

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0366.1
(1346 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
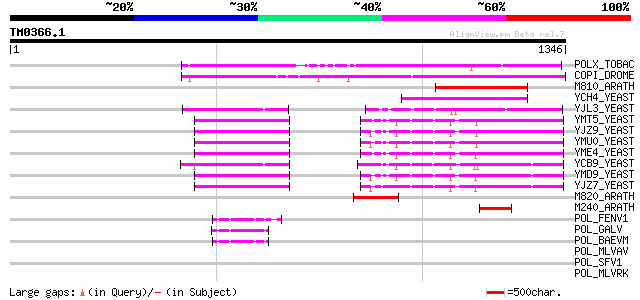
Score E
Sequences producing significant alignments: (bits) Value
POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from tran... 519 e-146
COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contain... 428 e-119
M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810... 202 7e-51
YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein 151 1e-35
YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein 110 2e-23
YMT5_YEAST (Q04214) Transposon Ty1 protein B 110 3e-23
YJZ9_YEAST (P47100) Transposon Ty1 protein B 108 1e-22
YMU0_YEAST (Q04670) Transposon Ty1 protein B 108 1e-22
YME4_YEAST (Q04711) Transposon Ty1 protein B 107 3e-22
YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B) 105 6e-22
YMD9_YEAST (Q03434) Transposon Ty1 protein B 104 1e-21
YJZ7_YEAST (P47098) Transposon Ty1 protein B 103 3e-21
M820_ARATH (P92520) Hypothetical mitochondrial protein AtMg00820... 96 5e-19
M240_ARATH (P93290) Hypothetical mitochondrial protein AtMg00240... 84 3e-15
POL_FENV1 (P31792) Pol polyprotein [Contains: Reverse transcript... 47 3e-04
POL_GALV (P21414) Pol polyprotein [Contains: Protease (EC 3.4.23... 46 8e-04
POL_BAEVM (P10272) Pol polyprotein [Contains: Protease (EC 3.4.2... 46 8e-04
POL_MLVAV (P03356) Pol polyprotein [Contains: Protease (EC 3.4.2... 44 0.003
POL_SFV1 (P23074) Pol polyprotein [Contains: Protease (EC 3.4.23... 44 0.004
POL_MLVRK (P31795) Pol polyprotein [Contains: Protease (EC 3.4.2... 44 0.004
>POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from
transposon TNT 1-94 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1328
Score = 519 bits (1336), Expect = e-146
Identities = 333/943 (35%), Positives = 515/943 (54%), Gaps = 71/943 (7%)
Query: 418 LWHFRLGHLSHDRLLALHTVQNSISVSKSIV---CDVCHLAKQKRKMFTVSVSKAQKCFD 474
LWH R+GH+S L L ++ IS +K CD C KQ R F S + D
Sbjct: 424 LWHKRMGHMSEKGLQIL-AKKSLISYAKGTTVKPCDYCLFGKQHRVSFQTSSERKLNILD 482
Query: 475 LVHMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVKTQFGQ 534
LV+ D+ GP+ S+ +KYF+T +DD SR +WV +L K +V Q + F LV+ + G+
Sbjct: 483 LVYSDVCGPMEIESMGGNKYFVTFIDDASRKLWVYILKTKDQVFQVFQKFHALVERETGR 542
Query: 535 IVKAIRSDNGPEFLLPAFY---SAQGIVHQKSCVSTPQQNGRVERKHQHILNIARALLFQ 591
+K +RSDNG E+ F S+ GI H+K+ TPQ NG ER ++ I+ R++L
Sbjct: 543 KLKRLRSDNGGEYTSREFEEYCSSHGIRHEKTVPGTPQHNGVAERMNRTIVEKVRSMLRM 602
Query: 592 SKLPKKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLCFATTLTN 651
+KLPK W +V A +L+NR PS L + P + + V +KVFG FA
Sbjct: 603 AKLPKSFWGEAVQTACYLINRSPSVPLAFEIPERVWTNKEVSYSHLKVFGCRAFAHVPKE 662
Query: 652 NRSKLDPRARKCIFLGYKQGMKGYVLMDLITQEIFISRDVIFHEHVLPYKGSNSPAWTCL 711
R+KLD ++ CIF+GY GY L D + +++ SRDV+F E
Sbjct: 663 QRTKLDDKSIPCIFIGYGDEEFGYRLWDPVKKKVIRSRDVVFRE---------------- 706
Query: 712 DLQTQAESTADKLNVASEIQSAKGDASVSTPCASIEDGVQHEMPSGASIEDEIPNTQTVD 771
++ + AD ++ ++++ V+ P S + S S DE+ +
Sbjct: 707 ---SEVRTAAD---MSEKVKNGIIPNFVTIPSTS------NNPTSAESTTDEVS-----E 749
Query: 772 EIPQPGSITAEGEVEVRRSTRPLKRPVHLANFQFSALPIRSSTAHSIVHHYSDERLSTAH 831
+ QPG + +GE + ++ H + P+R S + E
Sbjct: 750 QGEQPGEVIEQGE----QLDEGVEEVEHPTQGEEQHQPLRRSERPRV------ESRRYPS 799
Query: 832 RAYALNIAQDQEPSTFAEA---NKDLHWREAMQAEIKALEKNGTWKLVDLPEGVKPIGNK 888
Y L I+ D+EP + E + +AMQ E+++L+KNGT+KLV+LP+G +P+ K
Sbjct: 800 TEYVL-ISDDREPESLKEVLSHPEKNQLMKAMQEEMESLQKNGTYKLVELPKGKRPLKCK 858
Query: 889 WVYRVKRNVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVRVILALAASQNWHL 948
WV+++K++ D L RYKARLV KG+ Q +G+D+ + FSPV K+T++R IL+LAAS + +
Sbjct: 859 WVFKLKKDGDCKLVRYKARLVVKGFEQKKGIDFDEIFSPVVKMTSIRTILSLAASLDLEV 918
Query: 949 HQLDVDNAFLHGNLDEDVYMTIPAGVP-SVKPNQVCKLLKSLYGLKQASRKWYEKLSAHL 1007
QLDV AFLHG+L+E++YM P G + K + VCKL KSLYGLKQA R+WY K + +
Sbjct: 919 EQLDVKTAFLHGDLEEEIYMEQPEGFEVAGKKHMVCKLNKSLYGLKQAPRQWYMKFDSFM 978
Query: 1008 ETLGFKQTASDHSLFVK-FQGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDL 1066
++ + +T SD ++ K F ++F LL+YVDD+++ G +K L ++F +KDL
Sbjct: 979 KSQTYLKTYSDPCVYFKRFSENNFIILLLYVDDMLIVGKDKGLIAKLKGDLSKSFDMKDL 1038
Query: 1067 GVLKYFLGLEVAHSTSG--ISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLS------ 1118
G + LG+++ + + L Q KY +L+ +KPV+TPL ++LS
Sbjct: 1039 GPAQQILGMKIVRERTSRKLWLSQEKYIERVLERFNMKNAKPVSTPLAGHLKLSKKMCPT 1098
Query: 1119 --QEQGKPYDDPAAYRRLVGRLLY-LTTTRPDISHATQQLSQYMSNPMDGHFKAAQRVLR 1175
+E+G P Y VG L+Y + TRPDI+HA +S+++ NP H++A + +LR
Sbjct: 1099 TVEEKGNMAKVP--YSSAVGSLMYAMVCTRPDIAHAVGVVSRFLENPGKEHWEAVKWILR 1156
Query: 1176 YLKASPGLGLLFPRNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQTTV 1235
YL+ + G L F + I ++GY+DAD AG D R+S +GY F +SW++K Q V
Sbjct: 1157 YLRGTTGDCLCFGGSDPI-LKGYTDADMAGDIDNRKSSTGYLFTFSGGAISWQSKLQKCV 1215
Query: 1236 SRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTK 1295
+ S+ EAEY A E+ WL LQ+L + V++CD+QSA+ ++ N ++H RTK
Sbjct: 1216 ALSTTEAEYIAATETGKEMIWLKRFLQELGLH-QKEYVVYCDSQSAIDLSKNSMYHARTK 1274
Query: 1296 HLDIDCHVVREKLQAGILKLLPIPTTLQVADVFTKALQPRVFQ 1338
H+D+ H +RE + LK+L I T AD+ TK + F+
Sbjct: 1275 HIDVRYHWIREMVDDESLKVLKISTNENPADMLTKVVPRNKFE 1317
>COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contains:
