

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0370.4
(170 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
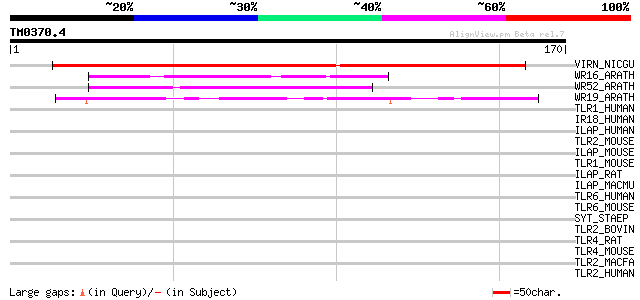
Score E
Sequences producing significant alignments: (bits) Value
VIRN_NICGU (Q40392) TMV resistance protein N 152 4e-37
WR16_ARATH (Q9FL92) Probable WRKY transcription factor 16 (WRKY ... 50 3e-06
WR52_ARATH (Q9FH83) Probable WRKY transcription factor 52 (WRKY ... 49 7e-06
WR19_ARATH (Q9SZ67) Probable WRKY transcription factor 19 (WRKY ... 44 2e-04
TLR1_HUMAN (Q15399) Toll-like receptor 1 precursor (Toll/interle... 37 0.017
IR18_HUMAN (Q13478) Interleukin-18 receptor 1 precursor (IL1 rec... 36 0.049
ILAP_HUMAN (Q9NPH3) Interleukin-1 receptor accessory protein pre... 34 0.14
TLR2_MOUSE (Q9QUN7) Toll-like receptor 2 precursor 33 0.24
ILAP_MOUSE (Q61730) Interleukin-1 receptor accessory protein pre... 33 0.32
TLR1_MOUSE (Q9EPQ1) Toll-like receptor 1 precursor (Toll/interle... 33 0.41
ILAP_RAT (Q63621) Interleukin-1 receptor accessory protein precu... 33 0.41
ILAP_MACMU (P59822) Interleukin-1 receptor accessory protein pre... 33 0.41
TLR6_HUMAN (Q9Y2C9) Toll-like receptor 6 precursor 32 0.54
TLR6_MOUSE (Q9EPW9) Toll-like receptor 6 precursor 32 0.70
SYT_STAEP (Q8CS74) Threonyl-tRNA synthetase (EC 6.1.1.3) (Threon... 32 0.70
TLR2_BOVIN (Q95LA9) Toll-like receptor 2 precursor 31 1.2
TLR4_RAT (Q9QX05) Toll-like receptor 4 precursor (Toll4) 30 2.1
TLR4_MOUSE (Q9QUK6) Toll-like receptor 4 precursor 30 2.1
TLR2_MACFA (Q95M53) Toll-like receptor 2 precursor 30 2.1
TLR2_HUMAN (O60603) Toll-like receptor 2 precursor (Toll/interle... 30 2.1
>VIRN_NICGU (Q40392) TMV resistance protein N
Length = 1144
Score = 152 bits (384), Expect = 4e-37
Identities = 78/145 (53%), Positives = 99/145 (67%), Gaps = 1/145 (0%)
Query: 14 FNYDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVL 73
++YDVF+SFRGEDTR TF L++ L KGI+TF DD+ L+ G I L KAIEES
Sbjct: 10 WSYDVFLSFRGEDTRKTFTSHLYEVLNDKGIKTFQDDKRLEYGATIPGELCKAIEESQFA 69
Query: 74 IIVLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHE 133
I+V S+NYA+S WCL+ELVKI+EC R +Q V PIFY V+PSHVR+QK + +A HE
Sbjct: 70 IVVFSENYATSRWCLNELVKIMECKTR-FKQTVIPIFYDVDPSHVRNQKESFAKAFEEHE 128
Query: 134 KKYGINSDRIRTWRSTLNEVCQLVG 158
KY + + I+ WR LNE L G
Sbjct: 129 TKYKDDVEGIQRWRIALNEAANLKG 153
>WR16_ARATH (Q9FL92) Probable WRKY transcription factor 16 (WRKY
DNA-binding protein 16)
Length = 1372
Score = 50.1 bits (118), Expect = 3e-06
Identities = 31/92 (33%), Positives = 52/92 (55%), Gaps = 8/92 (8%)
Query: 25 EDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVLIIVLSKNYASS 84
E+ R++FV L K L+RKG V+D + + + +S +E + V +++L N
