.3.3) (Mutarotase)]
Length = 699
Score = 122 bits (305), Expect = 2e-27
Identities = 96/330 (29%), Positives = 153/330 (46%), Gaps = 32/330 (9%)
Query: 46 SNWGATLVSLVLPDKHGKLGDVVLGYDSLKQYTN-DSTYFGTTAGRVANRIGGAQFTLNG 104
+N GA++V L + + VVLGY++ + Y N DS Y G T GR ANRI +F+L
Sbjct: 389 ANLGASIVDLKVNGQ-----SVVLGYENEEGYLNPDSAYIGATIGRYANRISKGKFSLCN 443
Query: 105 IHYKLIANEGNNTLHGGPRGFSDVLWKVERYVKEGDKPRITFSYHSFDGEEG--FPGDLL 162
Y+L N G N H F + + ++ K T Y D E+ FPGDLL
Sbjct: 444 KDYQLTVNNGVNANHSSIGSFHRKRF-LGPIIQNPSKDVFTAEYMLIDNEKDTEFPGDLL 502
Query: 163 VTVSYILG-SKNSLSIIMKAK-ALNKPTPVNLINHAYWNLGNHNSGNILGEVVQIFGSRI 220
VT+ Y + ++ SL ++ K K + TP+NL NH+Y+NL I G + + +
Sbjct: 503 VTIQYTVNVAQKSLEMVYKGKLTAGEATPINLTNHSYFNLNKPYGDTIEGTEIMVRSKKS 562
Query: 221 TVLDSHLIPTGEIASVKGTPYDFLKPQVVGARINQLPKTNGYDINYVLDGGKSKGLI--- 277
+D ++IPTG I + ++ KP V+G PK +D +V+D I
Sbjct: 563 VDVDKNMIPTGNIVDREIATFNSTKPTVLG------PKNPQFDCCFVVDENAKPSQINTL 616
Query: 278 --KLAAIV--MDKKSGRVMKLFTNAPGLQFYTANKVKNEKGKGGFVYQPRAGLCLESQVF 333
+L IV S +++ + P QFYT + + Y+ R G +E +
Sbjct: 617 NNELTLIVKAFHPDSNITLEVLSTEPTYQFYTGDFLSAG-------YEARQGFAIEPGRY 669
Query: 334 PDSVNHPNFPSSIVTPE-KPYKHVMLLKFS 362
D++N N+ + + Y ++ +FS
Sbjct: 670 IDAINQENWKDCVTLKNGETYGSKIVYRFS 699
>GALX_PACTA (P40801) GAL10 bifunctional protein [Includes:
UDP-glucose 4-epimerase (EC 5.1.3.2) (Galactowaldenase);
Aldose 1-epimerase (EC 5.1.3.3) (Mutarotase)]
Length = 689
Score = 108 bits (270), Expect = 2e-23
Identities = 103/325 (31%), Positives = 149/325 (45%), Gaps = 50/325 (15%)
Query: 29 DHIRLFELKKGDLSLKVSNWGATLVSLVLPDKHGKLGDVVL--GYDSLKQYTN-DSTYFG 85
D + G + +SN GAT+V+ KL + L G+D+LK Y D+ +FG
Sbjct: 362 DRLNTVRTFNGKFEVSISNHGATIVA-------AKLNGIKLNLGFDNLKGYLREDNPFFG 414
Query: 86 TTAGRVANRIGGAQFTLNGIHYKLIANEGNNT-LHGGPRGFSDVLWKVERYVKEGDKPRI 144
T GRVANRI +NG HY++ NE + T LHGG G++ + + VK +K +
Sbjct: 415 ATIGRVANRISKGDLLINGTHYQVGLNELHRTSLHGGTYGYNKRTF-LGPIVKTNEKEKE 473
Query: 145 T---FSYHSFDGEEGFPGDLLVTVSYIL---GSKNSLSIIMKAKALNK----PTPVNLIN 194
T F DG EG+PGD+ V Y + G L I +AK L + T V+L N
Sbjct: 474 TTMEFVLIDLDGTEGYPGDVETKVIYTVRDTGVGGELGIEYEAKLLEESGRDSTAVSLTN 533
Query: 195 HAYWNLGNHNSGNILGEVVQIFGSR---ITVLDSHLIPTGEIASVKGTPYDFLKPQVVGA 251
H+YWN+GN S I G +++ ++ LDS PTG+I V T D +G
Sbjct: 534 HSYWNIGNQPS--IEGTHIKLVSNKHLESNPLDS--TPTGKI--VTSTDLDSQNSAKLG- 586
Query: 252 RINQLPKTNGYDINYV------LDGGKSKGLIKLAAIVMDKKSGRVMKLFTNAPGLQFYT 305
P +D +V LD + +++ A K+ T P QFYT
Sbjct: 587 -----PDGPVFDYCFVTKQQDKLDTRNDE--LRVVATATHPKTRIAFTTLTTEPAFQFYT 639
