

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0372a.5
(392 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
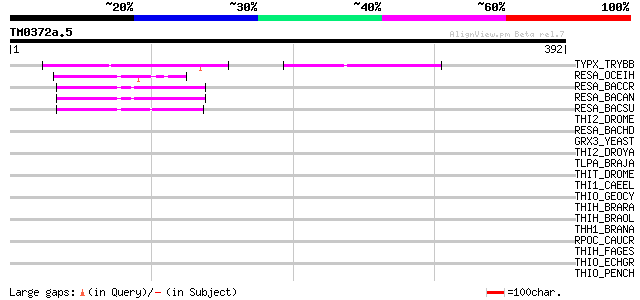
TYPX_TRYBB (O77404) Tryparedoxin 87 1e-16
RESA_OCEIH (Q8CXF3) Thiol-disulfide oxidoreductase resA 50 1e-05
RESA_BACCR (Q81FU5) Thiol-disulfide oxidoreductase resA 49 3e-05
RESA_BACAN (Q81SZ9) Thiol-disulfide oxidoreductase resA 49 3e-05
RESA_BACSU (P35160) Thiol-disulfide oxidoreductase resA 48 4e-05
THI2_DROME (Q9V429) Thioredoxin 2 (DmTrx-2) 42 0.004
RESA_BACHD (Q9KCJ4) Thiol-disulfide oxidoreductase resA 40 0.013
GRX3_YEAST (Q03835) Monothiol glutaredoxin 3 39 0.023
THI2_DROYA (Q6XHI1) Thioredoxin 2 37 0.066
TLPA_BRAJA (P43221) Thiol:disulfide interchange protein tlpA (Cy... 37 0.11
THIT_DROME (Q8IFW4) Thioredoxin T (ThioredoxinT) 36 0.15
THI1_CAEEL (Q09433) Probable thioredoxin B0228.5 36 0.15
THIO_GEOCY (O96952) Thioredoxin 36 0.19
THIH_BRARA (O64432) Thioredoxin H-type (TRX-H) 35 0.25
THIH_BRAOL (P68176) Thioredoxin H-type (TRX-H) (Pollen coat prot... 35 0.25
THH1_BRANA (P68177) Thioredoxin H-type 1 (TRX-H-1) 35 0.25
RPOC_CAUCR (Q9AAU1) DNA-directed RNA polymerase beta' chain (EC ... 35 0.25
THIH_FAGES (Q96419) Thioredoxin H-type (TRX-H) 35 0.33
THIO_ECHGR (O17486) Thioredoxin 34 0.73
THIO_PENCH (P34723) Thioredoxin 33 0.96
>TYPX_TRYBB (O77404) Tryparedoxin
Length = 144
Score = 86.7 bits (213), Expect = 1e-16
Identities = 50/134 (37%), Positives = 72/134 (53%), Gaps = 4/134 (2%)
Query: 24 GAEYLLSCEGKVPLSECDGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFV 83
GA LLS G+V L GK + L+FSA+WC PCR F P L E YE N E++ +
Sbjct: 10 GATNLLSKSGEVSLGSLVGKTVFLYFSASWCPPCRGFTPVLAEFYEK-HHVAKNFEVVLI 68
Query: 84 SFDREEDGFNEHLKSMPWLAVPFDV-NLHRRLIDRYQVDQIPSFIPLYSED--LIVEKNL 140
S+D E F+++ MPWLA+PFD + L + V+ IP+ I + ++ +I +
Sbjct: 69 SWDENESDFHDYYGKMPWLALPFDQRSTVSELGKTFGVESIPTLITINADTGAIIGTQAR 128
Query: 141 IECIEDYGADAFPF 154
IED FP+
Sbjct: 129 TRVIEDPDGANFPW 142
Score = 76.6 bits (187), Expect = 1e-13
Identities = 42/114 (36%), Positives = 61/114 (52%), Gaps = 4/114 (3%)
Query: 194 KVPVSELAGKTIGLYFCAYWSPPSRAFTAQLTDVYNNLKATKGNCFEIVLISTDRDHKEF 253
+V + L GKT+ LYF A W PP R FT L + Y K FE+VLIS D + +F
Sbjct: 20 EVSLGSLVGKTVFLYFSASWCPPCRGFTPVLAEFYEKHHVAKN--FEVVLISWDENESDF 77
Query: 254 NVNRSSTPWLAIPYEDR-TRHDLCRIYDIKGIPALVLIGPD-GKVISLNGKFMV 305
+ PWLA+P++ R T +L + + ++ IP L+ I D G +I + V