Sbjct: 14 EEVRYSFVSHLSKALQRKG----VNDVFIDSDDSLSNESQSMVERARVSVMILPGN---R 66
Query: 85 TWCLDELVKILECSRRGNQQLVYPIFYHVEPS 116
T LD+LVK+L+C ++ Q+V P+ Y V S
Sbjct: 67 TVSLDKLVKVLDC-QKNKDQVVVPVLYGVRSS 97
>WR52_ARATH (Q9FH83) Probable WRKY transcription factor 52 (WRKY
DNA-binding protein 52) (Disease resistance protein
RRS1) (Resistance to Ralstonia solanacearum 1 protein)
Length = 1378
Score = 48.5 bits (114), Expect = 7e-06
Identities = 29/87 (33%), Positives = 45/87 (51%), Gaps = 2/87 (2%)
Query: 25 EDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVLIIVLSKNYASS 84
E+ R++FV L + L+RKGI V D + +++ IE++ V ++VL N S
Sbjct: 17 EEVRYSFVSHLSEALRRKGINNVVVD--VDIDDLLFKESQAKIEKAGVSVMVLPGNCDPS 74
Query: 85 TWCLDELVKILECSRRGNQQLVYPIFY 111
LD+ K+LEC R Q V + Y
Sbjct: 75 EVWLDKFAKVLECQRNNKDQAVVSVLY 101
>WR19_ARATH (Q9SZ67) Probable WRKY transcription factor 19 (WRKY
DNA-binding protein 19)
Length = 1895
Score = 43.5 bits (101), Expect = 2e-04
Identities = 43/150 (28%), Positives = 69/150 (45%), Gaps = 29/150 (19%)
Query: 15 NYDVFISF-RGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVL 73
+YDV I + R + + F+ L L R+GI + K V A+ + VL
Sbjct: 667 DYDVVIRYGRADISNEDFISHLRASLCRRGISVYE-----KFNEV------DALPKCRVL 715
Query: 74 IIVLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEP-SHVRHQKGKYGEAMAAH 132
IIVL+ Y S L+ ILE + ++VYPIFY + P V + K +
Sbjct: 716 IIVLTSTYVPSN-----LLNILE-HQHTEDRVVYPIFYRLSPYDFVCNSKN--------Y 761
Query: 133 EKKYGINSDRIRTWRSTLNEVCQLVGHHIS 162
E+ Y D + W++ L E+ Q+ G+ ++
Sbjct: 762 ERFY--LQDEPKKWQAALKEITQMPGYTLT 789
>TLR1_HUMAN (Q15399) Toll-like receptor 1 precursor
(Toll/interleukin-1 receptor-like) (TIL)
Length = 786
Score = 37.4 bits (85), Expect = 0.017
Identities = 23/76 (30%), Positives = 38/76 (49%), Gaps = 1/76 (1%)
Query: 16 YDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVLII 75
+ FIS+ G D+ ++L L+++G++ + + G I + IE+S I
Sbjct: 637 FHAFISYSGHDSFWVKNELL-PNLEKEGMQICLHERNFVPGKSIVENIITCIEKSYKSIF 695
Query: 76 VLSKNYASSTWCLDEL 91
VLS N+ S WC EL
Sbjct: 696 VLSPNFVQSEWCHYEL 711
>IR18_HUMAN (Q13478) Interleukin-18 receptor 1 precursor (IL1
receptor-related protein) (IL-1Rrp)
Length = 541
Score = 35.8 bits (81), Expect = 0.049
Identities = 26/76 (34%), Positives = 41/76 (53%), Gaps = 7/76 (9%)
Query: 16 YDVFISFRGE-----DTRHTF-VKILHKELKRK-GIRTFVDDEMLKAGNVISPALSKAIE 68
YD F+S+ E HTF V+IL + L++ G + + + + G + + IE
Sbjct: 375 YDAFVSYLKECRPENGEEHTFAVEILPRVLEKHFGYKLCIFERDVVPGGAVVDEIHSLIE 434
Query: 69 ESMVLIIVLSKNYASS 84
+S LIIVLSK+Y S+
Sbjct: 435 KSRRLIIVLSKSYMSN 450
>ILAP_HUMAN (Q9NPH3) Interleukin-1 receptor accessory protein
precursor (IL-1 receptor accessory protein) (IL-1RAcP)