Copia VLP protein; Copia protease (EC 3.4.23.-)]
Length = 1409
Score = 428 bits (1100), Expect = e-119
Identities = 301/992 (30%), Positives = 482/992 (48%), Gaps = 71/992 (7%)
Query: 418 LWHFRLGHLSHDRLLAL--------HTVQNSISVSKSIVCDVCHLAKQKRKMFTVSVSKA 469
LWH R GH+S +LL + ++ N++ +S I C+ C KQ R F K
Sbjct: 417 LWHERFGHISDGKLLEIKRKNMFSDQSLLNNLELSCEI-CEPCLNGKQARLPFKQLKDKT 475
Query: 470 --QKCFDLVHMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITL 527
++ +VH D+ GP+ ++ + YF+ +D ++ + L+ K +V ++F+
Sbjct: 476 HIKRPLFVVHSDVCGPITPVTLDDKNYFVIFVDQFTHYCVTYLIKYKSDVFSMFQDFVAK 535
Query: 528 VKTQFGQIVKAIRSDNGPEFL---LPAFYSAQGIVHQKSCVSTPQQNGRVERKHQHILNI 584
+ F V + DNG E+L + F +GI + + TPQ NG ER + I
Sbjct: 536 SEAHFNLKVVYLYIDNGREYLSNEMRQFCVKKGISYHLTVPHTPQLNGVSERMIRTITEK 595
Query: 585 ARALLFQSKLPKKMWCYSVLHAVFLMNRIPSKLL--KNKSPYELLYREAVDLEMMKVFGS 642
AR ++ +KL K W +VL A +L+NRIPS+ L +K+PYE+ + + L+ ++VFG+
Sbjct: 596 ARTMVSGAKLDKSFWGEAVLTATYLINRIPSRALVDSSKTPYEMWHNKKPYLKHLRVFGA 655
Query: 643 LCFATTLTNNRSKLDPRARKCIFLGYKQGMKGYVLMDLITQEIFISRDVIFHEHVLPYKG 702
+ + N + K D ++ K IF+GY+ G+ L D + ++ ++RDV+ E +
Sbjct: 656 TVYVH-IKNKQGKFDDKSFKSIFVGYEPN--GFKLWDAVNEKFIVARDVVVDETNMV--N 710
Query: 703 SNSPAWTCLDLQTQAESTADKLNVASEIQSAKGDASVSTPCASIE------DGVQHEMPS 756
S + + + L+ ES S + S C +I+ + P+
Sbjct: 711 SRAVKFETVFLKDSKESENKNFPNDSRKIIQTEFPNESKECDNIQFLKDSKESENKNFPN 770
Query: 757 GAS--IEDEIPNTQTVDEIPQPGSITAEGEVEVRRSTRPLKRPVHLANFQFSALPIRSST 814
+ I+ E PN + Q + E ++ KR HL + S P S
Sbjct: 771 DSRKIIQTEFPNESKECDNIQFLKDSKESNKYFLNESKKRKRDDHLNESKGSGNPNESRE 830
Query: 815 AHSIVH-------------------------------HYSDERLSTAHRAYALNIAQDQE 843
+ + H Y++E S + +
Sbjct: 831 SETAEHLKEIGIDNPTKNDGIEIINRRSERLKTKPQISYNEEDNSLNKVVLNAHTIFNDV 890
Query: 844 PSTFAEA---NKDLHWREAMQAEIKALEKNGTWKLVDLPEGVKPIGNKWVYRVKRNVDGT 900
P++F E + W EA+ E+ A + N TW + PE + ++WV+ VK N G
Sbjct: 891 PNSFDEIQYRDDKSSWEEAINTELNAHKINNTWTITKRPENKNIVDSRWVFSVKYNELGN 950
Query: 901 LARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVRVILALAASQNWHLHQLDVDNAFLHG 960
RYKARLVA+G+ Q +DY +TF+PVA++++ R IL+L N +HQ+DV AFL+G
Sbjct: 951 PIRYKARLVARGFTQKYQIDYEETFAPVARISSFRFILSLVIQYNLKVHQMDVKTAFLNG 1010
Query: 961 NLDEDVYMTIPAGVPSVKPNQVCKLLKSLYGLKQASRKWYEKLSAHLETLGFKQTASDHS 1020
L E++YM +P G+ S + VCKL K++YGLKQA+R W+E L+ F ++ D
Sbjct: 1011 TLKEEIYMRLPQGI-SCNSDNVCKLNKAIYGLKQAARCWFEVFEQALKECEFVNSSVDRC 1069
Query: 1021 LFVKFQGSSFTGL--LVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDLGVLKYFLGLEVA 1078
+++ +G+ + L+YVDDV++ +T K L + F + DL +K+F+G+ +
Sbjct: 1070 IYILDKGNINENIYVLLYVDDVVIATGDMTRMNNFKRYLMEKFRMTDLNEIKHFIGIRIE 1129
Query: 1079 HSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQGKPYDDPAAYRRLVGRL 1138
I L Q Y +L + V+TPL I + D R L+G L
Sbjct: 1130 MQEDKIYLSQSAYVKKILSKFNMENCNAVSTPLPSKINY-ELLNSDEDCNTPCRSLIGCL 1188
Query: 1139 LY-LTTTRPDISHATQQLSQYMSNPMDGHFKAAQRVLRYLKASPGLGLLFPRNSTI--NI 1195
+Y + TRPD++ A LS+Y S ++ +RVLRYLK + + L+F +N I
Sbjct: 1189 MYIMLCTRPDLTTAVNILSRYSSKNNSELWQNLKRVLRYLKGTIDMKLIFKKNLAFENKI 1248
Query: 1196 QGYSDADWAGCPDTRRSISGYCF-YIGRSLVSWKAKKQTTVSRSSNEAEYRALAYATCEL 1254
GY D+DWAG R+S +GY F +L+ W K+Q +V+ SS EAEY AL A E
Sbjct: 1249 IGYVDSDWAGSEIDRKSTTGYLFKMFDFNLICWNTKRQNSVAASSTEAEYMALFEAVREA 1308
Query: 1255 QWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKHLDIDCHVVREKLQAGILK 1314
WL +LL + + ++ DNQ + IA NP H+R KH+DI H RE++Q ++
Sbjct: 1309 LWLKFLLTSINIKLENPIKIYEDNQGCISIANNPSCHKRAKHIDIKYHFAREQVQNNVIC 1368
Query: 1315 LLPIPTTLQVADVFTKALQPRVFQGFATKLAM 1346
L IPT Q+AD+FTK L F KL +
Sbjct: 1369 LEYIPTENQLADIFTKPLPAARFVELRDKLGL 1400
>M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810
(ORF240b)
Length = 240
Score = 202 bits (513), Expect = 7e-51
Identities = 102/224 (45%), Positives = 140/224 (61%), Gaps = 1/224 (0%)
Query: 1033 LLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDLGVLKYFLGLEVAHSTSGISLCQRKYC 1092
LL+YVDD++L G++ T ++ L F +KDLG + YFLG+++ SG+ L Q KY
Sbjct: 3 LLLYVDDILLTGSSNTLLNMLIFQLSSTFSMKDLGPVHYFLGIQIKTHPSGLFLSQTKYA 62
Query: 1093 LDLLQETGTLGSKPVATPLDPSIRLSQEQGKPYDDPAAYRRLVGRLLYLTTTRPDISHAT 1152
+L G L KP++TPL + S K Y DP+ +R +VG L YLT TRPDIS+A
Sbjct: 63 EQILNNAGMLDCKPMSTPLPLKLNSSVSTAK-YPDPSDFRSIVGALQYLTLTRPDISYAV 121
Query: 1153 QQLSQYMSNPMDGHFKAAQRVLRYLKASPGLGLLFPRNSTINIQGYSDADWAGCPDTRRS 1212
+ Q M P F +RVLRY+K + GL +NS +N+Q + D+DWAGC TRRS
Sbjct: 122 NIVCQRMHEPTLADFDLLKRVLRYVKGTIFHGLYIHKNSKLNVQAFCDSDWAGCTSTRRS 181
Query: 1213 ISGYCFYIGRSLVSWKAKKQTTVSRSSNEAEYRALAYATCELQW 1256
+G+C ++G +++SW AK+Q TVSRSS E EYRALA EL W
Sbjct: 182 TTGFCTFLGCNIISWSAKRQPTVSRSSTETEYRALALTAAELTW 225
>YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein
Length = 308
Score = 151 bits (381), Expect = 1e-35
Identities = 99/308 (32%), Positives = 155/308 (50%), Gaps = 4/308 (1%)
Query: 951 LDVDNAFLHGNLDEDVYMTIPAG-VPSVKPNQVCKLLKSLYGLKQASRKWYEKLSAHLET 1009
+DVD AFL+ +DE +Y+ P G V P+ V +L +YGLKQA W E ++ L+
Sbjct: 1 MDVDTAFLNSTMDEPIYVKQPPGFVNERNPDYVWELYGGMYGLKQAPLLWNEHINNTLKK 60
Query: 1010 LGFKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDLGVL 1069
+GF + +H L+ + + VYVDD+++ + + VK L + + +KDLG +
Sbjct: 61 IGFCRHEGEHGLYFRSTSDGPIYIGVYVDDLLVAAPSPKIYDRVKQELTKLYSMKDLGKV 120
Query: 1070 KYFLGLEVAHSTSG-ISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQGKPYDDP 1128