Sbjct: 78 HDYYGKMPWLALPFDQRSTVSELGKTFGVESIPTLITINADTGAIIGTQARTRV 131
>RESA_OCEIH (Q8CXF3) Thiol-disulfide oxidoreductase resA
Length = 192
Score = 50.1 bits (118), Expect = 1e-05
Identities = 34/98 (34%), Positives = 55/98 (55%), Gaps = 11/98 (11%)
Query: 32 EGKVPLSECDGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREE-- 89
+ V LS+ +GK + L F A WC PC+A +P + +LY +++G+ EI+ VS D E
Sbjct: 67 QSTVQLSDLEGKGVMLNFWATWCDPCKAEMPYMQDLYAEYKEKGV--EIVAVSLDGTELV 124
Query: 90 -DGF-NEHLKSMPWLAVPFDVNLHRRLIDRYQVDQIPS 125
D F +E+ + P VP D N + D Y++ +P+
Sbjct: 125 VDQFIDEYDLTFP---VPHDKN--GEVKDLYKIGPMPT 157
Score = 37.4 bits (85), Expect = 0.066
Identities = 32/107 (29%), Positives = 49/107 (44%), Gaps = 6/107 (5%)
Query: 191 DDRKVPVSELAGKTIGLYFCAYWSPPSRAFTAQLTDVYNNLKATKGNCFEIVLISTDRDH 250
D V +S+L GK + L F A W P +A + D+Y K KG EIV +S D
Sbjct: 66 DQSTVQLSDLEGKGVMLNFWATWCDPCKAEMPYMQDLYAEYK-EKG--VEIVAVSL--DG 120
Query: 251 KEFNVNRSSTPW-LAIPYEDRTRHDLCRIYDIKGIPALVLIGPDGKV 296
E V++ + L P ++ +Y I +P I P+G++
Sbjct: 121 TELVVDQFIDEYDLTFPVPHDKNGEVKDLYKIGPMPTTYFIKPNGEI 167
>RESA_BACCR (Q81FU5) Thiol-disulfide oxidoreductase resA
Length = 173
Score = 48.5 bits (114), Expect = 3e-05
Identities = 29/105 (27%), Positives = 54/105 (50%), Gaps = 3/105 (2%)
Query: 34 KVPLSECDGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREEDGFN 93
K+ L + GK + L F WC+PC +P + ELY +++G+ EII + D E D
Sbjct: 53 KIELKDLKGKGVFLNFWGTWCKPCEKEMPYMNELYPKYKEKGV--EIIALDAD-ETDIAV 109
Query: 94 EHLKSMPWLAVPFDVNLHRRLIDRYQVDQIPSFIPLYSEDLIVEK 138
++ + L P ++ +++I Y V +P+ + + +VE+
Sbjct: 110 KNFVNQYGLKFPVAIDKGQKIIGTYGVGPLPTSFLIDKDGKVVEQ 154
Score = 37.7 bits (86), Expect = 0.051
Identities = 31/128 (24%), Positives = 56/128 (43%), Gaps = 8/128 (6%)
Query: 173 ANLEEL-LAHEGRDFLISG-DDRKVPVSELAGKTIGLYFCAYWSPPSRAFTAQLTDVYNN 230
AN E++ + E +F+++ + +K+ + +L GK + L F W P + ++Y
Sbjct: 30 ANKEKMQIGKEAPNFIVTDLEGKKIELKDLKGKGVFLNFWGTWCKPCEKEMPYMNELYPK 89
Query: 231 LKATKGNCFEIVLISTDRDHKEFNVNRSSTPW-LAIPYEDRTRHDLCRIYDIKGIPALVL 289
K KG + +I+ D D + V + L P + Y + +P L
Sbjct: 90 YK-EKG----VEIIALDADETDIAVKNFVNQYGLKFPVAIDKGQKIIGTYGVGPLPTSFL 144
Query: 290 IGPDGKVI 297
I DGKV+
Sbjct: 145 IDKDGKVV 152
>RESA_BACAN (Q81SZ9) Thiol-disulfide oxidoreductase resA
Length = 173
Score = 48.5 bits (114), Expect = 3e-05
Identities = 29/105 (27%), Positives = 54/105 (50%), Gaps = 3/105 (2%)
Query: 34 KVPLSECDGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREEDGFN 93
