

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0367.7
(172 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
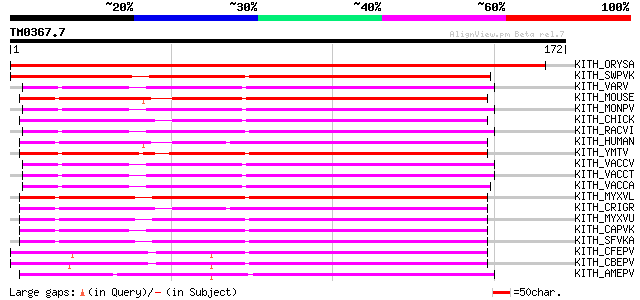
KITH_ORYSA (O81263) Thymidine kinase (EC 2.7.1.21) 270 1e-72
KITH_SWPVK (P23335) Thymidine kinase (EC 2.7.1.21) 131 7e-31
KITH_VARV (P04364) Thymidine kinase (EC 2.7.1.21) 117 1e-26
KITH_MOUSE (P04184) Thymidine kinase, cytosolic (EC 2.7.1.21) 117 1e-26
KITH_MONPV (P04363) Thymidine kinase (EC 2.7.1.21) 115 4e-26
KITH_CHICK (P04047) Thymidine kinase, cytosolic (EC 2.7.1.21) 115 5e-26
KITH_RACVI (Q90029) Thymidine kinase (EC 2.7.1.21) 115 7e-26
KITH_HUMAN (P04183) Thymidine kinase, cytosolic (EC 2.7.1.21) 115 7e-26
KITH_YMTV (Q90033) Thymidine kinase (EC 2.7.1.21) 114 1e-25
KITH_VACCV (P03297) Thymidine kinase (EC 2.7.1.21) 114 1e-25
KITH_VACCT (Q9JFB7) Thymidine kinase (EC 2.7.1.21) 114 1e-25
KITH_VACCA (O57203) Thymidine kinase (EC 2.7.1.21) 114 1e-25
KITH_MYXVL (Q9Q8N6) Thymidine kinase (EC 2.7.1.21) 113 2e-25
KITH_CRIGR (P09768) Thymidine kinase, cytosolic (EC 2.7.1.21) 113 2e-25
KITH_MYXVU (P28851) Thymidine kinase (EC 2.7.1.21) 111 7e-25
KITH_CAPVK (P16600) Thymidine kinase (EC 2.7.1.21) 107 2e-23
KITH_SFVKA (P07605) Thymidine kinase (EC 2.7.1.21) 105 4e-23
KITH_CFEPV (Q05880) Thymidine kinase (EC 2.7.1.21) 99 6e-21
KITH_CBEPV (Q05879) Thymidine kinase (EC 2.7.1.21) 96 5e-20
KITH_AMEPV (P28852) Thymidine kinase (EC 2.7.1.21) 94 1e-19
>KITH_ORYSA (O81263) Thymidine kinase (EC 2.7.1.21)
Length = 212
Score = 270 bits (690), Expect = 1e-72
Identities = 125/166 (75%), Positives = 147/166 (88%)
Query: 1 NVAIIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFF 60
NVA+IKS KD RYGLDS+VTHDGTK+PCWAL LSSF+ K G +AY+++DVIGIDEAQFF
Sbjct: 39 NVALIKSDKDNRYGLDSVVTHDGTKMPCWALPELSSFQDKLGTEAYDKVDVIGIDEAQFF 98
Query: 61 DDLYDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAF 120
DDL+DFC +AAD DGK VVVAGLDG+Y R FGSVLDIIPLADSVTKLTARCE+CG+ AF
Sbjct: 99 DDLHDFCCKAADRDGKIVVVAGLDGDYKRNKFGSVLDIIPLADSVTKLTARCELCGRRAF 158
Query: 121 FTLRKTQETQVELIGGVDVYMPVCRQHYVNGQVAMETTRLVLESQK 166
FTLRKT+ET+ ELIGG DVYMPVCRQHY++GQ+ +E TR+VL+ +K
Sbjct: 159 FTLRKTRETKTELIGGADVYMPVCRQHYLDGQIVIEATRIVLDLEK 204
>KITH_SWPVK (P23335) Thymidine kinase (EC 2.7.1.21)
Length = 177
Score = 131 bits (330), Expect = 7e-31
Identities = 66/149 (44%), Positives = 96/149 (64%), Gaps = 6/149 (4%)
Query: 1 NVAIIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFF 60
N +IK SKD RYG D++ THD + + +L K D + +D++GIDE QFF
Sbjct: 34 NCRVIKYSKDNRYGNDAVYTHDKCYISAVSTDSLFDIK-----DTLDDVDIVGIDEGQFF 88
Query: 61 DDLYDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAF 120
+D+ +FC A+ GK V+VA LDG Y R+ FG++L++IPL++ VTKL A C IC + A