FLGL + ST+G I+L + Y E+ K TPL S L + D
Sbjct: 121 DKFLGLNIHQSTNGDITLSLQDYIAKAASESEINTFKLTQTPLCNSKPLFETTSPHLKDI 180
Query: 1129 AAYRRLVGRLLYLTTT-RPDISHATQQLSQYMSNPMDGHFKAAQRVLRYLKASPGLGLLF 1187
Y+ +VG+LL+ T RPDIS+ LS+++ P H ++A+RVLRYL + + L +
Sbjct: 181 TPYQSIVGQLLFCANTGRPDISYPVSLLSRFLREPRAIHLESARRVLRYLYTTRSMCLKY 240
Query: 1188 PRNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAKK-QTTVSRSSNEAEYRA 1246
S + + Y DA D S GY + + V+W +KK + + S EAEY
Sbjct: 241 RSGSQVALTVYCDASHGAIHDLPHSTGGYVTLLAGAPVTWSSKKLKGVIPVPSTEAEYIT 300
Query: 1247 LAYATCEL 1254
+ E+
Sbjct: 301 ASETVMEI 308
>YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein
Length = 1803
Score = 110 bits (276), Expect = 2e-23
Identities = 113/504 (22%), Positives = 225/504 (44%), Gaps = 47/504 (9%)
Query: 863 EIKALEKNGTWKLVDLPEGVKPIGNKWVYRVKRNVDGTLARYKARLVAKGYNQVEGLDYF 922
++K + + + ++P+ + + ++ KRN YKAR+V +G Q
Sbjct: 1302 DMKVFDVDVKYSRSEIPDNLI-VPTNTIFTKKRN-----GIYKARIVCRGDTQSP----- 1350
Query: 923 DTFSPVAKLTT----VRVILALAASQNWHLHQLDVDNAFLHGNLDEDVYMTIPAGVPSVK 978
DT+S + + +++ L +A ++N + LD+++AFL+ L+E++Y+ P
Sbjct: 1351 DTYSVITTESLNHNHIKIFLMIANNRNMFMKTLDINHAFLYAKLEEEIYIPHPHD----- 1405
Query: 979 PNQVCKLLKSLYGLKQASRKWYEKLSAHLETLGFKQTASDHSLFVKFQGSSFTGLLVYVD 1038
V KL K+LYGLKQ+ ++W + L +L +G K + L+ + + VYVD
Sbjct: 1406 RRCVVKLNKALYGLKQSPKEWNDHLRQYLNGIGLKDNSYTPGLYQTEDKNLM--IAVYVD 1463
Query: 1039 DVILFGNTVTEFQLVKDSLHQAFGIKDLGVL------KYFLGLEVAHS----------TS 1082
D ++ + + L F +K G L LG+++ ++ S
Sbjct: 1464 DCVIAASNEQRLDEFINKLKSNFELKITGTLIDDVLDTDILGMDLVYNKRLGTIDLTLKS 1523
Query: 1083 GISLCQRKYCLDLLQETGTLGSKPVATPLDPS---IRLSQEQGKPYDDPAAYRRLVGRLL 1139
I+ +KY +L + + +DP +++S+E+ + ++L+G L
Sbjct: 1524 FINRMDKKYNEELKKIRKSSIPHMSTYKIDPKKDVLQMSEEEFR--QGVLKLQQLLGELN 1581
Query: 1140 YLT-TTRPDISHATQQLSQYMSNPMDGHFKAAQRVLRYLKASPGLGLLFPR--NSTINIQ 1196
Y+ R DI A +++++ ++ P + F ++++YL +G+ + R N +
Sbjct: 1582 YVRHKCRYDIEFAVKKVARLVNYPHERVFYMIYKIIQYLVRYKDIGIHYDRDCNKDKKVI 1641
Query: 1197 GYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQTTVSRSSNEAEYRALAYATCELQW 1256
+DA G +S G + G ++ + + K T SS EAE A+ + +
Sbjct: 1642 AITDAS-VGSEYDAQSRIGVILWYGMNIFNVYSNKSTNRCVSSTEAELHAIYEGYADSET 1700
Query: 1257 LLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKHLDIDCHVVREKLQAGILKLL 1316
L L++L V+ D++ A+ + K I +++EK++ +KLL
Sbjct: 1701 LKVTLKELGEGDNNDIVMITDSKPAIQGLNRSYQQPKEKFTWIKTEIIKEKIKEKSIKLL 1760
Query: 1317 PIPTTLQVADVFTKALQPRVFQGF 1340
I +AD+ TK + F+ F
Sbjct: 1761 KITGKGNIADLLTKPVSASDFKRF 1784
Score = 83.2 bits (204), Expect = 4e-15
Identities = 73/271 (26%), Positives = 120/271 (43%), Gaps = 16/271 (5%)
Query: 420 HFRLGHLSHDRL---LALHTVQNSISVSKS---IVCDVCHLAKQ-KRKMFTVSVSKAQKC 472
H R+GH ++ + + + S+ + K C C ++K KR +T S++
Sbjct: 562 HKRMGHTGIQQIENSIKHNHYEESLDLIKEPNEFWCQTCKISKATKRNHYTGSMNNHSTD 621
Query: 473 FDLVH---MDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGE--VQQQVKNFITL 527
+ MDI+GP++ ++ +Y L ++D+ +R+ NK + QV+ I
Sbjct: 622 HEPGSSWCMDIFGPVSSSNADTKRYMLIMVDNNTRYCMTSTHFNKNAETILAQVRKNIQY 681
Query: 528 VKTQFGQIVKAIRSDNGPEFL---LPAFYSAQGIVHQKSCVSTPQQNGRVERKHQHILNI 584
V+TQF + V+ I SD G EF + ++ ++GI H + NGR ER + I+
Sbjct: 682 VETQFDRKVREINSDRGTEFTNDQIEEYFISKGIHHILTSTQDHAANGRAERYIRTIITD 741
Query: 585 ARALLFQSKLPKKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLC 644
A LL QS L K W Y+V A + N + K K P + + R+ V + +M
Sbjct: 742 ATTLLRQSNLRVKFWEYAVTSATNIRNYLEHK-STGKLPLKAISRQPVTVRLMSFLPFGE 800
Query: 645 FATTLTNNRSKLDPRARKCIFLGYKQGMKGY 675
+N KL P I L GY
Sbjct: 801 KGIIWNHNHKKLKPSGLPSIILCKDPNSYGY 831
>YMT5_YEAST (Q04214) Transposon Ty1 protein B
Length = 1328
Score = 110 bits (275), Expect = 3e-23
Identities = 133/539 (24%), Positives = 239/539 (43%), Gaps = 73/539 (13%)
Query: 851 NKDLHWRE----AMQAEIKALEKNGTW---KLVDLPEGVKP---IGNKWVYRVKRNVDGT 900
NKD+ +E A E+ L K TW K D E + P I + +++ KR DGT
Sbjct: 814 NKDIKEKEKYIEAYHKEVNQLLKMKTWDTDKYYDRKE-IDPKRVINSMFIFNRKR--DGT 870
Query: 901 LARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVR-----VILALAASQNWHLHQLDVDN 955
+KAR VA+G + + DT+ + TV L+LA N+++ QLD+ +
Sbjct: 871 ---HKARFVARG-----DIQHPDTYDSGMQSNTVHHYALMTSLSLALDNNYYITQLDISS 922
Query: 956 AFLHGNLDEDVYMTIPAGVPSVKPN-QVCKLLKSLYGLKQASRKWYEKLSAHL-ETLGFK 1013
A+L+ ++ E++Y+ P P + N ++ +L KSLYGLKQ+ WYE + ++L + G +
Sbjct: 923 AYLYADIKEELYIRPP---PHLGMNDKLIRLKKSLYGLKQSGANWYETIKSYLIKQCGME 979
Query: 1014 QTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDLGV----- 1068
+ +F Q + ++VDD++LF + + + D L + K + +
Sbjct: 980 EVRGWSCVFENSQ----VTICLFVDDMVLFSKNLNSNKRIIDKLKMQYDTKIINLGESDE 1035
Query: 1069 -LKY-FLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQGKP-- 1124
++Y LGLE+ + + KY ++ + T + PL+P R G+P
Sbjct: 1036 EIQYDILGLEIKYQ-------RGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGL 1088
Query: 1125 YDDPA--------------AYRRLVGRLLYL-TTTRPDISHATQQLSQYMSNPMDGHFKA 1169
Y D ++L+G Y+ R D+ + L+Q++ P
Sbjct: 1089 YIDQQELELEEDDYKMKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSKQVLDM 1148
Query: 1170 AQRVLRYLKASPGLGLLF----PRNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLV 1225
+++++ + L++ P T + SDA + P + I G + + ++
Sbjct: 1149 TYELIQFIWNTRDKQLIWHKSKPVKPTNKLVVISDASYGNQPYYKSQI-GNIYLLNGKVI 1207