K+ L + GK + L F WC+PC +P + ELY +++G+ EII + D E D
Sbjct: 53 KIELKDLKGKGVFLNFWGTWCKPCEKEMPYMNELYPKYKEKGV--EIIALDAD-ETDIAV 109
Query: 94 EHLKSMPWLAVPFDVNLHRRLIDRYQVDQIPSFIPLYSEDLIVEK 138
++ + L P ++ +++I Y V +P+ + + +VE+
Sbjct: 110 KNFVNQYGLKFPVAIDKGQKIIGTYGVGPLPTSFLIDKDGKVVEQ 154
Score = 35.8 bits (81), Expect = 0.19
Identities = 28/121 (23%), Positives = 51/121 (42%), Gaps = 7/121 (5%)
Query: 179 LAHEGRDFLISG-DDRKVPVSELAGKTIGLYFCAYWSPPSRAFTAQLTDVYNNLKATKGN 237
+ E +F+++ + +K+ + +L GK + L F W P + ++Y K KG
Sbjct: 37 IGKEAPNFVVTDLEGKKIELKDLKGKGVFLNFWGTWCKPCEKEMPYMNELYPKYK-EKG- 94
Query: 238 CFEIVLISTDRDHKEFNVNRSSTPW-LAIPYEDRTRHDLCRIYDIKGIPALVLIGPDGKV 296
+ +I+ D D + V + L P + Y + +P LI DGKV
Sbjct: 95 ---VEIIALDADETDIAVKNFVNQYGLKFPVAIDKGQKIIGTYGVGPLPTSFLIDKDGKV 151
Query: 297 I 297
+
Sbjct: 152 V 152
>RESA_BACSU (P35160) Thiol-disulfide oxidoreductase resA
Length = 179
Score = 48.1 bits (113), Expect = 4e-05
Identities = 26/104 (25%), Positives = 52/104 (50%), Gaps = 3/104 (2%)
Query: 34 KVPLSECDGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREEDGFN 93
++ LS+ GK + L F WC PC+ P + Y+ + +G+ EI+ V+ + +
Sbjct: 54 RIELSDLKGKGVFLNFWGTWCEPCKKEFPYMANQYKHFKSQGV--EIVAVNVGESKIAVH 111
Query: 94 EHLKSMPWLAVPFDVNLHRRLIDRYQVDQIPSFIPLYSEDLIVE 137
+KS + P ++ R+++D Y V +P+ + E +V+
Sbjct: 112 NFMKSY-GVNFPVVLDTDRQVLDAYDVSPLPTTFLINPEGKVVK 154
Score = 37.4 bits (85), Expect = 0.066
Identities = 25/107 (23%), Positives = 47/107 (43%), Gaps = 4/107 (3%)
Query: 193 RKVPVSELAGKTIGLYFCAYWSPPSRAFTAQLTDVYNNLKATKGNCFEIVLISTDRDHKE 252
+++ +S+L GK + L F W P + + + Y + K+ EIV ++
Sbjct: 53 KRIELSDLKGKGVFLNFWGTWCEPCKKEFPYMANQYKHFKSQG---VEIVAVNVGESKIA 109
Query: 253 FNVNRSSTPWLAIPYEDRTRHDLCRIYDIKGIPALVLIGPDGKVISL 299
+ N + + P T + YD+ +P LI P+GKV+ +
Sbjct: 110 VH-NFMKSYGVNFPVVLDTDRQVLDAYDVSPLPTTFLINPEGKVVKV 155
>THI2_DROME (Q9V429) Thioredoxin 2 (DmTrx-2)
Length = 114
Score = 41.6 bits (96), Expect = 0.004
Identities = 23/65 (35%), Positives = 34/65 (51%), Gaps = 4/65 (6%)
Query: 37 LSECDGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREED-GFNEH 95
L++ GK++ L F A WC PC+ P+LVEL N+ ++ V D ED +
Sbjct: 23 LTKASGKLVVLDFFATWCGPCKMISPKLVELSTQFAD---NVVVLKVDVDECEDIAMEYN 79
Query: 96 LKSMP 100
+ SMP
Sbjct: 80 ISSMP 84
>RESA_BACHD (Q9KCJ4) Thiol-disulfide oxidoreductase resA
Length = 176
Score = 39.7 bits (91), Expect = 0.013