Sbjct: 89 NDIVEFCEYIANK-GKIVIVAALDGTYERKPFGNILNLIPLSEKVTKLNAICMICHRDAS 147
Query: 121 FTLRKTQETQVELIGGVDVYMPVCRQHYV 149
F+ R + E ++ELIGG + Y+ VCR Y+
Sbjct: 148 FSKRLSDEKEIELIGGKEKYLSVCRSCYL 176
>KITH_VARV (P04364) Thymidine kinase (EC 2.7.1.21)
Length = 177
Score = 117 bits (294), Expect = 1e-26
Identities = 63/146 (43%), Positives = 87/146 (59%), Gaps = 7/146 (4%)
Query: 5 IKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFDDLY 64
IK S D RYG + THD L ++A VIGIDE QFF D+
Sbjct: 38 IKYSNDNRYGT-GLWTHDKNNFEALEATKLCDV-----LEAITDFSVIGIDEGQFFPDIV 91
Query: 65 DFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFTLR 124
+FC A+ +GK V+VA LDG + R+ F ++LD+IPL++ V KLTA C C K A F+ R
Sbjct: 92 EFCERMAN-EGKIVIVAALDGTFQRKPFNNILDLIPLSEMVVKLTAVCMKCFKEASFSKR 150
Query: 125 KTQETQVELIGGVDVYMPVCRQHYVN 150
ET++E+IGG+D+Y VCR+ Y++
Sbjct: 151 LGTETKIEIIGGIDMYQSVCRKCYID 176
>KITH_MOUSE (P04184) Thymidine kinase, cytosolic (EC 2.7.1.21)
Length = 233
Score = 117 bits (293), Expect = 1e-26
Identities = 69/146 (47%), Positives = 89/146 (60%), Gaps = 9/146 (6%)
Query: 4 IIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQK-FGVDAYEQLDVIGIDEAQFFDD 62
+IK +KDTRY +S THD + L Q+ GV VIGIDE QFF D
Sbjct: 52 VIKYAKDTRYS-NSFSTHDRNTMDALPACMLRDVTQEALGVA------VIGIDEGQFFPD 104
Query: 63 LYDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFT 122
+ DFC A+ +GKTV+VA LDG + R+ FGS+L+++PLA+SV KLTA C C + A +T
Sbjct: 105 IVDFCEMMAN-EGKTVIVAALDGTFQRKAFGSILNLVPLAESVVKLTAVCMECFREAAYT 163
Query: 123 LRKTQETQVELIGGVDVYMPVCRQHY 148
R E +VE+IGG D Y VCR Y
Sbjct: 164 KRLGLEKEVEVIGGADKYHSVCRLCY 189
>KITH_MONPV (P04363) Thymidine kinase (EC 2.7.1.21)
Length = 177
Score = 115 bits (289), Expect = 4e-26
Identities = 63/146 (43%), Positives = 87/146 (59%), Gaps = 7/146 (4%)
Query: 5 IKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFDDLY 64
IK S D RYG + THD + L ++A VIGIDE QFF D+
Sbjct: 38 IKYSNDNRYGT-GLWTHDKNNFAALEVTKLCDV-----LEAITDFSVIGIDEGQFFPDIV 91
Query: 65 DFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFTLR 124
+FC A+ +GK V+VA LDG + RR F ++L++IPL++ V KLTA C C K A F+ R
Sbjct: 92 EFCERMAN-EGKIVIVAALDGTFQRRPFNNILNLIPLSEMVVKLTAVCMKCFKEASFSKR 150
Query: 125 KTQETQVELIGGVDVYMPVCRQHYVN 150
ET++E+IGG D+Y VCR+ Y++
Sbjct: 151 LGAETEIEIIGGNDMYQSVCRKCYID 176
>KITH_CHICK (P04047) Thymidine kinase, cytosolic (EC 2.7.1.21)
Length = 224
Score = 115 bits (288), Expect = 5e-26
Identities = 64/145 (44%), Positives = 86/145 (59%), Gaps = 6/145 (4%)
Query: 4 IIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFDDL 63
++K +KDTRY + THD + L Q+ A VIGIDE QFF D+
Sbjct: 52 LVKYAKDTRYCTTGVSTHDRNTMEARPACALQDVYQEALGSA-----VIGIDEGQFFPDI 106