Query: 306 ANKVKNEKGKGGFVYQPRAGLCLES 330
+ V V+ R+G CLE+
Sbjct: 640 GDGV-----DVAGVFTKRSGFCLEA 659
>GALX_SCHPO (Q9HDU3) GAL10 bifunctional protein [Includes:
UDP-glucose 4-epimerase (EC 5.1.3.2) (Galactowaldenase);
Aldose 1-epimerase (EC 5.1.3.3) (Mutarotase)]
Length = 713
Score = 106 bits (264), Expect = 1e-22
Identities = 96/333 (28%), Positives = 150/333 (44%), Gaps = 43/333 (12%)
Query: 32 RLFELKKGDLSLKVSNWGATLVSLVLPDKHGKLGDVVLGYDSLKQYT-NDSTYFGTTAGR 90
RL + DL + ++N+GA + ++ ++ +V G++ +Y ++ +FG T GR
Sbjct: 369 RLHTICFQDLEVSIANYGALVQAVRYKGRN-----LVNGFNDFSRYKLKENPFFGATIGR 423
Query: 91 VANRIGGAQFTLNGIHYKLIANEGN-NTLHGGPRGFSDVLWKVERYVKEGDKPRITFSYH 149
ANRI QF ++G Y L NE N TLHGG GF + + D + F
Sbjct: 424 FANRIANGQFEVDGHLYTLCKNENNKTTLHGGNNGFDKQFFLGPIARQYEDYNTLEFILV 483
Query: 150 SFDGEEGFPGDLLVTVSYILGSKNSL-----SIIMKAKALNKPTPVNLINHAYWNLGNHN 204
DG GFP DL V Y + NSL S+I + LN T VNL NH+YWNL + N
Sbjct: 484 DKDGNNGFPSDLETLVKYTI-KNNSLEIEYKSVIPEYSKLN-VTAVNLTNHSYWNLASPN 541
Query: 205 ---SGNILGEVVQIFGSRITVLDSHLIPTGEIASVKGTPYDFLKPQVVGARINQLPKTNG 261
G I+ ++ + V +PTG+I + D KP + I+
Sbjct: 542 KTIDGTIIKSTTNVY---LKVNSETSLPTGDIVEWQN---DITKPTKLDPNIS------- 588
Query: 262 YDINYVLDGGKSKGLI-----KLAAIVMDKKSGRVMKLF--TNAPGLQFYTANKVKNEKG 314
+D +++D SK + L IV +KL T P Q YT + G
Sbjct: 589 FDNCFIVDREASKFCLDTRKYSLKNIVEVIHPSVPVKLVVSTTEPAFQLYTGD------G 642
Query: 315 KGGFVYQPRAGLCLESQVFPDSVNHPNFPSSIV 347
+Q R+G C+E+ F +++N+ + ++
Sbjct: 643 NDICEFQSRSGFCVETGRFINALNNEKWSKQVI 675
>GALX_KLULA (P09609) GAL10 bifunctional protein [Includes:
UDP-glucose 4-epimerase (EC 5.1.3.2) (Galactowaldenase);
Aldose 1-epimerase (EC 5.1.3.3) (Mutarotase)]
Length = 688
Score = 101 bits (252), Expect = 3e-21
Identities = 102/352 (28%), Positives = 154/352 (42%), Gaps = 47/352 (13%)
Query: 32 RLFELKKG-DLSLKVSNWGATLVSLVLPDKHGKLGDVVLGYDSLKQYTNDST-YFGTTAG 89
R+ L KG +K++N GAT+V +V+ VV D+ +Y + S +FG T G
Sbjct: 363 RVVSLGKGTSFEVKLANLGATIVDIVVNGC-----SVVASLDNETEYKDQSNPFFGATVG 417
Query: 90 RVANRIGGAQFTLNGIHYKL-IANEGNNTLHGGPRGFSDVLWKVERYVKEGDKPRITFSY 148
ANRI F +NG +L ++ E N +H G + + + VK + T +
Sbjct: 418 PYANRIANGTFEINGKRIQLTVSKEDGNVVHSGINSYHSKKF-LGPIVKNPEAHIWTADF 476
Query: 149 HSFDGEEGFPGDLLVTVSYILGS-KNSLSIIMKAKALNK-PTPVNLINHAYWNLGNHNSG 206
D E FP L V Y + S K +L++ ++K + T N+ NH Y+NL N
Sbjct: 477 KYVDEETEFPARLSTLVRYKVDSEKKTLTVEYESKVAGQGSTGANITNHTYFNLNKFNEA 536
Query: 207 NILGEVVQIFGSRITVLDSHLIPTGEIASVKGTPYDFLKPQVVGARINQLPKTNGYDINY 266
I G VQ+ + +++ L+PTG LK A N P T +