Identities = 27/103 (26%), Positives = 50/103 (48%), Gaps = 3/103 (2%)
Query: 35 VPLSECDGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREEDGFNE 94
+ L E +GK + L F +C PC +P + +LY +++G+ EII V+ + E
Sbjct: 55 IELRELEGKGVFLNFWGTYCPPCEREMPHMEKLYGEYKEQGV--EIIAVNANEPELTVQR 112
Query: 95 HLKSMPWLAVPFDVNLHRRLIDRYQVDQIPSFIPLYSEDLIVE 137
+ L+ P ++ +ID Y + +P+ I + IV+
Sbjct: 113 FVDRY-GLSFPIVIDKGLNVIDAYGIRPLPTTILINEHGEIVK 154
>GRX3_YEAST (Q03835) Monothiol glutaredoxin 3
Length = 285
Score = 38.9 bits (89), Expect = 0.023
Identities = 21/73 (28%), Positives = 38/73 (51%), Gaps = 6/73 (8%)
Query: 43 KVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREEDGFNEHLKSMPWL 102
K+I L+F +W PC+A L +++E + N + F+S D +E+ L +
Sbjct: 58 KLIVLYFHTSWAEPCKA----LKQVFEAISNEPSNSNVSFLSIDADENSEISELFEIS-- 111
Query: 103 AVPFDVNLHRRLI 115
AVP+ + +H+ I
Sbjct: 112 AVPYFIIIHKGTI 124
>THI2_DROYA (Q6XHI1) Thioredoxin 2
Length = 106
Score = 37.4 bits (85), Expect = 0.066
Identities = 21/65 (32%), Positives = 33/65 (50%), Gaps = 4/65 (6%)
Query: 37 LSECDGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREED-GFNEH 95
L++ GK++ L F A WC PC+ P+L EL + + ++ V D ED +
Sbjct: 15 LTKASGKLVVLDFFATWCGPCKMISPKLAEL---STQYADTVVVLKVDVDECEDIAMEYN 71
Query: 96 LKSMP 100
+ SMP
Sbjct: 72 ISSMP 76
>TLPA_BRAJA (P43221) Thiol:disulfide interchange protein tlpA
(Cytochrome c biogenesis protein tlpA)
Length = 221
Score = 36.6 bits (83), Expect = 0.11
Identities = 19/53 (35%), Positives = 28/53 (51%), Gaps = 2/53 (3%)
Query: 37 LSECDGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREE 89
LS+ GK + + A WC PCR +P L EL L G N E++ ++ D +
Sbjct: 90 LSDFRGKTLLVNLWATWCVPCRKEMPALDELQGKL--SGPNFEVVAINIDTRD 140
Score = 30.8 bits (68), Expect = 6.2
Identities = 29/109 (26%), Positives = 45/109 (40%), Gaps = 8/109 (7%)
Query: 191 DDRKVPVSELAGKTIGLYFCAYWSPPSRAFTAQLTDVYNNLKATKGNCFEIVLISTD-RD 249
D + +S+ GKT+ + A W P R L ++ L G FE+V I+ D RD
Sbjct: 84 DGKPKKLSDFRGKTLLVNLWATWCVPCRKEMPALDELQGKL---SGPNFEVVAINIDTRD 140
Query: 250 HKEFNVNRSSTPWLAIPY----EDRTRHDLCRIYDIKGIPALVLIGPDG 294
++ + Y + + DL I G+P VL+ P G
Sbjct: 141 PEKPKTFLKEANLTRLGYFNDQKAKVFQDLKAIGRALGMPTSVLVDPQG 189
>THIT_DROME (Q8IFW4) Thioredoxin T (ThioredoxinT)
Length = 157
Score = 36.2 bits (82), Expect = 0.15
Identities = 20/61 (32%), Positives = 33/61 (53%), Gaps = 4/61 (6%)
Query: 41 DGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREEDGFNEH-LKSM 99