Query: 64 YDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFTL 123
+FC + A+ GKTV+VA LDG + R+ FGS+L+++PLA+SV KL A C C + A +T
Sbjct: 107 VEFCEKMAN-TGKTVIVAALDGTFQRKAFGSILNLVPLAESVVKLNAVCMECYREASYTK 165
Query: 124 RKTQETQVELIGGVDVYMPVCRQHY 148
R E +VE+IGG D Y VCR Y
Sbjct: 166 RLGAEREVEVIGGADKYHSVCRACY 190
>KITH_RACVI (Q90029) Thymidine kinase (EC 2.7.1.21)
Length = 177
Score = 115 bits (287), Expect = 7e-26
Identities = 63/146 (43%), Positives = 87/146 (59%), Gaps = 7/146 (4%)
Query: 5 IKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFDDLY 64
IK + DTRYG + THD L + +DA VIGIDE QFF D+
Sbjct: 38 IKYTNDTRYGT-GLWTHDKHNFSAMETTKLLNI-----IDAVTDFSVIGIDEGQFFPDIV 91
Query: 65 DFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFTLR 124
+FC A++ GK V+VA LDG + R+ F ++ ++IPL++ V KLTA C C K A F+ R
Sbjct: 92 EFCEYMANN-GKIVIVAALDGTFQRKPFTTISNLIPLSEMVVKLTAVCMKCFKEASFSKR 150
Query: 125 KTQETQVELIGGVDVYMPVCRQHYVN 150
ET++E+IGG D+Y VCR+ Y+N
Sbjct: 151 LGTETEIEIIGGEDMYQSVCRKCYIN 176
>KITH_HUMAN (P04183) Thymidine kinase, cytosolic (EC 2.7.1.21)
Length = 234
Score = 115 bits (287), Expect = 7e-26
Identities = 68/146 (46%), Positives = 88/146 (59%), Gaps = 9/146 (6%)
Query: 4 IIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQK-FGVDAYEQLDVIGIDEAQFFDD 62
+IK +KDTRY S THD + L Q+ GV VIGIDE QFF D
Sbjct: 52 VIKYAKDTRYS-SSFCTHDRNTMEALPACLLRDVAQEALGVA------VIGIDEGQFFPD 104
Query: 63 LYDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFT 122
+ +FC EA + GKTV+VA LDG + R+ FG++L+++PLA+SV KLTA C C + A +T
Sbjct: 105 IVEFC-EAMANAGKTVIVAALDGTFQRKPFGAILNLVPLAESVVKLTAVCMECFREAAYT 163
Query: 123 LRKTQETQVELIGGVDVYMPVCRQHY 148
R E +VE+IGG D Y VCR Y
Sbjct: 164 KRLGTEKEVEVIGGADKYHSVCRLCY 189
>KITH_YMTV (Q90033) Thymidine kinase (EC 2.7.1.21)
Length = 181
Score = 114 bits (285), Expect = 1e-25
Identities = 65/145 (44%), Positives = 92/145 (62%), Gaps = 7/145 (4%)
Query: 4 IIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFDDL 63
+IK SKD RYG +VTHD +P + +L+ ++A DVIGIDE QFF D+
Sbjct: 39 VIKYSKDERYGR-GLVTHDNDSVPAIPVNSLNEINCN-DINA----DVIGIDEGQFFPDI 92
Query: 64 YDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFTL 123
+FC A++ GK ++VA LDG +LR+ FG++L++IP A+ V KLTA C IC A F+
Sbjct: 93 VEFCDYMANN-GKILIVAALDGTFLRQPFGNILNLIPRAEYVLKLTAVCMICFGSASFSK 151
Query: 124 RKTQETQVELIGGVDVYMPVCRQHY 148
R + E ++E+IGG + Y VCR Y
Sbjct: 152 RISDEQEIEIIGGKEKYQSVCRVCY 176
>KITH_VACCV (P03297) Thymidine kinase (EC 2.7.1.21)
Length = 177
Score = 114 bits (284), Expect = 1e-25
Identities = 61/146 (41%), Positives = 87/146 (58%), Gaps = 7/146 (4%)
Query: 5 IKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFDDLY 64
IK S D RYG + THD L +++ VIGIDE QFF D+