Sbjct: 537 TIKGSKVQLIDNTGLEVNNKLLPTGN-----------LKKYTQVATFNDSP-TEITEKEP 584
Query: 267 VLDGGKSKGLIKLAAIVMDKKSGRVMKLF--------------TNAPGLQFYTANKVKNE 312
VLD GL +D +S + +F T P Q YT + V +
Sbjct: 585 VLDFCFVSGL----PAKLDTRSSPLTPVFKLSNEDAKLEVEVATTEPTFQVYTGDYV-DV 639
Query: 313 KGKGGFVYQPRAGLCLESQVFPDSVNHPNFPSSIVTPE-KPYKHVMLLKFST 363
K K Y+ RAG+C E + D+VN+P + SS+V P + Y H + F T
Sbjct: 640 KDK----YENRAGICCEPGRYIDAVNNPEWKSSVVLPAGETYGHKLSYTFRT 687
>YH10_YEAST (P38893) Hypothetical 37.9 kDa protein in TWT1-FLO5
intergenic region
Length = 341
Score = 94.4 bits (233), Expect = 4e-19
Identities = 93/347 (26%), Positives = 159/347 (45%), Gaps = 45/347 (12%)
Query: 21 NGSAKKEKDHIRLFELKKGDLSLKVSNWGATLVSLVLPDKHGKLGDVVLGYDSLKQYTND 80
N A E + I + + KK L +S GATL+ L + ++ +VLGY + Y +D
Sbjct: 3 NNKAGGEYEVITIGDAKK--LQATISELGATLLDLKVNNE-----SIVLGYPDIHGYISD 55
Query: 81 S-TYFGTTAGRVANRIGGAQFTLNGIHYKLIANEGNNTLHGGPRGFSDVLWKVERYVKEG 139
Y G T GR ANRI F++ ++L N NT H F +K +
Sbjct: 56 GYNYIGATVGRYANRIYKGMFSMEDGPHQLTVNNCGNTNHSSISSFHLKKYKASKVQNPL 115
Query: 140 DKPRITFSYHSFDGE---EGFPGDLLVTVSYILG-SKNSLSIIMKAKALN-KPTPVNLIN 194
D I + D FPGDL V + Y L + +L + +AK ++ + TP+N+ N
Sbjct: 116 DDLYIV-EFTLLDDRTLPNEFPGDLAVNLKYTLNVADMTLDLEYEAKLVSGEATPINMTN 174
Query: 195 HAYWNLG-NHNSGNILGEVVQIFGSR-ITVLDSHLIPTGEIASVKGTPYDFLKPQVVGAR 252
H Y+NL + +I G V++ + + V + LIPTG+I K +D KP ++
Sbjct: 175 HTYFNLNKTMDKKSISGTEVRLCSDKSLEVSEGALIPTGKIVQRKIATFDSSKPTIL--- 231
Query: 253 INQLPKTNG--YDINYVLDGGKSKGLIKLAAIVMDK----------KSGRVMKLFTNAPG 300
+ +G YD +++D ++K L ++ ++K S +++ T P
Sbjct: 232 -----QDDGPIYDYAFIVD--ENKNLKTTDSVSVNKLVPAFKAYHPASRLSLEVSTTEPT 284
Query: 301 LQFYTANKVKNEKGKGGFVYQPRAGLCLESQVFPDSVNHPNFPSSIV 347
+ FYT + + + GF PR+G +E + D++N + ++
Sbjct: 285 VLFYTGDNLCD-----GFT--PRSGFAVEQGRYVDAINRDGWRDCVL 324
>YN9A_YEAST (P53757) Hypothetical 37.9 kDa protein in BIO3-HXT17
intergenic region
Length = 342
Score = 92.4 bits (228), Expect = 2e-18
Identities = 86/316 (27%), Positives = 142/316 (44%), Gaps = 36/316 (11%)
Query: 49 GATLVSLVLPDKHGKLGDVVLGYDSLKQYTNDSTYFGTTAGRVANRIGGAQFTLNGIHYK 108
GATLV L + + VV GY +++ Y D G T GR ANRI F+L+ +K
Sbjct: 29 GATLVDLKVNGQ-----SVVQGYSNVQDYLTDGNMMGATVGRYANRIAKGVFSLDDGPHK 83
Query: 109 LIANEGNNTLHGGPRGFSDVLWKVERYVKEGDKPRITFSY-----HSFDGEEGFPGDLLV 163
L N NT H + +K V+ K + H+ FPGDL V
Sbjct: 84 LTVNNCGNTNHSSISSLNLKQYKASP-VENPSKGVYVVEFKLLDDHTQPNPNEFPGDLEV 142
Query: 164 TVSYILG-SKNSLSIIMKAKAL-NKPTPVNLINHAYWNLGNHNS-GNILGEVVQIFGSR- 219
TV Y L ++ +L + +A+ + TP+N+ NH+Y+NL S +I G V++ ++
