

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
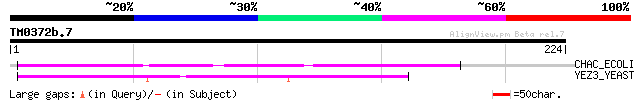
Query= TM0372b.7
(224 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CHAC_ECOLI (P39163) Cation transport protein chaC 80 4e-15
YEZ3_YEAST (P32656) Hypothetical 26.3 kDa protein in RAD4-CHD1 i... 77 3e-14
>CHAC_ECOLI (P39163) Cation transport protein chaC
Length = 231
Score = 80.1 bits (196), Expect = 4e-15
Identities = 56/179 (31%), Positives = 81/179 (44%), Gaps = 9/179 (5%)
Query: 4 WVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGEICW 63
W+FGYGSL+WNP E+ E G + + R F L RGT P R L KEG
Sbjct: 48 WIFGYGSLMWNPALEFTESCTGTLVGWHRAFCLRLTAGRGTAHQPGRMLAL--KEGGRTT 105
Query: 64 GAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPDKVN 123
G Y + +++L + + + E + + + D IVF P
Sbjct: 106 GVAYRLPEETLEQELTLLW----KREMITGCYLPTWCQLDLDDGRTVNAIVFIMDP---R 158
Query: 124 NKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHEDDMVIELAHEVRKVL 182
+ Y V+A IA A GP G N YLF LE+ + +G +DD + +L V+K+L
Sbjct: 159 HPEYESDTRAQVIAPLIAAASGPLGTNAQYLFSLEQELIKLGMQDDGLNDLLVSVKKLL 217
>YEZ3_YEAST (P32656) Hypothetical 26.3 kDa protein in RAD4-CHD1
intergenic region
Length = 232
Score = 77.4 bits (189), Expect = 3e-14
Identities = 55/198 (27%), Positives = 87/198 (43%), Gaps = 42/198 (21%)
Query: 4 WVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTL--------- 54
WV GYGSL++ P Y ++ I + R F + DHRGTP +P R TL
Sbjct: 9 WVLGYGSLIYKPPSHYTHRIPAIIHGFARRFWQSSTDHRGTPANPGRVATLIPYEDIIRQ 68
Query: 55 ----------------EEKEGEICWGAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDF 98
++ + + G VY + PE + V +YL RE V+
Sbjct: 69 TAFLKNVNLYSESAPIQDPDDLVTIGVVYYI--PPEHAQEVREYLNVREQNGYTLHEVEV 126
Query: 99 YKEGDSSHPALTG---------------VIVFTSTPDKVNNKYYLGPAPLDVMARQIATA 143
+ E + H A G V++ + ++N+ ++GP +D A+ IA +
Sbjct: 127 HLETNREHEAELGEALEQLPRHNKSGKRVLLTSVYIGTIDNEAFVGPETVDETAKVIAVS 186
Query: 144 HGPCGNNRDYLFLLEKAM 161
HGP G+N +YL LE+A+
Sbjct: 187 HGPSGSNYEYLAKLEQAL 204
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.141 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,284,554
Number of Sequences: 164201
Number of extensions: 1426929
Number of successful extensions: 2717
Number of sequences better than 10.0: 2
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 2713
Number of HSP's gapped (non-prelim): 2
length of query: 224
length of database: 59,974,054
effective HSP length: 106
effective length of query: 118
effective length of database: 42,568,748
effective search space: 5023112264
effective search space used: 5023112264
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0372b.7