Sbjct: 38 IKYSNDNRYGT-GLWTHDKNNFEALEATKLCDV-----LESITDFSVIGIDEGQFFPDIV 91
Query: 65 DFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFTLR 124
+FC A+ +GK V+VA LDG + R+ F ++L++IPL++ V KLTA C C K A F+ R
Sbjct: 92 EFCERMAN-EGKIVIVAALDGTFQRKPFNNILNLIPLSEMVVKLTAVCMKCFKEASFSKR 150
Query: 125 KTQETQVELIGGVDVYMPVCRQHYVN 150
+ET++E+IGG D+Y VCR+ Y++
Sbjct: 151 LGEETEIEIIGGNDMYQSVCRKCYID 176
>KITH_VACCT (Q9JFB7) Thymidine kinase (EC 2.7.1.21)
Length = 177
Score = 114 bits (284), Expect = 1e-25
Identities = 61/146 (41%), Positives = 87/146 (58%), Gaps = 7/146 (4%)
Query: 5 IKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFDDLY 64
IK S D RYG + THD L +++ VIGIDE QFF D+
Sbjct: 38 IKYSNDNRYGT-GLWTHDKNNFEALEATKLCDV-----LESITDFSVIGIDEGQFFPDIV 91
Query: 65 DFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFTLR 124
+FC A+ +GK V+VA LDG + R+ F ++L++IPL++ V KLTA C C K A F+ R
Sbjct: 92 EFCERMAN-EGKIVIVAALDGTFQRKPFNNILNLIPLSEMVVKLTAVCMKCFKEASFSKR 150
Query: 125 KTQETQVELIGGVDVYMPVCRQHYVN 150
+ET++E+IGG D+Y VCR+ Y++
Sbjct: 151 LGEETEIEIIGGNDMYQSVCRKCYID 176
>KITH_VACCA (O57203) Thymidine kinase (EC 2.7.1.21)
Length = 177
Score = 114 bits (284), Expect = 1e-25
Identities = 62/145 (42%), Positives = 86/145 (58%), Gaps = 7/145 (4%)
Query: 5 IKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFDDLY 64
IK S D RYG + THD L +++ VIGIDE QFF D+
Sbjct: 38 IKYSNDNRYGT-GLWTHDKNNFEALEATKLCDV-----LESITDFSVIGIDEGQFFPDIV 91
Query: 65 DFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFTLR 124
+FC A+ +GK V+VA LDG + R+ F ++L++IPL++ V KLTA C C K A F+ R
Sbjct: 92 EFCERMAN-EGKIVIVAALDGTFQRKPFNNILNLIPLSEMVVKLTAVCMKCFKEASFSKR 150
Query: 125 KTQETQVELIGGVDVYMPVCRQHYV 149
+ET++E+IGG D+Y VCR+ YV
Sbjct: 151 LGEETEIEIIGGNDMYQSVCRKCYV 175
>KITH_MYXVL (Q9Q8N6) Thymidine kinase (EC 2.7.1.21)
Length = 178
Score = 113 bits (283), Expect = 2e-25
Identities = 61/145 (42%), Positives = 88/145 (60%), Gaps = 7/145 (4%)
Query: 4 IIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFDDL 63
++K KD RYG + + THD T + A+L + E ++VIGIDE QFF ++
Sbjct: 39 VVKYEKDARYG-NGVRTHDNTCISAVPTASLDDVES-----ISEHVEVIGIDEGQFFPNI 92
Query: 64 YDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFTL 123
FC A+ GK ++VA LDG + R+ F ++ ++IPLA++VTKL A C C K A F+
Sbjct: 93 VSFCERMANA-GKVLIVAALDGTFQRKPFTNICELIPLAENVTKLNAVCMYCYKDASFSK 151
Query: 124 RKTQETQVELIGGVDVYMPVCRQHY 148
R ET++E+IGG D Y VCR+ Y
Sbjct: 152 RLGNETEIEIIGGSDKYKSVCRKCY 176
>KITH_CRIGR (P09768) Thymidine kinase, cytosolic (EC 2.7.1.21)
Length = 234
Score = 113 bits (282), Expect = 2e-25
Identities = 67/145 (46%), Positives = 86/145 (59%), Gaps = 7/145 (4%)
Query: 4 IIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFDDL 63