Query: 1226 SWKAKKQTTVSRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALH-I 1284
K+ K + S+ EAE A++ + L L YL+Q+L T L D++S + I
Sbjct: 1208 GGKSTKASLTCTSTTEAEIHAISESVPLLNNLSYLIQELDKK-PITKGLLTDSKSTISII 1266
Query: 1285 AANPVFHERTKHLDIDCHVVREKLQAGILKLLPIPTTLQVADVFTKALQPRVFQGFATK 1343
+N R + +R+++ L + I T +ADV TK L + F+ K
Sbjct: 1267 ISNNEEKFRNRFFGTKAMRLRDEVSGNHLHVCYIETKKNIADVMTKPLPIKTFKLLTNK 1325
Score = 65.5 bits (158), Expect = 1e-09
Identities = 56/238 (23%), Positives = 99/238 (41%), Gaps = 11/238 (4%)
Query: 449 CDVCHLAKQKR----KMFTVSVSKAQKCFDLVHMDIWGPLAQASVHNHKYFLTVLDDYSR 504
C C + K + K + + + F +H DI+GP+ YF++ D+ ++
Sbjct: 210 CPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSYFISFTDETTK 269
Query: 505 FVWVVLLNNKGE--VQQQVKNFITLVKTQFGQIVKAIRSDNGPEF---LLPAFYSAQGIV 559
F WV L+++ E + + +K QF V I+ D G E+ L F GI
Sbjct: 270 FRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHKFLEKNGIT 329
Query: 560 HQKSCVSTPQQNGRVERKHQHILNIARALLFQSKLPKKMWCYSVLHAVFLMNRIPSKLLK 619
+ + + +G ER ++ +L+ R L S LP +W ++ + + N + S K
Sbjct: 330 PCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRNSLASPKSK 389
Query: 620 NKSPYELLYREAVDLEMMKVFGSLCFATTLTNNRSKLDPRARKCIFLGYKQGMKGYVL 677
KS + +D+ + FG N SK+ PR L + GY++
Sbjct: 390 -KSARQHAGLAGLDISTLLPFGQPVIVND-HNPNSKIHPRGIPGYALHPSRNSYGYII 445
>YJZ9_YEAST (P47100) Transposon Ty1 protein B
Length = 1755
Score = 108 bits (270), Expect = 1e-22
Identities = 132/541 (24%), Positives = 240/541 (43%), Gaps = 77/541 (14%)
Query: 851 NKDLHWRE----AMQAEIKALEKNGTW--------KLVDLPEGVKPIGNKWVYRVKRNVD 898
NKD+ +E A E+ L K TW K +D P+ V I + +++ KR D
Sbjct: 1241 NKDIKEKEKYIEAYHKEVNQLLKMKTWDTDEYYDRKEID-PKRV--INSMFIFNKKR--D 1295
Query: 899 GTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVR-----VILALAASQNWHLHQLDV 953
GT +KAR VA+G + + DT+ + TV L+LA N+++ QLD+
Sbjct: 1296 GT---HKARFVARG-----DIQHPDTYDSGMQSNTVHHYALMTSLSLALDNNYYITQLDI 1347
Query: 954 DNAFLHGNLDEDVYMTIPAGVPSVKPN-QVCKLLKSLYGLKQASRKWYEKLSAHL-ETLG 1011
+A+L+ ++ E++Y+ P P + N ++ +L KSLYGLKQ+ WYE + ++L + G
Sbjct: 1348 SSAYLYADIKEELYIRPP---PHLGMNDKLIRLKKSLYGLKQSGANWYETIKSYLIQQCG 1404
Query: 1012 FKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDLGV--- 1068
++ +F Q + ++VDD++LF + + + + L + K + +
Sbjct: 1405 MEEVRGWSCVFKNSQ----VTICLFVDDMVLFSKNLNSNKRIIEKLKMQYDTKIINLGES 1460
Query: 1069 ---LKY-FLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQGKP 1124
++Y LGLE+ + + KY ++ + T + PL+P R G+P
Sbjct: 1461 DEEIQYDILGLEIKYQ-------RGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQP 1513
Query: 1125 --YDDPA--------------AYRRLVGRLLYL-TTTRPDISHATQQLSQYMSNPMDGHF 1167
Y D ++L+G Y+ R D+ + L+Q++ P
Sbjct: 1514 GLYIDQQELELEEDDYKMKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSKQVL 1573
Query: 1168 KAAQRVLRYLKASPGLGLLF----PRNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRS 1223
+++++ + L++ P T + SDA + P + I G + +
Sbjct: 1574 DMTYELIQFIWNTRDKQLIWHKSKPVKPTNKLVVISDASYGNQPYYKSQI-GNIYLLNGK 1632
Query: 1224 LVSWKAKKQTTVSRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALH 1283
++ K+ K + S+ EAE A++ + L L YL+Q+L T L D++S +
Sbjct: 1633 VIGGKSTKASLTCTSTTEAEIHAISESVPLLNNLSYLIQELDKK-PITKGLLTDSKSTIS 1691
Query: 1284 -IAANPVFHERTKHLDIDCHVVREKLQAGILKLLPIPTTLQVADVFTKALQPRVFQGFAT 1342
I +N R + +R+++ L + I T +ADV TK L + F+
Sbjct: 1692 IIISNNEEKFRNRFFGTKAMRLRDEVSGNHLHVCYIETKKNIADVMTKPLPIKTFKLLTN 1751
Query: 1343 K 1343
K
Sbjct: 1752 K 1752
Score = 65.5 bits (158), Expect = 1e-09
Identities = 56/238 (23%), Positives = 99/238 (41%), Gaps = 11/238 (4%)
Query: 449 CDVCHLAKQKR----KMFTVSVSKAQKCFDLVHMDIWGPLAQASVHNHKYFLTVLDDYSR 504
C C + K + K + + + F +H DI+GP+ YF++ D+ ++
Sbjct: 637 CPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSYFISFTDETTK 696
Query: 505 FVWVVLLNNKGE--VQQQVKNFITLVKTQFGQIVKAIRSDNGPEF---LLPAFYSAQGIV 559
F WV L+++ E + + +K QF V I+ D G E+ L F GI
Sbjct: 697 FRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHKFLEKNGIT 756
Query: 560 HQKSCVSTPQQNGRVERKHQHILNIARALLFQSKLPKKMWCYSVLHAVFLMNRIPSKLLK 619
+ + + +G ER ++ +L+ R L S LP +W ++ + + N + S K
Sbjct: 757 PCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRNSLASPKSK 816
Query: 620 NKSPYELLYREAVDLEMMKVFGSLCFATTLTNNRSKLDPRARKCIFLGYKQGMKGYVL 677
KS + +D+ + FG N SK+ PR L + GY++
Sbjct: 817 -KSARQHAGLAGLDISTLLPFGQPVIVND-HNPNSKIHPRGIPGYALHPSRNSYGYII 872
>YMU0_YEAST (Q04670) Transposon Ty1 protein B
Length = 1328
Score = 108 bits (269), Expect = 1e-22
Identities = 133/541 (24%), Positives = 242/541 (44%), Gaps = 77/541 (14%)
Query: 851 NKDLHWRE----AMQAEIKALEKNGTW--------KLVDLPEGVKPIGNKWVYRVKRNVD 898
NKD+ +E A E+ L K TW K +D P+ V I + +++ KR D
Sbjct: 814 NKDIKEKEKYIQAYHKEVNQLLKMKTWDTDRYYDRKEID-PKRV--INSMFIFNRKR--D 868
Query: 899 GTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVR-----VILALAASQNWHLHQLDV 953
GT +KAR VA+G + + DT+ P + TV L+LA N+++ QLD+
Sbjct: 869 GT---HKARFVARG-----DIQHPDTYDPGMQSNTVHHYALMTSLSLALDNNYYITQLDI 920
Query: 954 DNAFLHGNLDEDVYMTIPAGVPSVKPN-QVCKLLKSLYGLKQASRKWYEKLSAHL-ETLG 1011
+A+L+ ++ E++Y+ P P + N ++ +L KSLYGLKQ+ WYE + ++L + G
Sbjct: 921 SSAYLYADIKEELYIRPP---PHLGMNDKLIRLKKSLYGLKQSGANWYETIKSYLIKQCG 977
Query: 1012 FKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDLGV--- 1068
++ +F Q + ++VDD+ILF + + + +L + + K + +