Length = 570
Score = 34.3 bits (77), Expect = 0.14
Identities = 31/120 (25%), Positives = 55/120 (45%), Gaps = 9/120 (7%)
Query: 7 DGEELHIFNYDVFISFRGEDTRHTFVKILHKELKRK--GIRTFVDDEMLKAGNVISPALS 64
DG+E YD+++S+ FV + + + G + + D G +++
Sbjct: 401 DGKE-----YDIYVSYARNAEEEEFVLLTLRGVLENEFGYKLCIFDRDSLPGGIVTDETL 455
Query: 65 KAIEESMVLIIVLSKNYA-SSTWCLDELVKILE-CSRRGNQQLVYPIFYHVEPSHVRHQK 122
I++S L++VLS NY T L EL LE + RGN ++ + V+ + V+ K
Sbjct: 456 SFIQKSRRLLVVLSPNYVLQGTQALLELKAGLENMASRGNINVILVQYKAVKETKVKELK 515
>TLR2_MOUSE (Q9QUN7) Toll-like receptor 2 precursor
Length = 784
Score = 33.5 bits (75), Expect = 0.24
Identities = 21/78 (26%), Positives = 38/78 (47%), Gaps = 3/78 (3%)
Query: 16 YDVFISFRGEDTRHTFVKILHKELKRKG--IRTFVDDEMLKAGNVISPALSKAIEESMVL 73
YD F+S+ +D+ H ++ ++L+ + + G I + +IE+S
Sbjct: 641 YDAFVSYSEQDS-HWVENLMVQQLENSDPPFKLCLHKRDFVPGKWIIDNIIDSIEKSHKT 699
Query: 74 IIVLSKNYASSTWCLDEL 91
+ VLS+N+ S WC EL
Sbjct: 700 VFVLSENFVRSEWCKYEL 717
>ILAP_MOUSE (Q61730) Interleukin-1 receptor accessory protein
precursor (IL-1 receptor accessory protein) (IL-1RAcP)
Length = 570
Score = 33.1 bits (74), Expect = 0.32
Identities = 31/120 (25%), Positives = 54/120 (44%), Gaps = 9/120 (7%)
Query: 7 DGEELHIFNYDVFISFRGEDTRHTFVKILHKELKRK--GIRTFVDDEMLKAGNVISPALS 64
DG+E YD+++S+ FV + + + G + + D G +++
Sbjct: 401 DGKE-----YDIYVSYARNVEEEEFVLLTLRGVLENEFGYKLCIFDRDSLPGGIVTDETL 455
Query: 65 KAIEESMVLIIVLSKNYA-SSTWCLDELVKILE-CSRRGNQQLVYPIFYHVEPSHVRHQK 122
I++S L++VLS NY T L EL LE + RGN ++ + V+ V+ K
Sbjct: 456 SFIQKSRRLLVVLSPNYVLQGTQALLELKAGLENMASRGNINVILVQYKAVKDMKVKELK 515
>TLR1_MOUSE (Q9EPQ1) Toll-like receptor 1 precursor
(Toll/interleukin-1 receptor-like) (TIL)
Length = 795
Score = 32.7 bits (73), Expect = 0.41
Identities = 21/76 (27%), Positives = 36/76 (46%), Gaps = 1/76 (1%)
Query: 16 YDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVLII 75
+ F+S+ G D+ ++L L++ I+ + + G I + IE+S I
Sbjct: 640 FHAFVSYSGHDSAWVKNELL-PNLEKDDIQICLHERNFVPGKSIVENIINFIEKSYKSIF 698
Query: 76 VLSKNYASSTWCLDEL 91
VLS ++ S WC EL
Sbjct: 699 VLSPHFIQSEWCHYEL 714
>ILAP_RAT (Q63621) Interleukin-1 receptor accessory protein
precursor (IL-1 receptor accessory protein) (IL-1RAcP)
Length = 570
Score = 32.7 bits (73), Expect = 0.41
Identities = 31/120 (25%), Positives = 54/120 (44%), Gaps = 9/120 (7%)
Query: 7 DGEELHIFNYDVFISFRGEDTRHTFVKILHKELKRK--GIRTFVDDEMLKAGNVISPALS 64
DG+E YD+++S+ FV + + + G + + D G +++
Sbjct: 401 DGKE-----YDIYVSYARNAEEEEFVLLTLRGVLENEFGYKLCIFDRDSFPGGIVTDETL 455
Query: 65 KAIEESMVLIIVLSKNYA-SSTWCLDELVKILE-CSRRGNQQLVYPIFYHVEPSHVRHQK 122
I++S L++VLS NY T L EL LE + RGN ++ + V+ V+ K