Sbjct: 143 TVKYTLNVAEMTLDMEYQAQLVRGDATPINMTNHSYFNLNKVKSEKSIRGTEVKVCSNKS 202
Query: 220 ITVLDSHLIPTGEIASVKGTPYDFLKPQVVGARINQLPKTNGYDINYVLDGGK------S 273
+ V + L+PTG+I +D KP V+ T +D +++D K S
Sbjct: 203 LEVTEGALLPTGKIIERNIATFDSTKPTVLH------EDTPVFDCTFIIDANKDLKTTDS 256
Query: 274 KGLIKLAAI--VMDKKSGRVMKLFTNAPGLQFYTANKVKNEKGKGGFVYQPRAGLCLESQ 331
+ KL + +S ++ T P + YT + N GK + PR+G ++
Sbjct: 257 VSVNKLVPVFKAYHPESHIKFEVSTTEPTVHLYTGD---NLCGK----FVPRSGFAVQQG 309
Query: 332 VFPDSVNHPNFPSSIV 347
+ D++N + ++
Sbjct: 310 RYVDAINRDEWRGCVL 325
>GALM_LACHE (Q00053) Aldose 1-epimerase (EC 5.1.3.3) (Mutarotase)
(Fragment)
Length = 129
Score = 50.8 bits (120), Expect = 5e-06
Identities = 27/79 (34%), Positives = 45/79 (56%), Gaps = 5/79 (6%)
Query: 43 LKVSNWGATLVSLVLPDKHGKLGDVVLGYDSLKQYTNDSTYFGTTAGRVANRIGGAQFTL 102
+KV N+GATL ++L D++ ++L +S Y+ + Y G T GR+A R+ Q+
Sbjct: 29 VKVLNYGATLEKVLLNDEN-----MILSLNSPADYSQERNYLGGTVGRIAGRVRKGQWRH 83
Query: 103 NGIHYKLIANEGNNTLHGG 121
++L N+G N +HGG
Sbjct: 84 GLETHQLPINDGENHIHGG 102
>POLG_YEFV2 (P19901) Genome polyprotein [Contains: Capsid protein C
(Core protein); Matrix protein (Envelope protein M);
Major envelope protein E; Nonstructural proteins NS1,
NS2A, NS4A and NS4B; Flavivirin (EC 3.4.21.91) (NS2B/NS3
proteinase); RNA-direct
Length = 3411
Score = 32.3 bits (72), Expect = 2.0
Identities = 18/64 (28%), Positives = 32/64 (49%)
Query: 50 ATLVSLVLPDKHGKLGDVVLGYDSLKQYTNDSTYFGTTAGRVANRIGGAQFTLNGIHYKL 109
AT+ L L ++ G L + G + + TND+ + G V+ R+ + TL G YK+
Sbjct: 525 ATIRVLALGNQEGSLKTALTGAMRVTKDTNDNNLYKLHGGHVSCRVKLSALTLKGTSYKI 584
Query: 110 IANE 113
++
Sbjct: 585 CTDK 588
>POLG_YEFV1 (P03314) Genome polyprotein [Contains: Capsid protein C
(Core protein); Matrix protein (Envelope protein M);
Major envelope protein E; Nonstructural proteins NS1,
NS2A, NS4A and NS4B; Flavivirin (EC 3.4.21.91) (NS2B/NS3
proteinase); RNA-direct
Length = 3411
Score = 32.3 bits (72), Expect = 2.0
Identities = 18/64 (28%), Positives = 32/64 (49%)
Query: 50 ATLVSLVLPDKHGKLGDVVLGYDSLKQYTNDSTYFGTTAGRVANRIGGAQFTLNGIHYKL 109
AT+ L L ++ G L + G + + TND+ + G V+ R+ + TL G YK+
Sbjct: 525 ATIRVLALGNQEGSLKTALTGAMRVTKDTNDNNLYKLHGGHVSCRVKLSALTLKGTSYKI 584
Query: 110 IANE 113
++
Sbjct: 585 CTDK 588
>PIP_SULSO (Q97UA2) Proline iminopeptidase (EC 3.4.11.5) (PIP)
(Prolyl aminopeptidase) (PAP) (Tricorn protease
interacting factor F1)
Length = 310
Score = 30.4 bits (67), Expect = 7.5
Identities = 24/55 (43%), Positives = 29/55 (52%), Gaps = 7/55 (12%)