Sbjct: 978 MEEVRGWSCVFKNSQ----VTICLFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGES 1033
Query: 1069 ---LKY-FLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQGKP 1124
++Y LGLE+ + + KY ++ + T + PL+P R G+P
Sbjct: 1034 DNEIQYDILGLEIKYQ-------RGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQP 1086
Query: 1125 --YDDPA--------------AYRRLVGRLLYL-TTTRPDISHATQQLSQYMSNPMDGHF 1167
Y D ++L+G Y+ R D+ + L+Q++ P
Sbjct: 1087 GLYIDQQELELEEDDYKMKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSKQVL 1146
Query: 1168 KAAQRVLRYLKASPGLGLLF----PRNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRS 1223
+++++ + L++ P T + SDA + P + I G + +
Sbjct: 1147 DMTYELIQFIWNTRDKQLIWHKSKPVKPTNKLVVISDASYGNQPYYKSQI-GNIYLLNGK 1205
Query: 1224 LVSWKAKKQTTVSRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALH 1283
++ K+ K + S+ EAE A++ + L L +L+Q+L T L D++S +
Sbjct: 1206 VIGGKSTKASLTCTSTTEAEIHAISESVPLLNNLSHLVQELNKK-PITKGLLTDSKSTIS 1264
Query: 1284 -IAANPVFHERTKHLDIDCHVVREKLQAGILKLLPIPTTLQVADVFTKALQPRVFQGFAT 1342
I +N R + +R+++ L + I T +ADV TK L + F+
Sbjct: 1265 IIISNNEEKFRNRFFGTKAMRLRDEVSGNHLHVCYIETKKNIADVMTKPLPIKTFKLLTN 1324
Query: 1343 K 1343
K
Sbjct: 1325 K 1325
Score = 65.5 bits (158), Expect = 1e-09
Identities = 56/238 (23%), Positives = 99/238 (41%), Gaps = 11/238 (4%)
Query: 449 CDVCHLAKQKR----KMFTVSVSKAQKCFDLVHMDIWGPLAQASVHNHKYFLTVLDDYSR 504
C C + K + K + + + F +H DI+GP+ YF++ D+ ++
Sbjct: 210 CPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSYFISFTDETTK 269
Query: 505 FVWVVLLNNKGE--VQQQVKNFITLVKTQFGQIVKAIRSDNGPEF---LLPAFYSAQGIV 559
F WV L+++ E + + +K QF V I+ D G E+ L F GI
Sbjct: 270 FRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHKFLEKNGIT 329
Query: 560 HQKSCVSTPQQNGRVERKHQHILNIARALLFQSKLPKKMWCYSVLHAVFLMNRIPSKLLK 619
+ + + +G ER ++ +L+ R L S LP +W ++ + + N + S K
Sbjct: 330 PCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRNSLASPKSK 389
Query: 620 NKSPYELLYREAVDLEMMKVFGSLCFATTLTNNRSKLDPRARKCIFLGYKQGMKGYVL 677
KS + +D+ + FG N SK+ PR L + GY++
Sbjct: 390 -KSARQHAGLAGLDISTLLPFGQPVIVND-HNPNSKIHPRGIPGYALHPSRNSYGYII 445
>YME4_YEAST (Q04711) Transposon Ty1 protein B
Length = 1328
Score = 107 bits (266), Expect = 3e-22
Identities = 131/539 (24%), Positives = 240/539 (44%), Gaps = 73/539 (13%)
Query: 851 NKDLHWRE----AMQAEIKALEKNGTW---KLVDLPEGVKP---IGNKWVYRVKRNVDGT 900
NKD+ +E A E+ L K TW K D E + P I + +++ KR DGT
Sbjct: 814 NKDIKEKEKYIEAYHKEVNQLLKMNTWDTDKYYDRKE-IDPKRVINSMFIFNRKR--DGT 870
Query: 901 LARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVR-----VILALAASQNWHLHQLDVDN 955
+KAR VA+G + + DT+ + TV L+LA N+++ QLD+ +
Sbjct: 871 ---HKARFVARG-----DIQHPDTYDSGMQSNTVHHYALMTSLSLALDNNYYITQLDISS 922
Query: 956 AFLHGNLDEDVYMTIPAGVPSVKPN-QVCKLLKSLYGLKQASRKWYEKLSAHL-ETLGFK 1013
A+L+ ++ E++Y+ P P + N ++ +L KSLYGLKQ+ WYE + ++L + G +
Sbjct: 923 AYLYADIKEELYIRPP---PHLGMNDKLIRLKKSLYGLKQSGANWYETIKSYLIKQCGME 979
Query: 1014 QTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDLGV----- 1068
+ +F Q + ++VDD+ILF + + + +L + + K + +
Sbjct: 980 EVRGWSCVFKNSQ----VTICLFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESDN 1035
Query: 1069 -LKY-FLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQGKP-- 1124
++Y LGLE+ + + KY ++ + T + PL+P R G+P
Sbjct: 1036 EIQYDILGLEIKYQ-------RGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGL 1088
Query: 1125 YDDP--------------AAYRRLVGRLLYL-TTTRPDISHATQQLSQYMSNPMDGHFKA 1169
Y D ++L+G Y+ R D+ + L+Q++ P
Sbjct: 1089 YIDQDELEIDEDEYKEKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSRQVLDM 1148
Query: 1170 AQRVLRYLKASPGLGLLFPRNSTI----NIQGYSDADWAGCPDTRRSISGYCFYIGRSLV 1225
+++++ + L++ +N + SDA + P + I G + + ++
Sbjct: 1149 TYELIQFMWDTRDKQLIWHKNKPTEPDNKLVAISDASYGNQPYYKSQI-GNIYLLNGKVI 1207
Query: 1226 SWKAKKQTTVSRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALH-I 1284
K+ K + S+ EAE A++ + L L +L+Q+L T L D++S + I
Sbjct: 1208 GGKSTKASLTCTSTTEAEIHAISESVPLLNNLSHLVQELNKK-PITKGLLTDSKSTISII 1266
Query: 1285 AANPVFHERTKHLDIDCHVVREKLQAGILKLLPIPTTLQVADVFTKALQPRVFQGFATK 1343
+N R + +R+++ L + I T +ADV TK L + F+ K
Sbjct: 1267 ISNNEEKFRNRFFGTKAMRLRDEVSGNHLHVCYIETKKNIADVMTKPLPIKTFKLLTNK 1325
Score = 65.5 bits (158), Expect = 1e-09
Identities = 56/238 (23%), Positives = 99/238 (41%), Gaps = 11/238 (4%)
Query: 449 CDVCHLAKQKR----KMFTVSVSKAQKCFDLVHMDIWGPLAQASVHNHKYFLTVLDDYSR 504
C C + K + K + + + F +H DI+GP+ YF++ D+ ++
Sbjct: 210 CPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSYFISFTDETTK 269
Query: 505 FVWVVLLNNKGE--VQQQVKNFITLVKTQFGQIVKAIRSDNGPEF---LLPAFYSAQGIV 559
F WV L+++ E + + +K QF V I+ D G E+ L F GI
Sbjct: 270 FRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHKFLEKNGIT 329
Query: 560 HQKSCVSTPQQNGRVERKHQHILNIARALLFQSKLPKKMWCYSVLHAVFLMNRIPSKLLK 619
+ + + +G ER ++ +L+ R L S LP +W ++ + + N + S K
Sbjct: 330 PCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRNSLASPKSK 389
Query: 620 NKSPYELLYREAVDLEMMKVFGSLCFATTLTNNRSKLDPRARKCIFLGYKQGMKGYVL 677
KS + +D+ + FG N SK+ PR L + GY++
Sbjct: 390 -KSARQHAGLAGLDISTLLPFGQPVIVND-HNPNSKIHPRGIPGYALHPSRNSYGYII 445
>YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B)
Length = 1770
Score = 105 bits (263), Expect = 6e-22
Identities = 132/545 (24%), Positives = 243/545 (44%), Gaps = 72/545 (13%)