+IK +KDTRY S THD + L Q+ A VIGIDE QFF D+
Sbjct: 52 VIKYAKDTRYS-SSFSTHDRNTMDALPACLLRDVAQEALGAA-----VIGIDEGQFFPDI 105
Query: 64 YDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFTL 123
+FC E + GKTV+VA LDG + R+ FGS+L+++PLA+SV KLTA C C + A +T
Sbjct: 106 VEFC-EVMANAGKTVIVAALDGTFQRKAFGSILNLVPLAESVVKLTAVCMECFREAAYTK 164
Query: 124 RKTQETQVELIGGVDVYMPVCRQHY 148
R E +VE+IGG D Y VCR Y
Sbjct: 165 RLGLEKEVEVIGGADKYHSVCRVCY 189
>KITH_MYXVU (P28851) Thymidine kinase (EC 2.7.1.21)
Length = 178
Score = 111 bits (278), Expect = 7e-25
Identities = 60/145 (41%), Positives = 87/145 (59%), Gaps = 7/145 (4%)
Query: 4 IIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFDDL 63
++ KD RYG + + THD T + A+L + E ++VIGIDE QFF ++
Sbjct: 39 VVNYEKDARYG-NGVRTHDNTCISAVPTASLDDVES-----ISEHVEVIGIDEGQFFPNI 92
Query: 64 YDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFTL 123
FC A+ GK ++VA LDG + R+ F ++ ++IPLA++VTKL A C C K A F+
Sbjct: 93 VSFCERMANA-GKVLIVAALDGTFQRKPFTNICELIPLAENVTKLNAVCMYCYKDASFSK 151
Query: 124 RKTQETQVELIGGVDVYMPVCRQHY 148
R ET++E+IGG D Y VCR+ Y
Sbjct: 152 RLGNETEIEIIGGSDKYKSVCRKCY 176
>KITH_CAPVK (P16600) Thymidine kinase (EC 2.7.1.21)
Length = 177
Score = 107 bits (266), Expect = 2e-23
Identities = 59/145 (40%), Positives = 83/145 (56%), Gaps = 7/145 (4%)
Query: 4 IIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFDDL 63
++K KD RYG +S+ THD + + L VD D+IGIDE QFF D+
Sbjct: 37 VVKYLKDIRYG-NSVYTHDNNHVSAISTTLLYDV-----VDKIMNFDIIGIDEGQFFKDI 90
Query: 64 YDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFTL 123
F A+ GK +++A LD + R+ F +L +IPL++ VTKL A C C K A F+
Sbjct: 91 VSFSENMANM-GKIIIIAALDSTFQRKEFNDILKLIPLSEKVTKLNAVCMECYKDAAFSK 149
Query: 124 RKTQETQVELIGGVDVYMPVCRQHY 148
R T+E ++ELIGG + Y VCR+ Y
Sbjct: 150 RITKEKEIELIGGKEKYKSVCRKCY 174
>KITH_SFVKA (P07605) Thymidine kinase (EC 2.7.1.21)
Length = 176
Score = 105 bits (263), Expect = 4e-23
Identities = 56/145 (38%), Positives = 83/145 (56%), Gaps = 7/145 (4%)
Query: 4 IIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFDDL 63
++K KD RYG + + THD + +L E +VIGIDE QFF ++
Sbjct: 37 VVKYEKDIRYG-NGVCTHDNMSITAVCTPSLDKIDS-----VAENAEVIGIDEGQFFPNI 90
Query: 64 YDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFTL 123
FC A+ GK ++VA LDG + R+ F ++ ++IPLA++VTKL A C C K+ F+
Sbjct: 91 ATFCERMANR-GKVLIVAALDGTFQRKPFSNISELIPLAENVTKLNAVCMYCYKNGSFSK 149
Query: 124 RKTQETQVELIGGVDVYMPVCRQHY 148
R + ++E+IGG D Y VCR+ Y
Sbjct: 150 RLGDKMEIEVIGGSDKYKSVCRKCY 174
>KITH_CFEPV (Q05880) Thymidine kinase (EC 2.7.1.21)
Length = 185
Score = 98.6 bits (244), Expect = 6e-21
Identities = 56/151 (37%), Positives = 89/151 (58%), Gaps = 6/151 (3%)