Sbjct: 456 SFIQKSRRLLVVLSPNYVLQGTQALLELKAGLENMASRGNINVILVQYKAVKDLKVKELK 515
>ILAP_MACMU (P59822) Interleukin-1 receptor accessory protein
precursor (IL-1 receptor accessory protein) (IL-1RAcP)
Length = 570
Score = 32.7 bits (73), Expect = 0.41
Identities = 30/120 (25%), Positives = 55/120 (45%), Gaps = 9/120 (7%)
Query: 7 DGEELHIFNYDVFISFRGEDTRHTFVKILHKELKRK--GIRTFVDDEMLKAGNVISPALS 64
DG+E YD+++S+ FV + + + G + + D G +++
Sbjct: 401 DGKE-----YDIYVSYARNAEEEEFVLLTLRGVLENEFGYKLCIFDRDSLPGGIVTDETL 455
Query: 65 KAIEESMVLIIVLSKNYA-SSTWCLDELVKILE-CSRRGNQQLVYPIFYHVEPSHVRHQK 122
I++S L++VLS NY T L EL LE + +GN ++ + V+ + V+ K
Sbjct: 456 SFIQKSRRLLVVLSPNYVLQGTQALLELKAGLENMASQGNINVILVQYKAVKETKVKELK 515
>TLR6_HUMAN (Q9Y2C9) Toll-like receptor 6 precursor
Length = 796
Score = 32.3 bits (72), Expect = 0.54
Identities = 21/76 (27%), Positives = 36/76 (46%), Gaps = 1/76 (1%)
Query: 16 YDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVLII 75
+ FIS+ D+ +++ L+++ I+ + + G I + IE+S I
Sbjct: 642 FHAFISYSEHDSAWVKSELV-PYLEKEDIQICLHERNFVPGKSIVENIINCIEKSYKSIF 700
Query: 76 VLSKNYASSTWCLDEL 91
VLS N+ S WC EL
Sbjct: 701 VLSPNFVQSEWCHYEL 716
>TLR6_MOUSE (Q9EPW9) Toll-like receptor 6 precursor
Length = 795
Score = 32.0 bits (71), Expect = 0.70
Identities = 21/76 (27%), Positives = 35/76 (45%), Gaps = 1/76 (1%)
Query: 16 YDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVLII 75
+ F+S+ D+ ++L L++ IR + + G I + IE+S I
Sbjct: 642 FHAFVSYSEHDSAWVKNELL-PNLEKDDIRVCLHERNFVPGKSIVENIINFIEKSYKAIF 700
Query: 76 VLSKNYASSTWCLDEL 91
VLS ++ S WC EL
Sbjct: 701 VLSPHFIQSEWCHYEL 716
>SYT_STAEP (Q8CS74) Threonyl-tRNA synthetase (EC 6.1.1.3)
(Threonine--tRNA ligase) (ThrRS)
Length = 645
Score = 32.0 bits (71), Expect = 0.70
Identities = 16/48 (33%), Positives = 25/48 (51%)
Query: 26 DTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVL 73
D + + ++L ELK +G+R +DD K G I A K I +V+
Sbjct: 556 DLHYDYARLLQDELKSQGVRVEIDDRNEKMGYKIREAQMKKIPYQIVV 603
>TLR2_BOVIN (Q95LA9) Toll-like receptor 2 precursor
Length = 784
Score = 31.2 bits (69), Expect = 1.2
Identities = 22/78 (28%), Positives = 37/78 (47%), Gaps = 3/78 (3%)
Query: 16 YDVFISFRGEDTRHTFVKILHKELKRKG--IRTFVDDEMLKAGNVISPALSKAIEESMVL 73
YD F+S+ D+ + ++ +EL+ + + G I + +IE+S
Sbjct: 641 YDAFVSYSERDS-YWVENLMVQELEHFNPPFKLCLHKRDFIPGKWIIDNIIDSIEKSHKT 699
Query: 74 IIVLSKNYASSTWCLDEL 91
I VLS+N+ S WC EL
Sbjct: 700 IFVLSENFVKSEWCKYEL 717
>TLR4_RAT (Q9QX05) Toll-like receptor 4 precursor (Toll4)
Length = 835
Score = 30.4 bits (67), Expect = 2.1
Identities = 32/122 (26%), Positives = 57/122 (46%), Gaps = 21/122 (17%)
Query: 16 YDVFISFRGED---TRHTFVKILHKELKRKGI----RTFVDDEMLKAGNVISPALSKAIE 68