Query: 843 EPSTFAEANKDL-HWREAMQAEIKALEKNGTW---KLVDLPE--GVKPIGNKWVYRVKRN 896
E T+ + NK+ + EA EI L K TW K D + K I + +++ KR
Sbjct: 1251 EAITYNKDNKEKDRYVEAYHKEISQLLKMNTWDTNKYYDRNDIDPKKVINSMFIFNKKR- 1309
Query: 897 VDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVR-----VILALAASQNWHLHQL 951
DGT +KAR VA+G + + DT+ + TV L++A ++++ QL
Sbjct: 1310 -DGT---HKARFVARG-----DIQHPDTYDSDMQSNTVHHYALMTSLSIALDNDYYITQL 1360
Query: 952 DVDNAFLHGNLDEDVYMTIPAGVPSVKPN-QVCKLLKSLYGLKQASRKWYEKLSAHLETL 1010
D+ +A+L+ ++ E++Y+ P P + N ++ +L KSLYGLKQ+ WYE + ++L
Sbjct: 1361 DISSAYLYADIKEELYIRPP---PHLGLNDKLLRLRKSLYGLKQSGANWYETIKSYLINC 1417
Query: 1011 GFKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDLGV-- 1068
Q S K +S + ++VDD+ILF + + + +L + + K + +
Sbjct: 1418 CDMQEVRGWSCVFK---NSQVTICLFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGE 1474
Query: 1069 ----LKY-FLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQGK 1123
++Y LGLE+ + S KY ++++ T + PL+P + + G+
Sbjct: 1475 SDNEIQYDILGLEIKYQRS-------KYMKLGMEKSLTEKLPKLNVPLNPKGKKLRAPGQ 1527
Query: 1124 PYD---------DPAAYR-------RLVGRLLYL-TTTRPDISHATQQLSQYMSNPMDGH 1166
P D Y+ +L+G Y+ R D+ + L+Q++ P
Sbjct: 1528 PGHYIDQDELEIDEDEYKEKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSRQV 1587
Query: 1167 FKAAQRVLRYLKASPGLGLLFPRNSTI----NIQGYSDADWAGCPDTRRSISGYCFYIGR 1222
+++++ + L++ +N + SDA + P + I G F +
Sbjct: 1588 LDMTYELIQFMWDTRDKQLIWHKNKPTKPDNKLVAISDASYGNQPYYKSQI-GNIFLLNG 1646
Query: 1223 SLVSWKAKKQTTVSRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPV---LFCDNQ 1279
++ K+ K + S+ EAE A++ A L L +L+Q+L P+ L D++
Sbjct: 1647 KVIGGKSTKASLTCTSTTEAEIHAVSEAIPLLNNLSHLVQEL----NKKPIIKGLLTDSR 1702
Query: 1280 SALHIAANPVFHE-RTKHLDIDCHVVREKLQAGILKLLPIPTTLQVADVFTKALQPRVFQ 1338
S + I + + R + +R+++ L + I T +ADV TK L + F+
Sbjct: 1703 STISIIKSTNEEKFRNRFFGTKAMRLRDEVSGNNLYVYYIETKKNIADVMTKPLPIKTFK 1762
Query: 1339 GFATK 1343
K
Sbjct: 1763 LLTNK 1767
Score = 74.3 bits (181), Expect = 2e-12
Identities = 68/282 (24%), Positives = 121/282 (42%), Gaps = 22/282 (7%)
Query: 415 PGALWHFRLGHLSHDRLLALHTVQNSISVSK----------SIVCDVCHLAKQKR----K 460
P L H LGH + R + +N+++ K + C C + K + K
Sbjct: 590 PYPLIHRMLGHANF-RSIQKSLKKNAVTYLKESDIEWSNASTYQCPDCLIGKSTKHRHVK 648
Query: 461 MFTVSVSKAQKCFDLVHMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGE--VQ 518
+ ++ + F +H DI+GP+ YF++ D+ +RF WV L+++ E +
Sbjct: 649 GSRLKYQESYEPFQYLHTDIFGPVHHLPKSAPSYFISFTDEKTRFQWVYPLHDRREESIL 708
Query: 519 QQVKNFITLVKTQFGQIVKAIRSDNGPEF---LLPAFYSAQGIVHQKSCVSTPQQNGRVE 575
+ + +K QF V I+ D G E+ L F++ +GI + + + +G E
Sbjct: 709 NVFTSILAFIKNQFNARVLVIQMDRGSEYTNKTLHKFFTNRGITACYTTTADSRAHGVAE 768
Query: 576 RKHQHILNIARALLFQSKLPKKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLYREAVDLE 635
R ++ +LN R LL S LP +W +V + + N + S +KS + +D+
Sbjct: 769 RLNRTLLNDCRTLLHCSGLPNHLWFSAVEFSTIIRNSLVSP-KNDKSARQHAGLAGLDIT 827
Query: 636 MMKVFGSLCFATTLTNNRSKLDPRARKCIFLGYKQGMKGYVL 677
+ FG N SK+ PR L + GY++
Sbjct: 828 TILPFGQPVIVNN-HNPDSKIHPRGIPGYALHPSRNSYGYII 868
>YMD9_YEAST (Q03434) Transposon Ty1 protein B
Length = 1328
Score = 104 bits (260), Expect = 1e-21
Identities = 131/544 (24%), Positives = 242/544 (44%), Gaps = 83/544 (15%)
Query: 851 NKDLHWRE----AMQAEIKALEKNGTW--------KLVDLPEGVKPIGNKWVYRVKRNVD 898
NKD+ +E A E+ L K TW K +D P+ V I + +++ KR D
Sbjct: 814 NKDIKEKEKYIEAYHKEVNQLLKMKTWDTDEYYDRKEID-PKRV--INSMFIFNKKR--D 868
Query: 899 GTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVR-----VILALAASQNWHLHQLDV 953
GT +KAR VA+G + + DT+ + TV L+LA N+++ QLD+
Sbjct: 869 GT---HKARFVARG-----DIQHPDTYDSGMQSNTVHHYALMTSLSLALDNNYYITQLDI 920
Query: 954 DNAFLHGNLDEDVYMTIPAGVPSVKPN-QVCKLLKSLYGLKQASRKWYEKLSAHL-ETLG 1011
+A+L+ ++ E++Y+ P P + N ++ +L KSLYGLKQ+ WYE + ++L + G
Sbjct: 921 SSAYLYADIKEELYIRPP---PHLGMNDKLIRLKKSLYGLKQSGANWYETIKSYLIKQCG 977
Query: 1012 FKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDLGV--- 1068
++ +F Q + ++VDD+ILF + + + +L + + K + +
Sbjct: 978 MEEVRGWSCVFKNSQ----VTICLFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGES 1033
Query: 1069 ---LKY-FLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQGKP 1124
++Y LGLE+ + + KY ++ + T + PL+P R G+P
Sbjct: 1034 DNEIQYDILGLEIKYQ-------RGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQP 1086
Query: 1125 --YDDP--------------AAYRRLVGRLLYL-TTTRPDISHATQQLSQYMSNPMDGHF 1167
Y D ++L+G Y+ R D+ + L+Q++ P
Sbjct: 1087 GLYIDQDELEIDEDEYKEKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSRQVL 1146
Query: 1168 KAAQRVLRYLKASPGLGLLFPRNSTI----NIQGYSDADWAGCPDTRRSISGYCFYIGRS 1223
+++++ + L++ +N + SDA + P + I G + +
Sbjct: 1147 DMTYELIQFMWDTRDKQLIWHKNKPTEPDNKLVAISDASYGNQPYYKSQI-GNIYLLNGK 1205
Query: 1224 LVSWKAKKQTTVSRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPV---LFCDNQS 1280
++ K+ K + S+ EAE A++ + L L YL+Q+L P+ L D++S
Sbjct: 1206 VIGGKSTKASLTCTSTTEAEIHAISESVPLLNNLSYLIQEL----NKKPIIKGLLTDSRS 1261
Query: 1281 ALHIAANPVFHE-RTKHLDIDCHVVREKLQAGILKLLPIPTTLQVADVFTKALQPRVFQG 1339
+ I + + R + +R+++ L + I T +ADV TK L + F+
Sbjct: 1262 TISIIKSTNEEKFRNRFFGTKAMRLRDEVSGNNLYVYYIETKKNIADVMTKPLPIKTFKL 1321
Query: 1340 FATK 1343
K
Sbjct: 1322 LTNK 1325
Score = 65.5 bits (158), Expect = 1e-09
Identities = 56/238 (23%), Positives = 99/238 (41%), Gaps = 11/238 (4%)