+ K++ + F A+WC PC+ P+L EL R + ++ V+ D ED E+ + SM
Sbjct: 19 EDKLVVIDFYADWCGPCKIIAPKLDELAHEYSDRVV---VLKVNVDENEDITVEYNVNSM 75
Query: 100 P 100
P
Sbjct: 76 P 76
>THI1_CAEEL (Q09433) Probable thioredoxin B0228.5
Length = 115
Score = 36.2 bits (82), Expect = 0.15
Identities = 31/104 (29%), Positives = 41/104 (38%), Gaps = 29/104 (27%)
Query: 43 KVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREEDGFNEHLKSMPWL 102
K+I L F A WC PC+A P EL T + IIF D +E
Sbjct: 28 KIIILDFYATWCGPCKAIAPLYKELATT------HKGIIFCKVDVDE------------- 68
Query: 103 AVPFDVNLHRRLIDRYQVDQIPSFIPLYSEDLIVEKNLIECIED 146
L +Y V +P+FI + D I + L C+ED
Sbjct: 69 --------AEDLCSKYDVKMMPTFIFTKNGDAI--EALEGCVED 102
>THIO_GEOCY (O96952) Thioredoxin
Length = 106
Score = 35.8 bits (81), Expect = 0.19
Identities = 18/67 (26%), Positives = 34/67 (49%), Gaps = 9/67 (13%)
Query: 37 LSECDGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFD---REEDGFN 93
L + K++ + F+A+WC PC+ P+ VE+ + ++IF D +E
Sbjct: 15 LKDAGDKLVVIDFTASWCGPCQRIAPKYVEMAKEFP------DVIFYKVDVDENDETAEA 68
Query: 94 EHLKSMP 100
E +++MP
Sbjct: 69 EKIQAMP 75
>THIH_BRARA (O64432) Thioredoxin H-type (TRX-H)
Length = 123
Score = 35.4 bits (80), Expect = 0.25
Identities = 18/56 (32%), Positives = 30/56 (53%), Gaps = 6/56 (10%)
Query: 34 KVPLSECDGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREE 89
K+ ++ K+I + F+A WC PCR P VEL + +L+++F D +E
Sbjct: 25 KLKAAKESNKLIVIDFTAVWCPPCRFIAPIFVELAKK------HLDVVFFKVDVDE 74
>THIH_BRAOL (P68176) Thioredoxin H-type (TRX-H) (Pollen coat
protein)
Length = 123
Score = 35.4 bits (80), Expect = 0.25
Identities = 18/56 (32%), Positives = 30/56 (53%), Gaps = 6/56 (10%)
Query: 34 KVPLSECDGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREE 89
K+ ++ K+I + F+A WC PCR P VEL + +L+++F D +E
Sbjct: 25 KLKAAKESNKLIVIDFTAVWCPPCRFIAPIFVELAKK------HLDVVFFKVDVDE 74
>THH1_BRANA (P68177) Thioredoxin H-type 1 (TRX-H-1)
Length = 123
Score = 35.4 bits (80), Expect = 0.25
Identities = 18/56 (32%), Positives = 30/56 (53%), Gaps = 6/56 (10%)
Query: 34 KVPLSECDGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREE 89
K+ ++ K+I + F+A WC PCR P VEL + +L+++F D +E
Sbjct: 25 KLKAAKESNKLIVIDFTAVWCPPCRFIAPIFVELAKK------HLDVVFFKVDVDE 74
>RPOC_CAUCR (Q9AAU1) DNA-directed RNA polymerase beta' chain (EC
2.7.7.6) (RNAP beta' subunit) (Transcriptase beta'
chain) (RNA polymerase beta' subunit)
Length = 1396
Score = 35.4 bits (80), Expect = 0.25