Query: 81 STYFGTTAGRVANRIGGAQ--FTLNGIHYKLIANEGNN----TLHGGPRGFSDVL 129
S YF T G N G + F +N I+YKL +G+N TLHGGP G D L
Sbjct: 2 SYYFITMKGYSINVEEGYKRIFGIN-IYYKLYRVKGSNRNLVTLHGGPGGSHDYL 55
>YJ86_DEIRA (Q9RSY3) Hypothetical UPF0230 protein DR1986
Length = 281
Score = 30.0 bits (66), Expect = 9.9
Identities = 17/45 (37%), Positives = 23/45 (50%), Gaps = 6/45 (13%)
Query: 193 INHAYWNLGNHNSGNILGEV------VQIFGSRITVLDSHLIPTG 231
+ HA L H SG + G V Q FG R+TV+D+H + G
Sbjct: 77 LEHADEVLSLHISGQLSGTVGSARLAAQEFGGRVTVVDTHTVTLG 121
>SYMM_NEUCR (Q9C2H9) Probable methionyl-tRNA synthetase,
mitochondrial (EC 6.1.1.10) (Methionine--tRNA ligase)
(MetRS)
Length = 622
Score = 30.0 bits (66), Expect = 9.9
Identities = 18/66 (27%), Positives = 31/66 (46%), Gaps = 4/66 (6%)
Query: 183 ALNKPTPVNLINHAYWNLGNHNSGNILGEVVQIFGSR----ITVLDSHLIPTGEIASVKG 238
AL+ P P +++HA+W + +G VV F + I + ++I G IA+
Sbjct: 345 ALDLPLPKRILSHAHWTMEKKKMSKSIGNVVNPFYAMERFGIDTMRFYMIHNGGIANDAD 404
Query: 239 TPYDFL 244
D+L
Sbjct: 405 YSNDWL 410
>RRPO_BOOLV (Q992J0) RNA-directed RNA polymerase (EC 2.7.7.48)
(RdRp) (RNA replicase) (Protein A)
Length = 998
Score = 30.0 bits (66), Expect = 9.9
Identities = 16/47 (34%), Positives = 23/47 (48%), Gaps = 6/47 (12%)
Query: 320 YQPRAGLCLESQVFPDSVNHPNFPSSIVTPEKPYKHVMLLKFSTKAP 366
Y P GLC S+VF D +N P T + P + + L +T+ P
Sbjct: 720 YNPEVGLCFLSRVFVDPLNTP------TTIQDPLRTLRKLHITTRDP 760
>BGLR_MOUSE (P12265) Beta-glucuronidase precursor (EC 3.2.1.31)
Length = 648
Score = 30.0 bits (66), Expect = 9.9
Identities = 22/94 (23%), Positives = 39/94 (41%), Gaps = 8/94 (8%)
Query: 134 RYVKEGDKPRITFSYHSF---DGEEGFPGDLLVTVSYILGSKNSLSIIMKAKAL-----N 185
R E + + +YHS +E G+L+ + + +++ L +I K +
Sbjct: 551 RMFSEEYQKAVLENYHSVLDQKRKEYVVGELIWNFADFMTNQSPLRVIGNKKGIFTRQRQ 610
Query: 186 KPTPVNLINHAYWNLGNHNSGNILGEVVQIFGSR 219
T ++ YW + N G+ G Q FGSR
Sbjct: 611 PKTSAFILRERYWRIANETGGHGSGPRTQCFGSR 644
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.138 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 47,028,130
Number of Sequences: 164201
Number of extensions: 2157990
Number of successful extensions: 4467
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 4421
Number of HSP's gapped (non-prelim): 19
length of query: 372
length of database: 59,974,054
effective HSP length: 112
effective length of query: 260
effective length of database: 41,583,542
effective search space: 10811720920
effective search space used: 10811720920
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0377b.5