Query: 449 CDVCHLAKQKR----KMFTVSVSKAQKCFDLVHMDIWGPLAQASVHNHKYFLTVLDDYSR 504
C C + K + K + + + F +H DI+GP+ YF++ D+ ++
Sbjct: 210 CPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSYFISFTDETTK 269
Query: 505 FVWVVLLNNKGE--VQQQVKNFITLVKTQFGQIVKAIRSDNGPEF---LLPAFYSAQGIV 559
F WV L+++ E + + +K QF V I+ D G E+ L F GI
Sbjct: 270 FRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHKFLEKNGIT 329
Query: 560 HQKSCVSTPQQNGRVERKHQHILNIARALLFQSKLPKKMWCYSVLHAVFLMNRIPSKLLK 619
+ + + +G ER ++ +L+ R L S LP +W ++ + + N + S K
Sbjct: 330 PCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRNSLASPKSK 389
Query: 620 NKSPYELLYREAVDLEMMKVFGSLCFATTLTNNRSKLDPRARKCIFLGYKQGMKGYVL 677
KS + +D+ + FG N SK+ PR L + GY++
Sbjct: 390 -KSARQHAGLAGLDISTLLPFGQPVIVND-HNPNSKIHPRGIPGYALHPSRNSYGYII 445
>YJZ7_YEAST (P47098) Transposon Ty1 protein B
Length = 1755
Score = 103 bits (257), Expect = 3e-21
Identities = 131/541 (24%), Positives = 241/541 (44%), Gaps = 77/541 (14%)
Query: 851 NKDLHWRE----AMQAEIKALEKNGTW--------KLVDLPEGVKPIGNKWVYRVKRNVD 898
NKD+ +E A E+ L K TW K +D P+ V I + +++ KR D
Sbjct: 1241 NKDIKEKEKYIEAYHKEVNQLLKMKTWDTDEYYDRKEID-PKRV--INSMFIFNKKR--D 1295
Query: 899 GTLARYKARLVAKGYNQVEGLDYFDTF--SPVAKLTTVRVILALAASQNWHLHQLDVDNA 956
GT +KAR VA+G ++ D +DT S + L+LA N+++ QLD+ +A
Sbjct: 1296 GT---HKARFVARG--DIQHPDTYDTGMQSNTVHHYALMTSLSLALDNNYYITQLDISSA 1350
Query: 957 FLHGNLDEDVYMTIPAGVPSVKPN-QVCKLLKSLYGLKQASRKWYEKLSAHL-ETLGFKQ 1014
+L+ ++ E++Y+ P P + N ++ +L KS YGLKQ+ WYE + ++L + G ++
Sbjct: 1351 YLYADIKEELYIRPP---PHLGMNDKLIRLKKSHYGLKQSGANWYETIKSYLIKQCGMEE 1407
Query: 1015 TASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDLGV------ 1068
+F Q + ++VDD+ILF + + + +L + + K + +
Sbjct: 1408 VRGWSCVFKNSQ----VTICLFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESDNE 1463
Query: 1069 LKY-FLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQGKP--Y 1125
++Y LGLE+ + + KY ++ + T + PL+P R G+P Y
Sbjct: 1464 IQYDILGLEIKYQ-------RGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLY 1516
Query: 1126 DDP--------------AAYRRLVGRLLYL-TTTRPDISHATQQLSQYMSNPMDGHFKAA 1170
D ++L+G Y+ R D+ + L+Q++ P
Sbjct: 1517 IDQDELEIDEDEYKEKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSRQVLDMT 1576
Query: 1171 QRVLRYLKASPGLGLLFPRNSTI----NIQGYSDADWAGCPDTRRSISGYCFYIGRSLVS 1226
+++++ + L++ +N + SDA + P + I G F + ++
Sbjct: 1577 YELIQFMWDTRDKQLIWHKNKPTEPDNKLVAISDASYGNQPYYKSQI-GNIFLLNGKVIG 1635
Query: 1227 WKAKKQTTVSRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPV---LFCDNQSALH 1283
K+ K + S+ EAE A++ + L L YL+Q+L P+ L D++S +
Sbjct: 1636 GKSTKASLTCTSTTEAEIHAISESVPLLNNLSYLIQEL----NKKPIIKGLLTDSRSTIS 1691
Query: 1284 IAANPVFHE-RTKHLDIDCHVVREKLQAGILKLLPIPTTLQVADVFTKALQPRVFQGFAT 1342
I + + R + +R+++ L + I T +ADV TK L + F+
Sbjct: 1692 IIKSTNEEKFRNRFFGTKAMRLRDEVSGNNLYVYYIETKKNIADVMTKPLPIKTFKLLTN 1751
Query: 1343 K 1343
K
Sbjct: 1752 K 1752
Score = 65.5 bits (158), Expect = 1e-09
Identities = 56/238 (23%), Positives = 99/238 (41%), Gaps = 11/238 (4%)
Query: 449 CDVCHLAKQKR----KMFTVSVSKAQKCFDLVHMDIWGPLAQASVHNHKYFLTVLDDYSR 504
C C + K + K + + + F +H DI+GP+ YF++ D+ ++
Sbjct: 637 CPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSYFISFTDETTK 696
Query: 505 FVWVVLLNNKGE--VQQQVKNFITLVKTQFGQIVKAIRSDNGPEF---LLPAFYSAQGIV 559
F WV L+++ E + + +K QF V I+ D G E+ L F GI
Sbjct: 697 FRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHKFLEKNGIT 756
Query: 560 HQKSCVSTPQQNGRVERKHQHILNIARALLFQSKLPKKMWCYSVLHAVFLMNRIPSKLLK 619
+ + + +G ER ++ +L+ R L S LP +W ++ + + N + S K
Sbjct: 757 PCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRNSLASPKSK 816
Query: 620 NKSPYELLYREAVDLEMMKVFGSLCFATTLTNNRSKLDPRARKCIFLGYKQGMKGYVL 677
KS + +D+ + FG N SK+ PR L + GY++
Sbjct: 817 -KSARQHAGLAGLDISTLLPFGQPVIVND-HNPNSKIHPRGIPGYALHPSRNSYGYII 872
>M820_ARATH (P92520) Hypothetical mitochondrial protein AtMg00820
(ORF170)
Length = 170
Score = 96.3 bits (238), Expect = 5e-19
Identities = 51/110 (46%), Positives = 70/110 (63%), Gaps = 2/110 (1%)
Query: 834 YALNIAQD--QEPSTFAEANKDLHWREAMQAEIKALEKNGTWKLVDLPEGVKPIGNKWVY 891
Y+L I +EP + A KD W +AMQ E+ AL +N TW LV P +G KWV+
Sbjct: 16 YSLTITTTIKKEPKSVIFALKDPGWCQAMQEELDALSRNKTWILVPPPVNQNILGCKWVF 75
Query: 892 RVKRNVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVRVILALA 941
+ K + DGTL R KARLVAKG++Q EG+ + +T+SPV + T+R IL +A
Sbjct: 76 KTKLHSDGTLDRLKARLVAKGFHQEEGIYFVETYSPVVRTATIRTILNVA 125
>M240_ARATH (P93290) Hypothetical mitochondrial protein AtMg00240
(ORF111a)
Length = 111
Score = 83.6 bits (205), Expect = 3e-15
Identities = 34/79 (43%), Positives = 53/79 (67%)
Query: 1139 LYLTTTRPDISHATQQLSQYMSNPMDGHFKAAQRVLRYLKASPGLGLLFPRNSTINIQGY 1198
+YLT TRPD++ A +LSQ+ S +A +VL Y+K + G GL + S + ++ +
Sbjct: 1 MYLTITRPDLTFAVNRLSQFSSASRTAQMQAVYKVLHYVKGTVGQGLFYSATSDLQLKAF 60
Query: 1199 SDADWAGCPDTRRSISGYC 1217
+D+DWA CPDTRRS++G+C
Sbjct: 61 ADSDWASCPDTRRSVTGFC 79
>POL_FENV1 (P31792) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)] (Fragment)
Length = 1046
Score = 47.4 bits (111), Expect = 3e-04
Identities = 48/172 (27%), Positives = 73/172 (41%), Gaps = 23/172 (13%)
Query: 492 HKYFLTVLDDYSRFVWVVLLNNKGEVQQQV-KNFITLVKTQFGQIVKAIRSDNGPEFLLP 550
+KY L +D +S WV + E V K + + +FG + K I SDNGP F+