Identities = 37/132 (28%), Positives = 55/132 (41%), Gaps = 21/132 (15%)
Query: 68 YETLRKRGI-----NLEIIFVSFDREEDGFNE---------HLKSMPW-LAVPFDV---N 109
Y+ ++ +GI +E+ RE G E LKS+P +A+ D+ +
Sbjct: 76 YKRMKYKGIICEKCGVEVTLARVRRERMGHIELASPVAHIWFLKSLPSRIAMMLDMPLKD 135
Query: 110 LHRRLIDRYQVDQIPSFIPLYSEDLIVEKNLIECIEDYGADAFPF---TRKRHEELKAID 166
+ R L Y + P PL L+ E + + E+YG D+F LKAID
Sbjct: 136 IERVLYFEYYIVTEPGLTPLKQHQLLSEDDYMRAQEEYGDDSFTAEIGAEAIQNLLKAID 195
Query: 167 KRKREEANLEEL 178
K E EEL
Sbjct: 196 LEKEAERLREEL 207
>THIH_FAGES (Q96419) Thioredoxin H-type (TRX-H)
Length = 116
Score = 35.0 bits (79), Expect = 0.33
Identities = 24/89 (26%), Positives = 38/89 (41%), Gaps = 29/89 (32%)
Query: 42 GKVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREEDGFNEHLKSMPW 101
GK+I + F+A+WC PCR P + EL K P
Sbjct: 27 GKLIVIDFTASWCGPCRVITPYVSEL----------------------------AKKFPH 58
Query: 102 LA-VPFDVNLHRRLIDRYQVDQIPSFIPL 129
+A DV+ + + + Y+V+ +PSF+ L
Sbjct: 59 VAFFKVDVDDLKDVAEEYKVEAMPSFVIL 87
>THIO_ECHGR (O17486) Thioredoxin
Length = 107
Score = 33.9 bits (76), Expect = 0.73
Identities = 18/49 (36%), Positives = 28/49 (56%), Gaps = 7/49 (14%)
Query: 41 DGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREE 89
D ++C FF A WC PC++ P+L + + K N ++IFV D +E
Sbjct: 22 DKLLVCDFF-ATWCGPCKSLAPKL----DAMAKE--NEKVIFVKLDVDE 63
>THIO_PENCH (P34723) Thioredoxin
Length = 106
Score = 33.5 bits (75), Expect = 0.96
Identities = 16/39 (41%), Positives = 23/39 (58%), Gaps = 1/39 (2%)
Query: 32 EGKVPLSECDGKVICLFFSANWCRPCRAFVPRLVELYET 70
E K +++ G V+ + F A WC PC+A P L +L ET
Sbjct: 11 EYKEKVTDATGPVV-VDFHATWCGPCKAIAPALEKLSET 48
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.140 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,316,956
Number of Sequences: 164201
Number of extensions: 2123955
Number of successful extensions: 5927
Number of sequences better than 10.0: 76
Number of HSP's better than 10.0 without gapping: 34
Number of HSP's successfully gapped in prelim test: 42
Number of HSP's that attempted gapping in prelim test: 5851
Number of HSP's gapped (non-prelim): 114
length of query: 392
length of database: 59,974,054
effective HSP length: 112
effective length of query: 280
effective length of database: 41,583,542
effective search space: 11643391760
effective search space used: 11643391760
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0372a.5