Query: 1 NVAIIKSSKDTRYGLDSI-VTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQF 59
N II S D R+ D+I + HDG K+ + K + +++ ++IGIDE QF
Sbjct: 31 NCVIISHSIDNRHDDDNILINHDGFKISHDDFIKTNILINKIKI--FDKYEIIGIDECQF 88
Query: 60 FD--DLYDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGK 117
FD DL FC A++ GK ++VAGL+ ++ + F S++ +IP+++ +TKL + C C
Sbjct: 89 FDSNDLIIFCDTLANN-GKKIIVAGLNSDFNKNPFKSIIKLIPISEKITKLQSICNFCYN 147
Query: 118 HAFFTLRKTQETQVELIGGVDVYMPVCRQHY 148
A FT++K + + IGG D+Y+PVCR Y
Sbjct: 148 DATFTMKKFNKDIIIEIGGSDLYIPVCRICY 178
>KITH_CBEPV (Q05879) Thymidine kinase (EC 2.7.1.21)
Length = 186
Score = 95.5 bits (236), Expect = 5e-20
Identities = 55/152 (36%), Positives = 87/152 (57%), Gaps = 7/152 (4%)
Query: 1 NVAIIKSSKDTRYGLDS--IVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQ 58
N II S D R D ++ HDG K+ + K + +++ ++IGIDE Q
Sbjct: 31 NCVIISHSIDNRCEEDDNILINHDGFKISHDDFIKTNILINKIKI--FDKYEIIGIDECQ 88
Query: 59 FFD--DLYDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICG 116
FFD DL FC A++ GK ++VAGL+ ++ + F S++ +IP+++ +TKL + C C
Sbjct: 89 FFDSNDLMIFCDTLANN-GKKIIVAGLNSDFNKNPFKSIIKLIPISEKITKLQSICNFCY 147
Query: 117 KHAFFTLRKTQETQVELIGGVDVYMPVCRQHY 148
A FT++K + + IGG D+Y+PVCR Y
Sbjct: 148 NDATFTMKKFNKDIIIEIGGSDLYIPVCRICY 179
>KITH_AMEPV (P28852) Thymidine kinase (EC 2.7.1.21)
Length = 182
Score = 94.4 bits (233), Expect = 1e-19
Identities = 57/149 (38%), Positives = 86/149 (57%), Gaps = 4/149 (2%)
Query: 4 IIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFD-- 61
II + D R+ +I+ HDG L L + + ++ + D+IGIDE QFF+
Sbjct: 34 IITHNIDNRFINKNIINHDGNILNKEYLY-IKTNNLINEINIVDNYDIIGIDECQFFEEN 92
Query: 62 DLYDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFF 121
DL FC + A++ K V+VAGL+ ++ R F S+ +IP + + KL A C+ C K A F
Sbjct: 93 DLEQFCDKMANNK-KKVIVAGLNCDFNRNIFNSISKLIPKVEKIKKLQAICQFCYKDASF 151
Query: 122 TLRKTQETQVELIGGVDVYMPVCRQHYVN 150
T++K + Q+ IGG D+Y+PVCR Y N
Sbjct: 152 TIKKHNKNQIIEIGGQDLYVPVCRLCYNN 180
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.138 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,300,839
Number of Sequences: 164201
Number of extensions: 771412
Number of successful extensions: 1844
Number of sequences better than 10.0: 61
Number of HSP's better than 10.0 without gapping: 41
Number of HSP's successfully gapped in prelim test: 20
Number of HSP's that attempted gapping in prelim test: 1748
Number of HSP's gapped (non-prelim): 61
length of query: 172
length of database: 59,974,054
effective HSP length: 102
effective length of query: 70
effective length of database: 43,225,552
effective search space: 3025788640
effective search space used: 3025788640
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0367.7