Sbjct: 779 YKYLLVFVDTFSG--WVEAYPTRQETAHMVAKKILEEIFPRFG-LPKVIGSDNGPAFVSQ 835
Query: 551 AFYSAQ---GIVHQKSCVSTPQQNGRVERKHQHILNIARALLFQSKLPKKMWCYSVLHAV 607
GI + C PQ +G+VER ++ I L ++ L K W + A+
Sbjct: 836 VSQGLARTLGINWKLHCAYRPQSSGQVERMNRTIKETLTKLTLETGL--KDWRRLLSLAL 893
Query: 608 FLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLCFATTLTNNRSKLDPR 659
P++ +PYE+LY L +TL N+ S DP+
Sbjct: 894 LRARNTPNRF--GLTPYEILYGGPPPL------------STLLNSFSPSDPK 931
>POL_GALV (P21414) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1165
Score = 45.8 bits (107), Expect = 8e-04
Identities = 41/143 (28%), Positives = 63/143 (43%), Gaps = 11/143 (7%)
Query: 490 HNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQV-KNFITLVKTQFGQIVKAIRSDNGPEFL 548
+ +KY L +D +S WV K E V K + + +FG I K + SDNGP F+
Sbjct: 892 YGNKYLLVFIDTFSG--WVEAFPTKTETALIVCKKILEEILPRFG-IPKVLGSDNGPAFV 948
Query: 549 LPA---FYSAQGIVHQKSCVSTPQQNGRVERKHQHILNIARALLFQSKLPKKMWCYSVLH 605
+ GI + C PQ +G+VER ++ I L ++ K W +
Sbjct: 949 AQVSQGLATQLGINWKLHCAYRPQSSGQVERMNRTIKETLTKLALET--GGKDWVTLLPL 1006
Query: 606 AVFLMNRIPSKLLKNKSPYELLY 628
A+ P + +PYE+LY
Sbjct: 1007 ALLRARNTPGRF--GLTPYEILY 1027
>POL_BAEVM (P10272) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1189
Score = 45.8 bits (107), Expect = 8e-04
Identities = 41/141 (29%), Positives = 63/141 (44%), Gaps = 11/141 (7%)
Query: 492 HKYFLTVLDDYSRFVWVVLLNNKGEVQQQV-KNFITLVKTQFGQIVKAIRSDNGPEFLLP 550
+KY L +D +S WV + E V K + + +FG + K I SDNGP F+
Sbjct: 922 YKYLLVFVDTFSG--WVEAFPTRQETAHIVAKKILEEIFPRFG-LPKVIGSDNGPAFVSQ 978
Query: 551 AFYSAQ---GIVHQKSCVSTPQQNGRVERKHQHILNIARALLFQSKLPKKMWCYSVLHAV 607
GI + C PQ +G+VER ++ I L ++ L K W + A+
Sbjct: 979 VSQGLARILGINWKLHCAYRPQSSGQVERMNRTIKETLTKLTLETGL--KDWRRLLSLAL 1036
Query: 608 FLMNRIPSKLLKNKSPYELLY 628
P++ +PYE+LY
Sbjct: 1037 LRARNTPNRF--GLTPYEILY 1055
>POL_MLVAV (P03356) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1196
Score = 43.9 bits (102), Expect = 0.003
Identities = 41/156 (26%), Positives = 67/156 (42%), Gaps = 11/156 (7%)
Query: 477 HMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQV-KNFITLVKTQFGQI 535
H +I + ++ +KY L +D +S WV K E + V K + + +FG +
Sbjct: 912 HWEIDFTEVKPGLYGYKYLLVFVDTFSG--WVEAFPTKRETARVVSKKLLEEIFPRFG-M 968
Query: 536 VKAIRSDNGPEFLLPAFYSAQ---GIVHQKSCVSTPQQNGRVERKHQHILNIARALLFQS 592
+ + SDNGP F S GI + C PQ +G+VER ++ I L +
Sbjct: 969 PQVLGSDNGPAFTSQVSQSVADLLGIDWKLHCAYRPQSSGQVERMNRTIKETLTKLTLAA 1028
Query: 593 KLPKKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLY 628
+ W + A++ P +PYE+LY
Sbjct: 1029 --GTRDWVLLLPLALYRARNTPGP--HGLTPYEILY 1060
>POL_SFV1 (P23074) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1161
Score = 43.5 bits (101), Expect = 0.004
Identities = 47/165 (28%), Positives = 70/165 (41%), Gaps = 15/165 (9%)
Query: 468 KAQKCFDLVHMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKG-EVQQQVKNFIT 526
K K FD ++D GPL ++ + H L V+D + FVW+ + N +T
Sbjct: 879 KPLKPFDKFYIDYIGPLPPSNGYLH--VLVVVDSMTGFVWLYPTKAPSTSATVKALNMLT 936
Query: 527 LVKTQFGQIVKAIRSDNGPEFLLPAFYS---AQGIVHQKSCVSTPQQNGRVERKHQHILN 583
+ I K + SD G F F +GI + S PQ +G+VERK+ I
Sbjct: 937 SIA-----IPKVLHSDQGAAFTSSTFADWAKEKGIQLEFSTPYHPQSSGKVERKNSDIKR 991
Query: 584 IARALLFQSKLPKKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLY 628
+ LL P K W Y +L V L +P++LL+
Sbjct: 992 LLTKLLIGR--PAK-W-YDLLPVVQLALNNSYSPSSKYTPHQLLF 1032
>POL_MLVRK (P31795) Pol polyprotein [Contains: Protease (EC
3.4.23.-); Reverse transcriptase/ribonuclease H (EC
2.7.7.49) (EC 3.1.26.4) (RT); Integrase (IN)] (Fragment)
Length = 581
Score = 43.5 bits (101), Expect = 0.004
Identities = 56/225 (24%), Positives = 95/225 (41%), Gaps = 31/225 (13%)
Query: 422 RLGHLSHDRLLALHTVQNS----ISVSKSI--VCDVCHLAKQKRKMFTVSVSKAQKCFDL 475
RL HL + ++ AL S ++ K++ V D C + Q V+ SKA+ +
Sbjct: 234 RLTHLGYQKMKALLDRGESPYYMLNRDKTLQYVADSCTVCAQ------VNASKAKIGAGV 287
Query: 476 --------VHMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQV-KNFIT 526
H +I + ++ +KY L +D +S WV K E + V K +
Sbjct: 288 RVRGHRPGTHWEIDFTEVKPGLYGYKYLLVFVDTFSG--WVEAFPTKHETAKIVTKKLLE 345
Query: 527 LVKTQFGQIVKAIRSDNGPEFLLPAFYSAQ---GIVHQKSCVSTPQQNGRVERKHQHILN 583
+ +FG + + + +DNGP F+ S GI + C PQ +G+VER ++ I
Sbjct: 346 EIFPRFG-MPQVLGTDNGPAFVSQVSQSVAKLLGIDWKLHCAYRPQSSGQVERMNRTIKE 404
Query: 584 IARALLFQSKLPKKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLY 628
L + + W + A++ P +PYE+LY
Sbjct: 405 TLTKLTLAT--GTRDWVLLLPLALYRARNTPGP--HGLTPYEILY 445
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.335 0.144 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 140,027,485
Number of Sequences: 164201
Number of extensions: 5574437
Number of successful extensions: 20742
Number of sequences better than 10.0: 65
Number of HSP's better than 10.0 without gapping: 17
Number of HSP's successfully gapped in prelim test: 48
Number of HSP's that attempted gapping in prelim test: 20608
Number of HSP's gapped (non-prelim): 98
length of query: 1346
length of database: 59,974,054
effective HSP length: 122
effective length of query: 1224
effective length of database: 39,941,532
effective search space: 48888435168
effective search space used: 48888435168
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 72 (32.3 bits)
Lotus: description of TM0366.1