YD F+ + ++ R+ VK L + + R + R F+ + A N+I K
Sbjct: 672 YDAFVIYSSQNEDWVRNELVKNLEEGVPRFQLCLHYRDFIPGVAI-AANIIQEGFHK--- 727
Query: 69 ESMVLIIVLSKNYASSTWCLDELVKILE-----CSRRGNQQLVYPIFYHVEPSHVRHQKG 123
S +I+V+S+++ S WC+ E +I + SR G +++ + VE S +R Q
Sbjct: 728 -SRKVIVVVSRHFIQSRWCIFE-YEIAQTWQFLSSRSG---IIFIVLEKVEKSLLRQQVE 782
Query: 124 KY 125
Y
Sbjct: 783 LY 784
>TLR4_MOUSE (Q9QUK6) Toll-like receptor 4 precursor
Length = 835
Score = 30.4 bits (67), Expect = 2.1
Identities = 32/122 (26%), Positives = 57/122 (46%), Gaps = 21/122 (17%)
Query: 16 YDVFISFRGED---TRHTFVKILHKELKRKGI----RTFVDDEMLKAGNVISPALSKAIE 68
YD F+ + ++ R+ VK L + + R + R F+ + A N+I K
Sbjct: 672 YDAFVIYSSQNEDWVRNELVKNLEEGVPRFHLCLHYRDFIPGVAI-AANIIQEGFHK--- 727
Query: 69 ESMVLIIVLSKNYASSTWCLDELVKILE-----CSRRGNQQLVYPIFYHVEPSHVRHQKG 123
S +I+V+S+++ S WC+ E +I + SR G +++ + VE S +R Q
Sbjct: 728 -SRKVIVVVSRHFIQSRWCIFE-YEIAQTWQFLSSRSG---IIFIVLEKVEKSLLRQQVE 782
Query: 124 KY 125
Y
Sbjct: 783 LY 784
>TLR2_MACFA (Q95M53) Toll-like receptor 2 precursor
Length = 784
Score = 30.4 bits (67), Expect = 2.1
Identities = 21/78 (26%), Positives = 36/78 (45%), Gaps = 3/78 (3%)
Query: 16 YDVFISFRGEDTRHTFVKILHKELKRKG--IRTFVDDEMLKAGNVISPALSKAIEESMVL 73
YD F+S+ D + ++ +EL+ + + G I + +IE+S
Sbjct: 641 YDAFVSYSERDA-YWVENLMVQELENFNPPFKLCLHKRDFIPGKWIIDNIIDSIEKSHKT 699
Query: 74 IIVLSKNYASSTWCLDEL 91
+ VLS+N+ S WC EL
Sbjct: 700 VFVLSENFVKSEWCKYEL 717
>TLR2_HUMAN (O60603) Toll-like receptor 2 precursor
(Toll/interleukin 1 receptor-like protein 4)
Length = 784
Score = 30.4 bits (67), Expect = 2.1
Identities = 21/78 (26%), Positives = 36/78 (45%), Gaps = 3/78 (3%)
Query: 16 YDVFISFRGEDTRHTFVKILHKELKRKG--IRTFVDDEMLKAGNVISPALSKAIEESMVL 73
YD F+S+ D + ++ +EL+ + + G I + +IE+S
Sbjct: 641 YDAFVSYSERDA-YWVENLMVQELENFNPPFKLCLHKRDFIPGKWIIDNIIDSIEKSHKT 699
Query: 74 IIVLSKNYASSTWCLDEL 91
+ VLS+N+ S WC EL
Sbjct: 700 VFVLSENFVKSEWCKYEL 717
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.326 0.140 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,175,089
Number of Sequences: 164201
Number of extensions: 729591
Number of successful extensions: 2318
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 15
Number of HSP's successfully gapped in prelim test: 31
Number of HSP's that attempted gapping in prelim test: 2300
Number of HSP's gapped (non-prelim): 47
length of query: 170
length of database: 59,974,054
effective HSP length: 102
effective length of query: 68
effective length of database: 43,225,552
effective search space: 2939337536
effective search space used: 2939337536
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0370.4


